Project
PRJNA195909:zebrafish embryo and larva development
Navigation
Downloads
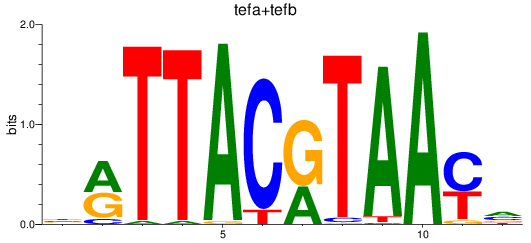
Results for tefa+tefb
Z-value: 0.90
Transcription factors associated with tefa+tefb
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
tefa
|
ENSDARG00000039117 | TEF transcription factor, PAR bZIP family member a |
|
tefb
|
ENSDARG00000098103 | TEF transcription factor, PAR bZIP family member b |
|
tefa
|
ENSDARG00000109532 | TEF transcription factor, PAR bZIP family member a |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| tefb | dr11_v1_chr3_-_5067585_5067585 | 0.89 | 1.2e-03 | Click! |
| tefa | dr11_v1_chr12_-_19151708_19151708 | -0.22 | 5.7e-01 | Click! |
Activity profile of tefa+tefb motif
Sorted Z-values of tefa+tefb motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr5_+_22974019 | 1.84 |
ENSDART00000147157
ENSDART00000020434 |
brwd3
|
bromodomain and WD repeat domain containing 3 |
| chr14_+_3507326 | 1.71 |
ENSDART00000159326
|
gstp1
|
glutathione S-transferase pi 1 |
| chr3_+_7763114 | 1.28 |
ENSDART00000057434
|
hook2
|
hook microtubule-tethering protein 2 |
| chr9_-_12888082 | 1.21 |
ENSDART00000133135
ENSDART00000134415 |
si:ch211-167j6.3
|
si:ch211-167j6.3 |
| chr16_+_39146696 | 1.12 |
ENSDART00000121756
ENSDART00000084381 |
sybu
|
syntabulin (syntaxin-interacting) |
| chr24_-_23784701 | 1.05 |
ENSDART00000090368
|
sgk3
|
serum/glucocorticoid regulated kinase family, member 3 |
| chr21_+_43178831 | 0.93 |
ENSDART00000151512
|
aff4
|
AF4/FMR2 family, member 4 |
| chr20_+_1121458 | 0.85 |
ENSDART00000064472
|
pnrc1
|
proline-rich nuclear receptor coactivator 1 |
| chr20_+_23083800 | 0.79 |
ENSDART00000132093
|
usp46
|
ubiquitin specific peptidase 46 |
| chr23_+_19590006 | 0.77 |
ENSDART00000021231
|
slmapb
|
sarcolemma associated protein b |
| chr16_-_35427060 | 0.75 |
ENSDART00000172294
|
ctps1b
|
CTP synthase 1b |
| chr18_+_25546227 | 0.71 |
ENSDART00000085824
|
pex11a
|
peroxisomal biogenesis factor 11 alpha |
| chr7_+_38380135 | 0.70 |
ENSDART00000174005
|
rhpn2
|
rhophilin, Rho GTPase binding protein 2 |
| chr7_-_40959667 | 0.68 |
ENSDART00000084070
|
rbm33a
|
RNA binding motif protein 33a |
| chr12_-_31484677 | 0.68 |
ENSDART00000066578
|
tectb
|
tectorin beta |
| chr20_-_44576949 | 0.66 |
ENSDART00000148639
|
ubxn2a
|
UBX domain protein 2A |
| chr5_-_30074332 | 0.63 |
ENSDART00000147963
|
bco2a
|
beta-carotene oxygenase 2a |
| chr12_-_23365737 | 0.63 |
ENSDART00000170376
|
mpp7a
|
membrane protein, palmitoylated 7a (MAGUK p55 subfamily member 7) |
| chr20_+_38201644 | 0.61 |
ENSDART00000022694
|
ehd3
|
EH-domain containing 3 |
| chr19_-_45650994 | 0.61 |
ENSDART00000163504
|
trps1
|
trichorhinophalangeal syndrome I |
| chr25_-_13490744 | 0.60 |
ENSDART00000056721
|
ldhd
|
lactate dehydrogenase D |
| chr12_-_26383242 | 0.59 |
ENSDART00000152941
|
usp54b
|
ubiquitin specific peptidase 54b |
| chr22_-_13165186 | 0.58 |
ENSDART00000105762
|
ahr2
|
aryl hydrocarbon receptor 2 |
| chr14_+_21699414 | 0.58 |
ENSDART00000169942
|
stx3a
|
syntaxin 3A |
| chr17_-_23709347 | 0.58 |
ENSDART00000124661
|
papss2a
|
3'-phosphoadenosine 5'-phosphosulfate synthase 2a |
| chr18_-_39288894 | 0.56 |
ENSDART00000186216
|
mapk6
|
mitogen-activated protein kinase 6 |
| chr4_-_14915268 | 0.56 |
ENSDART00000067040
|
si:dkey-180p18.9
|
si:dkey-180p18.9 |
| chr8_+_8712446 | 0.55 |
ENSDART00000158674
|
elk1
|
ELK1, member of ETS oncogene family |
| chr16_+_40954481 | 0.54 |
ENSDART00000058587
|
gbp
|
glycogen synthase kinase binding protein |
| chr23_+_9522781 | 0.52 |
ENSDART00000136486
|
osbpl2b
|
oxysterol binding protein-like 2b |
| chr7_+_34297271 | 0.51 |
ENSDART00000180342
|
bbox1
|
butyrobetaine (gamma), 2-oxoglutarate dioxygenase (gamma-butyrobetaine hydroxylase) 1 |
| chr18_-_43866526 | 0.51 |
ENSDART00000111309
|
treh
|
trehalase (brush-border membrane glycoprotein) |
| chr11_-_23459779 | 0.51 |
ENSDART00000183935
ENSDART00000184125 ENSDART00000193284 |
plekha6
|
pleckstrin homology domain containing, family A member 6 |
| chr9_-_13871935 | 0.51 |
ENSDART00000146597
|
raph1a
|
Ras association (RalGDS/AF-6) and pleckstrin homology domains 1a |
| chr9_+_17862858 | 0.51 |
ENSDART00000166566
|
dgkh
|
diacylglycerol kinase, eta |
| chr19_-_30810328 | 0.50 |
ENSDART00000184875
|
myclb
|
MYCL proto-oncogene, bHLH transcription factor b |
| chr25_-_12803723 | 0.50 |
ENSDART00000158787
|
ca5a
|
carbonic anhydrase Va |
| chr3_-_15144067 | 0.48 |
ENSDART00000127738
ENSDART00000060426 ENSDART00000180799 |
fam173a
|
family with sequence similarity 173, member A |
| chr5_-_54395488 | 0.47 |
ENSDART00000160781
|
zmynd19
|
zinc finger, MYND-type containing 19 |
| chr21_-_32781612 | 0.47 |
ENSDART00000031028
|
cnot6a
|
CCR4-NOT transcription complex, subunit 6a |
| chr9_-_5318873 | 0.46 |
ENSDART00000129308
|
ACVR1C
|
activin A receptor type 1C |
| chr1_-_354115 | 0.46 |
ENSDART00000141590
ENSDART00000098627 |
pros1
|
protein S |
| chr11_-_40147032 | 0.45 |
ENSDART00000052918
|
si:dkey-264d12.4
|
si:dkey-264d12.4 |
| chr7_+_52761841 | 0.45 |
ENSDART00000111444
|
ppip5k1a
|
diphosphoinositol pentakisphosphate kinase 1a |
| chr7_+_38278860 | 0.44 |
ENSDART00000016265
|
lrp3
|
low density lipoprotein receptor-related protein 3 |
| chr3_-_26184018 | 0.44 |
ENSDART00000191604
|
si:ch211-11k18.4
|
si:ch211-11k18.4 |
| chr19_+_7549854 | 0.44 |
ENSDART00000138866
ENSDART00000151758 |
pbxip1a
|
pre-B-cell leukemia homeobox interacting protein 1a |
| chr1_+_16600690 | 0.44 |
ENSDART00000162164
|
mtus1b
|
microtubule associated tumor suppressor 1b |
| chr22_+_15973122 | 0.44 |
ENSDART00000144545
|
rc3h1a
|
ring finger and CCCH-type domains 1a |
| chr3_+_43373867 | 0.43 |
ENSDART00000159455
ENSDART00000172425 |
zfand2a
|
zinc finger, AN1-type domain 2A |
| chr10_+_18911152 | 0.42 |
ENSDART00000030205
|
bnip3lb
|
BCL2 interacting protein 3 like b |
| chr8_+_21225064 | 0.42 |
ENSDART00000129210
|
cry1ba
|
cryptochrome circadian clock 1ba |
| chr13_-_42536642 | 0.40 |
ENSDART00000134533
|
btaf1
|
BTAF1 RNA polymerase II, B-TFIID transcription factor-associated |
| chr19_-_30811161 | 0.38 |
ENSDART00000103524
|
myclb
|
MYCL proto-oncogene, bHLH transcription factor b |
| chr8_-_29851706 | 0.37 |
ENSDART00000149297
|
slc20a2
|
solute carrier family 20 (phosphate transporter), member 2 |
| chr20_-_36617313 | 0.37 |
ENSDART00000172395
ENSDART00000152856 |
enah
|
enabled homolog (Drosophila) |
| chr3_-_26183699 | 0.37 |
ENSDART00000147517
ENSDART00000140731 |
si:ch211-11k18.4
|
si:ch211-11k18.4 |
| chr19_-_24757231 | 0.36 |
ENSDART00000128177
|
si:dkey-154b15.1
|
si:dkey-154b15.1 |
| chr1_+_19764995 | 0.35 |
ENSDART00000138276
|
si:ch211-42i9.8
|
si:ch211-42i9.8 |
| chr18_-_14879135 | 0.35 |
ENSDART00000099701
|
selenoo1
|
selenoprotein O1 |
| chr22_+_10606863 | 0.35 |
ENSDART00000147975
|
rad54l2
|
RAD54 like 2 |
| chr14_+_21699129 | 0.34 |
ENSDART00000073707
|
stx3a
|
syntaxin 3A |
| chr13_+_30035253 | 0.33 |
ENSDART00000181303
ENSDART00000057525 ENSDART00000136622 |
dnajb12a
|
DnaJ (Hsp40) homolog, subfamily B, member 12a |
| chr6_-_1553314 | 0.33 |
ENSDART00000077209
|
tpra1
|
transmembrane protein, adipocyte asscociated 1 |
| chr20_-_25533739 | 0.32 |
ENSDART00000063064
|
cyp2ad6
|
cytochrome P450, family 2, subfamily AD, polypeptide 6 |
| chr6_-_29612269 | 0.30 |
ENSDART00000104293
|
pex5la
|
peroxisomal biogenesis factor 5-like a |
| chr20_-_20270191 | 0.30 |
ENSDART00000009356
|
ppp2r5ea
|
protein phosphatase 2, regulatory subunit B', epsilon isoform a |
| chr18_-_2549198 | 0.30 |
ENSDART00000186516
|
CABZ01070631.1
|
|
| chr5_-_29238889 | 0.29 |
ENSDART00000143098
|
si:dkey-61l1.4
|
si:dkey-61l1.4 |
| chr25_+_15354095 | 0.29 |
ENSDART00000090397
|
kiaa1549la
|
KIAA1549-like a |
| chr15_+_28303161 | 0.29 |
ENSDART00000087926
|
myo1cb
|
myosin Ic, paralog b |
| chr7_+_17716601 | 0.25 |
ENSDART00000173792
ENSDART00000080825 |
rtn3
|
reticulon 3 |
| chr21_-_32467099 | 0.25 |
ENSDART00000186354
|
zgc:123105
|
zgc:123105 |
| chr7_+_38717624 | 0.24 |
ENSDART00000132522
|
syt13
|
synaptotagmin XIII |
| chr8_-_11324143 | 0.23 |
ENSDART00000008215
|
pip5k1bb
|
phosphatidylinositol-4-phosphate 5-kinase, type I, beta b |
| chr4_+_5341592 | 0.23 |
ENSDART00000123375
ENSDART00000067371 |
zgc:113263
|
zgc:113263 |
| chr10_+_26612321 | 0.23 |
ENSDART00000134322
|
fhl1b
|
four and a half LIM domains 1b |
| chr14_-_30905288 | 0.22 |
ENSDART00000173449
ENSDART00000173451 |
si:ch211-126c2.4
|
si:ch211-126c2.4 |
| chr6_+_48348415 | 0.21 |
ENSDART00000064826
|
mov10a
|
Mov10 RISC complex RNA helicase a |
| chr16_-_33806390 | 0.21 |
ENSDART00000160671
|
rspo1
|
R-spondin 1 |
| chr14_+_23717165 | 0.19 |
ENSDART00000006373
|
ndfip1
|
Nedd4 family interacting protein 1 |
| chr15_-_31419805 | 0.19 |
ENSDART00000060111
|
or111-11
|
odorant receptor, family D, subfamily 111, member 11 |
| chr23_-_21763598 | 0.18 |
ENSDART00000145408
|
vps13d
|
vacuolar protein sorting 13 homolog D (S. cerevisiae) |
| chr9_-_44953349 | 0.17 |
ENSDART00000135156
|
vil1
|
villin 1 |
| chr9_-_44953664 | 0.17 |
ENSDART00000188558
ENSDART00000185210 |
vil1
|
villin 1 |
| chr11_-_7078392 | 0.17 |
ENSDART00000112156
ENSDART00000188556 |
si:ch211-253b8.5
|
si:ch211-253b8.5 |
| chr15_+_21882419 | 0.16 |
ENSDART00000157216
|
si:dkey-103g5.4
|
si:dkey-103g5.4 |
| chr9_+_46644633 | 0.16 |
ENSDART00000160285
|
slc4a3
|
solute carrier family 4 (anion exchanger), member 3 |
| chr4_-_5302866 | 0.15 |
ENSDART00000138590
|
SNAP91 (1 of many)
|
si:ch211-214j24.9 |
| chr9_+_13682133 | 0.15 |
ENSDART00000175639
|
mpp4a
|
membrane protein, palmitoylated 4a (MAGUK p55 subfamily member 4) |
| chr5_-_22001003 | 0.14 |
ENSDART00000134393
ENSDART00000143878 |
arhgef9a
|
Cdc42 guanine nucleotide exchange factor (GEF) 9a |
| chr24_-_39772045 | 0.13 |
ENSDART00000087441
|
GFOD1
|
si:ch211-276f18.2 |
| chr22_+_10606573 | 0.12 |
ENSDART00000192638
|
rad54l2
|
RAD54 like 2 |
| chr5_-_21030934 | 0.12 |
ENSDART00000133461
ENSDART00000098667 |
camk2b1
|
calcium/calmodulin-dependent protein kinase (CaM kinase) II beta 1 |
| chr5_+_15350954 | 0.10 |
ENSDART00000140990
ENSDART00000137287 ENSDART00000061653 |
pebp1
|
phosphatidylethanolamine binding protein 1 |
| chr21_-_32467799 | 0.10 |
ENSDART00000007675
ENSDART00000133099 |
zgc:123105
|
zgc:123105 |
| chr16_-_19568388 | 0.08 |
ENSDART00000141616
|
abcb5
|
ATP-binding cassette, sub-family B (MDR/TAP), member 5 |
| chr11_+_25504215 | 0.07 |
ENSDART00000154213
|
tfe3b
|
transcription factor binding to IGHM enhancer 3b |
| chr14_+_36628131 | 0.07 |
ENSDART00000188625
ENSDART00000125345 |
TENM3
|
si:dkey-237h12.3 |
| chr12_+_36416173 | 0.06 |
ENSDART00000190278
|
unk
|
unkempt family zinc finger |
| chr23_+_4022620 | 0.06 |
ENSDART00000055099
|
b4galt5
|
UDP-Gal:betaGlcNAc beta 1,4- galactosyltransferase, polypeptide 5 |
| chr17_+_15845765 | 0.05 |
ENSDART00000130881
ENSDART00000074936 |
gabrr2a
|
gamma-aminobutyric acid (GABA) A receptor, rho 2a |
| chr22_-_10605045 | 0.05 |
ENSDART00000184812
|
bap1
|
BRCA1 associated protein-1 (ubiquitin carboxy-terminal hydrolase) |
| chr7_-_6604623 | 0.05 |
ENSDART00000172874
|
kcnj10a
|
potassium inwardly-rectifying channel, subfamily J, member 10a |
| chr23_-_18981595 | 0.04 |
ENSDART00000147617
|
bcl2l1
|
bcl2-like 1 |
| chr16_-_54800202 | 0.04 |
ENSDART00000059560
ENSDART00000161833 |
khdc4
|
KH domain containing 4, pre-mRNA splicing factor |
| chr5_-_65153731 | 0.04 |
ENSDART00000171024
|
sh2d3cb
|
SH2 domain containing 3Cb |
| chr15_+_31332552 | 0.03 |
ENSDART00000134933
ENSDART00000173915 |
or119-2
|
odorant receptor, family F, subfamily 119, member 2 |
| chr24_+_19592368 | 0.02 |
ENSDART00000180193
|
slco5a1a
|
solute carrier organic anion transporter family member 5A1a |
| chr15_-_16121496 | 0.01 |
ENSDART00000128624
|
sgk494a
|
uncharacterized serine/threonine-protein kinase SgK494a |
| chr15_+_24737599 | 0.00 |
ENSDART00000078024
|
crk
|
v-crk avian sarcoma virus CT10 oncogene homolog |
Network of associatons between targets according to the STRING database.
First level regulatory network of tefa+tefb
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.1 | GO:0060074 | synapse maturation(GO:0060074) |
| 0.2 | 0.6 | GO:0006089 | lactate metabolic process(GO:0006089) |
| 0.2 | 0.6 | GO:0050427 | sulfate assimilation(GO:0000103) purine ribonucleoside bisphosphate metabolic process(GO:0034035) purine ribonucleoside bisphosphate biosynthetic process(GO:0034036) 3'-phosphoadenosine 5'-phosphosulfate metabolic process(GO:0050427) 3'-phosphoadenosine 5'-phosphosulfate biosynthetic process(GO:0050428) |
| 0.2 | 0.7 | GO:0044210 | 'de novo' CTP biosynthetic process(GO:0044210) |
| 0.2 | 0.9 | GO:0098967 | exocytic insertion of neurotransmitter receptor to plasma membrane(GO:0098881) exocytic insertion of neurotransmitter receptor to postsynaptic membrane(GO:0098967) |
| 0.2 | 0.5 | GO:0005991 | trehalose metabolic process(GO:0005991) |
| 0.2 | 0.5 | GO:0042730 | fibrinolysis(GO:0042730) |
| 0.1 | 0.4 | GO:0070358 | actin polymerization-dependent cell motility(GO:0070358) |
| 0.1 | 0.7 | GO:0016559 | peroxisome fission(GO:0016559) |
| 0.1 | 0.8 | GO:0032228 | regulation of synaptic transmission, GABAergic(GO:0032228) |
| 0.1 | 0.3 | GO:2000392 | lamellipodium morphogenesis(GO:0072673) regulation of lamellipodium morphogenesis(GO:2000392) |
| 0.1 | 0.4 | GO:1903523 | negative regulation of heart contraction(GO:0045822) negative regulation of blood circulation(GO:1903523) |
| 0.1 | 0.5 | GO:0045329 | carnitine biosynthetic process(GO:0045329) |
| 0.1 | 0.4 | GO:2001106 | regulation of guanyl-nucleotide exchange factor activity(GO:1905097) regulation of Rho guanyl-nucleotide exchange factor activity(GO:2001106) |
| 0.1 | 0.6 | GO:0022602 | ovarian follicle development(GO:0001541) ovulation cycle process(GO:0022602) ovulation cycle(GO:0042698) |
| 0.0 | 1.7 | GO:0006749 | glutathione metabolic process(GO:0006749) |
| 0.0 | 0.7 | GO:0031468 | nuclear envelope reassembly(GO:0031468) |
| 0.0 | 0.5 | GO:0006020 | inositol metabolic process(GO:0006020) |
| 0.0 | 0.2 | GO:0045053 | protein retention in Golgi apparatus(GO:0045053) |
| 0.0 | 0.3 | GO:0022615 | protein import into peroxisome matrix, docking(GO:0016560) protein to membrane docking(GO:0022615) |
| 0.0 | 0.1 | GO:0050787 | response to mercury ion(GO:0046689) detoxification of mercury ion(GO:0050787) |
| 0.0 | 0.5 | GO:0007205 | protein kinase C-activating G-protein coupled receptor signaling pathway(GO:0007205) |
| 0.0 | 1.3 | GO:0031122 | cytoplasmic microtubule organization(GO:0031122) |
| 0.0 | 0.2 | GO:1904356 | regulation of telomere maintenance via telomere lengthening(GO:1904356) |
| 0.0 | 0.4 | GO:0035435 | phosphate ion transmembrane transport(GO:0035435) |
| 0.0 | 1.8 | GO:0008360 | regulation of cell shape(GO:0008360) |
| 0.0 | 0.9 | GO:0000288 | nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay(GO:0000288) |
| 0.0 | 0.3 | GO:0071218 | response to misfolded protein(GO:0051788) cellular response to misfolded protein(GO:0071218) |
| 0.0 | 0.3 | GO:0031952 | regulation of protein autophosphorylation(GO:0031952) |
| 0.0 | 0.6 | GO:0032456 | endocytic recycling(GO:0032456) |
| 0.0 | 0.1 | GO:0016121 | carotene metabolic process(GO:0016119) carotene catabolic process(GO:0016121) terpene metabolic process(GO:0042214) terpene catabolic process(GO:0046247) |
| 0.0 | 0.0 | GO:0035520 | monoubiquitinated protein deubiquitination(GO:0035520) monoubiquitinated histone deubiquitination(GO:0035521) monoubiquitinated histone H2A deubiquitination(GO:0035522) |
| 0.0 | 0.7 | GO:0048840 | otolith development(GO:0048840) |
| 0.0 | 0.2 | GO:0086001 | cardiac muscle cell action potential(GO:0086001) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.7 | GO:0097268 | cytoophidium(GO:0097268) |
| 0.1 | 0.9 | GO:0032783 | ELL-EAF complex(GO:0032783) |
| 0.1 | 0.7 | GO:0005779 | integral component of peroxisomal membrane(GO:0005779) intrinsic component of peroxisomal membrane(GO:0031231) |
| 0.1 | 0.3 | GO:0032433 | filopodium tip(GO:0032433) |
| 0.0 | 0.6 | GO:0034751 | aryl hydrocarbon receptor complex(GO:0034751) |
| 0.0 | 0.9 | GO:0048787 | presynaptic active zone membrane(GO:0048787) |
| 0.0 | 1.1 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.0 | 0.6 | GO:0060170 | ciliary membrane(GO:0060170) |
| 0.0 | 0.5 | GO:0030014 | CCR4-NOT complex(GO:0030014) |
| 0.0 | 0.2 | GO:0070187 | telosome(GO:0070187) |
| 0.0 | 0.4 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.6 | GO:0004779 | adenylylsulfate kinase activity(GO:0004020) sulfate adenylyltransferase activity(GO:0004779) sulfate adenylyltransferase (ATP) activity(GO:0004781) |
| 0.2 | 0.7 | GO:0003883 | CTP synthase activity(GO:0003883) |
| 0.2 | 0.5 | GO:0008336 | gamma-butyrobetaine dioxygenase activity(GO:0008336) |
| 0.2 | 0.5 | GO:0015927 | alpha,alpha-trehalase activity(GO:0004555) trehalase activity(GO:0015927) |
| 0.1 | 0.5 | GO:0033857 | inositol heptakisphosphate kinase activity(GO:0000829) diphosphoinositol-pentakisphosphate kinase activity(GO:0033857) |
| 0.1 | 0.6 | GO:0004457 | lactate dehydrogenase activity(GO:0004457) |
| 0.1 | 0.2 | GO:0032574 | 5'-3' RNA helicase activity(GO:0032574) |
| 0.1 | 0.4 | GO:0035613 | RNA stem-loop binding(GO:0035613) |
| 0.1 | 1.1 | GO:0019894 | kinesin binding(GO:0019894) |
| 0.0 | 0.2 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.0 | 0.3 | GO:0005052 | peroxisome matrix targeting signal-1 binding(GO:0005052) |
| 0.0 | 0.4 | GO:0005522 | profilin binding(GO:0005522) |
| 0.0 | 0.5 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.0 | 1.1 | GO:0015459 | potassium channel regulator activity(GO:0015459) |
| 0.0 | 0.4 | GO:0005315 | inorganic phosphate transmembrane transporter activity(GO:0005315) |
| 0.0 | 0.4 | GO:0070566 | adenylyltransferase activity(GO:0070566) |
| 0.0 | 0.3 | GO:0030544 | Hsp70 protein binding(GO:0030544) |
| 0.0 | 0.5 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.0 | 0.5 | GO:0015248 | sterol transporter activity(GO:0015248) |
| 0.0 | 0.1 | GO:1990715 | mRNA CDS binding(GO:1990715) |
| 0.0 | 0.1 | GO:0008489 | UDP-galactose:glucosylceramide beta-1,4-galactosyltransferase activity(GO:0008489) |
| 0.0 | 0.5 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.0 | 0.7 | GO:0051959 | dynein light intermediate chain binding(GO:0051959) |
| 0.0 | 0.3 | GO:0072542 | phosphatase activator activity(GO:0019211) protein phosphatase activator activity(GO:0072542) |
| 0.0 | 0.6 | GO:0004707 | MAP kinase activity(GO:0004707) |
| 0.0 | 0.9 | GO:0005484 | SNAP receptor activity(GO:0005484) |
| 0.0 | 0.1 | GO:0003834 | beta-carotene 15,15'-monooxygenase activity(GO:0003834) carotenoid dioxygenase activity(GO:0010436) |
| 0.0 | 0.4 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.0 | 1.4 | GO:0036459 | thiol-dependent ubiquitin-specific protease activity(GO:0004843) thiol-dependent ubiquitinyl hydrolase activity(GO:0036459) ubiquitinyl hydrolase activity(GO:0101005) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | PID TCR RAS PATHWAY | Ras signaling in the CD4+ TCR pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.7 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.1 | 0.5 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 0.1 | 0.5 | REACTOME DIGESTION OF DIETARY CARBOHYDRATE | Genes involved in Digestion of dietary carbohydrate |
| 0.0 | 0.5 | REACTOME SIGNALING BY NODAL | Genes involved in Signaling by NODAL |
| 0.0 | 0.5 | REACTOME REVERSIBLE HYDRATION OF CARBON DIOXIDE | Genes involved in Reversible Hydration of Carbon Dioxide |
| 0.0 | 0.4 | REACTOME GENERATION OF SECOND MESSENGER MOLECULES | Genes involved in Generation of second messenger molecules |
| 0.0 | 0.5 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.0 | 0.6 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |