Project
PRJNA195909:zebrafish embryo and larva development
Navigation
Downloads
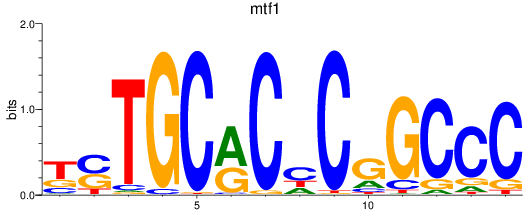
Results for mtf1
Z-value: 1.19
Transcription factors associated with mtf1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
mtf1
|
ENSDARG00000102898 | metal-regulatory transcription factor 1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| mtf1 | dr11_v1_chr16_+_4133519_4133519 | -0.90 | 1.1e-03 | Click! |
Activity profile of mtf1 motif
Sorted Z-values of mtf1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr11_-_497854 | 4.71 |
ENSDART00000104520
|
cnbpb
|
CCHC-type zinc finger, nucleic acid binding protein b |
| chr11_-_497680 | 3.77 |
ENSDART00000154888
|
cnbpb
|
CCHC-type zinc finger, nucleic acid binding protein b |
| chr2_-_7666021 | 3.27 |
ENSDART00000180007
|
CABZ01021592.1
|
|
| chr7_-_74090168 | 2.60 |
ENSDART00000050528
|
tyrp1a
|
tyrosinase-related protein 1a |
| chr17_-_125091 | 2.53 |
ENSDART00000158825
|
actc1b
|
actin, alpha, cardiac muscle 1b |
| chr8_+_31435452 | 2.51 |
ENSDART00000145282
|
selenop
|
selenoprotein P |
| chr7_+_49715750 | 2.50 |
ENSDART00000019446
|
ascl1b
|
achaete-scute family bHLH transcription factor 1b |
| chr2_+_33457310 | 2.18 |
ENSDART00000056657
|
zgc:113531
|
zgc:113531 |
| chr3_+_23221047 | 2.10 |
ENSDART00000009393
|
col1a1a
|
collagen, type I, alpha 1a |
| chr25_-_23526058 | 2.00 |
ENSDART00000191331
ENSDART00000062930 |
phlda2
|
pleckstrin homology-like domain, family A, member 2 |
| chr23_-_20309505 | 1.90 |
ENSDART00000130856
|
lamb2l
|
laminin, beta 2-like |
| chr1_-_35928942 | 1.89 |
ENSDART00000033566
|
smad1
|
SMAD family member 1 |
| chr1_+_44906517 | 1.79 |
ENSDART00000142820
|
wu:fc21g02
|
wu:fc21g02 |
| chr22_-_37349967 | 1.75 |
ENSDART00000104493
|
sox2
|
SRY (sex determining region Y)-box 2 |
| chr16_+_26449615 | 1.62 |
ENSDART00000039746
|
epb41b
|
erythrocyte membrane protein band 4.1b |
| chr8_+_1065458 | 1.59 |
ENSDART00000081432
|
sprb
|
sepiapterin reductase b |
| chr20_-_54259780 | 1.58 |
ENSDART00000172631
|
fkbp3
|
FK506 binding protein 3 |
| chr1_-_35929143 | 1.52 |
ENSDART00000185002
|
smad1
|
SMAD family member 1 |
| chr10_+_21789954 | 1.50 |
ENSDART00000157769
ENSDART00000171703 |
pcdh1gc5
|
protocadherin 1 gamma c 5 |
| chr1_-_59251874 | 1.48 |
ENSDART00000168919
|
olfm2b
|
olfactomedin 2b |
| chr7_+_25323742 | 1.46 |
ENSDART00000110347
|
cyp26b1
|
cytochrome P450, family 26, subfamily b, polypeptide 1 |
| chr1_-_12278056 | 1.43 |
ENSDART00000139336
ENSDART00000137463 |
cplx2l
|
complexin 2, like |
| chr1_-_12278522 | 1.42 |
ENSDART00000142122
ENSDART00000003825 |
cplx2l
|
complexin 2, like |
