Project
PRJNA195909:zebrafish embryo and larva development
Navigation
Downloads
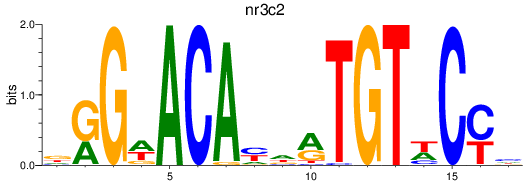
Results for nr3c2
Z-value: 0.73
Transcription factors associated with nr3c2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
nr3c2
|
ENSDARG00000102082 | nuclear receptor subfamily 3, group C, member 2 |
|
nr3c2
|
ENSDARG00000115513 | nuclear receptor subfamily 3, group C, member 2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| nr3c2 | dr11_v1_chr1_-_37087966_37087966 | -0.42 | 2.7e-01 | Click! |
Activity profile of nr3c2 motif
Sorted Z-values of nr3c2 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr20_+_35445462 | 1.53 |
ENSDART00000124497
|
tdrd6
|
tudor domain containing 6 |
| chr11_-_43473824 | 1.53 |
ENSDART00000179561
|
tmem63bb
|
transmembrane protein 63Bb |
| chr6_+_4255319 | 1.42 |
ENSDART00000170351
|
nbeal1
|
neurobeachin-like 1 |
| chr18_-_20608025 | 1.41 |
ENSDART00000090156
ENSDART00000151980 |
bcl2l13
|
BCL2 like 13 |
| chr10_+_33754967 | 1.39 |
ENSDART00000153442
|
rxfp2a
|
relaxin/insulin-like family peptide receptor 2a |
| chr14_-_46113321 | 1.28 |
ENSDART00000169040
ENSDART00000161475 ENSDART00000124925 |
si:ch211-235e9.8
|
si:ch211-235e9.8 |
| chr3_-_40051425 | 1.26 |
ENSDART00000146700
|
llgl1
|
lethal giant larvae homolog 1 (Drosophila) |
| chr17_-_6613458 | 1.16 |
ENSDART00000175024
|
si:ch211-189e2.3
|
si:ch211-189e2.3 |
| chr4_+_11723852 | 1.14 |
ENSDART00000028820
|
mkln1
|
muskelin 1, intracellular mediator containing kelch motifs |
| chr18_-_20608300 | 1.07 |
ENSDART00000140632
|
bcl2l13
|
BCL2 like 13 |
| chr11_+_34522554 | 1.03 |
ENSDART00000109833
|
zmat3
|
zinc finger, matrin-type 3 |
| chr24_-_23784701 | 0.95 |
ENSDART00000090368
|
sgk3
|
serum/glucocorticoid regulated kinase family, member 3 |
| chr24_-_20641000 | 0.87 |
ENSDART00000166135
|
zbtb47b
|
zinc finger and BTB domain containing 47b |
| chr22_-_34979139 | 0.86 |
ENSDART00000116455
ENSDART00000133537 |
arhgap19
|
Rho GTPase activating protein 19 |
| chr11_+_5880562 | 0.85 |
ENSDART00000129663
ENSDART00000130768 ENSDART00000160909 |
dazap1
|
DAZ associated protein 1 |
| chr24_-_38097305 | 0.82 |
ENSDART00000124321
|
crp2
|
C-reactive protein 2 |
| chr22_-_16275236 | 0.76 |
ENSDART00000149051
|
cdc14ab
|
cell division cycle 14Ab |
| chr8_+_23034718 | 0.75 |
ENSDART00000184512
|
ythdf1
|
YTH N(6)-methyladenosine RNA binding protein 1 |
| chr7_-_51749683 | 0.68 |
ENSDART00000083190
|
hdac8
|
histone deacetylase 8 |
| chr13_-_33700461 | 0.66 |
ENSDART00000160520
|
mad2l1bp
|
MAD2L1 binding protein |
| chr11_+_34523132 | 0.66 |
ENSDART00000192257
|
zmat3
|
zinc finger, matrin-type 3 |
| chr5_-_9540641 | 0.63 |
ENSDART00000124384
ENSDART00000160079 |
gak
|
cyclin G associated kinase |
| chr20_+_54336137 | 0.59 |
ENSDART00000113792
|
cipcb
|
CLOCK-interacting pacemaker b |
| chr15_+_23951560 | 0.49 |
ENSDART00000191133
|
myo18ab
|
myosin XVIIIAb |
| chr1_+_54199406 | 0.48 |
ENSDART00000176578
|
tsc2
|
TSC complex subunit 2 |
| chr16_+_11188810 | 0.47 |
ENSDART00000186011
|
cicb
|
capicua transcriptional repressor b |
| chr15_+_40188076 | 0.47 |
ENSDART00000063779
|
efhd1
|
EF-hand domain family, member D1 |
| chr21_+_6291027 | 0.46 |
ENSDART00000180467
ENSDART00000184952 ENSDART00000184006 |
fnbp1b
|
formin binding protein 1b |
| chr25_-_13408760 | 0.46 |
ENSDART00000154445
|
gins3
|
GINS complex subunit 3 |
| chr9_+_22388686 | 0.42 |
ENSDART00000182731
ENSDART00000181462 |
dgkg
|
diacylglycerol kinase, gamma |
| chr5_+_30518441 | 0.39 |
ENSDART00000132664
|
hmbsa
|
hydroxymethylbilane synthase a |
| chr8_+_39767915 | 0.39 |
ENSDART00000017153
|
hps4
|
Hermansky-Pudlak syndrome 4 |
| chr2_-_32512648 | 0.35 |
ENSDART00000170674
|
abcf2a
|
ATP-binding cassette, sub-family F (GCN20), member 2a |
| chr2_+_29257942 | 0.32 |
ENSDART00000184362
ENSDART00000025562 |
cdh18a
|
cadherin 18, type 2a |
| chr22_-_36519590 | 0.30 |
ENSDART00000129318
|
CABZ01045212.1
|
|
| chr21_+_6290566 | 0.28 |
ENSDART00000161647
|
fnbp1b
|
formin binding protein 1b |
| chr10_+_8554929 | 0.25 |
ENSDART00000190849
|
tbc1d10ab
|
TBC1 domain family, member 10Ab |
| chr10_-_24689725 | 0.24 |
ENSDART00000079566
|
si:ch211-287a12.9
|
si:ch211-287a12.9 |
| chr8_-_45834825 | 0.22 |
ENSDART00000132965
|
ogdha
|
oxoglutarate (alpha-ketoglutarate) dehydrogenase a (lipoamide) |
| chr20_-_39391833 | 0.21 |
ENSDART00000135149
|
si:dkey-217m5.8
|
si:dkey-217m5.8 |
| chr10_+_20364009 | 0.18 |
ENSDART00000186139
ENSDART00000080395 |
golga7
|
golgin A7 |
| chr8_-_45835056 | 0.18 |
ENSDART00000022242
|
ogdha
|
oxoglutarate (alpha-ketoglutarate) dehydrogenase a (lipoamide) |
| chr5_-_54792239 | 0.15 |
ENSDART00000056213
|
pik3r1
|
phosphoinositide-3-kinase, regulatory subunit 1 (alpha) |
| chr1_-_28473350 | 0.11 |
ENSDART00000190608
ENSDART00000148175 |
si:ch1073-440b2.1
|
si:ch1073-440b2.1 |
| chr10_+_29849497 | 0.08 |
ENSDART00000099994
ENSDART00000132212 |
hspa8
|
heat shock protein 8 |
| chr3_-_37476475 | 0.07 |
ENSDART00000148107
|
si:ch211-278a6.1
|
si:ch211-278a6.1 |
| chr16_-_11932923 | 0.04 |
ENSDART00000103975
|
cd4-1
|
CD4-1 molecule |
| chr25_+_10416583 | 0.04 |
ENSDART00000073907
|
ehf
|
ets homologous factor |
| chr19_+_7916790 | 0.01 |
ENSDART00000081584
|
si:dkey-266f7.1
|
si:dkey-266f7.1 |
| chr5_+_9382301 | 0.01 |
ENSDART00000124017
|
ugt2a7
|
UDP glucuronosyltransferase 2 family, polypeptide A7 |
| chr15_-_18429550 | 0.01 |
ENSDART00000136208
|
ncam1b
|
neural cell adhesion molecule 1b |
Network of associatons between targets according to the STRING database.
First level regulatory network of nr3c2
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | GO:0042754 | negative regulation of circadian rhythm(GO:0042754) |
| 0.1 | 1.5 | GO:0030719 | P granule organization(GO:0030719) |
| 0.1 | 1.4 | GO:0007190 | activation of adenylate cyclase activity(GO:0007190) positive regulation of cAMP metabolic process(GO:0030816) positive regulation of cAMP biosynthetic process(GO:0030819) positive regulation of adenylate cyclase activity(GO:0045762) |
| 0.1 | 0.6 | GO:0072318 | clathrin coat disassembly(GO:0072318) vesicle uncoating(GO:0072319) |
| 0.1 | 0.4 | GO:1903232 | melanosome assembly(GO:1903232) |
| 0.1 | 1.3 | GO:0032878 | regulation of establishment or maintenance of cell polarity(GO:0032878) |
| 0.1 | 0.7 | GO:0070933 | histone H4 deacetylation(GO:0070933) |
| 0.1 | 1.4 | GO:0007096 | regulation of exit from mitosis(GO:0007096) |
| 0.1 | 1.3 | GO:0008089 | anterograde axonal transport(GO:0008089) |
| 0.1 | 0.5 | GO:1902975 | cell cycle DNA replication initiation(GO:1902292) nuclear cell cycle DNA replication initiation(GO:1902315) mitotic DNA replication initiation(GO:1902975) |
| 0.1 | 0.4 | GO:0018160 | peptidyl-pyrromethane cofactor linkage(GO:0018160) |
| 0.1 | 0.7 | GO:0061157 | RNA destabilization(GO:0050779) mRNA destabilization(GO:0061157) |
| 0.0 | 0.5 | GO:0060575 | intestinal epithelial cell differentiation(GO:0060575) |
| 0.0 | 0.8 | GO:0048026 | positive regulation of mRNA splicing, via spliceosome(GO:0048026) |
| 0.0 | 0.4 | GO:0007205 | protein kinase C-activating G-protein coupled receptor signaling pathway(GO:0007205) |
| 0.0 | 0.1 | GO:1902946 | positive regulation of receptor-mediated endocytosis(GO:0048260) protein localization to early endosome(GO:1902946) |
| 0.0 | 0.5 | GO:0014866 | skeletal myofibril assembly(GO:0014866) |
| 0.0 | 0.5 | GO:0060319 | primitive erythrocyte differentiation(GO:0060319) |
| 0.0 | 0.2 | GO:0043550 | regulation of lipid kinase activity(GO:0043550) regulation of phosphatidylinositol 3-kinase activity(GO:0043551) |
| 0.0 | 0.4 | GO:0006099 | tricarboxylic acid cycle(GO:0006099) |
| 0.0 | 0.2 | GO:0043001 | Golgi to plasma membrane protein transport(GO:0043001) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.5 | GO:0033391 | chromatoid body(GO:0033391) |
| 0.1 | 0.8 | GO:1902636 | kinociliary basal body(GO:1902636) |
| 0.1 | 0.4 | GO:0031085 | BLOC-3 complex(GO:0031085) |
| 0.1 | 0.5 | GO:0033596 | TSC1-TSC2 complex(GO:0033596) |
| 0.1 | 0.5 | GO:0000811 | GINS complex(GO:0000811) |
| 0.0 | 0.4 | GO:0045252 | oxoglutarate dehydrogenase complex(GO:0045252) |
| 0.0 | 1.3 | GO:0030864 | cortical actin cytoskeleton(GO:0030864) |
| 0.0 | 0.2 | GO:0002178 | palmitoyltransferase complex(GO:0002178) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.8 | GO:0034046 | poly(G) binding(GO:0034046) |
| 0.1 | 0.7 | GO:1990247 | N6-methyladenosine-containing RNA binding(GO:1990247) |
| 0.1 | 1.3 | GO:0045159 | myosin II binding(GO:0045159) |
| 0.1 | 0.4 | GO:0004418 | hydroxymethylbilane synthase activity(GO:0004418) |
| 0.0 | 1.5 | GO:0005227 | calcium activated cation channel activity(GO:0005227) |
| 0.0 | 0.4 | GO:0004591 | oxoglutarate dehydrogenase (succinyl-transferring) activity(GO:0004591) |
| 0.0 | 0.7 | GO:0032041 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.0 | 1.0 | GO:0015459 | potassium channel regulator activity(GO:0015459) |
| 0.0 | 0.4 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.0 | 0.8 | GO:0015485 | cholesterol binding(GO:0015485) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |
| 0.0 | 0.4 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.0 | 0.7 | REACTOME NOTCH1 INTRACELLULAR DOMAIN REGULATES TRANSCRIPTION | Genes involved in NOTCH1 Intracellular Domain Regulates Transcription |
| 0.0 | 0.7 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.0 | 0.2 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |