Project
PRJNA195909:zebrafish embryo and larva development
Navigation
Downloads
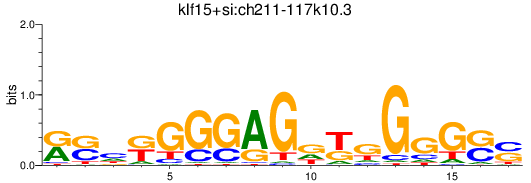
Results for klf15+si:ch211-117k10.3
Z-value: 0.56
Transcription factors associated with klf15+si:ch211-117k10.3
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
si_ch211-117k10.3
|
ENSDARG00000090914 | si_ch211-117k10.3 |
|
klf15
|
ENSDARG00000091127 | Kruppel-like factor 15 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| klf15 | dr11_v1_chr23_-_35195908_35195908 | -0.85 | 4.0e-03 | Click! |
| si:ch211-117k10.3 | dr11_v1_chr11_-_38510469_38510469 | 0.45 | 2.2e-01 | Click! |
Activity profile of klf15+si:ch211-117k10.3 motif
Sorted Z-values of klf15+si:ch211-117k10.3 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr9_-_8314028 | 1.61 |
ENSDART00000102739
|
si:ch211-145c1.1
|
si:ch211-145c1.1 |
| chr14_+_34495216 | 1.13 |
ENSDART00000147756
|
wnt8a
|
wingless-type MMTV integration site family, member 8a |
| chr5_-_30080332 | 0.89 |
ENSDART00000140049
|
bco2a
|
beta-carotene oxygenase 2a |
| chr18_-_26715655 | 0.89 |
ENSDART00000181497
|
malt3
|
MALT paracaspase 3 |
| chr5_+_22067570 | 0.84 |
ENSDART00000045574
|
shisa2a
|
shisa family member 2a |
| chr7_+_1449999 | 0.81 |
ENSDART00000173864
|
si:cabz01101003.1
|
si:cabz01101003.1 |
| chr19_-_46037835 | 0.74 |
ENSDART00000163815
|
nup153
|
nucleoporin 153 |
| chr11_+_6010177 | 0.74 |
ENSDART00000170047
ENSDART00000022526 ENSDART00000161001 ENSDART00000188999 |
gtpbp3
|
GTP binding protein 3, mitochondrial |
| chr5_-_30074332 | 0.72 |
ENSDART00000147963
|
bco2a
|
beta-carotene oxygenase 2a |
| chr14_-_24111292 | 0.71 |
ENSDART00000186611
|
cpeb4a
|
cytoplasmic polyadenylation element binding protein 4a |
| chr6_-_29377092 | 0.70 |
ENSDART00000078665
|
tmem131
|
transmembrane protein 131 |
| chr18_-_26715156 | 0.68 |
ENSDART00000142043
|
malt3
|
MALT paracaspase 3 |
| chr3_+_35611625 | 0.59 |
ENSDART00000190995
|
traf7
|
TNF receptor-associated factor 7 |
| chr8_-_52938439 | 0.57 |
ENSDART00000168185
|
nr6a1a
|
nuclear receptor subfamily 6, group A, member 1a |
| chr10_+_44719697 | 0.55 |
ENSDART00000158087
|
scarb1
|
scavenger receptor class B, member 1 |
| chr19_+_13994563 | 0.53 |
ENSDART00000164696
|
tmem222b
|
transmembrane protein 222b |
| chr19_-_2707048 | 0.52 |
ENSDART00000166112
|
il6
|
interleukin 6 (interferon, beta 2) |
| chr22_-_20403194 | 0.51 |
ENSDART00000010048
|
map2k2a
|
mitogen-activated protein kinase kinase 2a |
| chr6_-_53326421 | 0.50 |
ENSDART00000191740
|
gnb1b
|
guanine nucleotide binding protein (G protein), beta polypeptide 1b |
| chr10_-_25543227 | 0.49 |
ENSDART00000007778
|
grik1a
|
glutamate receptor, ionotropic, kainate 1a |
| chr2_+_44977889 | 0.48 |
ENSDART00000144024
|
alg3
|
asparagine-linked glycosylation 3 (alpha-1,3-mannosyltransferase) |
| chr6_+_46697710 | 0.45 |
ENSDART00000154969
|
tespa1
|
thymocyte expressed, positive selection associated 1 |
| chr8_-_13722682 | 0.44 |
ENSDART00000142052
|
si:dkey-258f14.2
|
si:dkey-258f14.2 |
| chr1_+_7414318 | 0.42 |
ENSDART00000127426
|
si:ch73-383l1.1
|
si:ch73-383l1.1 |
| chr2_-_7246338 | 0.42 |
ENSDART00000186735
|
zgc:153115
|
zgc:153115 |
| chr8_+_8947623 | 0.41 |
ENSDART00000131215
|
slc35a2
|
solute carrier family 35 (UDP-galactose transporter), member 2 |
| chr24_-_38110779 | 0.39 |
ENSDART00000147783
|
crp
|
c-reactive protein, pentraxin-related |
| chr10_+_1849874 | 0.39 |
ENSDART00000158897
ENSDART00000149956 |
apc
|
adenomatous polyposis coli |
| chr5_-_35264517 | 0.36 |
ENSDART00000114981
|
ptcd2
|
pentatricopeptide repeat domain 2 |
| chr24_+_15020402 | 0.36 |
ENSDART00000148102
|
dok6
|
docking protein 6 |
| chr16_+_7991274 | 0.36 |
ENSDART00000179704
|
ano10a
|
anoctamin 10a |
| chr7_-_55454406 | 0.35 |
ENSDART00000108646
|
piezo1
|
piezo-type mechanosensitive ion channel component 1 |
| chr4_+_9057427 | 0.34 |
ENSDART00000113867
|
slc41a2b
|
solute carrier family 41 (magnesium transporter), member 2b |
| chr16_+_9713850 | 0.33 |
ENSDART00000164103
|
ecm1b
|
extracellular matrix protein 1b |
| chr11_-_40504170 | 0.32 |
ENSDART00000165394
|
si:dkeyp-61b2.1
|
si:dkeyp-61b2.1 |
| chr22_-_5918670 | 0.32 |
ENSDART00000141373
|
si:rp71-36a1.1
|
si:rp71-36a1.1 |
| chr12_+_10443785 | 0.31 |
ENSDART00000029133
|
snu13b
|
SNU13 homolog, small nuclear ribonucleoprotein b (U4/U6.U5) |
| chr22_+_26703026 | 0.30 |
ENSDART00000158756
|
crebbpa
|
CREB binding protein a |
| chr4_+_55758103 | 0.30 |
ENSDART00000185964
|
CT583728.23
|
|
| chr9_-_50000144 | 0.29 |
ENSDART00000123416
|
scn1a
|
sodium channel, voltage-gated, type I, alpha |
| chr23_+_36308717 | 0.29 |
ENSDART00000131711
|
hnrnpa1b
|
heterogeneous nuclear ribonucleoprotein A1b |
| chr2_+_24781026 | 0.29 |
ENSDART00000145692
|
pde4ca
|
phosphodiesterase 4C, cAMP-specific a |
| chr8_+_7740004 | 0.29 |
ENSDART00000170184
ENSDART00000187811 |
fgd1
|
FYVE, RhoGEF and PH domain containing 1 |
| chr12_+_47448558 | 0.28 |
ENSDART00000185689
ENSDART00000105328 |
fmn2b
|
formin 2b |
| chr5_-_69621227 | 0.27 |
ENSDART00000178543
|
aldh2.2
|
aldehyde dehydrogenase 2 family (mitochondrial), tandem duplicate 2 |
| chr7_+_19552381 | 0.27 |
ENSDART00000169060
|
si:ch211-212k18.5
|
si:ch211-212k18.5 |
| chr14_-_15154695 | 0.26 |
ENSDART00000160677
|
uvssa
|
UV-stimulated scaffold protein A |
| chr6_+_52918537 | 0.26 |
ENSDART00000174229
|
or137-1
|
odorant receptor, family H, subfamily 137, member 1 |
| chr2_+_7192966 | 0.25 |
ENSDART00000142735
|
si:ch211-13f8.1
|
si:ch211-13f8.1 |
| chr24_+_42131564 | 0.25 |
ENSDART00000153854
|
wwp1
|
WW domain containing E3 ubiquitin protein ligase 1 |
| chr15_+_3125136 | 0.25 |
ENSDART00000130968
|
rcbtb2
|
regulator of chromosome condensation (RCC1) and BTB (POZ) domain containing protein 2 |
| chr18_+_3579829 | 0.25 |
ENSDART00000158763
ENSDART00000182850 ENSDART00000162754 ENSDART00000178789 ENSDART00000172656 |
lrch3
|
leucine-rich repeats and calponin homology (CH) domain containing 3 |
| chr12_-_14211293 | 0.23 |
ENSDART00000158399
|
avl9
|
AVL9 homolog (S. cerevisiase) |
| chr23_+_31245395 | 0.23 |
ENSDART00000053588
|
irak1bp1
|
interleukin-1 receptor-associated kinase 1 binding protein 1 |
| chr12_-_15002563 | 0.23 |
ENSDART00000108852
ENSDART00000141909 |
pkmyt1
|
protein kinase, membrane associated tyrosine/threonine 1 |
| chr10_-_40333319 | 0.23 |
ENSDART00000150479
|
taar20a
|
trace amine associated receptor 20a |
| chr5_+_27525477 | 0.22 |
ENSDART00000051491
|
sfrp1a
|
secreted frizzled-related protein 1a |
| chr17_-_2721336 | 0.21 |
ENSDART00000109582
ENSDART00000192691 ENSDART00000189381 |
spata7
|
spermatogenesis associated 7 |
| chr18_-_2727764 | 0.21 |
ENSDART00000160841
|
ARHGEF17
|
si:ch211-248g20.5 |
| chr10_-_9115383 | 0.21 |
ENSDART00000139324
|
si:dkeyp-41f9.3
|
si:dkeyp-41f9.3 |
| chr24_-_32582378 | 0.21 |
ENSDART00000066590
|
rdh12l
|
retinol dehydrogenase 12, like |
| chr9_+_29323163 | 0.20 |
ENSDART00000184908
|
oxgr1b
|
oxoglutarate (alpha-ketoglutarate) receptor 1b |
| chr10_+_42678520 | 0.19 |
ENSDART00000182496
|
rhobtb2b
|
Rho-related BTB domain containing 2b |
| chr8_+_35356944 | 0.19 |
ENSDART00000143243
|
si:dkeyp-14d3.1
|
si:dkeyp-14d3.1 |
| chr7_+_29167744 | 0.18 |
ENSDART00000076345
|
slc38a8b
|
solute carrier family 38, member 8b |
| chr17_-_4395373 | 0.18 |
ENSDART00000015923
|
klhl10a
|
kelch-like family member 10a |
| chr11_-_23322182 | 0.17 |
ENSDART00000111289
|
kiss1
|
KiSS-1 metastasis-suppressor |
| chr21_+_7188943 | 0.16 |
ENSDART00000172174
|
agpat2
|
1-acylglycerol-3-phosphate O-acyltransferase 2 (lysophosphatidic acid acyltransferase, beta) |
| chr7_+_23495986 | 0.16 |
ENSDART00000190739
ENSDART00000115299 ENSDART00000101423 ENSDART00000142401 |
zgc:109889
|
zgc:109889 |
| chr10_+_39476600 | 0.16 |
ENSDART00000135756
|
kirrel3a
|
kirre like nephrin family adhesion molecule 3a |
| chr18_+_32780450 | 0.15 |
ENSDART00000168399
|
olfcg7
|
olfactory receptor C family, g7 |
| chr5_-_36916790 | 0.14 |
ENSDART00000143827
|
kptn
|
kaptin (actin binding protein) |
| chr11_-_19501439 | 0.13 |
ENSDART00000158328
ENSDART00000103983 |
slc2a9l1
|
solute carrier family 2 (facilitated glucose transporter), member 9-like 1 |
| chr20_+_35279968 | 0.13 |
ENSDART00000168216
ENSDART00000153332 |
fam49a
|
family with sequence similarity 49, member A |
| chr7_-_5711197 | 0.13 |
ENSDART00000134367
|
si:dkey-10h3.2
|
si:dkey-10h3.2 |
| chr25_+_150570 | 0.12 |
ENSDART00000170892
|
adam10b
|
ADAM metallopeptidase domain 10b |
| chr14_-_23799345 | 0.12 |
ENSDART00000124144
ENSDART00000181179 ENSDART00000124401 |
nr3c1
|
nuclear receptor subfamily 3, group C, member 1 (glucocorticoid receptor) |
| chr21_+_15601100 | 0.12 |
ENSDART00000180558
ENSDART00000145454 |
smarcb1b
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily b, member 1b |
| chr16_-_12470034 | 0.12 |
ENSDART00000147483
|
ephb6
|
eph receptor B6 |
| chr3_-_36690348 | 0.12 |
ENSDART00000192513
|
myh11b
|
myosin, heavy chain 11b, smooth muscle |
| chr4_-_11064073 | 0.11 |
ENSDART00000150760
|
si:dkey-21h14.8
|
si:dkey-21h14.8 |
| chr7_+_13918349 | 0.11 |
ENSDART00000172772
ENSDART00000173384 |
si:cabz01059983.1
|
si:cabz01059983.1 |
| chr20_-_7080427 | 0.11 |
ENSDART00000140166
ENSDART00000023677 |
si:ch211-121a2.2
|
si:ch211-121a2.2 |
| chr4_+_65306260 | 0.11 |
ENSDART00000167558
|
si:dkey-14o6.8
|
si:dkey-14o6.8 |
| chr8_-_53488832 | 0.11 |
ENSDART00000191801
|
chdh
|
choline dehydrogenase |
| chr22_+_12477996 | 0.10 |
ENSDART00000177704
|
CR847870.3
|
|
| chr21_+_1143141 | 0.10 |
ENSDART00000178294
|
CABZ01086139.1
|
|
| chr20_-_1560724 | 0.09 |
ENSDART00000179193
ENSDART00000092508 ENSDART00000133369 |
ptprk
|
protein tyrosine phosphatase, receptor type, K |
| chr22_-_6219674 | 0.08 |
ENSDART00000182582
|
si:ch211-274k16.2
|
si:ch211-274k16.2 |
| chr9_-_39968820 | 0.08 |
ENSDART00000100311
|
IKZF2
|
si:zfos-1425h8.1 |
| chr24_-_11467004 | 0.08 |
ENSDART00000144316
|
pxdc1b
|
PX domain containing 1b |
| chr23_-_40817792 | 0.08 |
ENSDART00000136343
|
si:dkeyp-27c8.1
|
si:dkeyp-27c8.1 |
| chr15_-_18223769 | 0.08 |
ENSDART00000139966
|
or131-1
|
odorant receptor, family H, subfamily 131, member 1 |
| chr25_-_34284144 | 0.08 |
ENSDART00000188109
|
gcnt3
|
glucosaminyl (N-acetyl) transferase 3, mucin type |
| chr14_+_48960078 | 0.08 |
ENSDART00000165514
|
mif
|
macrophage migration inhibitory factor |
| chr7_-_71331690 | 0.08 |
ENSDART00000149682
|
dhx15
|
DEAH (Asp-Glu-Ala-His) box helicase 15 |
| chr4_-_75505048 | 0.08 |
ENSDART00000163665
|
si:dkey-71l4.5
|
si:dkey-71l4.5 |
| chr22_-_4872418 | 0.07 |
ENSDART00000132360
|
si:ch73-256j6.6
|
si:ch73-256j6.6 |
| chr23_+_7518294 | 0.07 |
ENSDART00000081536
|
hck
|
HCK proto-oncogene, Src family tyrosine kinase |
| chr14_+_45596583 | 0.07 |
ENSDART00000110429
|
zgc:194285
|
zgc:194285 |
| chr23_+_39331216 | 0.07 |
ENSDART00000160957
|
kcng1
|
potassium voltage-gated channel, subfamily G, member 1 |
| chr3_+_52467879 | 0.06 |
ENSDART00000156039
|
adgre5a
|
adhesion G protein-coupled receptor E5a |
| chr22_-_17474583 | 0.06 |
ENSDART00000148027
|
si:ch211-197g15.8
|
si:ch211-197g15.8 |
| chr10_-_24961310 | 0.05 |
ENSDART00000037303
|
si:ch211-214k5.3
|
si:ch211-214k5.3 |
| chr18_+_46151505 | 0.05 |
ENSDART00000015034
ENSDART00000141287 |
blvrb
|
biliverdin reductase B |
| chr20_-_46053720 | 0.04 |
ENSDART00000136254
|
taar12h
|
trace amine associated receptor 12h |
| chr10_+_39476432 | 0.04 |
ENSDART00000190375
ENSDART00000183954 |
kirrel3a
|
kirre like nephrin family adhesion molecule 3a |
| chr22_+_413349 | 0.04 |
ENSDART00000082453
|
celsr2
|
cadherin, EGF LAG seven-pass G-type receptor 2 |
| chr16_-_12914288 | 0.04 |
ENSDART00000184221
|
cacng8b
|
calcium channel, voltage-dependent, gamma subunit 8b |
| chr2_-_23479714 | 0.04 |
ENSDART00000167291
|
si:ch211-14p21.3
|
si:ch211-14p21.3 |
| chr6_-_43677125 | 0.04 |
ENSDART00000150128
|
foxp1b
|
forkhead box P1b |
| chr5_-_8636168 | 0.03 |
ENSDART00000134877
|
fyba
|
FYN binding protein a |
| chr7_-_71331499 | 0.03 |
ENSDART00000081245
|
dhx15
|
DEAH (Asp-Glu-Ala-His) box helicase 15 |
| chr21_-_3201027 | 0.03 |
ENSDART00000161256
|
zbtb7c
|
zinc finger and BTB domain containing 7C |
| chr12_-_16990896 | 0.03 |
ENSDART00000152402
|
ifit12
|
interferon-induced protein with tetratricopeptide repeats 12 |
| chr22_-_18112374 | 0.03 |
ENSDART00000191154
|
ncanb
|
neurocan b |
| chr10_+_5645887 | 0.02 |
ENSDART00000171426
|
pdzph1
|
PDZ and pleckstrin homology domains 1 |
| chr4_-_73572030 | 0.02 |
ENSDART00000121652
|
znf1015
|
zinc finger protein 1015 |
| chr15_-_28618502 | 0.01 |
ENSDART00000086902
|
slc6a4a
|
solute carrier family 6 (neurotransmitter transporter), member 4a |
| chr21_+_41649336 | 0.00 |
ENSDART00000164694
ENSDART00000181539 |
ppp2r2bb
|
protein phosphatase 2, regulatory subunit B, beta b |
Network of associatons between targets according to the STRING database.
First level regulatory network of klf15+si:ch211-117k10.3
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.1 | GO:0072111 | spinal cord anterior/posterior patterning(GO:0021512) cell proliferation involved in pronephros development(GO:0039015) cell proliferation involved in kidney development(GO:0072111) |
| 0.2 | 0.6 | GO:0043654 | recognition of apoptotic cell(GO:0043654) |
| 0.2 | 1.6 | GO:0042214 | carotene metabolic process(GO:0016119) carotene catabolic process(GO:0016121) terpene metabolic process(GO:0042214) terpene catabolic process(GO:0046247) |
| 0.1 | 0.4 | GO:0010387 | COP9 signalosome assembly(GO:0010387) |
| 0.1 | 0.7 | GO:0002931 | response to ischemia(GO:0002931) |
| 0.1 | 0.4 | GO:0061031 | endodermal digestive tract morphogenesis(GO:0061031) Wnt signaling pathway involved in somitogenesis(GO:0090244) negative regulation of canonical Wnt signaling pathway involved in heart development(GO:1905067) |
| 0.1 | 0.5 | GO:0006953 | acute-phase response(GO:0006953) |
| 0.1 | 0.3 | GO:0010481 | ventriculo bulbo valve development(GO:0003173) epidermal cell division(GO:0010481) regulation of epidermal cell division(GO:0010482) |
| 0.1 | 0.5 | GO:1902624 | positive regulation of neutrophil migration(GO:1902624) |
| 0.0 | 0.3 | GO:0006283 | transcription-coupled nucleotide-excision repair(GO:0006283) |
| 0.0 | 0.1 | GO:0019695 | choline metabolic process(GO:0019695) |
| 0.0 | 0.7 | GO:0002098 | tRNA wobble uridine modification(GO:0002098) |
| 0.0 | 0.1 | GO:0071387 | cellular response to cortisol stimulus(GO:0071387) response to dexamethasone(GO:0071548) |
| 0.0 | 0.4 | GO:0015781 | pyrimidine nucleotide-sugar transport(GO:0015781) pyrimidine nucleotide-sugar transmembrane transport(GO:0090481) |
| 0.0 | 0.3 | GO:0006198 | cAMP catabolic process(GO:0006198) |
| 0.0 | 0.2 | GO:0097499 | protein localization to nonmotile primary cilium(GO:0097499) |
| 0.0 | 0.7 | GO:0006405 | RNA export from nucleus(GO:0006405) |
| 0.0 | 0.3 | GO:0070167 | regulation of bone mineralization(GO:0030500) regulation of biomineral tissue development(GO:0070167) |
| 0.0 | 0.5 | GO:0000187 | activation of MAPK activity(GO:0000187) |
| 0.0 | 0.2 | GO:0051127 | positive regulation of actin nucleation(GO:0051127) positive regulation of Arp2/3 complex-mediated actin nucleation(GO:2000601) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.1 | GO:0097189 | apoptotic body(GO:0097189) |
| 0.1 | 0.4 | GO:0030877 | beta-catenin destruction complex(GO:0030877) |
| 0.1 | 0.3 | GO:0031428 | box C/D snoRNP complex(GO:0031428) |
| 0.0 | 0.7 | GO:1990124 | messenger ribonucleoprotein complex(GO:1990124) |
| 0.0 | 0.4 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.0 | 0.1 | GO:0035060 | brahma complex(GO:0035060) |
| 0.0 | 0.7 | GO:0005643 | nuclear pore(GO:0005643) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.6 | GO:0010436 | beta-carotene 15,15'-monooxygenase activity(GO:0003834) carotenoid dioxygenase activity(GO:0010436) |
| 0.1 | 0.5 | GO:0000033 | alpha-1,3-mannosyltransferase activity(GO:0000033) |
| 0.1 | 0.6 | GO:0030169 | low-density lipoprotein particle binding(GO:0030169) |
| 0.0 | 0.7 | GO:0000900 | translation repressor activity, nucleic acid binding(GO:0000900) |
| 0.0 | 0.1 | GO:1902945 | metalloendopeptidase activity involved in amyloid precursor protein catabolic process(GO:1902945) |
| 0.0 | 0.4 | GO:0005459 | UDP-galactose transmembrane transporter activity(GO:0005459) |
| 0.0 | 0.7 | GO:0008139 | nuclear localization sequence binding(GO:0008139) |
| 0.0 | 0.5 | GO:0031682 | G-protein gamma-subunit binding(GO:0031682) |
| 0.0 | 1.1 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.0 | 0.5 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.0 | 0.3 | GO:0004029 | aldehyde dehydrogenase (NAD) activity(GO:0004029) |
| 0.0 | 0.2 | GO:0071933 | Arp2/3 complex binding(GO:0071933) |
| 0.0 | 0.4 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.0 | 0.5 | GO:1904315 | neurotransmitter receptor activity involved in regulation of postsynaptic membrane potential(GO:0099529) transmitter-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1904315) |
| 0.0 | 0.3 | GO:0005080 | protein kinase C binding(GO:0005080) |
| 0.0 | 0.3 | GO:0004115 | 3',5'-cyclic-AMP phosphodiesterase activity(GO:0004115) |
| 0.0 | 0.2 | GO:0003841 | 1-acylglycerol-3-phosphate O-acyltransferase activity(GO:0003841) |
| 0.0 | 0.3 | GO:0008381 | mechanically-gated ion channel activity(GO:0008381) mechanically gated channel activity(GO:0022833) |
| 0.0 | 0.3 | GO:0000993 | RNA polymerase II core binding(GO:0000993) |
| 0.0 | 1.6 | GO:0004197 | cysteine-type endopeptidase activity(GO:0004197) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | ST STAT3 PATHWAY | STAT3 Pathway |
| 0.0 | 0.7 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.0 | 0.4 | PID RET PATHWAY | Signaling events regulated by Ret tyrosine kinase |
| 0.0 | 0.4 | PID WNT CANONICAL PATHWAY | Canonical Wnt signaling pathway |
| 0.0 | 1.1 | WNT SIGNALING | Genes related to Wnt-mediated signal transduction |
| 0.0 | 0.5 | PID IL6 7 PATHWAY | IL6-mediated signaling events |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | REACTOME IL 6 SIGNALING | Genes involved in Interleukin-6 signaling |
| 0.0 | 0.6 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.0 | 0.4 | REACTOME CREATION OF C4 AND C2 ACTIVATORS | Genes involved in Creation of C4 and C2 activators |
| 0.0 | 0.4 | REACTOME CTNNB1 PHOSPHORYLATION CASCADE | Genes involved in Beta-catenin phosphorylation cascade |
| 0.0 | 0.7 | REACTOME TRANSPORT OF RIBONUCLEOPROTEINS INTO THE HOST NUCLEUS | Genes involved in Transport of Ribonucleoproteins into the Host Nucleus |
| 0.0 | 0.4 | REACTOME TRANSPORT OF VITAMINS NUCLEOSIDES AND RELATED MOLECULES | Genes involved in Transport of vitamins, nucleosides, and related molecules |
| 0.0 | 0.5 | REACTOME BIOSYNTHESIS OF THE N GLYCAN PRECURSOR DOLICHOL LIPID LINKED OLIGOSACCHARIDE LLO AND TRANSFER TO A NASCENT PROTEIN | Genes involved in Biosynthesis of the N-glycan precursor (dolichol lipid-linked oligosaccharide, LLO) and transfer to a nascent protein |
| 0.0 | 0.2 | REACTOME CYCLIN A B1 ASSOCIATED EVENTS DURING G2 M TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 0.0 | 0.5 | REACTOME NRAGE SIGNALS DEATH THROUGH JNK | Genes involved in NRAGE signals death through JNK |