| chr14_+_5117072 | 1.37 |
ENSDART00000189628
|
nanos1
|
nanos homolog 1 |
| chr20_-_147574 | 1.32 |
ENSDART00000104762
ENSDART00000131635 |
slc16a10
|
solute carrier family 16 (aromatic amino acid transporter), member 10 |
| chr19_+_10832092 | 1.23 |
ENSDART00000191851
|
tomm40l
|
translocase of outer mitochondrial membrane 40 homolog, like |
| chr23_+_23485858 | 1.11 |
ENSDART00000114067
|
agrn
|
agrin |
| chr8_+_22355909 | 1.10 |
ENSDART00000146457
ENSDART00000142883 |
zgc:153631
|
zgc:153631 |
| chr6_+_42918933 | 1.04 |
ENSDART00000064896
|
gnat1
|
guanine nucleotide binding protein (G protein), alpha transducing activity polypeptide 1 |
| chr11_-_3334248 | 1.04 |
ENSDART00000154314
ENSDART00000121861 |
prph
|
peripherin |
| chr23_-_28239750 | 0.98 |
ENSDART00000003548
|
znf385a
|
zinc finger protein 385A |
| chr23_+_36178104 | 0.98 |
ENSDART00000103131
|
hoxc1a
|
homeobox C1a |
| chr12_-_45876387 | 0.97 |
ENSDART00000043210
ENSDART00000149044 |
pax2b
|
paired box 2b |
| chr23_-_45705525 | 0.93 |
ENSDART00000148959
|
ednrab
|
endothelin receptor type Ab |
| chr1_+_50085440 | 0.89 |
ENSDART00000018469
ENSDART00000134988 |
npnt
|
nephronectin |
| chr4_+_38981587 | 0.88 |
ENSDART00000142713
|
si:dkey-66k12.3
|
si:dkey-66k12.3 |
| chr17_-_36988455 | 0.87 |
ENSDART00000187180
ENSDART00000126823 |
dnmt3ab
|
DNA (cytosine-5-)-methyltransferase 3 alpha b |
| chr24_-_38384432 | 0.85 |
ENSDART00000140739
|
lrrc4bb
|
leucine rich repeat containing 4Bb |
| chr14_+_597532 | 0.84 |
ENSDART00000159805
|
LO018568.2
|
|
| chr24_-_37877743 | 0.83 |
ENSDART00000105658
|
tmem204
|
transmembrane protein 204 |
| chr19_+_37701450 | 0.82 |
ENSDART00000087694
|
thsd7aa
|
thrombospondin, type I, domain containing 7Aa |
| chr1_+_50085640 | 0.78 |
ENSDART00000158988
ENSDART00000165973 |
npnt
|
nephronectin |
| chr20_-_46362606 | 0.78 |
ENSDART00000153087
|
bmf2
|
BCL2 modifying factor 2 |
| chr17_-_12385308 | 0.74 |
ENSDART00000080927
|
snap25b
|
synaptosomal-associated protein, 25b |
| chr7_-_38861741 | 0.72 |
ENSDART00000173629
ENSDART00000037361 ENSDART00000173953 |
phf21aa
|
PHD finger protein 21Aa |
| chr24_-_4450238 | 0.68 |
ENSDART00000066835
|
fzd8a
|
frizzled class receptor 8a |
| chr23_+_20931030 | 0.67 |
ENSDART00000167014
|
pax7b
|
paired box 7b |
| chr19_+_31183495 | 0.67 |
ENSDART00000088618
|
meox2b
|
mesenchyme homeobox 2b |
| chr8_-_39977026 | 0.65 |
ENSDART00000141707
|
asphd2
|
aspartate beta-hydroxylase domain containing 2 |
| chr2_+_54641644 | 0.65 |
ENSDART00000027313
|
ndufv2
|
NADH dehydrogenase (ubiquinone) flavoprotein 2 |
| chr1_+_9004719 | 0.63 |
ENSDART00000006211
ENSDART00000137211 |
prkcba
|
protein kinase C, beta a |
| chr4_+_57099307 | 0.63 |
ENSDART00000131654
|
si:ch211-238e22.2
|
si:ch211-238e22.2 |
| chr24_+_10397865 | 0.61 |
ENSDART00000155557
|
si:ch211-69l10.4
|
si:ch211-69l10.4 |
| chr6_-_20875111 | 0.60 |
ENSDART00000115118
ENSDART00000159916 |
tns1a
|
tensin 1a |
| chr14_-_45394704 | 0.60 |
ENSDART00000173078
ENSDART00000125970 |
si:ch211-168f7.5
|
si:ch211-168f7.5 |
| chr10_+_22724059 | 0.59 |
ENSDART00000136123
|
kdm6bb
|
lysine (K)-specific demethylase 6B, b |
| chr19_-_33212023 | 0.55 |
ENSDART00000189209
|
trib1
|
tribbles pseudokinase 1 |
| chr24_+_28525507 | 0.55 |
ENSDART00000191121
|
arhgap29a
|
Rho GTPase activating protein 29a |
| chr13_+_52006253 | 0.55 |
ENSDART00000181259
|
CABZ01080135.1
|
|
| chr12_-_45875946 | 0.54 |
ENSDART00000149970
|
pax2b
|
paired box 2b |
| chr11_+_575665 | 0.53 |
ENSDART00000122133
|
mkrn2os.1
|
MKRN2 opposite strand, tandem duplicate 1 |
| chr2_-_51095743 | 0.53 |
ENSDART00000184646
|
THEM6
|
si:ch73-52e5.2 |
| chr9_+_156142 | 0.52 |
ENSDART00000171501
|
cldn10b
|
claudin 10b |
| chr8_+_8973425 | 0.52 |
ENSDART00000066107
|
bcap31
|
B cell receptor associated protein 31 |
| chr4_-_28334464 | 0.51 |
ENSDART00000123617
|
celsr1a
|
cadherin EGF LAG seven-pass G-type receptor 1a |
| chr15_-_30816370 | 0.51 |
ENSDART00000142982
ENSDART00000050649 ENSDART00000136901 ENSDART00000100194 |
msi2b
|
musashi RNA-binding protein 2b |
| chr10_-_26997344 | 0.51 |
ENSDART00000146595
|
zgc:193742
|
zgc:193742 |
| chr8_+_38417461 | 0.50 |
ENSDART00000132718
|
nkx6.3
|
NK6 homeobox 3 |
| chr4_-_28335184 | 0.47 |
ENSDART00000178149
ENSDART00000043737 |
celsr1a
|
cadherin EGF LAG seven-pass G-type receptor 1a |
| chr3_+_30500968 | 0.47 |
ENSDART00000103447
|
si:dkey-13n23.3
|
si:dkey-13n23.3 |
| chr23_+_45611649 | 0.45 |
ENSDART00000169521
|
dclk2b
|
doublecortin-like kinase 2b |
| chr7_+_59169081 | 0.45 |
ENSDART00000167980
|
ostc
|
oligosaccharyltransferase complex subunit |
| chr12_+_20149707 | 0.43 |
ENSDART00000181942
|
foxj1b
|
forkhead box J1b |
| chr20_-_1314537 | 0.42 |
ENSDART00000144288
|
pcmt
|
protein-L-isoaspartate (D-aspartate) O-methyltransferase |
| chr23_-_45682136 | 0.41 |
ENSDART00000164646
|
fam160a1b
|
family with sequence similarity 160, member A1b |
| chr23_-_3681026 | 0.40 |
ENSDART00000192128
ENSDART00000040086 |
pacsin1a
|
protein kinase C and casein kinase substrate in neurons 1a |
| chr3_+_30501135 | 0.40 |
ENSDART00000165869
|
si:dkey-13n23.3
|
si:dkey-13n23.3 |
| chr23_+_45611980 | 0.38 |
ENSDART00000181582
|
dclk2b
|
doublecortin-like kinase 2b |
| chr16_-_29452039 | 0.38 |
ENSDART00000148960
|
si:ch211-113g11.6
|
si:ch211-113g11.6 |
| chr4_-_68564988 | 0.37 |
ENSDART00000191212
|
BX548011.3
|
|
| chr21_+_21621042 | 0.37 |
ENSDART00000134907
|
tgfb1b
|
transforming growth factor, beta 1b |
| chr16_+_11029762 | 0.37 |
ENSDART00000091183
|
erfl3
|
Ets2 repressor factor like 3 |
| chr12_+_22745618 | 0.36 |
ENSDART00000181024
ENSDART00000183993 ENSDART00000077548 |
ablim2
|
actin binding LIM protein family, member 2 |
| chr3_-_18756076 | 0.36 |
ENSDART00000055766
|
zgc:113333
|
zgc:113333 |
| chr20_-_1314355 | 0.33 |
ENSDART00000152436
|
pcmt
|
protein-L-isoaspartate (D-aspartate) O-methyltransferase |
| chr16_-_7692568 | 0.32 |
ENSDART00000148980
|
pimr209
|
Pim proto-oncogene, serine/threonine kinase, related 209 |
| chr20_-_5267600 | 0.32 |
ENSDART00000099258
|
cyp46a1.4
|
cytochrome P450, family 46, subfamily A, polypeptide 1, tandem duplicate 4 |
| chr17_+_25481466 | 0.31 |
ENSDART00000139451
|
aim1a
|
crystallin beta-gamma domain containing 1a |
| chr24_-_37877554 | 0.30 |
ENSDART00000132219
|
tmem204
|
transmembrane protein 204 |
| chr6_+_612330 | 0.30 |
ENSDART00000166872
ENSDART00000191758 |
kynu
|
kynureninase |
| chr25_-_36020344 | 0.27 |
ENSDART00000181448
|
rbl2
|
retinoblastoma-like 2 (p130) |
| chr15_+_3219134 | 0.26 |
ENSDART00000113532
|
LO017656.1
|
|
| chr15_-_18429550 | 0.25 |
ENSDART00000136208
|
ncam1b
|
neural cell adhesion molecule 1b |
| chr14_-_3070613 | 0.25 |
ENSDART00000193729
ENSDART00000090196 |
slc35a4
|
solute carrier family 35, member A4 |
| chr12_-_44070043 | 0.24 |
ENSDART00000180059
|
si:ch73-329n5.3
|
si:ch73-329n5.3 |
| chr17_+_48831579 | 0.24 |
ENSDART00000154824
ENSDART00000178697 ENSDART00000192703 ENSDART00000180378 |
daam2
|
dishevelled associated activator of morphogenesis 2 |
| chr3_-_50124413 | 0.23 |
ENSDART00000189920
|
cldnk
|
claudin k |
| chr17_+_51499789 | 0.22 |
ENSDART00000187701
|
CABZ01067581.1
|
|
| chr9_-_128036 | 0.20 |
ENSDART00000165960
|
FQ377903.2
|
|
| chr24_-_3477103 | 0.19 |
ENSDART00000143723
|
idi1
|
isopentenyl-diphosphate delta isomerase 1 |
| chr3_+_23029934 | 0.18 |
ENSDART00000110343
|
nags
|
N-acetylglutamate synthase |
| chr10_+_24445698 | 0.18 |
ENSDART00000146370
|
slc7a1
|
solute carrier family 7 (cationic amino acid transporter, y+ system), member 1 |
| chr4_+_53268458 | 0.17 |
ENSDART00000165335
|
si:dkey-250k10.1
|
si:dkey-250k10.1 |
| chr22_-_38607504 | 0.16 |
ENSDART00000164609
|
si:ch211-126j24.1
|
si:ch211-126j24.1 |
| chr6_+_31684 | 0.15 |
ENSDART00000188853
ENSDART00000184553 |
CZQB01141835.2
|
|
| chr4_-_49582108 | 0.15 |
ENSDART00000154999
|
si:dkey-159n16.2
|
si:dkey-159n16.2 |
| chr12_+_41991635 | 0.15 |
ENSDART00000186161
ENSDART00000192510 |
TCERG1L
|
transcription elongation regulator 1 like |
| chr5_-_24270989 | 0.14 |
ENSDART00000146251
|
si:ch211-137i24.12
|
si:ch211-137i24.12 |
| chr22_-_30678518 | 0.13 |
ENSDART00000138282
|
si:dkey-42l16.1
|
si:dkey-42l16.1 |
| chr7_+_21275152 | 0.13 |
ENSDART00000173612
|
serpinh2
|
serine (or cysteine) peptidase inhibitor, clade H, member 2 |
| chr12_-_48312647 | 0.12 |
ENSDART00000114415
|
ascc1
|
activating signal cointegrator 1 complex subunit 1 |
| chr19_-_11336782 | 0.11 |
ENSDART00000131014
|
sept7a
|
septin 7a |
| chr2_+_41876595 | 0.11 |
ENSDART00000112243
|
crlf1a
|
cytokine receptor-like factor 1a |
| chr12_+_48851381 | 0.11 |
ENSDART00000187241
ENSDART00000187796 |
dlg5b.1
|
discs, large homolog 5b (Drosophila), tandem duplicate 1 |
| chr1_-_16394814 | 0.11 |
ENSDART00000013024
|
fgf20a
|
fibroblast growth factor 20a |
| chr9_-_48397702 | 0.10 |
ENSDART00000147169
|
zgc:172182
|
zgc:172182 |
| chr4_-_72513569 | 0.10 |
ENSDART00000174130
|
CR788316.4
|
|
| chr19_-_35400819 | 0.10 |
ENSDART00000148080
|
rnf19b
|
ring finger protein 19B |
| chr18_-_50947868 | 0.09 |
ENSDART00000174276
|
st7
|
suppression of tumorigenicity 7 |
| chr5_-_14211487 | 0.08 |
ENSDART00000135731
|
npffr1l2
|
neuropeptide FF receptor 1 like 2 |
| chr7_+_74150839 | 0.08 |
ENSDART00000160195
|
ppp1cbl
|
protein phosphatase 1, catalytic subunit, beta isoform, like |
| chr10_+_4807913 | 0.06 |
ENSDART00000186099
|
palm2
|
paralemmin 2 |
| chr2_+_33796207 | 0.05 |
ENSDART00000124647
|
kiss1ra
|
KISS1 receptor a |
| chr3_+_53052769 | 0.05 |
ENSDART00000156935
|
pbx4
|
pre-B-cell leukemia transcription factor 4 |
| chr4_+_58667348 | 0.04 |
ENSDART00000186596
ENSDART00000180673 |
CR388148.2
|
|
| chr15_-_37719679 | 0.04 |
ENSDART00000184025
|
si:dkey-117a8.1
|
si:dkey-117a8.1 |
| chr6_+_10450000 | 0.03 |
ENSDART00000151288
ENSDART00000187431 ENSDART00000192474 ENSDART00000188214 ENSDART00000184766 ENSDART00000190082 |
kcnh7
|
potassium channel, voltage gated eag related subfamily H, member 7 |
| chr15_+_2559875 | 0.03 |
ENSDART00000178505
|
SH2B2
|
SH2B adaptor protein 2 |
| chr20_+_13894123 | 0.01 |
ENSDART00000007744
|
slc30a1a
|
solute carrier family 30 (zinc transporter), member 1a |
| chr6_+_7553085 | 0.01 |
ENSDART00000150939
ENSDART00000151114 |
myh10
|
myosin, heavy chain 10, non-muscle |
| chr22_+_569565 | 0.01 |
ENSDART00000037069
|
usp49
|
ubiquitin specific peptidase 49 |
| chr8_+_26059677 | 0.00 |
ENSDART00000009178
|
impdh2
|
IMP (inosine 5'-monophosphate) dehydrogenase 2 |
| chr12_-_25294096 | 0.00 |
ENSDART00000183398
|
hcar1-4
|
hydroxycarboxylic acid receptor 1-4 |
Network of associatons between targets according to the STRING database.
First level regulatory network of mtf1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.7 | 8.5 | GO:2000767 | positive regulation of cytoplasmic translation(GO:2000767) |
| 0.5 | 2.1 | GO:0035630 | bone mineralization involved in bone maturation(GO:0035630) |
| 0.5 | 1.5 | GO:0035992 | tendon formation(GO:0035992) |
| 0.4 | 2.5 | GO:0001887 | selenium compound metabolic process(GO:0001887) |
| 0.4 | 1.1 | GO:1902571 | regulation of serine-type peptidase activity(GO:1902571) |
| 0.3 | 2.6 | GO:0042438 | melanin biosynthetic process(GO:0042438) |
| 0.3 | 1.8 | GO:0021982 | pineal gland development(GO:0021982) |
| 0.2 | 0.7 | GO:0021531 | spinal cord radial glial cell differentiation(GO:0021531) eye field cell fate commitment involved in camera-type eye formation(GO:0060898) |
| 0.2 | 1.6 | GO:0046146 | tetrahydrobiopterin biosynthetic process(GO:0006729) tetrahydrobiopterin metabolic process(GO:0046146) |
| 0.2 | 0.9 | GO:0003319 | cardioblast migration to the midline involved in heart rudiment formation(GO:0003319) |
| 0.1 | 0.7 | GO:0050938 | regulation of xanthophore differentiation(GO:0050938) |
| 0.1 | 0.5 | GO:0070973 | protein localization to endoplasmic reticulum exit site(GO:0070973) |
| 0.1 | 0.6 | GO:1900108 | negative regulation of nodal signaling pathway(GO:1900108) |
| 0.1 | 1.0 | GO:0035588 | adenosine receptor signaling pathway(GO:0001973) purinergic receptor signaling pathway(GO:0035587) G-protein coupled purinergic receptor signaling pathway(GO:0035588) |
| 0.1 | 1.1 | GO:2000406 | positive regulation of lymphocyte migration(GO:2000403) positive regulation of T cell migration(GO:2000406) |
| 0.1 | 0.6 | GO:0071557 | histone H3-K27 demethylation(GO:0071557) |
| 0.1 | 0.2 | GO:0014014 | negative regulation of gliogenesis(GO:0014014) |
| 0.1 | 0.2 | GO:0097052 | L-kynurenine metabolic process(GO:0097052) L-kynurenine catabolic process(GO:0097053) |
| 0.1 | 1.4 | GO:0008354 | germ cell migration(GO:0008354) |
| 0.1 | 1.2 | GO:0030150 | protein import into mitochondrial matrix(GO:0030150) |
| 0.1 | 1.9 | GO:0070831 | basement membrane assembly(GO:0070831) |
| 0.1 | 3.8 | GO:0060395 | SMAD protein signal transduction(GO:0060395) |
| 0.1 | 2.9 | GO:0046928 | regulation of neurotransmitter secretion(GO:0046928) |
| 0.1 | 0.7 | GO:0045162 | clustering of voltage-gated sodium channels(GO:0045162) |
| 0.0 | 0.2 | GO:0006526 | arginine biosynthetic process(GO:0006526) |
| 0.0 | 2.2 | GO:0030514 | negative regulation of BMP signaling pathway(GO:0030514) |
| 0.0 | 2.5 | GO:0021575 | hindbrain morphogenesis(GO:0021575) |
| 0.0 | 0.5 | GO:0021772 | olfactory bulb development(GO:0021772) olfactory lobe development(GO:0021988) |
| 0.0 | 0.2 | GO:0009240 | isopentenyl diphosphate biosynthetic process(GO:0009240) isopentenyl diphosphate metabolic process(GO:0046490) |
| 0.0 | 0.7 | GO:0006120 | mitochondrial electron transport, NADH to ubiquinone(GO:0006120) |
| 0.0 | 0.8 | GO:0010507 | negative regulation of autophagy(GO:0010507) |
| 0.0 | 0.3 | GO:0006707 | cholesterol catabolic process(GO:0006707) sterol catabolic process(GO:0016127) |
| 0.0 | 0.9 | GO:0006305 | DNA alkylation(GO:0006305) DNA methylation(GO:0006306) |
| 0.0 | 0.4 | GO:0060088 | auditory receptor cell stereocilium organization(GO:0060088) |
| 0.0 | 1.6 | GO:0030866 | cortical actin cytoskeleton organization(GO:0030866) |
| 0.0 | 0.8 | GO:0072015 | glomerular visceral epithelial cell development(GO:0072015) glomerular epithelial cell development(GO:0072310) |
| 0.0 | 1.0 | GO:0045104 | intermediate filament-based process(GO:0045103) intermediate filament cytoskeleton organization(GO:0045104) |
| 0.0 | 0.4 | GO:0097320 | membrane tubulation(GO:0097320) |
| 0.0 | 0.3 | GO:2000134 | negative regulation of cell cycle G1/S phase transition(GO:1902807) negative regulation of G1/S transition of mitotic cell cycle(GO:2000134) |
| 0.0 | 0.3 | GO:0015781 | pyrimidine nucleotide-sugar transport(GO:0015781) pyrimidine nucleotide-sugar transmembrane transport(GO:0090481) |
| 0.0 | 0.4 | GO:0030032 | lamellipodium assembly(GO:0030032) |
| 0.0 | 2.0 | GO:0043068 | positive regulation of cell death(GO:0010942) positive regulation of apoptotic process(GO:0043065) positive regulation of programmed cell death(GO:0043068) |
| 0.0 | 0.1 | GO:0046850 | regulation of bone remodeling(GO:0046850) |
| 0.0 | 0.7 | GO:0032436 | positive regulation of proteasomal ubiquitin-dependent protein catabolic process(GO:0032436) positive regulation of proteasomal protein catabolic process(GO:1901800) |
| 0.0 | 1.5 | GO:0007422 | peripheral nervous system development(GO:0007422) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 2.6 | GO:0045009 | melanosome membrane(GO:0033162) chitosome(GO:0045009) |
| 0.2 | 3.4 | GO:0071144 | SMAD2-SMAD3 protein complex(GO:0071144) |
| 0.2 | 2.5 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.1 | 0.7 | GO:0070032 | synaptobrevin 2-SNAP-25-syntaxin-1a-complexin I complex(GO:0070032) |
| 0.1 | 2.9 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.1 | 1.2 | GO:0005742 | mitochondrial outer membrane translocase complex(GO:0005742) |
| 0.1 | 1.9 | GO:0043256 | basal lamina(GO:0005605) laminin complex(GO:0043256) |
| 0.0 | 2.2 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.0 | 1.4 | GO:0060293 | germ plasm(GO:0060293) |
| 0.0 | 0.1 | GO:0097058 | CRLF-CLCF1 complex(GO:0097058) |
| 0.0 | 0.4 | GO:0008250 | oligosaccharyltransferase complex(GO:0008250) |
| 0.0 | 0.6 | GO:0044666 | MLL3/4 complex(GO:0044666) |
| 0.0 | 1.2 | GO:0030964 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.0 | 2.1 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.0 | 2.5 | GO:0090575 | RNA polymerase II transcription factor complex(GO:0090575) |
| 0.0 | 1.0 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.0 | 0.5 | GO:0005844 | polysome(GO:0005844) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 2.5 | GO:0008430 | selenium binding(GO:0008430) |
| 0.5 | 1.5 | GO:0008401 | retinoic acid 4-hydroxylase activity(GO:0008401) |
| 0.3 | 1.3 | GO:0015349 | thyroid hormone transmembrane transporter activity(GO:0015349) |
| 0.2 | 9.9 | GO:0045182 | translation regulator activity(GO:0045182) |
| 0.2 | 0.6 | GO:0031434 | mitogen-activated protein kinase kinase binding(GO:0031434) |
| 0.2 | 3.4 | GO:0070411 | I-SMAD binding(GO:0070411) |
| 0.1 | 0.9 | GO:0004962 | endothelin receptor activity(GO:0004962) |
| 0.1 | 0.7 | GO:0004719 | protein-L-isoaspartate (D-aspartate) O-methyltransferase activity(GO:0004719) |
| 0.1 | 3.6 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.1 | 1.2 | GO:0008320 | protein transmembrane transporter activity(GO:0008320) |
| 0.1 | 0.9 | GO:0003886 | DNA (cytosine-5-)-methyltransferase activity(GO:0003886) |
| 0.1 | 0.4 | GO:0034714 | type III transforming growth factor beta receptor binding(GO:0034714) |
| 0.1 | 0.2 | GO:0034618 | arginine binding(GO:0034618) |
| 0.1 | 0.6 | GO:0071558 | histone demethylase activity (H3-K27 specific)(GO:0071558) |
| 0.1 | 0.3 | GO:0033781 | cholesterol 24-hydroxylase activity(GO:0033781) |
| 0.0 | 1.1 | GO:0043236 | laminin binding(GO:0043236) |
| 0.0 | 0.4 | GO:0001217 | bacterial-type RNA polymerase transcription factor activity, sequence-specific DNA binding(GO:0001130) bacterial-type RNA polymerase transcriptional repressor activity, sequence-specific DNA binding(GO:0001217) |
| 0.0 | 0.2 | GO:0016822 | hydrolase activity, acting on acid carbon-carbon bonds(GO:0016822) hydrolase activity, acting on acid carbon-carbon bonds, in ketonic substances(GO:0016823) |
| 0.0 | 0.4 | GO:0004563 | beta-N-acetylhexosaminidase activity(GO:0004563) |
| 0.0 | 1.0 | GO:0031683 | G-protein beta/gamma-subunit complex binding(GO:0031683) |
| 0.0 | 4.0 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.0 | 0.2 | GO:0016863 | intramolecular oxidoreductase activity, transposing C=C bonds(GO:0016863) |
| 0.0 | 1.2 | GO:0003755 | peptidyl-prolyl cis-trans isomerase activity(GO:0003755) |
| 0.0 | 0.6 | GO:0004697 | protein kinase C activity(GO:0004697) calcium-dependent protein kinase C activity(GO:0004698) |
| 0.0 | 0.3 | GO:0001102 | RNA polymerase II activating transcription factor binding(GO:0001102) |
| 0.0 | 0.7 | GO:0051537 | 2 iron, 2 sulfur cluster binding(GO:0051537) |
| 0.0 | 2.0 | GO:1901981 | phosphatidylinositol phosphate binding(GO:1901981) |
| 0.0 | 0.3 | GO:0015165 | pyrimidine nucleotide-sugar transmembrane transporter activity(GO:0015165) |
| 0.0 | 1.6 | GO:0016616 | oxidoreductase activity, acting on the CH-OH group of donors, NAD or NADP as acceptor(GO:0016616) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 3.4 | PID ALK2 PATHWAY | ALK2 signaling events |
| 0.1 | 2.8 | NABA BASEMENT MEMBRANES | Genes encoding structural components of basement membranes |
| 0.0 | 1.0 | PID RHODOPSIN PATHWAY | Visual signal transduction: Rods |
| 0.0 | 1.6 | PID HDAC CLASSI PATHWAY | Signaling events mediated by HDAC Class I |
| 0.0 | 0.3 | SA PTEN PATHWAY | PTEN is a tumor suppressor that dephosphorylates the lipid messenger phosphatidylinositol triphosphate. |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 3.4 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |
| 0.1 | 1.1 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
| 0.1 | 1.5 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.1 | 1.4 | REACTOME CYTOCHROME P450 ARRANGED BY SUBSTRATE TYPE | Genes involved in Cytochrome P450 - arranged by substrate type |
| 0.0 | 1.0 | REACTOME G PROTEIN ACTIVATION | Genes involved in G-protein activation |
| 0.0 | 0.5 | REACTOME APOPTOTIC CLEAVAGE OF CELLULAR PROTEINS | Genes involved in Apoptotic cleavage of cellular proteins |
| 0.0 | 0.2 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 0.0 | 0.7 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |