Project
PRJNA195909:zebrafish embryo and larva development
Navigation
Downloads
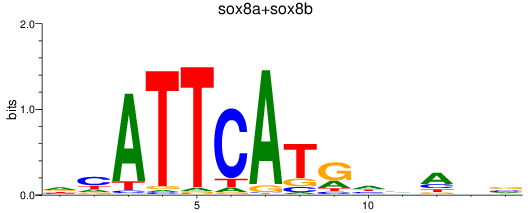
Results for sox8a+sox8b
Z-value: 0.94
Transcription factors associated with sox8a+sox8b
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
sox8b
|
ENSDARG00000037782 | SRY-box transcription factor 8b |
|
sox8a
|
ENSDARG00000105301 | SRY-box transcription factor 8a |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| sox8a | dr11_v1_chr12_-_1085227_1085227 | -0.85 | 3.9e-03 | Click! |
| sox8b | dr11_v1_chr3_-_62403550_62403550 | 0.13 | 7.4e-01 | Click! |
Activity profile of sox8a+sox8b motif
Sorted Z-values of sox8a+sox8b motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr9_-_50001606 | 2.08 |
ENSDART00000161648
ENSDART00000168514 |
scn1a
|
sodium channel, voltage-gated, type I, alpha |
| chr17_-_33707589 | 1.82 |
ENSDART00000124788
|
si:dkey-84k17.2
|
si:dkey-84k17.2 |
| chr3_-_3413669 | 1.65 |
ENSDART00000113517
ENSDART00000179861 ENSDART00000115331 |
zgc:171446
|
zgc:171446 |
| chr20_-_39789036 | 1.43 |
ENSDART00000086405
ENSDART00000098253 |
rnf217
|
ring finger protein 217 |
| chr3_-_5067585 | 1.31 |
ENSDART00000169609
|
tefb
|
thyrotrophic embryonic factor b |
| chr13_+_8892784 | 1.27 |
ENSDART00000075054
ENSDART00000143705 |
thada
|
thyroid adenoma associated |
| chr9_+_50001746 | 1.23 |
ENSDART00000058892
|
slc38a11
|
solute carrier family 38, member 11 |
| chr12_-_33359052 | 1.22 |
ENSDART00000135943
|
slc16a3
|
solute carrier family 16 (monocarboxylate transporter), member 3 |
| chr12_-_33359654 | 1.19 |
ENSDART00000001907
|
slc16a3
|
solute carrier family 16 (monocarboxylate transporter), member 3 |
| chr19_-_27448606 | 1.08 |
ENSDART00000079251
|
vars2
|
valyl-tRNA synthetase 2, mitochondrial (putative) |
| chr1_-_23308225 | 1.06 |
ENSDART00000137567
ENSDART00000008201 |
smim14
|
small integral membrane protein 14 |
| chr9_+_34952203 | 1.03 |
ENSDART00000121828
ENSDART00000142347 |
tfdp1a
|
transcription factor Dp-1, a |
| chr19_+_27448625 | 0.97 |
ENSDART00000052352
|
ppp1r11
|
protein phosphatase 1, regulatory (inhibitor) subunit 11 |
| chr24_-_41220538 | 0.95 |
ENSDART00000150207
|
acvr2ba
|
activin A receptor type 2Ba |
| chr8_+_10862353 | 0.94 |
ENSDART00000140717
|
brpf3b
|
bromodomain and PHD finger containing, 3b |
| chr11_+_30254556 | 0.91 |
ENSDART00000182316
|
ppef1
|
protein phosphatase, EF-hand calcium binding domain 1 |
| chr5_+_45139196 | 0.89 |
ENSDART00000113738
|
smarca2
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 |
| chr19_+_11432330 | 0.85 |
ENSDART00000188065
|
PQLC1
|
PQ loop repeat containing 1 |
| chr21_+_39673157 | 0.84 |
ENSDART00000100240
|
mrm3b
|
mitochondrial rRNA methyltransferase 3b |
| chr6_-_37743724 | 0.81 |
ENSDART00000149068
|
nipa2
|
non imprinted in Prader-Willi/Angelman syndrome 2 (human) |
| chr21_+_42717424 | 0.80 |
ENSDART00000166936
ENSDART00000172135 |
sh3pxd2b
|
SH3 and PX domains 2B |
| chr8_-_7603516 | 0.77 |
ENSDART00000179826
ENSDART00000190153 |
irak1
|
interleukin-1 receptor-associated kinase 1 |
| chr10_+_5268054 | 0.76 |
ENSDART00000114491
|
ror2
|
receptor tyrosine kinase-like orphan receptor 2 |
| chr25_-_37248795 | 0.75 |
ENSDART00000087247
ENSDART00000154045 |
glg1a
|
golgi glycoprotein 1a |
| chr3_+_32663865 | 0.75 |
ENSDART00000190240
|
si:dkey-16l2.16
|
si:dkey-16l2.16 |
| chr22_-_547748 | 0.74 |
ENSDART00000037455
ENSDART00000140101 |
ccnd3
|
cyclin D3 |
| chr7_+_52753067 | 0.72 |
ENSDART00000053814
|
mtfmt
|
mitochondrial methionyl-tRNA formyltransferase |
| chr11_+_13176568 | 0.71 |
ENSDART00000125371
ENSDART00000123257 |
mknk1
|
MAP kinase interacting serine/threonine kinase 1 |
| chr9_-_8979468 | 0.70 |
ENSDART00000134646
|
ing1
|
inhibitor of growth family, member 1 |
| chr5_+_45138934 | 0.70 |
ENSDART00000041412
ENSDART00000136002 |
smarca2
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily a, member 2 |
| chr2_+_24770435 | 0.70 |
ENSDART00000078854
|
mpv17l2
|
MPV17 mitochondrial membrane protein-like 2 |
| chr12_-_11593436 | 0.69 |
ENSDART00000138954
|
plekha1b
|
pleckstrin homology domain containing, family A (phosphoinositide binding specific) member 1b |
| chr25_+_4787431 | 0.69 |
ENSDART00000170640
|
myo5c
|
myosin VC |
| chr16_-_21047483 | 0.68 |
ENSDART00000136235
|
cbx3b
|
chromobox homolog 3b |
| chr8_-_7603700 | 0.68 |
ENSDART00000137975
|
irak1
|
interleukin-1 receptor-associated kinase 1 |
| chr10_+_9595575 | 0.66 |
ENSDART00000091780
ENSDART00000184287 |
rc3h2
|
ring finger and CCCH-type domains 2 |
| chr11_-_18015534 | 0.64 |
ENSDART00000181953
|
qrich1
|
glutamine-rich 1 |
| chr3_+_41726360 | 0.62 |
ENSDART00000154401
|
chst12a
|
carbohydrate (chondroitin 4) sulfotransferase 12a |
| chr13_-_8892514 | 0.61 |
ENSDART00000144553
|
plekhh2
|
pleckstrin homology domain containing, family H (with MyTH4 domain) member 2 |
| chr25_+_4787607 | 0.61 |
ENSDART00000159422
|
myo5c
|
myosin VC |
| chr9_-_3934963 | 0.60 |
ENSDART00000062336
|
ubr3
|
ubiquitin protein ligase E3 component n-recognin 3 |
| chr4_+_6833735 | 0.60 |
ENSDART00000136355
|
dock4b
|
dedicator of cytokinesis 4b |
| chr21_+_34088377 | 0.60 |
ENSDART00000170070
|
mtmr1b
|
myotubularin related protein 1b |
| chr22_+_9522971 | 0.60 |
ENSDART00000110048
|
strip1
|
striatin interacting protein 1 |
| chr21_+_43328685 | 0.59 |
ENSDART00000109620
ENSDART00000139668 |
sept8a
|
septin 8a |
| chr7_+_25003851 | 0.58 |
ENSDART00000144526
|
naa40
|
N(alpha)-acetyltransferase 40, NatD catalytic subunit, homolog (S. cerevisiae) |
| chr3_-_59981162 | 0.57 |
ENSDART00000128790
|
cdr2l
|
cerebellar degeneration-related protein 2-like |
| chr2_+_23222939 | 0.57 |
ENSDART00000026800
|
kifap3b
|
kinesin-associated protein 3b |
| chr15_+_3125136 | 0.55 |
ENSDART00000130968
|
rcbtb2
|
regulator of chromosome condensation (RCC1) and BTB (POZ) domain containing protein 2 |
| chr8_+_42718135 | 0.55 |
ENSDART00000158900
|
zgc:194007
|
zgc:194007 |
| chr2_+_15048410 | 0.52 |
ENSDART00000058484
|
cnn3b
|
calponin 3, acidic b |
| chr18_+_45666489 | 0.51 |
ENSDART00000180147
ENSDART00000151351 |
prrg4
|
proline rich Gla (G-carboxyglutamic acid) 4 (transmembrane) |
| chr2_+_29925561 | 0.50 |
ENSDART00000163118
ENSDART00000183800 ENSDART00000180738 ENSDART00000139604 |
htr5aa
|
5-hydroxytryptamine (serotonin) receptor 5A, genome duplicate a |
| chr3_+_62327332 | 0.48 |
ENSDART00000158414
|
BX470259.1
|
|
| chr21_+_34088110 | 0.47 |
ENSDART00000145123
ENSDART00000029599 ENSDART00000147519 |
mtmr1b
|
myotubularin related protein 1b |
| chr18_+_5917625 | 0.47 |
ENSDART00000169100
|
glg1b
|
golgi glycoprotein 1b |
| chr19_-_11950434 | 0.45 |
ENSDART00000024861
|
cpsf1
|
cleavage and polyadenylation specific factor 1 |
| chr19_-_11949996 | 0.44 |
ENSDART00000163478
ENSDART00000167299 |
cpsf1
|
cleavage and polyadenylation specific factor 1 |
| chr23_-_36418708 | 0.44 |
ENSDART00000132273
|
znf740b
|
zinc finger protein 740b |
| chr20_-_3276339 | 0.43 |
ENSDART00000166831
|
rps6kc1
|
ribosomal protein S6 kinase polypeptide 1 |
| chr16_+_46410520 | 0.42 |
ENSDART00000131072
|
rpz2
|
rapunzel 2 |
| chr22_-_817479 | 0.41 |
ENSDART00000123487
|
zgc:153675
|
zgc:153675 |
| chr6_+_7322587 | 0.40 |
ENSDART00000065500
|
abcc4
|
ATP-binding cassette, sub-family C (CFTR/MRP), member 4 |
| chr11_-_43226255 | 0.40 |
ENSDART00000172929
|
sptbn1
|
spectrin, beta, non-erythrocytic 1 |
| chr2_-_22492289 | 0.40 |
ENSDART00000168022
|
gtf2b
|
general transcription factor IIB |
| chr10_+_15967643 | 0.38 |
ENSDART00000136709
|
apbb1
|
amyloid beta (A4) precursor protein-binding, family B, member 1 (Fe65) |
| chr7_+_65121743 | 0.36 |
ENSDART00000108471
|
ipo7
|
importin 7 |
| chr12_+_34258139 | 0.36 |
ENSDART00000153127
|
socs3b
|
suppressor of cytokine signaling 3b |
| chr10_-_7386475 | 0.36 |
ENSDART00000167963
|
nrg1
|
neuregulin 1 |
| chr9_-_8979154 | 0.34 |
ENSDART00000145266
|
ing1
|
inhibitor of growth family, member 1 |
| chr4_+_6833583 | 0.34 |
ENSDART00000165179
ENSDART00000186134 ENSDART00000174507 |
dock4b
|
dedicator of cytokinesis 4b |
| chr11_-_2478374 | 0.33 |
ENSDART00000173205
|
si:ch73-267c23.10
|
si:ch73-267c23.10 |
| chr5_-_6567464 | 0.32 |
ENSDART00000184985
|
tnks1bp1
|
tankyrase 1 binding protein 1 |
| chr17_+_24604373 | 0.32 |
ENSDART00000082193
|
zmpste24
|
zinc metallopeptidase, STE24 homolog |
| chr9_-_43538328 | 0.32 |
ENSDART00000140526
|
znf385b
|
zinc finger protein 385B |
| chr5_+_31214341 | 0.32 |
ENSDART00000133432
|
atp2a3
|
ATPase sarcoplasmic/endoplasmic reticulum Ca2+ transporting 3 |
| chr10_+_11398111 | 0.29 |
ENSDART00000064210
|
cwc27
|
CWC27 spliceosome-associated protein homolog (S. cerevisiae) |
| chr7_+_17096281 | 0.28 |
ENSDART00000035558
|
htatip2
|
HIV-1 Tat interactive protein 2 |
| chr3_+_39579393 | 0.28 |
ENSDART00000055170
|
cln3
|
ceroid-lipofuscinosis, neuronal 3 |
| chr3_-_59981476 | 0.28 |
ENSDART00000035878
ENSDART00000124038 |
cdr2l
|
cerebellar degeneration-related protein 2-like |
| chr15_-_9419472 | 0.27 |
ENSDART00000122117
|
sacs
|
sacsin molecular chaperone |
| chr16_-_21047872 | 0.26 |
ENSDART00000131582
|
cbx3b
|
chromobox homolog 3b |
| chr24_-_3304664 | 0.24 |
ENSDART00000183168
|
sugct
|
succinyl-CoA:glutarate-CoA transferase |
| chr20_+_25936370 | 0.22 |
ENSDART00000141265
|
znf106b
|
zinc finger protein 106b |
| chr19_-_9648542 | 0.21 |
ENSDART00000172628
|
clcn1a
|
chloride channel, voltage-sensitive 1a |
| chr20_-_39391833 | 0.20 |
ENSDART00000135149
|
si:dkey-217m5.8
|
si:dkey-217m5.8 |
| chr5_+_41146786 | 0.20 |
ENSDART00000175766
|
sub1b
|
SUB1 homolog, transcriptional regulator b |
| chr1_+_6646529 | 0.18 |
ENSDART00000144641
ENSDART00000103701 ENSDART00000138919 |
ube2f
|
ubiquitin-conjugating enzyme E2F (putative) |
| chr14_+_31721171 | 0.18 |
ENSDART00000173034
|
adgrg4a
|
adhesion G protein-coupled receptor G4a |
| chr10_+_7709724 | 0.16 |
ENSDART00000097670
|
ggcx
|
gamma-glutamyl carboxylase |
| chr20_-_39391492 | 0.16 |
ENSDART00000184886
|
si:dkey-217m5.8
|
si:dkey-217m5.8 |
| chr8_-_46386024 | 0.14 |
ENSDART00000136602
ENSDART00000060919 ENSDART00000137472 |
qars
|
glutaminyl-tRNA synthetase |
| chr22_+_38301365 | 0.13 |
ENSDART00000137339
|
traf5
|
Tnf receptor-associated factor 5 |
| chr9_+_17423941 | 0.13 |
ENSDART00000112884
ENSDART00000155233 |
kbtbd7
|
kelch repeat and BTB (POZ) domain containing 7 |
| chr22_+_24673168 | 0.13 |
ENSDART00000135257
|
si:rp71-23d18.8
|
si:rp71-23d18.8 |
| chr12_+_18544159 | 0.12 |
ENSDART00000162900
|
mlst8
|
MTOR associated protein, LST8 homolog (S. cerevisiae) |
| chr2_+_51645164 | 0.10 |
ENSDART00000169600
|
abhd4
|
abhydrolase domain containing 4 |
| chr9_-_3034064 | 0.08 |
ENSDART00000115138
ENSDART00000144046 |
rapgef4
|
Rap guanine nucleotide exchange factor (GEF) 4 |
| chr19_-_17208728 | 0.06 |
ENSDART00000151228
|
stmn1a
|
stathmin 1a |
| chr24_-_33873451 | 0.04 |
ENSDART00000159840
|
asic1c
|
acid-sensing (proton-gated) ion channel 1c |
| chr19_-_24198139 | 0.03 |
ENSDART00000138864
ENSDART00000125350 |
prf1.7
|
perforin 1.7 |
| chr1_-_16688965 | 0.03 |
ENSDART00000079001
ENSDART00000178599 |
frg1
|
FSHD region gene 1 |
| chr10_-_14545658 | 0.02 |
ENSDART00000057865
|
ier3ip1
|
immediate early response 3 interacting protein 1 |
| chr10_-_22249444 | 0.00 |
ENSDART00000148831
|
fgf11b
|
fibroblast growth factor 11b |
Network of associatons between targets according to the STRING database.
First level regulatory network of sox8a+sox8b
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.6 | GO:0031507 | heterochromatin assembly(GO:0031507) |
| 0.2 | 0.7 | GO:0061668 | mitochondrial ribosome assembly(GO:0061668) |
| 0.2 | 1.4 | GO:0070498 | interleukin-1-mediated signaling pathway(GO:0070498) |
| 0.1 | 0.8 | GO:0071800 | podosome assembly(GO:0071800) |
| 0.1 | 1.0 | GO:0032515 | negative regulation of phosphoprotein phosphatase activity(GO:0032515) |
| 0.1 | 0.6 | GO:0071596 | ubiquitin-dependent protein catabolic process via the N-end rule pathway(GO:0071596) |
| 0.1 | 0.4 | GO:0060959 | activation of transmembrane receptor protein tyrosine kinase activity(GO:0007171) Schwann cell migration(GO:0036135) cardiac neuron differentiation(GO:0060945) cardiac neuron development(GO:0060959) |
| 0.1 | 0.3 | GO:0071586 | CAAX-box protein processing(GO:0071586) CAAX-box protein maturation(GO:0080120) |
| 0.1 | 0.9 | GO:0044154 | histone H3-K14 acetylation(GO:0044154) |
| 0.1 | 1.3 | GO:1990798 | pancreas regeneration(GO:1990798) |
| 0.0 | 0.2 | GO:0060261 | positive regulation of transcription initiation from RNA polymerase II promoter(GO:0060261) positive regulation of DNA-templated transcription, initiation(GO:2000144) |
| 0.0 | 0.7 | GO:1900087 | positive regulation of G1/S transition of mitotic cell cycle(GO:1900087) |
| 0.0 | 0.8 | GO:0015693 | magnesium ion transport(GO:0015693) |
| 0.0 | 1.1 | GO:0006418 | tRNA aminoacylation for protein translation(GO:0006418) |
| 0.0 | 0.6 | GO:0006474 | N-terminal protein amino acid acetylation(GO:0006474) |
| 0.0 | 0.3 | GO:0015809 | arginine transport(GO:0015809) |
| 0.0 | 0.9 | GO:0006378 | mRNA polyadenylation(GO:0006378) RNA polyadenylation(GO:0043631) |
| 0.0 | 0.4 | GO:0051123 | RNA polymerase II transcriptional preinitiation complex assembly(GO:0051123) |
| 0.0 | 0.8 | GO:0014068 | positive regulation of phosphatidylinositol 3-kinase signaling(GO:0014068) |
| 0.0 | 2.4 | GO:0015718 | monocarboxylic acid transport(GO:0015718) |
| 0.0 | 0.2 | GO:0017187 | peptidyl-glutamic acid carboxylation(GO:0017187) protein carboxylation(GO:0018214) |
| 0.0 | 1.4 | GO:0032436 | positive regulation of proteasomal ubiquitin-dependent protein catabolic process(GO:0032436) positive regulation of proteasomal protein catabolic process(GO:1901800) |
| 0.0 | 1.3 | GO:0099515 | vesicle transport along actin filament(GO:0030050) actin filament-based transport(GO:0099515) |
| 0.0 | 1.2 | GO:0003333 | amino acid transmembrane transport(GO:0003333) |
| 0.0 | 0.5 | GO:0007198 | adenylate cyclase-inhibiting serotonin receptor signaling pathway(GO:0007198) |
| 0.0 | 0.3 | GO:0006658 | phosphatidylserine metabolic process(GO:0006658) |
| 0.0 | 0.4 | GO:0098754 | detoxification(GO:0098754) |
| 0.0 | 1.1 | GO:0046856 | phosphatidylinositol dephosphorylation(GO:0046856) |
| 0.0 | 1.0 | GO:0032924 | activin receptor signaling pathway(GO:0032924) |
| 0.0 | 1.3 | GO:0071482 | cellular response to light stimulus(GO:0071482) |
| 0.0 | 0.9 | GO:0050906 | detection of stimulus involved in sensory perception(GO:0050906) |
| 0.0 | 0.6 | GO:0032456 | endocytic recycling(GO:0032456) |
| 0.0 | 0.2 | GO:0045116 | protein neddylation(GO:0045116) |
| 0.0 | 0.7 | GO:0000288 | nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay(GO:0000288) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0008091 | spectrin(GO:0008091) |
| 0.1 | 1.0 | GO:0048179 | activin receptor complex(GO:0048179) |
| 0.1 | 0.8 | GO:0002102 | podosome(GO:0002102) |
| 0.1 | 0.9 | GO:0005847 | mRNA cleavage and polyadenylation specificity factor complex(GO:0005847) |
| 0.1 | 0.9 | GO:0070775 | H3 histone acetyltransferase complex(GO:0070775) MOZ/MORF histone acetyltransferase complex(GO:0070776) |
| 0.0 | 0.4 | GO:0097550 | transcriptional preinitiation complex(GO:0097550) |
| 0.0 | 1.6 | GO:0016514 | SWI/SNF complex(GO:0016514) |
| 0.0 | 0.3 | GO:0071005 | U2-type precatalytic spliceosome(GO:0071005) |
| 0.0 | 0.7 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
| 0.0 | 0.6 | GO:0035869 | ciliary transition zone(GO:0035869) |
| 0.0 | 0.7 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.0 | 0.6 | GO:0032156 | septin ring(GO:0005940) septin complex(GO:0031105) septin cytoskeleton(GO:0032156) |
| 0.0 | 0.1 | GO:0031931 | TORC1 complex(GO:0031931) |
| 0.0 | 0.3 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.0 | 0.4 | GO:0055038 | recycling endosome membrane(GO:0055038) |
| 0.0 | 1.3 | GO:0016459 | myosin complex(GO:0016459) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 2.4 | GO:0015129 | lactate transmembrane transporter activity(GO:0015129) |
| 0.1 | 0.6 | GO:0043998 | H2A histone acetyltransferase activity(GO:0043998) |
| 0.1 | 0.4 | GO:0015562 | efflux transmembrane transporter activity(GO:0015562) |
| 0.1 | 1.1 | GO:0002161 | aminoacyl-tRNA editing activity(GO:0002161) |
| 0.1 | 1.0 | GO:0004865 | protein serine/threonine phosphatase inhibitor activity(GO:0004865) |
| 0.1 | 1.1 | GO:0052629 | phosphatidylinositol-3,5-bisphosphate 3-phosphatase activity(GO:0052629) |
| 0.1 | 0.7 | GO:0035613 | RNA stem-loop binding(GO:0035613) |
| 0.1 | 0.8 | GO:0016175 | superoxide-generating NADPH oxidase activity(GO:0016175) superoxide-generating NADPH oxidase activator activity(GO:0016176) |
| 0.1 | 0.7 | GO:0016742 | hydroxymethyl-, formyl- and related transferase activity(GO:0016742) |
| 0.1 | 0.9 | GO:0043994 | H3 histone acetyltransferase activity(GO:0010484) histone acetyltransferase activity (H3-K23 specific)(GO:0043994) |
| 0.1 | 0.4 | GO:0001046 | core promoter sequence-specific DNA binding(GO:0001046) core promoter binding(GO:0001047) |
| 0.1 | 1.0 | GO:0017002 | activin-activated receptor activity(GO:0017002) |
| 0.1 | 1.2 | GO:0017134 | fibroblast growth factor binding(GO:0017134) |
| 0.0 | 0.8 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.0 | 0.2 | GO:0008488 | gamma-glutamyl carboxylase activity(GO:0008488) |
| 0.0 | 1.4 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
| 0.0 | 0.4 | GO:0030296 | protein tyrosine kinase activator activity(GO:0030296) |
| 0.0 | 0.2 | GO:0008410 | CoA-transferase activity(GO:0008410) |
| 0.0 | 1.0 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.0 | 0.6 | GO:0019894 | kinesin binding(GO:0019894) |
| 0.0 | 1.3 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.0 | 0.8 | GO:0017147 | Wnt-protein binding(GO:0017147) |
| 0.0 | 1.6 | GO:0008094 | DNA-dependent ATPase activity(GO:0008094) |
| 0.0 | 0.5 | GO:0043176 | amine binding(GO:0043176) serotonin binding(GO:0051378) |
| 0.0 | 0.8 | GO:0030145 | manganese ion binding(GO:0030145) |
| 0.0 | 0.3 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.0 | 0.7 | GO:0009931 | calcium-dependent protein serine/threonine kinase activity(GO:0009931) |
| 0.0 | 0.3 | GO:0030544 | Hsp70 protein binding(GO:0030544) |
| 0.0 | 0.5 | GO:0030165 | PDZ domain binding(GO:0030165) |
| 0.0 | 0.4 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.0 | 0.8 | GO:0008173 | RNA methyltransferase activity(GO:0008173) |
| 0.0 | 0.2 | GO:0005247 | voltage-gated chloride channel activity(GO:0005247) |
| 0.0 | 0.7 | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity(GO:0016538) |
| 0.0 | 0.4 | GO:0046935 | 1-phosphatidylinositol-3-kinase regulator activity(GO:0046935) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.4 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.0 | 0.8 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
| 0.0 | 0.4 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.0 | 1.6 | PID AR TF PATHWAY | Regulation of Androgen receptor activity |
| 0.0 | 0.7 | PID IL2 STAT5 PATHWAY | IL2 signaling events mediated by STAT5 |
| 0.0 | 0.7 | ST ERK1 ERK2 MAPK PATHWAY | ERK1/ERK2 MAPK Pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.4 | REACTOME NFKB IS ACTIVATED AND SIGNALS SURVIVAL | Genes involved in NF-kB is activated and signals survival |
| 0.2 | 2.4 | REACTOME BILE SALT AND ORGANIC ANION SLC TRANSPORTERS | Genes involved in Bile salt and organic anion SLC transporters |
| 0.1 | 1.2 | REACTOME MITOCHONDRIAL TRNA AMINOACYLATION | Genes involved in Mitochondrial tRNA aminoacylation |
| 0.1 | 0.7 | REACTOME SPRY REGULATION OF FGF SIGNALING | Genes involved in Spry regulation of FGF signaling |
| 0.0 | 0.4 | REACTOME DOWNREGULATION OF ERBB2 ERBB3 SIGNALING | Genes involved in Downregulation of ERBB2:ERBB3 signaling |
| 0.0 | 0.9 | REACTOME PROCESSING OF INTRONLESS PRE MRNAS | Genes involved in Processing of Intronless Pre-mRNAs |
| 0.0 | 0.4 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.0 | 0.7 | REACTOME G1 PHASE | Genes involved in G1 Phase |
| 0.0 | 0.3 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.0 | 0.2 | REACTOME GAMMA CARBOXYLATION TRANSPORT AND AMINO TERMINAL CLEAVAGE OF PROTEINS | Genes involved in Gamma-carboxylation, transport, and amino-terminal cleavage of proteins |
| 0.0 | 0.3 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.0 | 0.4 | REACTOME ABC FAMILY PROTEINS MEDIATED TRANSPORT | Genes involved in ABC-family proteins mediated transport |
| 0.0 | 0.1 | REACTOME MTORC1 MEDIATED SIGNALLING | Genes involved in mTORC1-mediated signalling |