Project
PRJNA195909:zebrafish embryo and larva development
Navigation
Downloads
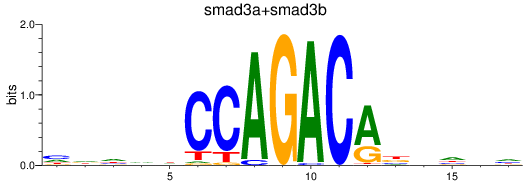
Results for smad3a+smad3b
Z-value: 0.90
Transcription factors associated with smad3a+smad3b
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
smad3b
|
ENSDARG00000010207 | SMAD family member 3b |
|
smad3a
|
ENSDARG00000036096 | SMAD family member 3a |
|
smad3a
|
ENSDARG00000117146 | SMAD family member 3a |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| smad3b | dr11_v1_chr18_+_19772874_19772874 | 0.26 | 5.0e-01 | Click! |
| smad3a | dr11_v1_chr7_-_34149263_34149263 | 0.10 | 7.9e-01 | Click! |
Activity profile of smad3a+smad3b motif
Sorted Z-values of smad3a+smad3b motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr12_+_16949391 | 1.81 |
ENSDART00000152635
|
zgc:174153
|
zgc:174153 |
| chr5_+_24305877 | 1.50 |
ENSDART00000144226
|
ctsll
|
cathepsin L, like |
| chr2_-_20599315 | 1.25 |
ENSDART00000114199
|
si:ch211-267e7.3
|
si:ch211-267e7.3 |
| chr2_-_7666021 | 1.09 |
ENSDART00000180007
|
CABZ01021592.1
|
|
| chr23_-_21463788 | 1.02 |
ENSDART00000079265
|
her4.4
|
hairy-related 4, tandem duplicate 4 |
| chr8_-_13541514 | 1.02 |
ENSDART00000063834
|
zgc:86586
|
zgc:86586 |
| chr14_+_38786298 | 0.96 |
ENSDART00000164440
|
si:ch211-195b11.3
|
si:ch211-195b11.3 |
| chr23_+_21455152 | 0.95 |
ENSDART00000158511
ENSDART00000161321 ENSDART00000160731 ENSDART00000137573 |
her4.2
|
hairy-related 4, tandem duplicate 2 |
| chr1_-_52494122 | 0.93 |
ENSDART00000131407
|
acy3.2
|
aspartoacylase (aminocyclase) 3, tandem duplicate 2 |
| chr23_-_21453614 | 0.87 |
ENSDART00000079274
|
her4.1
|
hairy-related 4, tandem duplicate 1 |
| chr19_-_3167729 | 0.79 |
ENSDART00000110763
ENSDART00000145710 ENSDART00000074620 ENSDART00000105174 |
stm
|
starmaker |
| chr3_-_61203203 | 0.78 |
ENSDART00000171787
|
pvalb1
|
parvalbumin 1 |
| chr8_-_23194408 | 0.75 |
ENSDART00000141090
|
si:ch211-196c10.15
|
si:ch211-196c10.15 |
| chr23_-_35694461 | 0.73 |
ENSDART00000185884
|
tuba1c
|
tubulin, alpha 1c |
| chr12_-_17479078 | 0.72 |
ENSDART00000079115
|
papss2b
|
3'-phosphoadenosine 5'-phosphosulfate synthase 2b |
| chr24_-_33291784 | 0.68 |
ENSDART00000124938
|
si:ch1073-406l10.2
|
si:ch1073-406l10.2 |
| chr4_-_11577253 | 0.66 |
ENSDART00000144452
|
net1
|
neuroepithelial cell transforming 1 |
| chr4_-_11737939 | 0.64 |
ENSDART00000150299
|
podxl
|
podocalyxin-like |
| chr11_+_30310170 | 0.64 |
ENSDART00000127797
|
ugt1b3
|
UDP glucuronosyltransferase 1 family, polypeptide B3 |
| chr17_-_27273296 | 0.61 |
ENSDART00000077087
|
id3
|
inhibitor of DNA binding 3 |
| chr3_-_58644920 | 0.61 |
ENSDART00000155953
|
dhrs7ca
|
dehydrogenase/reductase (SDR family) member 7Ca |
| chr21_+_26071874 | 0.61 |
ENSDART00000003001
ENSDART00000146573 |
rpl23a
|
ribosomal protein L23a |
| chr24_+_9744012 | 0.59 |
ENSDART00000129656
|
tmem108
|
transmembrane protein 108 |
| chr2_+_42236118 | 0.58 |
ENSDART00000114162
ENSDART00000141199 |
ftr02
|
finTRIM family, member 2 |
| chr21_-_26071773 | 0.58 |
ENSDART00000141382
|
rab34b
|
RAB34, member RAS oncogene family b |
| chr19_-_5332784 | 0.56 |
ENSDART00000010373
|
krt1-19d
|
keratin, type 1, gene 19d |
| chr11_+_30306606 | 0.54 |
ENSDART00000128276
ENSDART00000190222 |
ugt1b4
|
UDP glucuronosyltransferase 1 family, polypeptide B4 |
| chr21_+_18353703 | 0.54 |
ENSDART00000181396
ENSDART00000166359 |
si:ch73-287m6.1
|
si:ch73-287m6.1 |
| chr20_-_4883673 | 0.50 |
ENSDART00000145540
ENSDART00000053877 |
zdhhc14
|
zinc finger, DHHC-type containing 14 |
| chr23_+_42304602 | 0.48 |
ENSDART00000166004
|
cyp2aa11
|
cytochrome P450, family 2, subfamily AA, polypeptide 11 |
| chr6_+_6491013 | 0.46 |
ENSDART00000140827
|
bcl11ab
|
B cell CLL/lymphoma 11Ab |
| chr23_-_2099350 | 0.45 |
ENSDART00000091532
|
ndnf
|
neuron-derived neurotrophic factor |
| chr1_-_30366824 | 0.45 |
ENSDART00000125759
|
si:dkey-22o12.2
|
si:dkey-22o12.2 |
| chr20_+_46213553 | 0.44 |
ENSDART00000100532
|
stx7l
|
syntaxin 7-like |
| chr1_+_54043563 | 0.44 |
ENSDART00000149760
|
triobpa
|
TRIO and F-actin binding protein a |
| chr23_+_36052944 | 0.44 |
ENSDART00000103149
|
hoxc13a
|
homeobox C13a |
| chr22_+_661505 | 0.44 |
ENSDART00000149460
|
elf3
|
E74-like factor 3 (ets domain transcription factor, epithelial-specific ) |
| chr14_-_28052474 | 0.42 |
ENSDART00000172948
ENSDART00000135337 |
TSC22D3 (1 of many)
zgc:64189
|
si:ch211-220e11.3 zgc:64189 |
| chr20_-_46694573 | 0.42 |
ENSDART00000182557
|
AL929435.2
|
|
| chr8_-_37043900 | 0.42 |
ENSDART00000139567
|
renbp
|
renin binding protein |
| chr1_-_39943596 | 0.42 |
ENSDART00000149730
|
stox2a
|
storkhead box 2a |
| chr1_+_59073203 | 0.41 |
ENSDART00000149937
ENSDART00000162201 |
MFAP4 (1 of many)
zgc:173915
|
si:zfos-2330d3.3 zgc:173915 |
| chr1_+_21731382 | 0.41 |
ENSDART00000054395
|
pax5
|
paired box 5 |
| chr11_-_15001639 | 0.40 |
ENSDART00000170627
ENSDART00000168222 ENSDART00000171210 ENSDART00000163518 ENSDART00000168180 ENSDART00000157970 ENSDART00000169113 ENSDART00000180049 ENSDART00000188961 ENSDART00000183952 ENSDART00000193904 ENSDART00000193450 ENSDART00000186508 ENSDART00000169476 ENSDART00000162938 |
elavl1b
|
ELAV like RNA binding protein 1b |
| chr22_-_28373698 | 0.40 |
ENSDART00000157592
|
si:ch211-213c4.5
|
si:ch211-213c4.5 |
| chr15_+_47525073 | 0.39 |
ENSDART00000067583
|
sidt2
|
SID1 transmembrane family, member 2 |
| chr18_+_17611627 | 0.39 |
ENSDART00000046891
|
cetp
|
cholesteryl ester transfer protein, plasma |
| chr11_-_23332592 | 0.38 |
ENSDART00000125024
|
golt1a
|
golgi transport 1A |
| chr16_+_5774977 | 0.38 |
ENSDART00000134202
|
ccka
|
cholecystokinin a |
| chr15_+_46344655 | 0.37 |
ENSDART00000155893
|
si:ch1073-340i21.2
|
si:ch1073-340i21.2 |
| chr23_-_24682244 | 0.37 |
ENSDART00000104035
|
ctdsp2
|
CTD (carboxy-terminal domain, RNA polymerase II, polypeptide A) small phosphatase 2 |
| chr5_+_22459087 | 0.37 |
ENSDART00000134781
|
BX546499.1
|
|
| chr22_-_28871787 | 0.37 |
ENSDART00000104828
|
gtpbp2b
|
GTP binding protein 2b |
| chr10_+_17088261 | 0.36 |
ENSDART00000132103
|
si:dkey-106l3.7
|
si:dkey-106l3.7 |
| chr20_+_17581881 | 0.36 |
ENSDART00000182832
|
cdh2
|
cadherin 2, type 1, N-cadherin (neuronal) |
| chr8_+_39760258 | 0.36 |
ENSDART00000037914
|
cox6a1
|
cytochrome c oxidase subunit VIa polypeptide 1 |
| chr19_-_15281996 | 0.36 |
ENSDART00000103784
|
edn2
|
endothelin 2 |
| chr1_+_23784905 | 0.36 |
ENSDART00000171951
ENSDART00000188521 ENSDART00000183029 ENSDART00000187183 |
slit2
|
slit homolog 2 (Drosophila) |
| chr5_+_4806851 | 0.35 |
ENSDART00000067599
|
angptl2a
|
angiopoietin-like 2a |
| chr9_+_23900703 | 0.35 |
ENSDART00000127859
|
trim63b
|
tripartite motif containing 63b |
| chr2_-_59265521 | 0.35 |
ENSDART00000146341
ENSDART00000097799 |
ftr33
|
finTRIM family, member 33 |
| chr1_+_52563298 | 0.35 |
ENSDART00000142465
|
abca1a
|
ATP-binding cassette, sub-family A (ABC1), member 1A |
| chr3_-_28872756 | 0.34 |
ENSDART00000138840
|
eef2kmt
|
eukaryotic elongation factor 2 lysine methyltransferase |
| chr25_-_2081371 | 0.34 |
ENSDART00000104915
ENSDART00000156925 |
wnt7bb
|
wingless-type MMTV integration site family, member 7Bb |
| chr2_-_59157790 | 0.34 |
ENSDART00000192303
ENSDART00000159362 |
ftr32
|
finTRIM family, member 32 |
| chr25_-_29415369 | 0.34 |
ENSDART00000110774
ENSDART00000019183 |
ugt5a2
ugt5a1
|
UDP glucuronosyltransferase 5 family, polypeptide A2 UDP glucuronosyltransferase 5 family, polypeptide A1 |
| chr22_+_661711 | 0.34 |
ENSDART00000113795
|
elf3
|
E74-like factor 3 (ets domain transcription factor, epithelial-specific ) |
| chr4_-_18954001 | 0.33 |
ENSDART00000144814
|
si:dkey-31f5.8
|
si:dkey-31f5.8 |
| chr20_-_52199296 | 0.32 |
ENSDART00000131806
ENSDART00000143668 |
hlx1
|
H2.0-like homeo box 1 (Drosophila) |
| chr5_+_52067723 | 0.32 |
ENSDART00000166902
|
setbp1
|
SET binding protein 1 |
| chr19_+_627899 | 0.32 |
ENSDART00000148508
|
tert
|
telomerase reverse transcriptase |
| chr2_+_54641644 | 0.32 |
ENSDART00000027313
|
ndufv2
|
NADH dehydrogenase (ubiquinone) flavoprotein 2 |
| chr3_+_24190207 | 0.31 |
ENSDART00000034762
|
prr15la
|
proline rich 15-like a |
| chr19_-_46775562 | 0.29 |
ENSDART00000187144
|
abrab
|
actin binding Rho activating protein b |
| chr4_-_75616197 | 0.28 |
ENSDART00000157778
|
si:dkey-71l4.3
|
si:dkey-71l4.3 |
| chr25_-_173165 | 0.28 |
ENSDART00000193594
|
CABZ01114053.1
|
|
| chr14_+_6546516 | 0.28 |
ENSDART00000097214
|
adam19b
|
ADAM metallopeptidase domain 19b |
| chr5_+_7564644 | 0.28 |
ENSDART00000192173
|
CABZ01039096.1
|
|
| chr1_-_59104145 | 0.27 |
ENSDART00000132495
ENSDART00000152457 |
MFAP4 (1 of many)
si:zfos-2330d3.7
|
si:zfos-2330d3.1 si:zfos-2330d3.7 |
| chr1_-_22861348 | 0.27 |
ENSDART00000139412
|
SMIM18
|
si:dkey-92j12.6 |
| chr14_-_7128980 | 0.27 |
ENSDART00000171311
|
si:ch73-43g23.1
|
si:ch73-43g23.1 |
| chr18_-_8885792 | 0.27 |
ENSDART00000143619
|
si:dkey-95h12.1
|
si:dkey-95h12.1 |
| chr13_+_17694845 | 0.26 |
ENSDART00000079778
|
ifit8
|
interferon-induced protein with tetratricopeptide repeats 8 |
| chr19_-_8962884 | 0.26 |
ENSDART00000172582
ENSDART00000104657 |
mrps21
|
mitochondrial ribosomal protein S21 |
| chr1_-_59130383 | 0.26 |
ENSDART00000171552
|
FP015850.1
|
|
| chr17_+_14886828 | 0.25 |
ENSDART00000010507
ENSDART00000131052 |
ptger2a
|
prostaglandin E receptor 2a (subtype EP2) |
| chr24_-_6038025 | 0.25 |
ENSDART00000077819
ENSDART00000139216 |
ftr61
|
finTRIM family, member 61 |
| chr15_-_25556227 | 0.24 |
ENSDART00000156445
|
mmp20a
|
matrix metallopeptidase 20a (enamelysin) |
| chr24_+_42948 | 0.24 |
ENSDART00000122785
|
tmx3b
|
thioredoxin related transmembrane protein 3b |
| chr10_+_25982212 | 0.24 |
ENSDART00000128292
ENSDART00000108808 |
frem2a
|
Fras1 related extracellular matrix protein 2a |
| chr17_-_45386823 | 0.24 |
ENSDART00000156002
|
tmem206
|
transmembrane protein 206 |
| chr17_-_45386546 | 0.23 |
ENSDART00000182647
|
tmem206
|
transmembrane protein 206 |
| chr4_+_33654247 | 0.23 |
ENSDART00000192537
|
si:dkey-84h14.1
|
si:dkey-84h14.1 |
| chr2_-_59376399 | 0.22 |
ENSDART00000137134
|
ftr38
|
finTRIM family, member 38 |
| chr1_-_59139599 | 0.22 |
ENSDART00000152233
|
si:ch1073-110a20.3
|
si:ch1073-110a20.3 |
| chr4_-_64141714 | 0.22 |
ENSDART00000128628
|
BX914205.3
|
|
| chr12_-_44016898 | 0.22 |
ENSDART00000175304
|
si:dkey-201i2.4
|
si:dkey-201i2.4 |
| chr15_-_41734639 | 0.22 |
ENSDART00000154230
ENSDART00000167443 |
ftr90
|
finTRIM family, member 90 |
| chr18_-_20560007 | 0.22 |
ENSDART00000141367
ENSDART00000090186 |
si:ch211-238n5.4
|
si:ch211-238n5.4 |
| chr7_+_50448478 | 0.21 |
ENSDART00000098815
|
serinc4
|
serine incorporator 4 |
| chr18_+_13164325 | 0.21 |
ENSDART00000189057
|
tat
|
tyrosine aminotransferase |
| chr16_+_28932038 | 0.21 |
ENSDART00000149480
|
npr1b
|
natriuretic peptide receptor 1b |
| chr4_-_76488581 | 0.20 |
ENSDART00000174291
|
ftr51
|
finTRIM family, member 51 |
| chr5_+_71999996 | 0.20 |
ENSDART00000179933
ENSDART00000187070 |
PLPP7 (1 of many)
|
phospholipid phosphatase 7 (inactive) |
| chr3_-_58189429 | 0.19 |
ENSDART00000156092
|
si:ch211-256e16.11
|
si:ch211-256e16.11 |
| chr15_-_25556074 | 0.19 |
ENSDART00000124677
|
mmp20a
|
matrix metallopeptidase 20a (enamelysin) |
| chr4_+_50401133 | 0.19 |
ENSDART00000150644
|
si:dkey-156k2.7
|
si:dkey-156k2.7 |
| chr9_-_7378566 | 0.18 |
ENSDART00000144003
|
slc23a3
|
solute carrier family 23, member 3 |
| chr6_-_50404732 | 0.18 |
ENSDART00000055510
|
romo1
|
reactive oxygen species modulator 1 |
| chr16_-_32204470 | 0.17 |
ENSDART00000143928
|
rwdd1
|
RWD domain containing 1 |
| chr3_+_24189804 | 0.16 |
ENSDART00000134723
|
prr15la
|
proline rich 15-like a |
| chr4_-_71267461 | 0.16 |
ENSDART00000161219
ENSDART00000151838 |
si:dkeyp-51g9.4
|
si:dkeyp-51g9.4 |
| chr24_+_36373588 | 0.16 |
ENSDART00000079101
|
map3k14b
|
mitogen-activated protein kinase kinase kinase 14b |
| chr14_-_10762629 | 0.16 |
ENSDART00000061928
|
fgf16
|
fibroblast growth factor 16 |
| chr2_+_19578446 | 0.15 |
ENSDART00000164758
|
pimr50
|
Pim proto-oncogene, serine/threonine kinase, related 50 |
| chr5_-_24124118 | 0.15 |
ENSDART00000051550
|
capga
|
capping protein (actin filament), gelsolin-like a |
| chr16_-_53259409 | 0.15 |
ENSDART00000157080
|
si:ch211-269k10.4
|
si:ch211-269k10.4 |
| chr5_-_31716713 | 0.15 |
ENSDART00000131443
|
dpm2
|
dolichyl-phosphate mannosyltransferase polypeptide 2, regulatory subunit |
| chr21_+_25533908 | 0.15 |
ENSDART00000185225
|
nlrc3l1
|
NLR family, CARD domain containing 3-like 1 |
| chr22_+_19478140 | 0.14 |
ENSDART00000135291
ENSDART00000136576 |
si:dkey-78l4.8
|
si:dkey-78l4.8 |
| chr1_+_59073436 | 0.14 |
ENSDART00000161642
|
MFAP4 (1 of many)
|
si:zfos-2330d3.3 |
| chr8_+_23194212 | 0.14 |
ENSDART00000028946
ENSDART00000186842 ENSDART00000137815 ENSDART00000136914 ENSDART00000144855 ENSDART00000182689 |
tpd52l2a
|
tumor protein D52-like 2a |
| chr4_-_30440116 | 0.14 |
ENSDART00000159935
|
si:dkey-199m13.5
|
si:dkey-199m13.5 |
| chr3_-_48980319 | 0.14 |
ENSDART00000165319
|
ftr42
|
finTRIM family, member 42 |
| chr9_-_16853462 | 0.14 |
ENSDART00000160273
|
CT573248.2
|
|
| chr14_+_17003757 | 0.14 |
ENSDART00000170208
|
limch1b
|
LIM and calponin homology domains 1b |
| chr18_-_31227615 | 0.14 |
ENSDART00000003210
|
mc1r
|
melanocortin 1 receptor |
| chr21_-_17603182 | 0.13 |
ENSDART00000020048
ENSDART00000177270 |
gsna
|
gelsolin a |
| chr11_-_8208464 | 0.13 |
ENSDART00000161283
|
pimr203
|
Pim proto-oncogene, serine/threonine kinase, related 203 |
| chr7_-_1504382 | 0.13 |
ENSDART00000172770
|
si:zfos-405g10.4
|
si:zfos-405g10.4 |
| chr22_+_1526040 | 0.13 |
ENSDART00000164089
ENSDART00000159126 |
si:ch211-255f4.6
|
si:ch211-255f4.6 |
| chr20_+_46897504 | 0.13 |
ENSDART00000158124
|
si:ch73-21k16.1
|
si:ch73-21k16.1 |
| chr25_-_20268027 | 0.13 |
ENSDART00000138763
|
dnajb9a
|
DnaJ (Hsp40) homolog, subfamily B, member 9a |
| chr7_-_24390879 | 0.12 |
ENSDART00000036680
|
ptgr1
|
prostaglandin reductase 1 |
| chr3_-_27784457 | 0.12 |
ENSDART00000019098
|
dnase1
|
deoxyribonuclease I |
| chr2_-_19520324 | 0.12 |
ENSDART00000079877
|
pimr52
|
Pim proto-oncogene, serine/threonine kinase, related 52 |
| chr6_-_39051319 | 0.12 |
ENSDART00000155093
|
tns2b
|
tensin 2b |
| chr2_-_59285407 | 0.12 |
ENSDART00000181616
|
ftr34
|
finTRIM family, member 34 |
| chr1_-_57172294 | 0.12 |
ENSDART00000063774
|
rac1l
|
Rac family small GTPase 1, like |
| chr10_+_21718468 | 0.12 |
ENSDART00000126622
|
pcdh1gb9
|
protocadherin 1 gamma b 9 |
| chr2_+_47927026 | 0.11 |
ENSDART00000143023
|
ftr25
|
finTRIM family, member 25 |
| chr2_-_47904043 | 0.11 |
ENSDART00000185328
ENSDART00000126740 |
ftr22
|
finTRIM family, member 22 |
| chr6_+_49926115 | 0.11 |
ENSDART00000018523
|
ahcy
|
adenosylhomocysteinase |
| chr20_-_54377933 | 0.11 |
ENSDART00000182664
|
entpd5b
|
ectonucleoside triphosphate diphosphohydrolase 5b |
| chr4_-_46915024 | 0.11 |
ENSDART00000169913
|
si:ch211-134c10.1
|
si:ch211-134c10.1 |
| chr4_-_75869738 | 0.11 |
ENSDART00000170963
|
si:dkey-261j11.2
|
si:dkey-261j11.2 |
| chr13_-_21650404 | 0.11 |
ENSDART00000078352
|
tspan14
|
tetraspanin 14 |
| chr5_-_63218919 | 0.11 |
ENSDART00000149979
|
tecta
|
tectorin alpha |
| chr17_-_23609210 | 0.11 |
ENSDART00000064003
|
ifit15
|
interferon-induced protein with tetratricopeptide repeats 15 |
| chr15_-_37774544 | 0.11 |
ENSDART00000156119
|
si:dkey-42l23.9
|
si:dkey-42l23.9 |
| chr4_-_77636185 | 0.10 |
ENSDART00000152915
ENSDART00000131964 |
si:dkey-61p9.9
|
si:dkey-61p9.9 |
| chr4_+_47736069 | 0.10 |
ENSDART00000193329
|
si:ch211-196f19.1
|
si:ch211-196f19.1 |
| chr19_+_25478203 | 0.10 |
ENSDART00000132934
|
col28a1a
|
collagen, type XXVIII, alpha 1a |
| chr11_+_12175162 | 0.10 |
ENSDART00000125446
|
si:ch211-156l18.7
|
si:ch211-156l18.7 |
| chr19_+_1831911 | 0.10 |
ENSDART00000166653
|
ptk2aa
|
protein tyrosine kinase 2aa |
| chr13_-_3370638 | 0.10 |
ENSDART00000029649
|
prkn
|
parkin RBR E3 ubiquitin protein ligase |
| chr11_+_8170948 | 0.10 |
ENSDART00000112127
ENSDART00000186267 |
dnase2b
|
deoxyribonuclease II beta |
| chr8_-_18203274 | 0.10 |
ENSDART00000134078
ENSDART00000180235 ENSDART00000080006 ENSDART00000125418 ENSDART00000142114 |
elovl8b
|
ELOVL fatty acid elongase 8b |
| chr1_+_47178529 | 0.10 |
ENSDART00000158432
ENSDART00000074450 ENSDART00000137448 |
morc3b
|
MORC family CW-type zinc finger 3b |
| chr4_+_71014655 | 0.10 |
ENSDART00000170837
|
si:dkeyp-80d11.14
|
si:dkeyp-80d11.14 |
| chr5_+_38673168 | 0.10 |
ENSDART00000135267
|
si:dkey-58f10.12
|
si:dkey-58f10.12 |
| chr17_+_12285285 | 0.09 |
ENSDART00000154336
|
pimr174
|
Pim proto-oncogene, serine/threonine kinase, related 174 |
| chr4_+_35576329 | 0.09 |
ENSDART00000171191
ENSDART00000161611 |
si:dkeyp-4c4.2
|
si:dkeyp-4c4.2 |
| chr5_+_38752287 | 0.09 |
ENSDART00000133571
|
cxcl11.8
|
chemokine (C-X-C motif) ligand 11, duplicate 8 |
| chr21_+_25533531 | 0.09 |
ENSDART00000134052
|
nlrc3l1
|
NLR family, CARD domain containing 3-like 1 |
| chr6_-_8498676 | 0.09 |
ENSDART00000148627
|
pglyrp2
|
peptidoglycan recognition protein 2 |
| chr4_+_28997595 | 0.09 |
ENSDART00000133357
|
si:dkey-13e3.1
|
si:dkey-13e3.1 |
| chr4_-_75615597 | 0.09 |
ENSDART00000187794
|
si:dkey-71l4.3
|
si:dkey-71l4.3 |
| chr12_-_44122412 | 0.09 |
ENSDART00000169094
|
si:ch73-329n5.3
|
si:ch73-329n5.3 |
| chr16_-_22747125 | 0.09 |
ENSDART00000179865
|
tlr19
|
toll-like receptor 19 |
| chr19_-_24218942 | 0.09 |
ENSDART00000189198
|
BX547993.2
|
|
| chr11_+_28107632 | 0.08 |
ENSDART00000177721
|
nmur3
|
neuromedin U receptor 3 |
| chr23_-_10914275 | 0.08 |
ENSDART00000112965
|
pdzrn3a
|
PDZ domain containing RING finger 3a |
| chr3_-_33029426 | 0.08 |
ENSDART00000181287
|
CT583642.1
|
|
| chr20_-_43495506 | 0.08 |
ENSDART00000170679
|
pimr132
|
Pim proto-oncogene, serine/threonine kinase, related 132 |
| chr9_-_41818760 | 0.08 |
ENSDART00000140601
|
si:dkeyp-30e7.2
|
si:dkeyp-30e7.2 |
| chr10_+_23441488 | 0.08 |
ENSDART00000100783
|
si:ch73-122g19.1
|
si:ch73-122g19.1 |
| chr14_-_2270973 | 0.07 |
ENSDART00000180729
|
pcdh2ab9
|
protocadherin 2 alpha b 9 |
| chr4_+_16787488 | 0.07 |
ENSDART00000143006
|
golt1ba
|
golgi transport 1Ba |
| chr8_-_3346692 | 0.07 |
ENSDART00000057874
|
FUT9 (1 of many)
|
zgc:103510 |
| chr23_+_13928346 | 0.07 |
ENSDART00000155326
|
si:dkey-90a13.10
|
si:dkey-90a13.10 |
| chr16_-_22729119 | 0.07 |
ENSDART00000132944
|
tlr19
|
toll-like receptor 19 |
| chr15_-_37834433 | 0.07 |
ENSDART00000189748
|
si:dkey-238d18.3
|
si:dkey-238d18.3 |
| chr14_-_48348973 | 0.06 |
ENSDART00000185822
|
CABZ01080056.1
|
|
| chr13_+_2830574 | 0.06 |
ENSDART00000138143
|
si:ch211-233m11.1
|
si:ch211-233m11.1 |
| chr9_-_43375205 | 0.06 |
ENSDART00000138436
|
znf385b
|
zinc finger protein 385B |
| chr23_-_19230627 | 0.06 |
ENSDART00000007122
|
guca1b
|
guanylate cyclase activator 1B |
| chr4_-_62503772 | 0.05 |
ENSDART00000108891
|
si:dkey-165b20.1
|
si:dkey-165b20.1 |
| chr7_+_66822229 | 0.05 |
ENSDART00000112109
|
lyve1a
|
lymphatic vessel endothelial hyaluronic receptor 1a |
| chr10_+_21726853 | 0.05 |
ENSDART00000162869
|
pcdh1g14
|
protocadherin 1 gamma 14 |
| chr16_+_30961822 | 0.05 |
ENSDART00000059187
|
gstk2
|
glutathione S-transferase kappa 2 |
| chr1_+_24366110 | 0.05 |
ENSDART00000139913
|
pimr178
|
Pim proto-oncogene, serine/threonine kinase, related 178 |
| chr6_-_39649504 | 0.05 |
ENSDART00000179960
ENSDART00000190951 |
larp4ab
|
La ribonucleoprotein domain family, member 4Ab |
| chr3_+_13603272 | 0.04 |
ENSDART00000185084
|
hspbp1
|
HSPA (heat shock 70kDa) binding protein, cytoplasmic cochaperone 1 |
| chr15_-_41486793 | 0.04 |
ENSDART00000138525
|
si:ch211-187g4.1
|
si:ch211-187g4.1 |
| chr8_-_36287046 | 0.04 |
ENSDART00000162877
|
si:busm1-194e12.11
|
si:busm1-194e12.11 |
| chr17_+_12265095 | 0.04 |
ENSDART00000153812
|
pimr168
|
Pim proto-oncogene, serine/threonine kinase, related 168 |
| chr12_-_44010532 | 0.04 |
ENSDART00000183875
|
si:ch211-182p11.1
|
si:ch211-182p11.1 |
| chr4_-_25812329 | 0.04 |
ENSDART00000146658
|
tmcc3
|
transmembrane and coiled-coil domain family 3 |
| chr17_-_24564674 | 0.04 |
ENSDART00000105435
ENSDART00000135086 |
abch1
|
ATP-binding cassette, sub-family H, member 1 |
Network of associatons between targets according to the STRING database.
First level regulatory network of smad3a+smad3b
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.7 | GO:0050427 | sulfate assimilation(GO:0000103) purine ribonucleoside bisphosphate metabolic process(GO:0034035) purine ribonucleoside bisphosphate biosynthetic process(GO:0034036) 3'-phosphoadenosine 5'-phosphosulfate metabolic process(GO:0050427) 3'-phosphoadenosine 5'-phosphosulfate biosynthetic process(GO:0050428) |
| 0.2 | 0.6 | GO:0033632 | cell-cell adhesion mediated by integrin(GO:0033631) regulation of cell-cell adhesion mediated by integrin(GO:0033632) |
| 0.1 | 0.4 | GO:0034375 | high-density lipoprotein particle remodeling(GO:0034375) |
| 0.1 | 2.8 | GO:0048936 | peripheral nervous system neuron axonogenesis(GO:0048936) |
| 0.1 | 0.4 | GO:0090247 | hindbrain structural organization(GO:0021577) dorsal fin morphogenesis(GO:0035142) cell motility involved in somitogenic axis elongation(GO:0090247) cell migration involved in somitogenic axis elongation(GO:0090248) |
| 0.1 | 0.4 | GO:0007414 | axonal defasciculation(GO:0007414) |
| 0.1 | 0.5 | GO:2000171 | negative regulation of dendrite development(GO:2000171) |
| 0.1 | 0.8 | GO:0045299 | otolith mineralization(GO:0045299) |
| 0.1 | 0.4 | GO:0021828 | pallium development(GO:0021543) cerebral cortex cell migration(GO:0021795) cerebral cortex tangential migration(GO:0021800) cerebral cortex tangential migration using cell-axon interactions(GO:0021824) substrate-dependent cerebral cortex tangential migration(GO:0021825) gonadotrophin-releasing hormone neuronal migration to the hypothalamus(GO:0021828) hypothalamic tangential migration using cell-axon interactions(GO:0021856) cerebral cortex development(GO:0021987) |
| 0.1 | 0.3 | GO:0007571 | age-dependent general metabolic decline(GO:0007571) |
| 0.1 | 0.3 | GO:0010874 | regulation of cholesterol efflux(GO:0010874) |
| 0.1 | 0.6 | GO:0097106 | postsynaptic density organization(GO:0097106) |
| 0.1 | 0.6 | GO:0090385 | phagosome-lysosome fusion(GO:0090385) |
| 0.1 | 0.4 | GO:0060585 | regulation of systemic arterial blood pressure by endothelin(GO:0003100) vein smooth muscle contraction(GO:0014826) regulation of prostaglandin-endoperoxide synthase activity(GO:0060584) positive regulation of prostaglandin-endoperoxide synthase activity(GO:0060585) |
| 0.1 | 0.4 | GO:0019262 | N-acetylneuraminate catabolic process(GO:0019262) |
| 0.1 | 0.2 | GO:0018171 | peptidyl-cysteine oxidation(GO:0018171) |
| 0.1 | 0.9 | GO:0001867 | complement activation, lectin pathway(GO:0001867) |
| 0.1 | 0.4 | GO:1900026 | positive regulation of substrate adhesion-dependent cell spreading(GO:1900026) |
| 0.1 | 0.3 | GO:0071380 | response to prostaglandin E(GO:0034695) cellular response to prostaglandin E stimulus(GO:0071380) |
| 0.0 | 0.4 | GO:0060005 | reflex(GO:0060004) vestibular reflex(GO:0060005) vestibular receptor cell differentiation(GO:0060114) vestibular receptor cell development(GO:0060118) |
| 0.0 | 0.7 | GO:0050994 | regulation of lipid catabolic process(GO:0050994) |
| 0.0 | 0.7 | GO:1900107 | regulation of nodal signaling pathway(GO:1900107) |
| 0.0 | 0.2 | GO:2000379 | protein import into mitochondrial inner membrane(GO:0045039) positive regulation of reactive oxygen species metabolic process(GO:2000379) |
| 0.0 | 0.2 | GO:0006572 | tyrosine catabolic process(GO:0006572) |
| 0.0 | 0.2 | GO:0019348 | dolichol metabolic process(GO:0019348) |
| 0.0 | 0.4 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
| 0.0 | 0.2 | GO:0021846 | cell proliferation in forebrain(GO:0021846) |
| 0.0 | 0.6 | GO:0000027 | ribosomal large subunit assembly(GO:0000027) |
| 0.0 | 0.5 | GO:0007274 | neuromuscular synaptic transmission(GO:0007274) |
| 0.0 | 0.6 | GO:0071542 | dopaminergic neuron differentiation(GO:0071542) |
| 0.0 | 0.5 | GO:0048512 | rhythmic behavior(GO:0007622) circadian behavior(GO:0048512) |
| 0.0 | 0.4 | GO:0006120 | mitochondrial electron transport, NADH to ubiquinone(GO:0006120) |
| 0.0 | 0.2 | GO:0071678 | olfactory bulb axon guidance(GO:0071678) |
| 0.0 | 0.1 | GO:0033353 | S-adenosylmethionine cycle(GO:0033353) |
| 0.0 | 0.1 | GO:0098581 | detection of biotic stimulus(GO:0009595) detection of bacterium(GO:0016045) detection of other organism(GO:0098543) detection of external biotic stimulus(GO:0098581) |
| 0.0 | 0.1 | GO:0035889 | otolith tethering(GO:0035889) |
| 0.0 | 0.1 | GO:0006309 | apoptotic DNA fragmentation(GO:0006309) cellular component disassembly involved in execution phase of apoptosis(GO:0006921) apoptotic nuclear changes(GO:0030262) |
| 0.0 | 0.5 | GO:0018230 | peptidyl-L-cysteine S-palmitoylation(GO:0018230) peptidyl-S-diacylglycerol-L-cysteine biosynthetic process from peptidyl-cysteine(GO:0018231) |
| 0.0 | 0.4 | GO:0044243 | collagen catabolic process(GO:0030574) multicellular organism catabolic process(GO:0044243) |
| 0.0 | 0.3 | GO:0035775 | pronephric glomerulus morphogenesis(GO:0035775) |
| 0.0 | 0.1 | GO:0000737 | DNA catabolic process, endonucleolytic(GO:0000737) |
| 0.0 | 0.1 | GO:0007045 | cell-substrate adherens junction assembly(GO:0007045) focal adhesion assembly(GO:0048041) regulation of focal adhesion assembly(GO:0051893) regulation of cell-substrate junction assembly(GO:0090109) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0000333 | telomerase catalytic core complex(GO:0000333) |
| 0.1 | 0.6 | GO:0031528 | microvillus membrane(GO:0031528) |
| 0.1 | 0.2 | GO:0033185 | dolichol-phosphate-mannose synthase complex(GO:0033185) |
| 0.0 | 0.1 | GO:0016460 | myosin II complex(GO:0016460) |
| 0.0 | 0.4 | GO:0014704 | intercalated disc(GO:0014704) |
| 0.0 | 0.4 | GO:0034364 | high-density lipoprotein particle(GO:0034364) |
| 0.0 | 0.4 | GO:0005751 | mitochondrial respiratory chain complex IV(GO:0005751) |
| 0.0 | 0.2 | GO:0005744 | mitochondrial inner membrane presequence translocase complex(GO:0005744) |
| 0.0 | 0.0 | GO:0043190 | ATP-binding cassette (ABC) transporter complex(GO:0043190) |
| 0.0 | 3.5 | GO:0005764 | lysosome(GO:0005764) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.7 | GO:0004020 | adenylylsulfate kinase activity(GO:0004020) sulfate adenylyltransferase activity(GO:0004779) sulfate adenylyltransferase (ATP) activity(GO:0004781) |
| 0.1 | 0.3 | GO:0003721 | telomerase RNA reverse transcriptase activity(GO:0003721) |
| 0.1 | 0.9 | GO:0004046 | aminoacylase activity(GO:0004046) |
| 0.1 | 0.4 | GO:0031708 | endothelin B receptor binding(GO:0031708) |
| 0.1 | 0.3 | GO:0090554 | phospholipid-translocating ATPase activity(GO:0004012) phosphatidylcholine-translocating ATPase activity(GO:0090554) phosphatidylserine-translocating ATPase activity(GO:0090556) |
| 0.0 | 0.1 | GO:0004980 | melanocyte-stimulating hormone receptor activity(GO:0004980) |
| 0.0 | 0.4 | GO:0008420 | CTD phosphatase activity(GO:0008420) |
| 0.0 | 0.1 | GO:0004013 | adenosylhomocysteinase activity(GO:0004013) trialkylsulfonium hydrolase activity(GO:0016802) |
| 0.0 | 0.2 | GO:0016972 | thiol oxidase activity(GO:0016972) |
| 0.0 | 0.1 | GO:0032034 | myosin head/neck binding(GO:0032028) myosin II head/neck binding(GO:0032034) myosin II heavy chain binding(GO:0032038) |
| 0.0 | 0.1 | GO:0001607 | neuromedin U receptor activity(GO:0001607) |
| 0.0 | 1.5 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.0 | 0.2 | GO:0016941 | natriuretic peptide receptor activity(GO:0016941) |
| 0.0 | 0.3 | GO:0004957 | prostaglandin E receptor activity(GO:0004957) |
| 0.0 | 0.4 | GO:0022884 | macromolecule transmembrane transporter activity(GO:0022884) |
| 0.0 | 0.9 | GO:0003823 | antigen binding(GO:0003823) |
| 0.0 | 0.4 | GO:0048495 | Roundabout binding(GO:0048495) |
| 0.0 | 0.4 | GO:0016857 | racemase and epimerase activity, acting on carbohydrates and derivatives(GO:0016857) |
| 0.0 | 0.1 | GO:0016019 | peptidoglycan receptor activity(GO:0016019) |
| 0.0 | 2.5 | GO:0004197 | cysteine-type endopeptidase activity(GO:0004197) |
| 0.0 | 0.3 | GO:0031210 | phosphatidylcholine binding(GO:0031210) |
| 0.0 | 0.4 | GO:0016675 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors(GO:0016675) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.0 | 2.0 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.0 | 0.1 | GO:0004530 | deoxyribonuclease I activity(GO:0004530) |
| 0.0 | 0.6 | GO:0019843 | rRNA binding(GO:0019843) |
| 0.0 | 0.7 | GO:0030295 | kinase activator activity(GO:0019209) protein kinase activator activity(GO:0030295) |
| 0.0 | 0.5 | GO:0019707 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.0 | 0.7 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
| 0.0 | 0.4 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.7 | REACTOME POST CHAPERONIN TUBULIN FOLDING PATHWAY | Genes involved in Post-chaperonin tubulin folding pathway |
| 0.0 | 0.4 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.0 | 0.4 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.0 | 0.7 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |