Project
PRJNA195909:zebrafish embryo and larva development
Navigation
Downloads
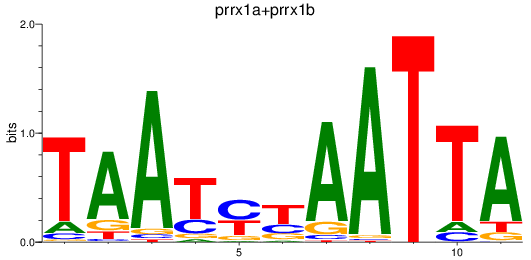
Results for prrx1a+prrx1b
Z-value: 2.05
Transcription factors associated with prrx1a+prrx1b
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
prrx1a
|
ENSDARG00000033971 | paired related homeobox 1a |
|
prrx1b
|
ENSDARG00000042027 | paired related homeobox 1b |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| prrx1a | dr11_v1_chr2_-_23172708_23172708 | 0.88 | 1.5e-03 | Click! |
| prrx1b | dr11_v1_chr20_+_34512130_34512130 | 0.86 | 2.6e-03 | Click! |
Activity profile of prrx1a+prrx1b motif
Sorted Z-values of prrx1a+prrx1b motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr25_+_31276842 | 21.89 |
ENSDART00000187238
|
tnni2a.4
|
troponin I type 2a (skeletal, fast), tandem duplicate 4 |
| chr4_+_9669717 | 6.93 |
ENSDART00000004604
|
si:dkey-153k10.9
|
si:dkey-153k10.9 |
| chr21_+_6556635 | 5.46 |
ENSDART00000139598
|
col5a1
|
procollagen, type V, alpha 1 |
| chr3_+_28939759 | 4.70 |
ENSDART00000141904
|
lgals1l1
|
lectin, galactoside-binding, soluble, 1 (galectin 1)-like 1 |
| chr16_+_23984755 | 4.25 |
ENSDART00000145328
|
apoc2
|
apolipoprotein C-II |
| chr19_-_3167729 | 3.90 |
ENSDART00000110763
ENSDART00000145710 ENSDART00000074620 ENSDART00000105174 |
stm
|
starmaker |
| chr25_+_31277415 | 3.79 |
ENSDART00000036275
|
tnni2a.4
|
troponin I type 2a (skeletal, fast), tandem duplicate 4 |
| chr16_-_21140097 | 3.69 |
ENSDART00000145837
ENSDART00000146500 |
si:dkey-271j15.3
|
si:dkey-271j15.3 |
| chr1_+_17676745 | 3.68 |
ENSDART00000030665
|
slc25a4
|
solute carrier family 25 (mitochondrial carrier; adenine nucleotide translocator), member 4 |
| chr9_-_22339582 | 3.58 |
ENSDART00000134805
|
crygm2d1
|
crystallin, gamma M2d1 |
| chr20_+_31287356 | 3.46 |
ENSDART00000007688
|
slc22a16
|
solute carrier family 22 (organic cation/carnitine transporter), member 16 |
| chr16_-_31717851 | 3.33 |
ENSDART00000169109
|
rbp5
|
retinol binding protein 1a, cellular |
| chr25_+_18964782 | 3.31 |
ENSDART00000017299
|
tdg.1
|
thymine DNA glycosylase, tandem duplicate 1 |
| chr1_-_42289704 | 3.26 |
ENSDART00000150124
|
si:ch211-71k14.1
|
si:ch211-71k14.1 |
| chr21_+_26390549 | 3.02 |
ENSDART00000185643
|
tmsb
|
thymosin, beta |
| chr7_-_48667056 | 2.89 |
ENSDART00000006378
|
cdkn1ca
|
cyclin-dependent kinase inhibitor 1Ca |
| chr2_+_16781015 | 2.84 |
ENSDART00000155147
ENSDART00000003845 |
tfa
|
transferrin-a |
| chr10_-_28835771 | 2.76 |
ENSDART00000192220
ENSDART00000188436 |
alcama
|
activated leukocyte cell adhesion molecule a |
| chr10_+_32104305 | 2.73 |
ENSDART00000099880
|
wnt11r
|
wingless-type MMTV integration site family, member 11, related |
| chr1_-_30979707 | 2.72 |
ENSDART00000008469
|
dlx2b
|
distal-less homeobox 2b |
| chr21_+_27382893 | 2.71 |
ENSDART00000005682
|
actn3a
|
actinin alpha 3a |
| chr16_-_31718013 | 2.70 |
ENSDART00000190716
|
rbp5
|
retinol binding protein 1a, cellular |
| chr15_-_14884332 | 2.66 |
ENSDART00000165237
|
si:ch211-24o8.4
|
si:ch211-24o8.4 |
| chr20_-_43775495 | 2.66 |
ENSDART00000100610
ENSDART00000149001 ENSDART00000148809 ENSDART00000100608 |
matn3a
|
matrilin 3a |
| chr13_+_24279021 | 2.63 |
ENSDART00000058629
|
acta1b
|
actin, alpha 1b, skeletal muscle |
| chr19_-_3240605 | 2.61 |
ENSDART00000105168
|
si:ch211-133n4.4
|
si:ch211-133n4.4 |
| chr12_-_5728755 | 2.61 |
ENSDART00000105887
|
dlx4b
|
distal-less homeobox 4b |
| chr6_+_9130989 | 2.58 |
ENSDART00000162588
|
rgn
|
regucalcin |
| chr20_-_44496245 | 2.56 |
ENSDART00000012229
|
fkbp1b
|
FK506 binding protein 1b |
| chr7_-_28148310 | 2.55 |
ENSDART00000044208
|
lmo1
|
LIM domain only 1 |
| chr16_-_45058919 | 2.53 |
ENSDART00000177134
|
gapdhs
|
glyceraldehyde-3-phosphate dehydrogenase, spermatogenic |
| chr1_-_40911332 | 2.50 |
ENSDART00000027463
|
hmx4
|
H6 family homeobox 4 |
| chr23_-_18913032 | 2.49 |
ENSDART00000136678
|
si:ch211-209j10.6
|
si:ch211-209j10.6 |
| chr13_+_25449681 | 2.44 |
ENSDART00000101328
|
atoh7
|
atonal bHLH transcription factor 7 |
| chr14_-_17068511 | 2.39 |
ENSDART00000163766
|
phox2bb
|
paired-like homeobox 2bb |
| chr21_+_7582036 | 2.31 |
ENSDART00000135485
ENSDART00000027268 |
otpa
|
orthopedia homeobox a |
| chr12_-_990149 | 2.29 |
ENSDART00000054367
|
kdelr2b
|
KDEL (Lys-Asp-Glu-Leu) endoplasmic reticulum protein retention receptor 2b |
| chr5_-_65037525 | 2.28 |
ENSDART00000158856
|
anxa1b
|
annexin A1b |
| chr7_+_26629084 | 2.28 |
ENSDART00000101044
ENSDART00000173765 |
hsbp1a
|
heat shock factor binding protein 1a |
| chr18_-_47662696 | 2.28 |
ENSDART00000184260
|
CABZ01073963.1
|
|
| chr23_+_45282858 | 2.26 |
ENSDART00000162353
|
CABZ01073265.1
|
|
| chr18_+_5490668 | 2.23 |
ENSDART00000167035
|
mibp2
|
muscle-specific beta 1 integrin binding protein 2 |
| chr11_-_45138857 | 2.22 |
ENSDART00000166501
|
cant1b
|
calcium activated nucleotidase 1b |
| chr25_+_29160102 | 2.18 |
ENSDART00000162854
|
pkmb
|
pyruvate kinase M1/2b |
| chr17_+_30704068 | 2.15 |
ENSDART00000062793
|
apoba
|
apolipoprotein Ba |
| chr25_+_20089986 | 2.14 |
ENSDART00000143441
ENSDART00000184073 |
tnni4b.2
|
troponin I4b, tandem duplicate 2 |
| chr6_+_23887314 | 2.10 |
ENSDART00000163188
|
znf648
|
zinc finger protein 648 |
| chr20_-_27225876 | 2.08 |
ENSDART00000149204
ENSDART00000149732 |
si:dkey-85n7.7
|
si:dkey-85n7.7 |
| chr3_+_28953274 | 2.07 |
ENSDART00000133528
ENSDART00000103602 |
lgals2a
|
lectin, galactoside-binding, soluble, 2a |
| chr21_+_13861589 | 2.06 |
ENSDART00000015629
ENSDART00000171306 |
stxbp1a
|
syntaxin binding protein 1a |
| chr15_-_2640966 | 2.06 |
ENSDART00000063320
|
cldne
|
claudin e |
| chr10_-_26744131 | 2.01 |
ENSDART00000020096
ENSDART00000162710 ENSDART00000179853 |
fgf13b
|
fibroblast growth factor 13b |
| chr2_+_56463167 | 2.00 |
ENSDART00000123392
|
rab11bb
|
RAB11B, member RAS oncogene family, b |
| chr14_-_24410673 | 1.99 |
ENSDART00000125923
|
cxcl14
|
chemokine (C-X-C motif) ligand 14 |
| chr12_-_558201 | 1.98 |
ENSDART00000168586
ENSDART00000158355 |
bsk146
|
brain specific kinase 146 |
| chr2_+_38039857 | 1.97 |
ENSDART00000159951
|
casq1a
|
calsequestrin 1a |
| chr22_-_23668356 | 1.96 |
ENSDART00000167106
ENSDART00000159622 ENSDART00000163228 |
cfh
|
complement factor H |
| chr9_-_1959917 | 1.95 |
ENSDART00000082359
|
hoxd3a
|
homeobox D3a |
| chr4_-_9909371 | 1.95 |
ENSDART00000102656
|
si:dkey-22l11.6
|
si:dkey-22l11.6 |
| chr24_-_21923930 | 1.91 |
ENSDART00000131944
|
tagln3b
|
transgelin 3b |
| chr8_-_14091886 | 1.90 |
ENSDART00000137857
|
si:ch211-229n2.7
|
si:ch211-229n2.7 |
| chr14_-_34044369 | 1.86 |
ENSDART00000149396
ENSDART00000123607 ENSDART00000190746 |
cyfip2
|
cytoplasmic FMR1 interacting protein 2 |
| chr12_+_15666949 | 1.85 |
ENSDART00000079803
|
nmt1b
|
N-myristoyltransferase 1b |
| chr12_-_30338779 | 1.84 |
ENSDART00000192511
|
vwa2
|
von Willebrand factor A domain containing 2 |
| chr7_-_30367650 | 1.84 |
ENSDART00000075519
|
aldh1a2
|
aldehyde dehydrogenase 1 family, member A2 |
| chr12_+_42436920 | 1.82 |
ENSDART00000177303
|
ebf3a
|
early B cell factor 3a |
| chr20_+_34320635 | 1.82 |
ENSDART00000153207
|
ivns1abpa
|
influenza virus NS1A binding protein a |
| chr9_+_34641237 | 1.80 |
ENSDART00000133996
|
shox
|
short stature homeobox |
| chr24_+_22485710 | 1.77 |
ENSDART00000146058
|
si:dkey-40h20.1
|
si:dkey-40h20.1 |
| chr5_-_31901468 | 1.76 |
ENSDART00000147814
ENSDART00000141446 |
coro1cb
|
coronin, actin binding protein, 1Cb |
| chr24_-_38657683 | 1.74 |
ENSDART00000154843
|
si:ch1073-164k15.3
|
si:ch1073-164k15.3 |
| chr23_-_12345764 | 1.74 |
ENSDART00000133956
|
phactr3a
|
phosphatase and actin regulator 3a |
| chr8_+_41647539 | 1.73 |
ENSDART00000136492
ENSDART00000138799 ENSDART00000134404 |
si:ch211-158d24.4
|
si:ch211-158d24.4 |
| chr4_-_16345227 | 1.73 |
ENSDART00000079521
|
kera
|
keratocan |
| chr14_-_21123551 | 1.73 |
ENSDART00000171679
ENSDART00000165882 |
si:dkey-74k8.4
|
si:dkey-74k8.4 |
| chr23_+_11285662 | 1.72 |
ENSDART00000111028
|
chl1a
|
cell adhesion molecule L1-like a |
| chr8_+_15254564 | 1.72 |
ENSDART00000024433
|
slc5a9
|
solute carrier family 5 (sodium/sugar cotransporter), member 9 |
| chr12_+_18681477 | 1.68 |
ENSDART00000127981
ENSDART00000143979 |
rgs9b
|
regulator of G protein signaling 9b |
| chr12_+_42436328 | 1.65 |
ENSDART00000167324
|
ebf3a
|
early B cell factor 3a |
| chr9_-_34300707 | 1.64 |
ENSDART00000049805
|
ildr2
|
immunoglobulin-like domain containing receptor 2 |
| chr3_-_24980067 | 1.64 |
ENSDART00000048871
|
desi1a
|
desumoylating isopeptidase 1a |
| chr13_-_31622195 | 1.63 |
ENSDART00000057432
|
six1a
|
SIX homeobox 1a |
| chr25_-_13381854 | 1.62 |
ENSDART00000164621
ENSDART00000169129 |
ndrg4
|
NDRG family member 4 |
| chr1_-_45049603 | 1.62 |
ENSDART00000023336
|
rps6
|
ribosomal protein S6 |
| chr21_-_44104600 | 1.58 |
ENSDART00000044599
|
oatx
|
organic anion transporter X |
| chr6_+_52350443 | 1.55 |
ENSDART00000151612
ENSDART00000151349 |
si:ch211-239j9.1
|
si:ch211-239j9.1 |
| chr22_-_10121880 | 1.54 |
ENSDART00000002348
|
rdh5
|
retinol dehydrogenase 5 (11-cis/9-cis) |
| chr5_+_34407763 | 1.53 |
ENSDART00000188849
ENSDART00000145127 |
lamc3
|
laminin, gamma 3 |
| chr2_+_30916188 | 1.52 |
ENSDART00000137012
|
myom1a
|
myomesin 1a (skelemin) |
| chr1_+_25801648 | 1.52 |
ENSDART00000129471
|
gucy1b1
|
guanylate cyclase 1 soluble subunit beta 1 |
| chr15_+_35043007 | 1.52 |
ENSDART00000086954
|
sesn3
|
sestrin 3 |
| chr22_-_32507966 | 1.51 |
ENSDART00000104693
|
pcbp4
|
poly(rC) binding protein 4 |
| chr3_-_37759147 | 1.51 |
ENSDART00000151067
|
si:dkey-260c8.6
|
si:dkey-260c8.6 |
| chr15_-_21165237 | 1.50 |
ENSDART00000157069
|
A2ML1 (1 of many)
|
si:ch211-212c13.8 |
| chr9_-_21918963 | 1.50 |
ENSDART00000090782
|
lmo7a
|
LIM domain 7a |
| chr7_-_17028015 | 1.49 |
ENSDART00000022441
|
dbx1a
|
developing brain homeobox 1a |
| chr14_-_17068712 | 1.47 |
ENSDART00000170277
|
phox2bb
|
paired-like homeobox 2bb |
| chr19_+_31771270 | 1.46 |
ENSDART00000147474
|
stmn2b
|
stathmin 2b |
| chr7_-_69636502 | 1.46 |
ENSDART00000126739
|
tspan5a
|
tetraspanin 5a |
| chr21_-_37973819 | 1.45 |
ENSDART00000133405
|
ripply1
|
ripply transcriptional repressor 1 |
| chr18_+_17611627 | 1.45 |
ENSDART00000046891
|
cetp
|
cholesteryl ester transfer protein, plasma |
| chr10_-_36808348 | 1.45 |
ENSDART00000099320
|
dhrs13a.1
|
dehydrogenase/reductase (SDR family) member 13a, tandem duplicate 1 |
| chr24_+_21346796 | 1.45 |
ENSDART00000126519
|
shisa2b
|
shisa family member 2b |
| chr6_-_8311044 | 1.45 |
ENSDART00000129674
|
slc44a2
|
solute carrier family 44 (choline transporter), member 2 |
| chr14_-_49063157 | 1.43 |
ENSDART00000021260
|
sept8b
|
septin 8b |
| chr15_-_17813680 | 1.43 |
ENSDART00000158556
|
CT573342.2
|
|
| chr7_+_21859337 | 1.41 |
ENSDART00000159626
|
si:dkey-85k7.7
|
si:dkey-85k7.7 |
| chr8_+_3085120 | 1.41 |
ENSDART00000148020
ENSDART00000136250 |
lhx2b
|
LIM homeobox 2b |
| chr1_-_669717 | 1.39 |
ENSDART00000160564
|
cyyr1
|
cysteine/tyrosine-rich 1 |
| chr21_+_26720803 | 1.38 |
ENSDART00000053797
|
slc3a2b
|
solute carrier family 3 (amino acid transporter heavy chain), member 2b |
| chr5_-_57655092 | 1.38 |
ENSDART00000074290
|
mia
|
melanoma inhibitory activity |
| chr20_-_30920356 | 1.37 |
ENSDART00000022951
|
kif25
|
kinesin family member 25 |
| chr18_-_46256560 | 1.36 |
ENSDART00000171375
|
si:dkey-244a7.1
|
si:dkey-244a7.1 |
| chr12_+_10631266 | 1.34 |
ENSDART00000161455
|
csf3a
|
colony stimulating factor 3 (granulocyte) a |
| chr12_+_316238 | 1.33 |
ENSDART00000187492
|
rcvrnb
|
recoverin b |
| chr24_-_23942722 | 1.33 |
ENSDART00000080810
|
arxa
|
aristaless related homeobox a |
| chr3_+_12440099 | 1.30 |
ENSDART00000158060
|
vasnb
|
vasorin b |
| chr13_+_30903816 | 1.30 |
ENSDART00000191727
|
ercc6
|
excision repair cross-complementation group 6 |
| chr4_-_16001118 | 1.30 |
ENSDART00000041070
ENSDART00000125389 |
mest
|
mesoderm specific transcript |
| chr15_-_44512461 | 1.29 |
ENSDART00000155456
|
gria4a
|
glutamate receptor, ionotropic, AMPA 4a |
| chr22_-_24284447 | 1.28 |
ENSDART00000149894
|
si:ch211-117l17.4
|
si:ch211-117l17.4 |
| chr1_-_52494122 | 1.26 |
ENSDART00000131407
|
acy3.2
|
aspartoacylase (aminocyclase) 3, tandem duplicate 2 |
| chr25_+_5044780 | 1.26 |
ENSDART00000153980
|
parvb
|
parvin, beta |
| chr20_-_7072487 | 1.25 |
ENSDART00000145954
|
si:ch211-121a2.2
|
si:ch211-121a2.2 |
| chr6_+_6924637 | 1.25 |
ENSDART00000065551
ENSDART00000151393 |
zak
|
sterile alpha motif and leucine zipper containing kinase AZK |
| chr25_-_15040369 | 1.25 |
ENSDART00000159342
ENSDART00000166490 |
pax6a
|
paired box 6a |
| chr3_-_30158395 | 1.23 |
ENSDART00000103502
|
si:ch211-152f23.5
|
si:ch211-152f23.5 |
| chr4_+_6032640 | 1.23 |
ENSDART00000157487
|
tfec
|
transcription factor EC |
| chr24_-_7697274 | 1.22 |
ENSDART00000186077
|
syt5b
|
synaptotagmin Vb |
| chr22_+_9872323 | 1.22 |
ENSDART00000129240
|
si:dkey-253d23.4
|
si:dkey-253d23.4 |
| chr7_+_15872357 | 1.22 |
ENSDART00000165757
|
pax6b
|
paired box 6b |
| chr19_+_10339538 | 1.20 |
ENSDART00000151808
ENSDART00000151235 |
rcvrn3
|
recoverin 3 |
| chr7_+_25858380 | 1.20 |
ENSDART00000148780
ENSDART00000079218 |
mtmr1a
|
myotubularin related protein 1a |
| chr4_+_72723304 | 1.20 |
ENSDART00000186791
ENSDART00000158902 ENSDART00000191925 |
rab3ip
|
RAB3A interacting protein (rabin3) |
| chr23_+_26733232 | 1.20 |
ENSDART00000035080
|
zgc:158263
|
zgc:158263 |
| chr8_+_2487250 | 1.19 |
ENSDART00000081325
|
dynll1
|
dynein, light chain, LC8-type 1 |
| chr11_+_27543093 | 1.18 |
ENSDART00000023889
|
barx1
|
BARX homeobox 1 |
| chr6_+_37894220 | 1.16 |
ENSDART00000087311
|
oca2
|
oculocutaneous albinism II |
| chr23_+_28770225 | 1.16 |
ENSDART00000132179
ENSDART00000142273 |
masp2
|
mannan-binding lectin serine peptidase 2 |
| chr22_-_910926 | 1.16 |
ENSDART00000180075
|
FP016205.1
|
|
| chr24_+_25069609 | 1.15 |
ENSDART00000115165
|
amer2
|
APC membrane recruitment protein 2 |
| chr16_+_31921812 | 1.15 |
ENSDART00000176928
ENSDART00000193733 |
rps9
|
ribosomal protein S9 |
| chr1_-_10647484 | 1.12 |
ENSDART00000164541
ENSDART00000188958 ENSDART00000190904 |
si:dkey-31e10.1
|
si:dkey-31e10.1 |
| chr4_-_4834347 | 1.12 |
ENSDART00000141803
|
coa6
|
cytochrome c oxidase assembly factor 6 |
| chr7_+_66822229 | 1.12 |
ENSDART00000112109
|
lyve1a
|
lymphatic vessel endothelial hyaluronic receptor 1a |
| chr20_-_22476255 | 1.11 |
ENSDART00000103510
|
pdgfra
|
platelet-derived growth factor receptor, alpha polypeptide |
| chr17_-_37214196 | 1.10 |
ENSDART00000128715
|
kif3cb
|
kinesin family member 3Cb |
| chr11_-_42554290 | 1.10 |
ENSDART00000130573
|
atp6ap1la
|
ATPase H+ transporting accessory protein 1 like a |
| chr24_-_17067284 | 1.10 |
ENSDART00000111237
|
armc3
|
armadillo repeat containing 3 |
| chr4_-_28373909 | 1.09 |
ENSDART00000056132
|
tprkb
|
Tp53rk binding protein |
| chr19_-_28130658 | 1.08 |
ENSDART00000079114
|
irx1b
|
iroquois homeobox 1b |
| chr14_-_4145594 | 1.08 |
ENSDART00000077348
|
casp3b
|
caspase 3, apoptosis-related cysteine peptidase b |
| chr24_-_38384432 | 1.08 |
ENSDART00000140739
|
lrrc4bb
|
leucine rich repeat containing 4Bb |
| chr19_+_9174166 | 1.07 |
ENSDART00000104637
ENSDART00000150968 |
si:ch211-81a5.8
|
si:ch211-81a5.8 |
| chr5_-_51484156 | 1.07 |
ENSDART00000162064
|
CR388055.1
|
|
| chr11_-_3860534 | 1.06 |
ENSDART00000082425
|
gata2a
|
GATA binding protein 2a |
| chr16_-_27174373 | 1.06 |
ENSDART00000166681
|
frrs1l
|
ferric-chelate reductase 1-like |
| chr2_-_30135446 | 1.06 |
ENSDART00000141906
|
trpa1a
|
transient receptor potential cation channel, subfamily A, member 1a |
| chr1_+_37391141 | 1.06 |
ENSDART00000083593
ENSDART00000168647 |
sparcl1
|
SPARC-like 1 |
| chr7_+_56098590 | 1.05 |
ENSDART00000098453
|
cdh15
|
cadherin 15, type 1, M-cadherin (myotubule) |
| chr23_+_45584223 | 1.05 |
ENSDART00000149367
|
si:ch73-290k24.5
|
si:ch73-290k24.5 |
| chr14_+_11430796 | 1.04 |
ENSDART00000165275
|
si:ch211-153b23.3
|
si:ch211-153b23.3 |
| chr15_-_17869115 | 1.04 |
ENSDART00000112838
|
atf5b
|
activating transcription factor 5b |
| chr18_-_16801033 | 1.04 |
ENSDART00000100100
|
admb
|
adrenomedullin b |
| chr25_+_34641536 | 1.03 |
ENSDART00000167033
|
CABZ01079011.1
|
|
| chr3_-_32337653 | 1.03 |
ENSDART00000156918
ENSDART00000156551 |
si:dkey-16p21.8
|
si:dkey-16p21.8 |
| chr1_-_50859053 | 1.03 |
ENSDART00000132779
ENSDART00000137648 |
si:dkeyp-123h10.2
|
si:dkeyp-123h10.2 |
| chr15_-_35246742 | 1.03 |
ENSDART00000131479
|
mff
|
mitochondrial fission factor |
| chr7_+_20147202 | 1.02 |
ENSDART00000011398
|
si:ch73-335l21.1
|
si:ch73-335l21.1 |
| chr8_-_38477817 | 1.01 |
ENSDART00000075989
|
inpp5l
|
inositol polyphosphate-5-phosphatase L |
| chr1_-_46706639 | 1.01 |
ENSDART00000074519
|
kpna3
|
karyopherin alpha 3 (importin alpha 4) |
| chr17_+_38476300 | 1.01 |
ENSDART00000123298
|
stard9
|
StAR-related lipid transfer (START) domain containing 9 |
| chr17_+_18117029 | 1.01 |
ENSDART00000154646
ENSDART00000179739 |
bcl11ba
|
B cell CLL/lymphoma 11Ba |
| chr13_+_30228077 | 1.00 |
ENSDART00000100813
ENSDART00000147729 ENSDART00000133404 |
rps24
|
ribosomal protein S24 |
| chr10_+_5414224 | 0.99 |
ENSDART00000138821
|
nfil3
|
nuclear factor, interleukin 3 regulated |
| chr4_-_77432218 | 0.99 |
ENSDART00000158683
|
slco1d1
|
solute carrier organic anion transporter family, member 1D1 |
| chr20_-_8094718 | 0.99 |
ENSDART00000122902
|
si:ch211-232i5.3
|
si:ch211-232i5.3 |
| chr20_+_5106568 | 0.98 |
ENSDART00000028039
|
cyp46a1.1
|
cytochrome P450, family 46, subfamily A, polypeptide 1, tandem duplicate 1 |
| chr10_-_31440500 | 0.98 |
ENSDART00000024778
|
robo3
|
roundabout, axon guidance receptor, homolog 3 (Drosophila) |
| chr2_-_10188598 | 0.98 |
ENSDART00000189122
|
dmbx1a
|
diencephalon/mesencephalon homeobox 1a |
| chr2_-_9489611 | 0.97 |
ENSDART00000146715
|
st6galnac3
|
ST6 (alpha-N-acetyl-neuraminyl-2,3-beta-galactosyl-1,3)-N-acetylgalactosaminide alpha-2,6-sialyltransferase 3 |
| chr24_+_9475809 | 0.96 |
ENSDART00000132688
|
si:ch211-285f17.1
|
si:ch211-285f17.1 |
| chr16_+_22654481 | 0.96 |
ENSDART00000179762
|
chrnb2b
|
cholinergic receptor, nicotinic, beta 2b |
| chr1_+_23784905 | 0.94 |
ENSDART00000171951
ENSDART00000188521 ENSDART00000183029 ENSDART00000187183 |
slit2
|
slit homolog 2 (Drosophila) |
| chr15_+_19797918 | 0.94 |
ENSDART00000113314
|
si:ch211-229d2.5
|
si:ch211-229d2.5 |
| chr19_+_37701450 | 0.94 |
ENSDART00000087694
|
thsd7aa
|
thrombospondin, type I, domain containing 7Aa |
| chr20_-_9462433 | 0.94 |
ENSDART00000152674
ENSDART00000040557 |
zgc:101840
|
zgc:101840 |
| chr15_-_17868870 | 0.93 |
ENSDART00000170950
|
atf5b
|
activating transcription factor 5b |
| chr18_-_21271373 | 0.93 |
ENSDART00000060001
|
pnp6
|
purine nucleoside phosphorylase 6 |
| chr18_-_15932704 | 0.92 |
ENSDART00000127769
|
plekhg7
|
pleckstrin homology domain containing, family G (with RhoGef domain) member 7 |
| chr17_-_26926577 | 0.91 |
ENSDART00000050202
|
rcan3
|
regulator of calcineurin 3 |
| chr6_+_43426599 | 0.91 |
ENSDART00000056457
|
mitfa
|
microphthalmia-associated transcription factor a |
| chr1_-_10647307 | 0.90 |
ENSDART00000103548
|
si:dkey-31e10.1
|
si:dkey-31e10.1 |
| chr5_+_40224938 | 0.90 |
ENSDART00000142897
|
si:dkey-193c22.2
|
si:dkey-193c22.2 |
| chr25_+_31227747 | 0.90 |
ENSDART00000033872
|
tnni2a.1
|
troponin I type 2a (skeletal, fast), tandem duplicate 1 |
| chr18_-_8885792 | 0.89 |
ENSDART00000143619
|
si:dkey-95h12.1
|
si:dkey-95h12.1 |
| chr1_-_46981134 | 0.89 |
ENSDART00000130607
|
pknox1.2
|
pbx/knotted 1 homeobox 1.2 |
| chr4_-_390431 | 0.89 |
ENSDART00000067482
ENSDART00000138500 |
dynlt1
|
dynein, light chain, Tctex-type 1 |
| chr10_-_10607118 | 0.88 |
ENSDART00000101089
|
dbh
|
dopamine beta-hydroxylase (dopamine beta-monooxygenase) |
| chr5_+_48683447 | 0.88 |
ENSDART00000008043
|
adgrv1
|
adhesion G protein-coupled receptor V1 |
Network of associatons between targets according to the STRING database.
First level regulatory network of prrx1a+prrx1b
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 3.9 | GO:0061549 | sympathetic ganglion development(GO:0061549) |
| 0.9 | 3.7 | GO:0015859 | intracellular nucleoside transport(GO:0015859) purine nucleoside transmembrane transport(GO:0015860) purine nucleotide transport(GO:0015865) ATP transport(GO:0015867) purine ribonucleotide transport(GO:0015868) adenine nucleotide transport(GO:0051503) mitochondrial ATP transmembrane transport(GO:1990544) |
| 0.9 | 2.6 | GO:0009912 | auditory receptor cell fate commitment(GO:0009912) inner ear receptor cell fate commitment(GO:0060120) |
| 0.9 | 2.6 | GO:0019852 | L-ascorbic acid metabolic process(GO:0019852) water-soluble vitamin biosynthetic process(GO:0042364) |
| 0.8 | 2.4 | GO:0015695 | organic cation transport(GO:0015695) |
| 0.8 | 3.0 | GO:0043152 | induction of bacterial agglutination(GO:0043152) |
| 0.7 | 2.0 | GO:0014722 | regulation of skeletal muscle contraction by calcium ion signaling(GO:0014722) regulation of skeletal muscle contraction by regulation of release of sequestered calcium ion(GO:0014809) |
| 0.6 | 1.8 | GO:0006499 | N-terminal protein myristoylation(GO:0006499) N-terminal peptidyl-glycine N-myristoylation(GO:0018008) protein myristoylation(GO:0018377) |
| 0.6 | 28.8 | GO:0003009 | skeletal muscle contraction(GO:0003009) |
| 0.6 | 3.9 | GO:0045299 | otolith mineralization(GO:0045299) |
| 0.5 | 1.5 | GO:0034375 | high-density lipoprotein particle remodeling(GO:0034375) |
| 0.5 | 1.8 | GO:0061113 | pancreas morphogenesis(GO:0061113) |
| 0.4 | 2.1 | GO:0006642 | triglyceride mobilization(GO:0006642) |
| 0.4 | 2.9 | GO:1904030 | negative regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0045736) negative regulation of cyclin-dependent protein kinase activity(GO:1904030) |
| 0.4 | 1.6 | GO:0014857 | skeletal muscle cell proliferation(GO:0014856) regulation of skeletal muscle cell proliferation(GO:0014857) |
| 0.4 | 2.8 | GO:0019731 | antibacterial humoral response(GO:0019731) |
| 0.4 | 1.2 | GO:2000648 | positive regulation of stem cell proliferation(GO:2000648) |
| 0.4 | 2.3 | GO:0021767 | mammillary body development(GO:0021767) social behavior(GO:0035176) intraspecies interaction between organisms(GO:0051703) |
| 0.4 | 2.3 | GO:2001279 | neutrophil apoptotic process(GO:0001781) regulation of T-helper 1 type immune response(GO:0002825) positive regulation of T-helper 1 type immune response(GO:0002827) negative regulation of type 2 immune response(GO:0002829) inflammatory cell apoptotic process(GO:0006925) myoblast migration involved in skeletal muscle regeneration(GO:0014839) regulation of prostaglandin biosynthetic process(GO:0031392) positive regulation of prostaglandin biosynthetic process(GO:0031394) regulation of phospholipase A2 activity(GO:0032429) interleukin-2 production(GO:0032623) regulation of interleukin-2 production(GO:0032663) positive regulation of interleukin-2 production(GO:0032743) myeloid cell apoptotic process(GO:0033028) regulation of neutrophil apoptotic process(GO:0033029) positive regulation of neutrophil apoptotic process(GO:0033031) regulation of myeloid cell apoptotic process(GO:0033032) positive regulation of myeloid cell apoptotic process(GO:0033034) T-helper 1 type immune response(GO:0042088) positive regulation of CD4-positive, alpha-beta T cell differentiation(GO:0043372) T-helper 1 cell differentiation(GO:0045063) positive regulation of T-helper cell differentiation(GO:0045624) regulation of T-helper 1 cell differentiation(GO:0045625) positive regulation of T-helper 1 cell differentiation(GO:0045627) regulation of T-helper 2 cell differentiation(GO:0045628) negative regulation of T-helper 2 cell differentiation(GO:0045629) positive regulation of fatty acid biosynthetic process(GO:0045723) positive regulation of alpha-beta T cell differentiation(GO:0046638) myoblast migration(GO:0051451) neutrophil clearance(GO:0097350) negative regulation of phospholipase A2 activity(GO:1900138) positive regulation of CD4-positive, alpha-beta T cell activation(GO:2000516) regulation of unsaturated fatty acid biosynthetic process(GO:2001279) positive regulation of unsaturated fatty acid biosynthetic process(GO:2001280) |
| 0.4 | 1.9 | GO:0070292 | N-acylphosphatidylethanolamine metabolic process(GO:0070292) |
| 0.4 | 1.5 | GO:0099543 | trans-synaptic signaling by soluble gas(GO:0099543) trans-synaptic signaling by nitric oxide(GO:0099548) |
| 0.4 | 0.7 | GO:1900620 | acetylcholine biosynthetic process(GO:0008292) acetate ester biosynthetic process(GO:1900620) |
| 0.4 | 1.4 | GO:0015871 | choline transport(GO:0015871) |
| 0.4 | 1.1 | GO:1902893 | pri-miRNA transcription from RNA polymerase II promoter(GO:0061614) regulation of pri-miRNA transcription from RNA polymerase II promoter(GO:1902893) positive regulation of pri-miRNA transcription from RNA polymerase II promoter(GO:1902895) |
| 0.3 | 1.3 | GO:0003322 | pancreatic A cell development(GO:0003322) |
| 0.3 | 2.3 | GO:0006621 | protein retention in ER lumen(GO:0006621) |
| 0.3 | 0.9 | GO:0007414 | axonal defasciculation(GO:0007414) |
| 0.3 | 3.7 | GO:0006285 | base-excision repair, AP site formation(GO:0006285) |
| 0.3 | 1.5 | GO:0033119 | negative regulation of RNA splicing(GO:0033119) negative regulation of mRNA splicing, via spliceosome(GO:0048025) |
| 0.3 | 2.1 | GO:0006283 | transcription-coupled nucleotide-excision repair(GO:0006283) |
| 0.3 | 1.8 | GO:2000576 | positive regulation of microtubule motor activity(GO:2000576) regulation of ATP-dependent microtubule motor activity, plus-end-directed(GO:2000580) positive regulation of ATP-dependent microtubule motor activity, plus-end-directed(GO:2000582) |
| 0.3 | 0.9 | GO:0046333 | octopamine biosynthetic process(GO:0006589) dopamine catabolic process(GO:0042420) norepinephrine biosynthetic process(GO:0042421) octopamine metabolic process(GO:0046333) |
| 0.3 | 0.9 | GO:0097264 | self proteolysis(GO:0097264) |
| 0.3 | 1.1 | GO:0045903 | positive regulation of translational fidelity(GO:0045903) |
| 0.3 | 1.1 | GO:0050960 | thermoception(GO:0050955) detection of temperature stimulus involved in thermoception(GO:0050960) detection of temperature stimulus involved in sensory perception(GO:0050961) |
| 0.3 | 3.9 | GO:0044872 | lipoprotein transport(GO:0042953) lipoprotein localization(GO:0044872) |
| 0.3 | 2.1 | GO:0090303 | positive regulation of wound healing(GO:0090303) |
| 0.3 | 1.5 | GO:0009110 | vitamin biosynthetic process(GO:0009110) |
| 0.2 | 1.4 | GO:0071632 | optomotor response(GO:0071632) |
| 0.2 | 0.7 | GO:0060292 | long term synaptic depression(GO:0060292) |
| 0.2 | 1.1 | GO:0003261 | cardiac muscle progenitor cell migration to the midline involved in heart field formation(GO:0003261) |
| 0.2 | 1.1 | GO:0002949 | tRNA threonylcarbamoyladenosine modification(GO:0002949) |
| 0.2 | 1.5 | GO:1901031 | response to leucine(GO:0043201) cellular response to leucine(GO:0071233) regulation of response to reactive oxygen species(GO:1901031) |
| 0.2 | 3.8 | GO:0051764 | actin crosslink formation(GO:0051764) |
| 0.2 | 2.5 | GO:0030656 | regulation of isoprenoid metabolic process(GO:0019747) regulation of vitamin metabolic process(GO:0030656) regulation of retinoic acid biosynthetic process(GO:1900052) |
| 0.2 | 2.3 | GO:0034605 | cellular response to heat(GO:0034605) |
| 0.2 | 1.8 | GO:0044550 | secondary metabolite biosynthetic process(GO:0044550) |
| 0.2 | 1.2 | GO:0021877 | forebrain neuron fate commitment(GO:0021877) |
| 0.2 | 1.2 | GO:0060221 | retinal rod cell differentiation(GO:0060221) |
| 0.2 | 1.2 | GO:0090104 | pancreatic D cell differentiation(GO:0003311) pancreatic epsilon cell differentiation(GO:0090104) |
| 0.2 | 1.0 | GO:0006450 | regulation of translational fidelity(GO:0006450) |
| 0.2 | 0.9 | GO:0071380 | response to prostaglandin E(GO:0034695) cellular response to prostaglandin E stimulus(GO:0071380) |
| 0.2 | 0.7 | GO:0060092 | inhibitory postsynaptic potential(GO:0060080) regulation of synaptic transmission, glycinergic(GO:0060092) |
| 0.2 | 0.3 | GO:0046635 | positive regulation of alpha-beta T cell activation(GO:0046635) |
| 0.2 | 0.5 | GO:0032290 | peripheral nervous system myelin formation(GO:0032290) semicircular canal fusion(GO:0060879) |
| 0.1 | 2.0 | GO:1905150 | regulation of voltage-gated sodium channel activity(GO:1905150) |
| 0.1 | 5.5 | GO:0030199 | collagen fibril organization(GO:0030199) |
| 0.1 | 0.8 | GO:0099525 | synaptic vesicle docking(GO:0016081) presynaptic dense core granule exocytosis(GO:0099525) |
| 0.1 | 1.9 | GO:0001964 | startle response(GO:0001964) |
| 0.1 | 0.4 | GO:0021559 | trigeminal nerve development(GO:0021559) |
| 0.1 | 1.5 | GO:0002031 | G-protein coupled receptor internalization(GO:0002031) |
| 0.1 | 1.6 | GO:0045162 | clustering of voltage-gated sodium channels(GO:0045162) |
| 0.1 | 2.0 | GO:0071425 | hematopoietic stem cell proliferation(GO:0071425) |
| 0.1 | 1.2 | GO:0036368 | cone photoresponse recovery(GO:0036368) |
| 0.1 | 0.9 | GO:0042554 | superoxide anion generation(GO:0042554) |
| 0.1 | 0.9 | GO:0032889 | regulation of vacuole fusion, non-autophagic(GO:0032889) |
| 0.1 | 0.3 | GO:0060898 | eye field cell fate commitment involved in camera-type eye formation(GO:0060898) |
| 0.1 | 1.9 | GO:0008535 | respiratory chain complex IV assembly(GO:0008535) |
| 0.1 | 0.5 | GO:0045604 | regulation of epidermal cell differentiation(GO:0045604) |
| 0.1 | 1.1 | GO:0030216 | keratinocyte differentiation(GO:0030216) |
| 0.1 | 1.0 | GO:0016127 | cholesterol catabolic process(GO:0006707) sterol catabolic process(GO:0016127) |
| 0.1 | 0.3 | GO:0097052 | L-kynurenine metabolic process(GO:0097052) L-kynurenine catabolic process(GO:0097053) |
| 0.1 | 1.0 | GO:0006607 | NLS-bearing protein import into nucleus(GO:0006607) |
| 0.1 | 0.3 | GO:2001046 | positive regulation of integrin-mediated signaling pathway(GO:2001046) |
| 0.1 | 2.6 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.1 | 2.2 | GO:0070831 | basement membrane assembly(GO:0070831) |
| 0.1 | 1.1 | GO:0016199 | axon midline choice point recognition(GO:0016199) |
| 0.1 | 0.2 | GO:1904158 | axonemal central apparatus assembly(GO:1904158) |
| 0.1 | 1.5 | GO:0042574 | retinal metabolic process(GO:0042574) |
| 0.1 | 1.4 | GO:0061074 | regulation of neural retina development(GO:0061074) |
| 0.1 | 1.0 | GO:0001574 | ganglioside biosynthetic process(GO:0001574) |
| 0.1 | 0.4 | GO:0006545 | glycine biosynthetic process(GO:0006545) |
| 0.1 | 0.7 | GO:1904071 | presynaptic active zone assembly(GO:1904071) |
| 0.1 | 1.2 | GO:0001867 | complement activation, lectin pathway(GO:0001867) |
| 0.1 | 1.1 | GO:0043248 | proteasome assembly(GO:0043248) |
| 0.1 | 1.0 | GO:0071545 | phosphorylated carbohydrate dephosphorylation(GO:0046838) inositol phosphate dephosphorylation(GO:0046855) inositol phosphate catabolic process(GO:0071545) |
| 0.1 | 1.6 | GO:0010883 | regulation of lipid storage(GO:0010883) |
| 0.1 | 2.8 | GO:0060536 | cartilage morphogenesis(GO:0060536) |
| 0.1 | 3.7 | GO:0006096 | glycolytic process(GO:0006096) |
| 0.1 | 2.2 | GO:0031532 | actin cytoskeleton reorganization(GO:0031532) |
| 0.1 | 1.2 | GO:0007398 | ectoderm development(GO:0007398) |
| 0.1 | 0.5 | GO:0046959 | learning(GO:0007612) nonassociative learning(GO:0046958) habituation(GO:0046959) |
| 0.1 | 0.2 | GO:0098908 | regulation of neuronal action potential(GO:0098908) |
| 0.1 | 1.2 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
| 0.1 | 0.2 | GO:0045198 | establishment of epithelial cell apical/basal polarity(GO:0045198) |
| 0.1 | 2.1 | GO:0006904 | vesicle docking involved in exocytosis(GO:0006904) |
| 0.1 | 0.7 | GO:0046051 | UTP biosynthetic process(GO:0006228) UTP metabolic process(GO:0046051) |
| 0.0 | 0.8 | GO:0071594 | T cell differentiation in thymus(GO:0033077) thymocyte aggregation(GO:0071594) |
| 0.0 | 0.3 | GO:0006086 | acetyl-CoA biosynthetic process from pyruvate(GO:0006086) |
| 0.0 | 0.6 | GO:0071378 | growth hormone receptor signaling pathway(GO:0060396) response to growth hormone(GO:0060416) cellular response to growth hormone stimulus(GO:0071378) |
| 0.0 | 0.8 | GO:0006825 | copper ion transport(GO:0006825) |
| 0.0 | 0.9 | GO:0051496 | positive regulation of stress fiber assembly(GO:0051496) |
| 0.0 | 0.1 | GO:0001774 | microglial cell activation(GO:0001774) |
| 0.0 | 0.6 | GO:0051452 | vacuolar acidification(GO:0007035) pH reduction(GO:0045851) intracellular pH reduction(GO:0051452) |
| 0.0 | 2.1 | GO:0070830 | apical junction assembly(GO:0043297) bicellular tight junction assembly(GO:0070830) |
| 0.0 | 1.6 | GO:0051904 | melanosome transport(GO:0032402) pigment granule transport(GO:0051904) |
| 0.0 | 1.8 | GO:0010669 | epithelial structure maintenance(GO:0010669) |
| 0.0 | 0.7 | GO:0005979 | regulation of glycogen biosynthetic process(GO:0005979) regulation of glucan biosynthetic process(GO:0010962) |
| 0.0 | 0.2 | GO:0060142 | regulation of syncytium formation by plasma membrane fusion(GO:0060142) |
| 0.0 | 0.4 | GO:0048641 | regulation of skeletal muscle tissue development(GO:0048641) |
| 0.0 | 0.4 | GO:0046887 | positive regulation of hormone secretion(GO:0046887) |
| 0.0 | 2.6 | GO:0014904 | myotube cell development(GO:0014904) skeletal muscle fiber development(GO:0048741) |
| 0.0 | 0.6 | GO:0060088 | auditory receptor cell stereocilium organization(GO:0060088) |
| 0.0 | 0.5 | GO:0030206 | chondroitin sulfate biosynthetic process(GO:0030206) |
| 0.0 | 1.2 | GO:0046856 | phosphatidylinositol dephosphorylation(GO:0046856) |
| 0.0 | 0.7 | GO:0043252 | sodium-independent organic anion transport(GO:0043252) |
| 0.0 | 0.1 | GO:0021846 | cell proliferation in forebrain(GO:0021846) |
| 0.0 | 0.3 | GO:0034058 | endosomal vesicle fusion(GO:0034058) |
| 0.0 | 0.6 | GO:0032981 | mitochondrial electron transport, NADH to ubiquinone(GO:0006120) NADH dehydrogenase complex assembly(GO:0010257) mitochondrial respiratory chain complex I assembly(GO:0032981) mitochondrial respiratory chain complex I biogenesis(GO:0097031) |
| 0.0 | 0.4 | GO:0036158 | outer dynein arm assembly(GO:0036158) |
| 0.0 | 3.6 | GO:0002088 | lens development in camera-type eye(GO:0002088) |
| 0.0 | 1.2 | GO:0036269 | swimming behavior(GO:0036269) |
| 0.0 | 0.1 | GO:0016998 | cell wall macromolecule catabolic process(GO:0016998) |
| 0.0 | 0.2 | GO:1901673 | regulation of mitotic spindle assembly(GO:1901673) |
| 0.0 | 1.1 | GO:0046928 | regulation of neurotransmitter secretion(GO:0046928) |
| 0.0 | 0.9 | GO:0060037 | pharyngeal system development(GO:0060037) |
| 0.0 | 0.5 | GO:0036092 | phosphatidylinositol-3-phosphate biosynthetic process(GO:0036092) |
| 0.0 | 1.9 | GO:0033339 | pectoral fin development(GO:0033339) |
| 0.0 | 1.1 | GO:1900449 | regulation of glutamate receptor signaling pathway(GO:1900449) |
| 0.0 | 0.3 | GO:0060122 | inner ear receptor stereocilium organization(GO:0060122) |
| 0.0 | 0.3 | GO:0010717 | regulation of epithelial to mesenchymal transition(GO:0010717) |
| 0.0 | 1.5 | GO:0021515 | cell differentiation in spinal cord(GO:0021515) |
| 0.0 | 0.8 | GO:0006376 | mRNA splice site selection(GO:0006376) |
| 0.0 | 0.6 | GO:0008277 | regulation of G-protein coupled receptor protein signaling pathway(GO:0008277) |
| 0.0 | 1.4 | GO:0018107 | peptidyl-threonine phosphorylation(GO:0018107) |
| 0.0 | 0.9 | GO:0045197 | establishment or maintenance of epithelial cell apical/basal polarity(GO:0045197) |
| 0.0 | 0.7 | GO:0055064 | chloride ion homeostasis(GO:0055064) |
| 0.0 | 1.7 | GO:0007492 | endoderm development(GO:0007492) |
| 0.0 | 0.4 | GO:0030150 | protein import into mitochondrial matrix(GO:0030150) |
| 0.0 | 0.1 | GO:0050802 | circadian sleep/wake cycle process(GO:0022410) melatonin metabolic process(GO:0030186) circadian sleep/wake cycle(GO:0042745) regulation of circadian sleep/wake cycle(GO:0042749) regulation of circadian sleep/wake cycle, sleep(GO:0045187) circadian sleep/wake cycle, sleep(GO:0050802) |
| 0.0 | 0.5 | GO:1903845 | negative regulation of transforming growth factor beta receptor signaling pathway(GO:0030512) negative regulation of cellular response to transforming growth factor beta stimulus(GO:1903845) |
| 0.0 | 1.5 | GO:0008045 | motor neuron axon guidance(GO:0008045) |
| 0.0 | 0.1 | GO:0030431 | sleep(GO:0030431) |
| 0.0 | 0.6 | GO:0036297 | interstrand cross-link repair(GO:0036297) |
| 0.0 | 1.0 | GO:0050852 | T cell receptor signaling pathway(GO:0050852) |
| 0.0 | 0.2 | GO:0038203 | TORC2 signaling(GO:0038203) |
| 0.0 | 0.3 | GO:1902017 | regulation of cilium assembly(GO:1902017) |
| 0.0 | 0.8 | GO:0007019 | microtubule depolymerization(GO:0007019) |
| 0.0 | 1.1 | GO:1902600 | hydrogen ion transmembrane transport(GO:1902600) |
| 0.0 | 0.4 | GO:0030497 | fatty acid elongation(GO:0030497) |
| 0.0 | 0.8 | GO:0071277 | cellular response to calcium ion(GO:0071277) |
| 0.0 | 0.3 | GO:1990798 | pancreas regeneration(GO:1990798) |
| 0.0 | 0.6 | GO:0042149 | cellular response to glucose starvation(GO:0042149) |
| 0.0 | 0.5 | GO:0006099 | tricarboxylic acid cycle(GO:0006099) |
| 0.0 | 0.4 | GO:0051482 | positive regulation of cytosolic calcium ion concentration involved in phospholipase C-activating G-protein coupled signaling pathway(GO:0051482) |
| 0.0 | 0.6 | GO:0032012 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.0 | 2.0 | GO:0060326 | cell chemotaxis(GO:0060326) |
| 0.0 | 1.0 | GO:0060216 | definitive hemopoiesis(GO:0060216) |
| 0.0 | 0.5 | GO:0071880 | adenylate cyclase-activating adrenergic receptor signaling pathway(GO:0071880) |
| 0.0 | 0.3 | GO:0038061 | NIK/NF-kappaB signaling(GO:0038061) |
| 0.0 | 0.2 | GO:0019433 | triglyceride catabolic process(GO:0019433) |
| 0.0 | 1.5 | GO:0007601 | visual perception(GO:0007601) |
| 0.0 | 0.1 | GO:0099558 | maintenance of presynaptic active zone structure(GO:0048790) maintenance of synapse structure(GO:0099558) presynaptic active zone organization(GO:1990709) |
| 0.0 | 0.3 | GO:0033334 | fin morphogenesis(GO:0033334) |
| 0.0 | 1.7 | GO:0030155 | regulation of cell adhesion(GO:0030155) |
| 0.0 | 1.0 | GO:0061640 | cytoskeleton-dependent cytokinesis(GO:0061640) |
| 0.0 | 0.4 | GO:0050731 | positive regulation of peptidyl-tyrosine phosphorylation(GO:0050731) |
| 0.0 | 0.3 | GO:0099500 | synaptic vesicle fusion to presynaptic active zone membrane(GO:0031629) vesicle fusion to plasma membrane(GO:0099500) |
| 0.0 | 1.1 | GO:0006979 | response to oxidative stress(GO:0006979) |
| 0.0 | 0.2 | GO:0032958 | inositol phosphate biosynthetic process(GO:0032958) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.4 | 5.5 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 0.7 | 28.8 | GO:0005861 | troponin complex(GO:0005861) |
| 0.6 | 6.4 | GO:0034362 | low-density lipoprotein particle(GO:0034362) |
| 0.3 | 1.2 | GO:0070319 | Golgi to plasma membrane transport vesicle(GO:0070319) |
| 0.3 | 2.0 | GO:0033018 | sarcoplasmic reticulum lumen(GO:0033018) |
| 0.2 | 0.9 | GO:0060171 | stereocilium membrane(GO:0060171) |
| 0.2 | 1.5 | GO:0061700 | GATOR2 complex(GO:0061700) |
| 0.2 | 1.1 | GO:0033181 | plasma membrane proton-transporting V-type ATPase complex(GO:0033181) |
| 0.2 | 0.5 | GO:0042709 | succinate-CoA ligase complex(GO:0042709) |
| 0.2 | 2.6 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.2 | 0.6 | GO:0070195 | growth hormone receptor complex(GO:0070195) |
| 0.2 | 1.1 | GO:0000408 | EKC/KEOPS complex(GO:0000408) |
| 0.1 | 1.5 | GO:0034364 | high-density lipoprotein particle(GO:0034364) |
| 0.1 | 2.2 | GO:0016328 | lateral plasma membrane(GO:0016328) |
| 0.1 | 1.8 | GO:0005868 | cytoplasmic dynein complex(GO:0005868) |
| 0.1 | 2.3 | GO:0005801 | cis-Golgi network(GO:0005801) |
| 0.1 | 1.9 | GO:0031209 | SCAR complex(GO:0031209) |
| 0.1 | 0.9 | GO:0043020 | NADPH oxidase complex(GO:0043020) |
| 0.1 | 0.9 | GO:0000015 | phosphopyruvate hydratase complex(GO:0000015) |
| 0.1 | 1.2 | GO:0031045 | dense core granule(GO:0031045) |
| 0.1 | 0.4 | GO:0035301 | Hedgehog signaling complex(GO:0035301) |
| 0.1 | 0.6 | GO:0005851 | eukaryotic translation initiation factor 2B complex(GO:0005851) |
| 0.1 | 1.9 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.1 | 4.3 | GO:0030426 | growth cone(GO:0030426) |
| 0.1 | 0.7 | GO:0098982 | GABA-ergic synapse(GO:0098982) |
| 0.1 | 1.5 | GO:0031430 | M band(GO:0031430) |
| 0.1 | 1.5 | GO:0001917 | photoreceptor inner segment(GO:0001917) |
| 0.1 | 0.4 | GO:0005854 | nascent polypeptide-associated complex(GO:0005854) |
| 0.1 | 0.7 | GO:0005832 | chaperonin-containing T-complex(GO:0005832) |
| 0.1 | 0.6 | GO:0000220 | vacuolar proton-transporting V-type ATPase, V0 domain(GO:0000220) |
| 0.1 | 0.5 | GO:0005943 | phosphatidylinositol 3-kinase complex, class IA(GO:0005943) |
| 0.0 | 0.3 | GO:0005967 | mitochondrial pyruvate dehydrogenase complex(GO:0005967) |
| 0.0 | 0.4 | GO:0034045 | pre-autophagosomal structure membrane(GO:0034045) |
| 0.0 | 4.4 | GO:0055037 | recycling endosome(GO:0055037) |
| 0.0 | 0.4 | GO:0030122 | AP-2 adaptor complex(GO:0030122) |
| 0.0 | 0.7 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.0 | 3.7 | GO:0005604 | basement membrane(GO:0005604) |
| 0.0 | 0.3 | GO:0030897 | HOPS complex(GO:0030897) |
| 0.0 | 0.4 | GO:0005742 | mitochondrial outer membrane translocase complex(GO:0005742) |
| 0.0 | 0.4 | GO:0036157 | outer dynein arm(GO:0036157) |
| 0.0 | 2.9 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.0 | 0.2 | GO:1990130 | Iml1 complex(GO:1990130) |
| 0.0 | 0.4 | GO:0005892 | acetylcholine-gated channel complex(GO:0005892) |
| 0.0 | 1.3 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.0 | 0.2 | GO:0035060 | brahma complex(GO:0035060) |
| 0.0 | 0.3 | GO:0014704 | intercalated disc(GO:0014704) |
| 0.0 | 1.0 | GO:0005940 | septin ring(GO:0005940) septin complex(GO:0031105) septin cytoskeleton(GO:0032156) |
| 0.0 | 1.8 | GO:0030141 | secretory granule(GO:0030141) |
| 0.0 | 1.0 | GO:0005758 | mitochondrial intermembrane space(GO:0005758) organelle envelope lumen(GO:0031970) |
| 0.0 | 1.1 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.0 | 0.3 | GO:0043679 | axon terminus(GO:0043679) |
| 0.0 | 2.0 | GO:0030018 | Z disc(GO:0030018) |
| 0.0 | 1.0 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.0 | 0.5 | GO:0033116 | endoplasmic reticulum-Golgi intermediate compartment membrane(GO:0033116) |
| 0.0 | 1.5 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.0 | 0.3 | GO:0031594 | neuromuscular junction(GO:0031594) |
| 0.0 | 2.1 | GO:0005923 | bicellular tight junction(GO:0005923) occluding junction(GO:0070160) |
| 0.0 | 2.9 | GO:0005840 | ribosome(GO:0005840) |
| 0.0 | 0.8 | GO:0016342 | catenin complex(GO:0016342) |
| 0.0 | 0.3 | GO:0045180 | basal cortex(GO:0045180) |
| 0.0 | 0.0 | GO:0031417 | NatC complex(GO:0031417) |
| 0.0 | 0.6 | GO:0045271 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.0 | 0.2 | GO:0097546 | ciliary base(GO:0097546) |
| 0.0 | 0.1 | GO:0030891 | VCB complex(GO:0030891) |
| 0.0 | 0.7 | GO:0032580 | Golgi cisterna membrane(GO:0032580) |
| 0.0 | 1.3 | GO:0016324 | apical plasma membrane(GO:0016324) |
| 0.0 | 0.3 | GO:0046930 | pore complex(GO:0046930) |
| 0.0 | 0.2 | GO:0044447 | axoneme part(GO:0044447) |
| 0.0 | 0.2 | GO:0016010 | dystrophin-associated glycoprotein complex(GO:0016010) glycoprotein complex(GO:0090665) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 3.7 | GO:0005471 | ATP:ADP antiporter activity(GO:0005471) |
| 0.8 | 2.3 | GO:0046923 | ER retention sequence binding(GO:0046923) |
| 0.7 | 6.8 | GO:0016936 | galactoside binding(GO:0016936) |
| 0.6 | 1.9 | GO:0008970 | phosphatidylcholine 1-acylhydrolase activity(GO:0008970) |
| 0.6 | 2.5 | GO:0004365 | glyceraldehyde-3-phosphate dehydrogenase (NAD+) (phosphorylating) activity(GO:0004365) glyceraldehyde-3-phosphate dehydrogenase (NAD(P)+) (phosphorylating) activity(GO:0043891) |
| 0.6 | 1.8 | GO:0004379 | glycylpeptide N-tetradecanoyltransferase activity(GO:0004379) |
| 0.5 | 5.4 | GO:0016918 | retinal binding(GO:0016918) |
| 0.4 | 3.7 | GO:0008263 | mismatch base pair DNA N-glycosylase activity(GO:0000700) pyrimidine-specific mismatch base pair DNA N-glycosylase activity(GO:0008263) |
| 0.4 | 1.2 | GO:0001729 | ceramide kinase activity(GO:0001729) |
| 0.4 | 2.4 | GO:0005412 | glucose:sodium symporter activity(GO:0005412) |
| 0.4 | 2.3 | GO:0019834 | phospholipase A2 inhibitor activity(GO:0019834) |
| 0.4 | 2.2 | GO:0004743 | pyruvate kinase activity(GO:0004743) |
| 0.4 | 2.9 | GO:0004861 | cyclin-dependent protein serine/threonine kinase inhibitor activity(GO:0004861) |
| 0.4 | 1.4 | GO:0015220 | choline transmembrane transporter activity(GO:0015220) |
| 0.3 | 2.2 | GO:0050262 | ribosylnicotinamide kinase activity(GO:0050262) |
| 0.3 | 0.9 | GO:0004500 | dopamine beta-monooxygenase activity(GO:0004500) |
| 0.3 | 3.5 | GO:0015101 | organic cation transmembrane transporter activity(GO:0015101) |
| 0.3 | 1.1 | GO:0005017 | platelet-derived growth factor-activated receptor activity(GO:0005017) platelet-derived growth factor binding(GO:0048407) |
| 0.2 | 1.0 | GO:0015349 | thyroid hormone transmembrane transporter activity(GO:0015349) |
| 0.2 | 2.1 | GO:0070325 | low-density lipoprotein particle receptor binding(GO:0050750) lipoprotein particle receptor binding(GO:0070325) |
| 0.2 | 0.7 | GO:0031716 | calcitonin receptor binding(GO:0031716) |
| 0.2 | 1.3 | GO:0050431 | transforming growth factor beta binding(GO:0050431) |
| 0.2 | 1.5 | GO:0070728 | leucine binding(GO:0070728) |
| 0.2 | 1.0 | GO:0016531 | copper chaperone activity(GO:0016531) |
| 0.2 | 0.7 | GO:0072571 | ADP-D-ribose binding(GO:0072570) mono-ADP-D-ribose binding(GO:0072571) |
| 0.2 | 0.5 | GO:0004774 | succinate-CoA ligase activity(GO:0004774) succinate-CoA ligase (ADP-forming) activity(GO:0004775) succinate-CoA ligase (GDP-forming) activity(GO:0004776) |
| 0.2 | 1.0 | GO:0033781 | cholesterol 24-hydroxylase activity(GO:0033781) |
| 0.2 | 0.6 | GO:0004903 | growth hormone receptor activity(GO:0004903) |
| 0.2 | 0.9 | GO:0008597 | calcium-dependent protein serine/threonine phosphatase regulator activity(GO:0008597) |
| 0.2 | 3.0 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.2 | 0.6 | GO:0015105 | arsenite transmembrane transporter activity(GO:0015105) |
| 0.1 | 1.5 | GO:0052650 | NADP-retinol dehydrogenase activity(GO:0052650) |
| 0.1 | 0.5 | GO:0018812 | 3-hydroxyacyl-CoA dehydratase activity(GO:0018812) 3-hydroxy-behenoyl-CoA dehydratase activity(GO:0102344) 3-hydroxy-lignoceroyl-CoA dehydratase activity(GO:0102345) |
| 0.1 | 1.8 | GO:0004029 | aldehyde dehydrogenase (NAD) activity(GO:0004029) |
| 0.1 | 1.0 | GO:0001665 | alpha-N-acetylgalactosaminide alpha-2,6-sialyltransferase activity(GO:0001665) |
| 0.1 | 1.2 | GO:0052629 | phosphatidylinositol-3,5-bisphosphate 3-phosphatase activity(GO:0052629) |
| 0.1 | 1.9 | GO:0000340 | RNA 7-methylguanosine cap binding(GO:0000340) |
| 0.1 | 1.3 | GO:0004046 | aminoacylase activity(GO:0004046) |
| 0.1 | 0.7 | GO:0043023 | ribosomal large subunit binding(GO:0043023) |
| 0.1 | 1.0 | GO:0052659 | inositol-1,3,4,5-tetrakisphosphate 5-phosphatase activity(GO:0052659) |
| 0.1 | 2.7 | GO:0017110 | nucleoside-diphosphatase activity(GO:0017110) |
| 0.1 | 0.3 | GO:0031434 | mitogen-activated protein kinase kinase binding(GO:0031434) |
| 0.1 | 0.9 | GO:0004634 | phosphopyruvate hydratase activity(GO:0004634) |
| 0.1 | 0.9 | GO:0004731 | purine-nucleoside phosphorylase activity(GO:0004731) |
| 0.1 | 0.5 | GO:1990380 | Lys48-specific deubiquitinase activity(GO:1990380) |
| 0.1 | 0.5 | GO:0004104 | cholinesterase activity(GO:0004104) |
| 0.1 | 0.9 | GO:0004957 | prostaglandin E receptor activity(GO:0004957) |
| 0.1 | 1.3 | GO:0004971 | AMPA glutamate receptor activity(GO:0004971) |
| 0.1 | 1.0 | GO:0005158 | insulin receptor binding(GO:0005158) |
| 0.1 | 1.5 | GO:0004383 | guanylate cyclase activity(GO:0004383) |
| 0.1 | 0.7 | GO:0008273 | calcium, potassium:sodium antiporter activity(GO:0008273) |
| 0.1 | 0.4 | GO:0039706 | co-receptor binding(GO:0039706) |
| 0.1 | 2.8 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.1 | 0.5 | GO:0004459 | L-lactate dehydrogenase activity(GO:0004459) |
| 0.1 | 0.2 | GO:0008440 | inositol-1,4,5-trisphosphate 3-kinase activity(GO:0008440) |
| 0.1 | 6.8 | GO:0008201 | heparin binding(GO:0008201) |
| 0.1 | 0.7 | GO:0004157 | dihydropyrimidinase activity(GO:0004157) |
| 0.1 | 1.5 | GO:0031210 | phosphatidylcholine binding(GO:0031210) |
| 0.1 | 1.0 | GO:0061608 | nuclear import signal receptor activity(GO:0061608) |
| 0.1 | 0.3 | GO:0004739 | pyruvate dehydrogenase (acetyl-transferring) activity(GO:0004739) |
| 0.1 | 0.5 | GO:0047238 | glucuronosyl-N-acetylgalactosaminyl-proteoglycan 4-beta-N-acetylgalactosaminyltransferase activity(GO:0047238) |
| 0.1 | 0.7 | GO:0016413 | O-acetyltransferase activity(GO:0016413) |
| 0.1 | 1.8 | GO:0045505 | dynein intermediate chain binding(GO:0045505) |
| 0.1 | 0.7 | GO:0010851 | cyclase regulator activity(GO:0010851) |
| 0.1 | 0.3 | GO:0015288 | porin activity(GO:0015288) |
| 0.1 | 1.7 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.1 | 0.4 | GO:0031705 | bombesin receptor binding(GO:0031705) |
| 0.1 | 1.5 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.1 | 2.6 | GO:0003755 | peptidyl-prolyl cis-trans isomerase activity(GO:0003755) |
| 0.0 | 0.8 | GO:0042910 | xenobiotic transporter activity(GO:0042910) |
| 0.0 | 0.5 | GO:0052812 | phosphatidylinositol-4,5-bisphosphate 3-kinase activity(GO:0046934) phosphatidylinositol-3,4-bisphosphate 5-kinase activity(GO:0052812) |
| 0.0 | 1.1 | GO:0097153 | cysteine-type endopeptidase activity involved in apoptotic process(GO:0097153) |
| 0.0 | 0.7 | GO:0015187 | glycine transmembrane transporter activity(GO:0015187) |
| 0.0 | 0.7 | GO:2001069 | glycogen binding(GO:2001069) |
| 0.0 | 0.8 | GO:0098882 | structural constituent of presynaptic active zone(GO:0098882) |
| 0.0 | 3.6 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.0 | 1.0 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.0 | 0.6 | GO:0051117 | ATPase binding(GO:0051117) |
| 0.0 | 3.8 | GO:0003777 | microtubule motor activity(GO:0003777) |
| 0.0 | 0.3 | GO:0042834 | peptidoglycan binding(GO:0042834) |
| 0.0 | 0.2 | GO:0004999 | vasoactive intestinal polypeptide receptor activity(GO:0004999) |
| 0.0 | 0.2 | GO:0008281 | sulfonylurea receptor activity(GO:0008281) |
| 0.0 | 2.3 | GO:0019905 | syntaxin binding(GO:0019905) |
| 0.0 | 1.5 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.0 | 0.9 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.0 | 2.0 | GO:0008009 | chemokine activity(GO:0008009) |
| 0.0 | 0.4 | GO:0045503 | dynein light chain binding(GO:0045503) |
| 0.0 | 0.6 | GO:0015026 | coreceptor activity(GO:0015026) |
| 0.0 | 0.1 | GO:0004174 | electron-transferring-flavoprotein dehydrogenase activity(GO:0004174) oxidoreductase activity, acting on the CH-NH group of donors, quinone or similar compound as acceptor(GO:0016649) |
| 0.0 | 1.0 | GO:0098631 | protein binding involved in cell adhesion(GO:0098631) protein binding involved in cell-cell adhesion(GO:0098632) |
| 0.0 | 1.0 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.0 | 0.1 | GO:0005035 | tumor necrosis factor-activated receptor activity(GO:0005031) death receptor activity(GO:0005035) |
| 0.0 | 0.5 | GO:0016832 | aldehyde-lyase activity(GO:0016832) |
| 0.0 | 1.3 | GO:0008013 | beta-catenin binding(GO:0008013) |
| 0.0 | 0.4 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 0.0 | 0.7 | GO:0004550 | nucleoside diphosphate kinase activity(GO:0004550) |
| 0.0 | 0.4 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.0 | 0.6 | GO:0016628 | oxidoreductase activity, acting on the CH-CH group of donors, NAD or NADP as acceptor(GO:0016628) |
| 0.0 | 0.2 | GO:0001642 | group III metabotropic glutamate receptor activity(GO:0001642) |
| 0.0 | 1.0 | GO:0000049 | tRNA binding(GO:0000049) |
| 0.0 | 0.1 | GO:0003796 | lysozyme activity(GO:0003796) |
| 0.0 | 0.3 | GO:0070403 | NAD+ binding(GO:0070403) |
| 0.0 | 4.1 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.0 | 0.1 | GO:0004833 | tryptophan 2,3-dioxygenase activity(GO:0004833) |
| 0.0 | 0.2 | GO:0031628 | opioid receptor binding(GO:0031628) |
| 0.0 | 0.4 | GO:0035615 | clathrin adaptor activity(GO:0035615) endocytic adaptor activity(GO:0098748) |
| 0.0 | 0.2 | GO:0016671 | oxidoreductase activity, acting on a sulfur group of donors, disulfide as acceptor(GO:0016671) |
| 0.0 | 1.8 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.0 | 0.4 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.0 | 0.4 | GO:0003954 | NADH dehydrogenase activity(GO:0003954) |
| 0.0 | 33.3 | GO:0000981 | RNA polymerase II transcription factor activity, sequence-specific DNA binding(GO:0000981) |
| 0.0 | 0.8 | GO:0005544 | calcium-dependent phospholipid binding(GO:0005544) |
| 0.0 | 1.5 | GO:0052689 | carboxylic ester hydrolase activity(GO:0052689) |
| 0.0 | 5.1 | GO:0051015 | actin filament binding(GO:0051015) |
| 0.0 | 0.2 | GO:0046920 | alpha-(1->3)-fucosyltransferase activity(GO:0046920) |
| 0.0 | 0.4 | GO:0030971 | receptor tyrosine kinase binding(GO:0030971) |
| 0.0 | 0.2 | GO:0030552 | cAMP binding(GO:0030552) |
| 0.0 | 1.7 | GO:0001664 | G-protein coupled receptor binding(GO:0001664) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 6.2 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.1 | 2.2 | PID CONE PATHWAY | Visual signal transduction: Cones |
| 0.1 | 1.2 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.1 | 0.7 | PID NFKAPPAB ATYPICAL PATHWAY | Atypical NF-kappaB pathway |
| 0.1 | 2.0 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.1 | 1.1 | PID PDGFRA PATHWAY | PDGFR-alpha signaling pathway |
| 0.1 | 1.6 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 0.9 | PID GLYPICAN 1PATHWAY | Glypican 1 network |
| 0.0 | 1.6 | PID IL2 PI3K PATHWAY | IL2 signaling events mediated by PI3K |
| 0.0 | 0.4 | PID VEGFR1 PATHWAY | VEGFR1 specific signals |
| 0.0 | 0.4 | PID RHODOPSIN PATHWAY | Visual signal transduction: Rods |
| 0.0 | 1.0 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.0 | 1.4 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 0.0 | 2.9 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 0.3 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.0 | 0.4 | SIG BCR SIGNALING PATHWAY | Members of the BCR signaling pathway |
| 0.0 | 0.6 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.0 | 0.4 | PID HEDGEHOG GLI PATHWAY | Hedgehog signaling events mediated by Gli proteins |
| 0.0 | 0.7 | PID HDAC CLASSI PATHWAY | Signaling events mediated by HDAC Class I |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 6.5 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.5 | 3.5 | REACTOME ORGANIC CATION ANION ZWITTERION TRANSPORT | Genes involved in Organic cation/anion/zwitterion transport |
| 0.5 | 2.0 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 0.2 | 1.8 | REACTOME ACTIVATION OF BH3 ONLY PROTEINS | Genes involved in Activation of BH3-only proteins |
| 0.2 | 1.5 | REACTOME MTORC1 MEDIATED SIGNALLING | Genes involved in mTORC1-mediated signalling |
| 0.2 | 5.6 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.1 | 1.6 | REACTOME KERATAN SULFATE DEGRADATION | Genes involved in Keratan sulfate degradation |
| 0.1 | 1.6 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.1 | 2.1 | REACTOME AMINE COMPOUND SLC TRANSPORTERS | Genes involved in Amine compound SLC transporters |
| 0.1 | 1.0 | REACTOME TERMINATION OF O GLYCAN BIOSYNTHESIS | Genes involved in Termination of O-glycan biosynthesis |
| 0.1 | 1.2 | REACTOME CREATION OF C4 AND C2 ACTIVATORS | Genes involved in Creation of C4 and C2 activators |
| 0.1 | 3.7 | REACTOME INTERACTIONS OF VPR WITH HOST CELLULAR PROTEINS | Genes involved in Interactions of Vpr with host cellular proteins |
| 0.1 | 0.9 | REACTOME AMINE DERIVED HORMONES | Genes involved in Amine-derived hormones |
| 0.1 | 2.1 | REACTOME RNA POL I TRANSCRIPTION INITIATION | Genes involved in RNA Polymerase I Transcription Initiation |
| 0.1 | 0.7 | REACTOME CRMPS IN SEMA3A SIGNALING | Genes involved in CRMPs in Sema3A signaling |
| 0.1 | 0.4 | REACTOME PHOSPHORYLATION OF CD3 AND TCR ZETA CHAINS | Genes involved in Phosphorylation of CD3 and TCR zeta chains |
| 0.0 | 0.8 | REACTOME BILE SALT AND ORGANIC ANION SLC TRANSPORTERS | Genes involved in Bile salt and organic anion SLC transporters |
| 0.0 | 0.9 | REACTOME LATENT INFECTION OF HOMO SAPIENS WITH MYCOBACTERIUM TUBERCULOSIS | Genes involved in Latent infection of Homo sapiens with Mycobacterium tuberculosis |
| 0.0 | 1.0 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.0 | 2.1 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.0 | 0.4 | REACTOME RETROGRADE NEUROTROPHIN SIGNALLING | Genes involved in Retrograde neurotrophin signalling |
| 0.0 | 1.0 | REACTOME ANTIVIRAL MECHANISM BY IFN STIMULATED GENES | Genes involved in Antiviral mechanism by IFN-stimulated genes |
| 0.0 | 0.7 | REACTOME FORMATION OF TUBULIN FOLDING INTERMEDIATES BY CCT TRIC | Genes involved in Formation of tubulin folding intermediates by CCT/TriC |
| 0.0 | 0.5 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.0 | 0.7 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.0 | 0.9 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.0 | 0.6 | REACTOME GLUCONEOGENESIS | Genes involved in Gluconeogenesis |
| 0.0 | 1.0 | REACTOME CROSS PRESENTATION OF SOLUBLE EXOGENOUS ANTIGENS ENDOSOMES | Genes involved in Cross-presentation of soluble exogenous antigens (endosomes) |
| 0.0 | 0.6 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.0 | 0.6 | REACTOME TRANSPORT OF GLUCOSE AND OTHER SUGARS BILE SALTS AND ORGANIC ACIDS METAL IONS AND AMINE COMPOUNDS | Genes involved in Transport of glucose and other sugars, bile salts and organic acids, metal ions and amine compounds |
| 0.0 | 0.4 | REACTOME NEGATIVE REGULATORS OF RIG I MDA5 SIGNALING | Genes involved in Negative regulators of RIG-I/MDA5 signaling |
| 0.0 | 0.3 | REACTOME ACTIVATION OF NF KAPPAB IN B CELLS | Genes involved in Activation of NF-kappaB in B Cells |
| 0.0 | 0.3 | REACTOME REGULATION OF SIGNALING BY CBL | Genes involved in Regulation of signaling by CBL |
| 0.0 | 1.1 | REACTOME DOWNSTREAM SIGNAL TRANSDUCTION | Genes involved in Downstream signal transduction |
| 0.0 | 0.2 | REACTOME TRAFFICKING AND PROCESSING OF ENDOSOMAL TLR | Genes involved in Trafficking and processing of endosomal TLR |
| 0.0 | 0.1 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.0 | 0.4 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |