Project
PRJNA195909:zebrafish embryo and larva development
Navigation
Downloads
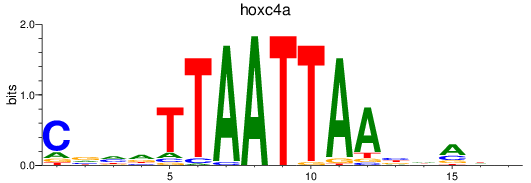
Results for hoxc4a
Z-value: 0.61
Transcription factors associated with hoxc4a
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
hoxc4a
|
ENSDARG00000070338 | homeobox C4a |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| hoxc4a | dr11_v1_chr23_+_36130883_36130883 | 0.76 | 1.7e-02 | Click! |
Activity profile of hoxc4a motif
Sorted Z-values of hoxc4a motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr9_-_22099536 | 1.70 |
ENSDART00000101923
|
CR391987.1
|
|
| chr12_-_35830625 | 1.43 |
ENSDART00000180028
|
CU459056.1
|
|
| chr16_+_23431189 | 1.39 |
ENSDART00000004679
|
icn
|
ictacalcin |
| chr19_+_10855158 | 1.35 |
ENSDART00000172219
ENSDART00000170826 |
apoea
|
apolipoprotein Ea |
| chr4_+_9669717 | 1.25 |
ENSDART00000004604
|
si:dkey-153k10.9
|
si:dkey-153k10.9 |
| chr18_-_48983690 | 1.17 |
ENSDART00000182359
|
FO681288.3
|
|
| chr23_-_21453614 | 1.05 |
ENSDART00000079274
|
her4.1
|
hairy-related 4, tandem duplicate 1 |
| chr16_-_42894628 | 1.03 |
ENSDART00000045600
|
hfe2
|
hemochromatosis type 2 |
| chr9_-_14504834 | 0.94 |
ENSDART00000056103
|
nrp2b
|
neuropilin 2b |
| chr24_-_25691020 | 0.88 |
ENSDART00000015391
|
chrnd
|
cholinergic receptor, nicotinic, delta (muscle) |
| chr22_-_16042243 | 0.88 |
ENSDART00000062633
|
s1pr1
|
sphingosine-1-phosphate receptor 1 |
| chr1_-_669717 | 0.87 |
ENSDART00000160564
|
cyyr1
|
cysteine/tyrosine-rich 1 |
| chr21_+_25236297 | 0.85 |
ENSDART00000112783
|
tmem45b
|
transmembrane protein 45B |
| chr5_-_66702479 | 0.83 |
ENSDART00000129197
|
mn1b
|
meningioma 1b |
| chr17_+_41992054 | 0.80 |
ENSDART00000182878
ENSDART00000111537 |
kiz
|
kizuna centrosomal protein |
| chr13_-_29420885 | 0.79 |
ENSDART00000024225
|
chata
|
choline O-acetyltransferase a |
| chr2_-_30668580 | 0.74 |
ENSDART00000087270
|
ctnnd2b
|
catenin (cadherin-associated protein), delta 2b |
| chr3_-_50443607 | 0.68 |
ENSDART00000074036
|
rcvrna
|
recoverin a |
| chr22_+_12770877 | 0.68 |
ENSDART00000044683
|
ftcd
|
formimidoyltransferase cyclodeaminase |
| chr16_+_46111849 | 0.65 |
ENSDART00000172232
|
sv2a
|
synaptic vesicle glycoprotein 2A |
| chr5_-_38451082 | 0.64 |
ENSDART00000136428
|
chrne
|
cholinergic receptor, nicotinic, epsilon |
| chr24_+_24461558 | 0.63 |
ENSDART00000182424
|
bhlhe22
|
basic helix-loop-helix family, member e22 |
| chr24_-_38657683 | 0.61 |
ENSDART00000154843
|
si:ch1073-164k15.3
|
si:ch1073-164k15.3 |
| chr24_+_24461341 | 0.60 |
ENSDART00000147658
|
bhlhe22
|
basic helix-loop-helix family, member e22 |
| chr18_+_3037998 | 0.58 |
ENSDART00000185844
ENSDART00000162657 |
rps3
|
ribosomal protein S3 |
| chr19_+_43297546 | 0.58 |
ENSDART00000168002
|
laptm5
|
lysosomal protein transmembrane 5 |
| chr14_-_413273 | 0.58 |
ENSDART00000163976
ENSDART00000179907 |
FAT4
|
FAT atypical cadherin 4 |
| chr4_+_57881965 | 0.57 |
ENSDART00000162234
|
si:dkeyp-44b5.4
|
si:dkeyp-44b5.4 |
| chr25_-_6011034 | 0.57 |
ENSDART00000075197
ENSDART00000136054 |
snx22
|
sorting nexin 22 |
| chr19_+_43359075 | 0.57 |
ENSDART00000148287
ENSDART00000149856 ENSDART00000188236 ENSDART00000136695 ENSDART00000193859 |
yrk
|
Yes-related kinase |
| chr12_-_31794908 | 0.56 |
ENSDART00000105583
ENSDART00000153449 |
nat9
|
N-acetyltransferase 9 (GCN5-related, putative) |
| chr5_+_11840905 | 0.54 |
ENSDART00000030444
|
tesca
|
tescalcin a |
| chr2_+_11685742 | 0.54 |
ENSDART00000138562
|
greb1l
|
growth regulation by estrogen in breast cancer-like |
| chr4_-_9891874 | 0.53 |
ENSDART00000067193
|
adm2a
|
adrenomedullin 2a |
| chr10_-_14556978 | 0.51 |
ENSDART00000126643
|
zgc:153395
|
zgc:153395 |
| chr9_+_37152564 | 0.51 |
ENSDART00000189497
|
gli2a
|
GLI family zinc finger 2a |
| chr4_+_16715267 | 0.49 |
ENSDART00000143849
|
pkp2
|
plakophilin 2 |
| chr10_+_21650828 | 0.49 |
ENSDART00000160754
|
pcdh1g1
|
protocadherin 1 gamma 1 |
| chr9_-_22834860 | 0.48 |
ENSDART00000146486
|
neb
|
nebulin |
| chr14_-_34771371 | 0.46 |
ENSDART00000160598
ENSDART00000150413 ENSDART00000168910 |
ablim3
|
actin binding LIM protein family, member 3 |
| chr1_+_21731382 | 0.46 |
ENSDART00000054395
|
pax5
|
paired box 5 |
| chr11_+_30057762 | 0.45 |
ENSDART00000164139
|
nhsb
|
Nance-Horan syndrome b (congenital cataracts and dental anomalies) |
| chr9_-_1986014 | 0.44 |
ENSDART00000142842
|
hoxd12a
|
homeobox D12a |
| chr11_+_34760628 | 0.43 |
ENSDART00000087216
|
si:dkey-202e22.2
|
si:dkey-202e22.2 |
| chr13_+_12299997 | 0.43 |
ENSDART00000108535
|
gabrb1
|
gamma-aminobutyric acid (GABA) A receptor, beta 1 |
| chr3_+_36284986 | 0.43 |
ENSDART00000059533
|
wipi1
|
WD repeat domain, phosphoinositide interacting 1 |
| chr22_-_12160283 | 0.42 |
ENSDART00000146785
ENSDART00000128176 |
tmem163b
|
transmembrane protein 163b |
| chr23_+_7471072 | 0.42 |
ENSDART00000135551
|
si:ch211-200e2.1
|
si:ch211-200e2.1 |
| chr12_-_35054354 | 0.41 |
ENSDART00000075351
|
zgc:112285
|
zgc:112285 |
| chr17_-_12336987 | 0.39 |
ENSDART00000172001
|
snap25b
|
synaptosomal-associated protein, 25b |
| chr6_-_11768198 | 0.39 |
ENSDART00000183463
|
march7
|
membrane-associated ring finger (C3HC4) 7 |
| chr8_-_18537866 | 0.39 |
ENSDART00000148802
ENSDART00000148962 ENSDART00000149506 |
nexn
|
nexilin (F actin binding protein) |
| chr11_+_43431289 | 0.39 |
ENSDART00000192526
|
slc29a1b
|
solute carrier family 29 (equilibrative nucleoside transporter), member 1b |
| chr16_-_12173554 | 0.39 |
ENSDART00000110567
ENSDART00000155935 |
clstn3
|
calsyntenin 3 |
| chr1_-_31534089 | 0.37 |
ENSDART00000007770
|
lbx1b
|
ladybird homeobox 1b |
| chr18_+_13162728 | 0.37 |
ENSDART00000101472
|
tat
|
tyrosine aminotransferase |
| chr24_+_21540842 | 0.36 |
ENSDART00000091529
|
wasf3b
|
WAS protein family, member 3b |
| chr15_-_14552101 | 0.36 |
ENSDART00000171169
|
numbl
|
numb homolog (Drosophila)-like |
| chr9_-_42418470 | 0.35 |
ENSDART00000144353
|
calcrla
|
calcitonin receptor-like a |
| chr8_-_12867434 | 0.35 |
ENSDART00000081657
|
slc2a6
|
solute carrier family 2 (facilitated glucose transporter), member 6 |
| chr6_+_25257728 | 0.33 |
ENSDART00000162581
|
kyat3
|
kynurenine aminotransferase 3 |
| chr2_+_2223837 | 0.33 |
ENSDART00000101038
ENSDART00000129354 |
tmie
|
transmembrane inner ear |
| chr4_+_45148652 | 0.33 |
ENSDART00000150798
|
si:dkey-51d8.9
|
si:dkey-51d8.9 |
| chr16_+_12836143 | 0.33 |
ENSDART00000067741
|
cacng6b
|
calcium channel, voltage-dependent, gamma subunit 6b |
| chr3_-_16039619 | 0.32 |
ENSDART00000143324
|
spsb3a
|
splA/ryanodine receptor domain and SOCS box containing 3a |
| chr3_-_3798755 | 0.31 |
ENSDART00000170535
|
gucy2c
|
guanylate cyclase 2C |
| chr11_-_45141309 | 0.31 |
ENSDART00000181736
|
cant1b
|
calcium activated nucleotidase 1b |
| chr14_-_451555 | 0.29 |
ENSDART00000190906
|
FAT4
|
FAT atypical cadherin 4 |
| chr14_-_858985 | 0.29 |
ENSDART00000148687
ENSDART00000149375 |
slc34a1a
|
solute carrier family 34 (type II sodium/phosphate cotransporter), member 1a |
| chr1_+_6172786 | 0.28 |
ENSDART00000126468
|
prkag3a
|
protein kinase, AMP-activated, gamma 3a non-catalytic subunit |
| chr12_+_31735159 | 0.28 |
ENSDART00000185442
|
RNF157
|
si:dkey-49c17.3 |
| chr20_+_98179 | 0.28 |
ENSDART00000022725
|
si:ch1073-155h21.1
|
si:ch1073-155h21.1 |
| chr16_-_12173399 | 0.28 |
ENSDART00000142574
|
clstn3
|
calsyntenin 3 |
| chr23_-_16485190 | 0.28 |
ENSDART00000155038
|
si:dkeyp-100a5.4
|
si:dkeyp-100a5.4 |
| chr15_-_17618800 | 0.27 |
ENSDART00000157185
|
adamts15b
|
ADAM metallopeptidase with thrombospondin type 1 motif, 15b |
| chr6_+_15373153 | 0.27 |
ENSDART00000155865
|
tmtops2a
|
teleost multiple tissue opsin 2a |
| chr14_-_15956921 | 0.27 |
ENSDART00000190500
|
flt4
|
fms-related tyrosine kinase 4 |
| chr19_-_1929310 | 0.27 |
ENSDART00000144113
|
si:ch211-149a19.3
|
si:ch211-149a19.3 |
| chr23_-_17657348 | 0.26 |
ENSDART00000054736
|
bhlhe23
|
basic helix-loop-helix family, member e23 |
| chr2_+_36701322 | 0.26 |
ENSDART00000002510
|
golim4b
|
golgi integral membrane protein 4b |
| chr4_+_53119675 | 0.26 |
ENSDART00000167470
|
si:dkey-8o9.2
|
si:dkey-8o9.2 |
| chr23_+_20669149 | 0.25 |
ENSDART00000138936
|
arhgef3l
|
Rho guanine nucleotide exchange factor (GEF) 3, like |
| chr15_-_46779934 | 0.24 |
ENSDART00000085136
|
clcn2c
|
chloride channel 2c |
| chr5_+_71802014 | 0.24 |
ENSDART00000124939
ENSDART00000097164 |
LHX3
|
LIM homeobox 3 |
| chr19_-_8877469 | 0.24 |
ENSDART00000193951
|
celf3a
|
cugbp, Elav-like family member 3a |
| chr12_+_7497882 | 0.23 |
ENSDART00000134608
|
phyhiplb
|
phytanoyl-CoA 2-hydroxylase interacting protein-like b |
| chr9_-_12811936 | 0.23 |
ENSDART00000188490
|
myo10l3
|
myosin X-like 3 |
| chr3_-_29941357 | 0.23 |
ENSDART00000147732
ENSDART00000137973 ENSDART00000103523 |
grna
|
granulin a |
| chr15_-_17619306 | 0.22 |
ENSDART00000184011
|
adamts15b
|
ADAM metallopeptidase with thrombospondin type 1 motif, 15b |
| chr3_+_5575313 | 0.22 |
ENSDART00000134693
ENSDART00000101807 |
si:ch211-106h11.3
|
si:ch211-106h11.3 |
| chr23_-_1660708 | 0.21 |
ENSDART00000175138
|
CU693481.1
|
|
| chr12_+_46742805 | 0.20 |
ENSDART00000192236
|
plaub
|
plasminogen activator, urokinase b |
| chr22_-_19552796 | 0.20 |
ENSDART00000148088
ENSDART00000105485 |
si:dkey-78l4.14
|
si:dkey-78l4.14 |
| chr8_-_30979494 | 0.20 |
ENSDART00000138959
|
si:ch211-251j10.3
|
si:ch211-251j10.3 |
| chr6_+_24420523 | 0.20 |
ENSDART00000185461
|
tgfbr3
|
transforming growth factor, beta receptor III |
| chr2_+_20605925 | 0.20 |
ENSDART00000191510
|
olfml2bb
|
olfactomedin-like 2Bb |
| chr13_+_7292061 | 0.19 |
ENSDART00000179504
|
CABZ01072077.1
|
Danio rerio neuroblast differentiation-associated protein AHNAK-like (LOC795051), mRNA. |
| chr7_+_38811800 | 0.19 |
ENSDART00000052322
|
zgc:110699
|
zgc:110699 |
| chr15_+_5360407 | 0.19 |
ENSDART00000110420
|
or112-1
|
odorant receptor, family A, subfamily 112, member 1 |
| chr2_-_17044959 | 0.18 |
ENSDART00000090260
|
clcn2a
|
chloride channel, voltage-sensitive 2a |
| chr8_+_30664077 | 0.18 |
ENSDART00000138750
|
adora2aa
|
adenosine A2a receptor a |
| chr2_+_37140448 | 0.18 |
ENSDART00000045016
ENSDART00000142940 |
pex19
|
peroxisomal biogenesis factor 19 |
| chr5_+_34623107 | 0.18 |
ENSDART00000184126
|
enc1
|
ectodermal-neural cortex 1 |
| chr17_-_20558961 | 0.18 |
ENSDART00000155993
|
sh3pxd2ab
|
SH3 and PX domains 2Ab |
| chr5_-_16218777 | 0.18 |
ENSDART00000141698
|
kremen1
|
kringle containing transmembrane protein 1 |
| chr7_+_34620418 | 0.17 |
ENSDART00000081338
|
slc9a5
|
solute carrier family 9, subfamily A (NHE5, cation proton antiporter 5), member 5 |
| chr1_+_51721851 | 0.17 |
ENSDART00000040397
|
prdx2
|
peroxiredoxin 2 |
| chr10_-_5581487 | 0.17 |
ENSDART00000141943
|
syk
|
spleen tyrosine kinase |
| chr8_-_45760087 | 0.16 |
ENSDART00000025620
|
ppiaa
|
peptidylprolyl isomerase Aa (cyclophilin A) |
| chr17_+_51682429 | 0.16 |
ENSDART00000004379
|
nol10
|
nucleolar protein 10 |
| chr14_+_22591624 | 0.16 |
ENSDART00000108987
|
gfra4b
|
GDNF family receptor alpha 4b |
| chr10_+_43039947 | 0.16 |
ENSDART00000193434
|
atg10
|
ATG10 autophagy related 10 homolog (S. cerevisiae) |
| chr1_-_19502322 | 0.16 |
ENSDART00000181888
ENSDART00000044030 |
kitb
|
v-kit Hardy-Zuckerman 4 feline sarcoma viral oncogene homolog b |
| chr11_-_2838699 | 0.16 |
ENSDART00000066189
|
lhfpl5a
|
LHFPL tetraspan subfamily member 5a |
| chr20_-_23842631 | 0.15 |
ENSDART00000153079
|
si:dkey-15j16.3
|
si:dkey-15j16.3 |
| chr12_-_47648538 | 0.15 |
ENSDART00000108477
|
fh
|
fumarate hydratase |
| chr18_-_17485419 | 0.15 |
ENSDART00000018764
|
foxl1
|
forkhead box L1 |
| chr9_+_11281969 | 0.15 |
ENSDART00000110691
|
wnt6b
|
wingless-type MMTV integration site family, member 6b |
| chr2_-_37140423 | 0.15 |
ENSDART00000144220
|
tspan37
|
tetraspanin 37 |
| chr12_+_20641102 | 0.15 |
ENSDART00000152964
|
calcoco2
|
calcium binding and coiled-coil domain 2 |
| chr12_-_26153101 | 0.14 |
ENSDART00000076051
|
opn4b
|
opsin 4b |
| chr21_-_20939488 | 0.14 |
ENSDART00000039043
|
rgs7bpb
|
regulator of G protein signaling 7 binding protein b |
| chr15_+_31899312 | 0.14 |
ENSDART00000155315
|
frya
|
furry homolog a (Drosophila) |
| chr1_+_11882186 | 0.13 |
ENSDART00000193419
|
snx8b
|
sorting nexin 8b |
| chr13_+_29462249 | 0.13 |
ENSDART00000147903
|
lrit1a
|
leucine-rich repeat, immunoglobulin-like and transmembrane domains 1a |
| chr17_+_50524573 | 0.13 |
ENSDART00000187974
|
CR382385.1
|
|
| chr24_+_13316737 | 0.13 |
ENSDART00000191658
|
SBSPON
|
somatomedin B and thrombospondin type 1 domain containing |
| chr9_-_30576522 | 0.12 |
ENSDART00000101085
|
morc3a
|
MORC family CW-type zinc finger 3a |
| chr5_-_23280098 | 0.12 |
ENSDART00000126540
ENSDART00000051533 |
plp1b
|
proteolipid protein 1b |
| chr14_+_1170968 | 0.12 |
ENSDART00000125203
ENSDART00000193575 |
hopx
|
HOP homeobox |
| chr16_+_40024883 | 0.12 |
ENSDART00000110100
|
hint3
|
histidine triad nucleotide binding protein 3 |
| chr10_+_42423318 | 0.12 |
ENSDART00000134282
|
npy8ar
|
neuropeptide Y receptor Y8a |
| chr4_-_13548806 | 0.12 |
ENSDART00000067155
|
il22
|
interleukin 22 |
| chr12_+_20641471 | 0.12 |
ENSDART00000133654
|
calcoco2
|
calcium binding and coiled-coil domain 2 |
| chr7_-_52388734 | 0.12 |
ENSDART00000174186
|
wdr93
|
WD repeat domain 93 |
| chr8_-_12867128 | 0.11 |
ENSDART00000142201
|
slc2a6
|
solute carrier family 2 (facilitated glucose transporter), member 6 |
| chr8_-_44926077 | 0.11 |
ENSDART00000084417
|
timm17b
|
translocase of inner mitochondrial membrane 17 homolog B (yeast) |
| chr1_-_513762 | 0.11 |
ENSDART00000148162
ENSDART00000144606 |
trmt10c
|
tRNA methyltransferase 10C, mitochondrial RNase P subunit |
| chr4_+_70151160 | 0.11 |
ENSDART00000111816
|
si:dkey-3h2.4
|
si:dkey-3h2.4 |
| chr9_-_32177117 | 0.10 |
ENSDART00000078568
|
sf3b1
|
splicing factor 3b, subunit 1 |
| chr21_+_25802190 | 0.10 |
ENSDART00000128987
|
nf2b
|
neurofibromin 2b (merlin) |
| chr1_+_51039558 | 0.10 |
ENSDART00000024743
|
dpy30
|
dpy-30 histone methyltransferase complex regulatory subunit |
| chr3_-_13921173 | 0.10 |
ENSDART00000159177
ENSDART00000165174 |
si:dkey-61n16.5
|
si:dkey-61n16.5 |
| chr1_+_57787642 | 0.09 |
ENSDART00000127091
|
si:dkey-1c7.2
|
si:dkey-1c7.2 |
| chr10_-_29236860 | 0.09 |
ENSDART00000111620
|
ccdc83
|
coiled-coil domain containing 83 |
| chr17_-_10122204 | 0.09 |
ENSDART00000160751
|
BX088587.1
|
|
| chr9_-_47472998 | 0.08 |
ENSDART00000134480
|
tns1b
|
tensin 1b |
| chr17_-_26604946 | 0.08 |
ENSDART00000087062
|
fam149b1
|
family with sequence similarity 149, member B1 |
| chr2_+_3201345 | 0.08 |
ENSDART00000130349
|
wnt9a
|
wingless-type MMTV integration site family, member 9A |
| chr4_+_69191065 | 0.08 |
ENSDART00000170595
|
znf1075
|
zinc finger protein 1075 |
| chr18_+_17827149 | 0.07 |
ENSDART00000190237
ENSDART00000189345 |
ZNF423
|
si:ch211-216l23.1 |
| chr10_+_29850330 | 0.07 |
ENSDART00000168898
|
hspa8
|
heat shock protein 8 |
| chr19_+_19032141 | 0.07 |
ENSDART00000192432
|
cpne4b
|
copine IVb |
| chr22_-_36530902 | 0.07 |
ENSDART00000056188
|
polr2h
|
info polymerase (RNA) II (DNA directed) polypeptide H |
| chr6_+_36821621 | 0.06 |
ENSDART00000104157
|
tmem45a
|
transmembrane protein 45a |
| chr2_+_38373272 | 0.06 |
ENSDART00000113111
|
psmb5
|
proteasome subunit beta 5 |
| chr8_+_53064920 | 0.06 |
ENSDART00000164823
|
nadka
|
NAD kinase a |
| chr23_+_12134839 | 0.06 |
ENSDART00000128551
ENSDART00000141204 |
ttll9
|
tubulin tyrosine ligase-like family, member 9 |
| chr1_-_50438247 | 0.06 |
ENSDART00000114098
|
dkk2
|
dickkopf WNT signaling pathway inhibitor 2 |
| chr22_+_19552987 | 0.05 |
ENSDART00000105315
|
hsd11b1la
|
hydroxysteroid (11-beta) dehydrogenase 1-like a |
| chr20_-_9095105 | 0.05 |
ENSDART00000140792
|
oma1
|
OMA1 zinc metallopeptidase |
| chr20_+_40457599 | 0.05 |
ENSDART00000017553
|
serinc1
|
serine incorporator 1 |
| chr10_+_35358675 | 0.05 |
ENSDART00000193263
|
si:dkey-259j3.5
|
si:dkey-259j3.5 |
| chr7_+_4474880 | 0.04 |
ENSDART00000143528
|
si:dkey-83f18.14
|
si:dkey-83f18.14 |
| chr6_+_11249706 | 0.04 |
ENSDART00000186547
ENSDART00000193287 |
atg9a
|
ATG9 autophagy related 9 homolog A (S. cerevisiae) |
| chr3_+_23029484 | 0.03 |
ENSDART00000187900
|
nags
|
N-acetylglutamate synthase |
| chr20_+_34717403 | 0.03 |
ENSDART00000034252
|
pnocb
|
prepronociceptin b |
| chr3_+_3681116 | 0.03 |
ENSDART00000109618
|
art4
|
ADP-ribosyltransferase 4 (Dombrock blood group) |
| chr22_+_26853254 | 0.03 |
ENSDART00000182487
|
tmem186
|
transmembrane protein 186 |
| chr17_+_31914877 | 0.03 |
ENSDART00000177801
|
FAM196A (1 of many)
|
family with sequence similarity 196 member A |
| chr11_+_24815667 | 0.02 |
ENSDART00000141730
|
rabif
|
RAB interacting factor |
| chr9_+_23895711 | 0.02 |
ENSDART00000034686
|
cops8
|
COP9 signalosome subunit 8 |
| chr9_+_30211038 | 0.02 |
ENSDART00000190847
|
senp7a
|
SUMO1/sentrin specific peptidase 7a |
| chr7_-_49594995 | 0.02 |
ENSDART00000174161
ENSDART00000109147 |
brsk2b
|
BR serine/threonine kinase 2b |
| chr10_+_26747755 | 0.02 |
ENSDART00000100329
|
f9b
|
coagulation factor IXb |
| chr11_+_44617021 | 0.02 |
ENSDART00000158887
|
rbm34
|
RNA binding motif protein 34 |
| chr2_-_37043905 | 0.02 |
ENSDART00000056514
|
gng7
|
guanine nucleotide binding protein (G protein), gamma 7 |
| chr17_+_2162916 | 0.01 |
ENSDART00000103775
|
pak6a
|
p21 protein (Cdc42/Rac)-activated kinase 6a |
| chr24_+_744713 | 0.01 |
ENSDART00000067764
|
stk17a
|
serine/threonine kinase 17a |
| chr13_-_12602920 | 0.00 |
ENSDART00000102311
|
lrit3b
|
leucine-rich repeat, immunoglobulin-like and transmembrane domains 3b |
| chr6_-_43677125 | 0.00 |
ENSDART00000150128
|
foxp1b
|
forkhead box P1b |
| chr1_+_9086942 | 0.00 |
ENSDART00000055023
|
cacng3a
|
calcium channel, voltage-dependent, gamma subunit 3a |
| chr3_-_34136778 | 0.00 |
ENSDART00000131951
|
clpp
|
caseinolytic mitochondrial matrix peptidase proteolytic subunit |
Network of associatons between targets according to the STRING database.
First level regulatory network of hoxc4a
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.4 | GO:0097242 | regulation of nitric-oxide synthase activity(GO:0050999) positive regulation of nitric-oxide synthase activity(GO:0051000) beta-amyloid clearance(GO:0097242) regulation of beta-amyloid clearance(GO:1900221) |
| 0.3 | 0.9 | GO:0003376 | sphingosine-1-phosphate signaling pathway(GO:0003376) |
| 0.3 | 0.8 | GO:0008292 | acetylcholine biosynthetic process(GO:0008292) acetate ester biosynthetic process(GO:1900620) |
| 0.2 | 0.7 | GO:0019557 | formate metabolic process(GO:0015942) histidine catabolic process to glutamate and formamide(GO:0019556) histidine catabolic process to glutamate and formate(GO:0019557) formamide metabolic process(GO:0043606) |
| 0.1 | 0.5 | GO:0048618 | post-embryonic foregut morphogenesis(GO:0048618) |
| 0.1 | 0.9 | GO:0071678 | olfactory bulb axon guidance(GO:0071678) |
| 0.1 | 0.5 | GO:0071691 | cardiac muscle thin filament assembly(GO:0071691) |
| 0.1 | 0.7 | GO:0051965 | positive regulation of synapse assembly(GO:0051965) |
| 0.1 | 0.5 | GO:0045110 | intermediate filament bundle assembly(GO:0045110) |
| 0.1 | 0.3 | GO:1902514 | calcium ion transmembrane transport via high voltage-gated calcium channel(GO:0061577) regulation of calcium ion transmembrane transport via high voltage-gated calcium channel(GO:1902514) |
| 0.1 | 0.4 | GO:0015864 | uridine transport(GO:0015862) pyrimidine nucleoside transport(GO:0015864) |
| 0.1 | 0.7 | GO:0036368 | cone photoresponse recovery(GO:0036368) |
| 0.1 | 0.4 | GO:0006572 | tyrosine catabolic process(GO:0006572) |
| 0.1 | 0.2 | GO:0045579 | positive regulation of B cell differentiation(GO:0045579) |
| 0.1 | 0.4 | GO:0048739 | cardiac muscle fiber development(GO:0048739) |
| 0.1 | 0.3 | GO:0060836 | lymphatic endothelial cell differentiation(GO:0060836) |
| 0.1 | 0.5 | GO:0060004 | reflex(GO:0060004) vestibular reflex(GO:0060005) vestibular receptor cell differentiation(GO:0060114) vestibular receptor cell development(GO:0060118) |
| 0.0 | 0.3 | GO:0044341 | sodium-dependent phosphate transport(GO:0044341) |
| 0.0 | 1.0 | GO:0048936 | peripheral nervous system neuron axonogenesis(GO:0048936) |
| 0.0 | 0.2 | GO:0061469 | regulation of type B pancreatic cell proliferation(GO:0061469) |
| 0.0 | 0.2 | GO:0015919 | peroxisomal membrane transport(GO:0015919) protein import into peroxisome membrane(GO:0045046) |
| 0.0 | 0.2 | GO:0048903 | anterior lateral line neuromast hair cell differentiation(GO:0048903) |
| 0.0 | 0.2 | GO:0038093 | Fc receptor signaling pathway(GO:0038093) |
| 0.0 | 0.2 | GO:0071340 | skeletal muscle acetylcholine-gated channel clustering(GO:0071340) |
| 0.0 | 0.4 | GO:0045162 | clustering of voltage-gated sodium channels(GO:0045162) |
| 0.0 | 0.6 | GO:2001235 | positive regulation of apoptotic signaling pathway(GO:2001235) |
| 0.0 | 0.3 | GO:0034497 | protein localization to pre-autophagosomal structure(GO:0034497) |
| 0.0 | 0.4 | GO:0007271 | synaptic transmission, cholinergic(GO:0007271) |
| 0.0 | 0.1 | GO:0090646 | mitochondrial tRNA processing(GO:0090646) |
| 0.0 | 0.6 | GO:0048246 | macrophage chemotaxis(GO:0048246) |
| 0.0 | 0.5 | GO:0030032 | lamellipodium assembly(GO:0030032) |
| 0.0 | 0.2 | GO:0033628 | regulation of cell adhesion mediated by integrin(GO:0033628) |
| 0.0 | 0.5 | GO:0060997 | dendritic spine morphogenesis(GO:0060997) |
| 0.0 | 0.3 | GO:0060088 | auditory receptor cell stereocilium organization(GO:0060088) |
| 0.0 | 0.2 | GO:0000303 | response to superoxide(GO:0000303) response to oxygen radical(GO:0000305) removal of superoxide radicals(GO:0019430) cellular response to oxygen radical(GO:0071450) cellular response to superoxide(GO:0071451) cellular oxidant detoxification(GO:0098869) cellular detoxification(GO:1990748) |
| 0.0 | 0.2 | GO:1990118 | sodium ion import(GO:0097369) sodium ion import across plasma membrane(GO:0098719) sodium ion import into cell(GO:1990118) |
| 0.0 | 0.4 | GO:0070831 | basement membrane assembly(GO:0070831) |
| 0.0 | 0.2 | GO:0006108 | malate metabolic process(GO:0006108) |
| 0.0 | 0.4 | GO:0048844 | artery morphogenesis(GO:0048844) |
| 0.0 | 0.4 | GO:0006829 | zinc II ion transport(GO:0006829) |
| 0.0 | 0.2 | GO:0031295 | lymphocyte costimulation(GO:0031294) T cell costimulation(GO:0031295) |
| 0.0 | 0.2 | GO:0048026 | positive regulation of mRNA splicing, via spliceosome(GO:0048026) |
| 0.0 | 0.1 | GO:0006348 | chromatin silencing at telomere(GO:0006348) |
| 0.0 | 0.2 | GO:0006801 | superoxide metabolic process(GO:0006801) |
| 0.0 | 0.1 | GO:1902946 | positive regulation of receptor-mediated endocytosis(GO:0048260) protein localization to early endosome(GO:1902946) |
| 0.0 | 0.2 | GO:0060122 | inner ear receptor stereocilium organization(GO:0060122) |
| 0.0 | 0.4 | GO:0060319 | primitive erythrocyte differentiation(GO:0060319) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.4 | GO:0034363 | intermediate-density lipoprotein particle(GO:0034363) |
| 0.1 | 1.5 | GO:0005892 | acetylcholine-gated channel complex(GO:0005892) |
| 0.1 | 0.4 | GO:0070032 | synaptobrevin 2-SNAP-25-syntaxin-1a-complexin I complex(GO:0070032) |
| 0.1 | 0.4 | GO:0034045 | pre-autophagosomal structure membrane(GO:0034045) |
| 0.0 | 0.3 | GO:1990454 | L-type voltage-gated calcium channel complex(GO:1990454) |
| 0.0 | 0.5 | GO:0014704 | intercalated disc(GO:0014704) |
| 0.0 | 0.4 | GO:0098563 | integral component of synaptic vesicle membrane(GO:0030285) intrinsic component of synaptic vesicle membrane(GO:0098563) |
| 0.0 | 0.3 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.0 | 0.4 | GO:0005605 | basal lamina(GO:0005605) laminin complex(GO:0043256) |
| 0.0 | 0.3 | GO:0005903 | brush border(GO:0005903) |
| 0.0 | 0.1 | GO:0042645 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.0 | 0.4 | GO:1902711 | GABA receptor complex(GO:1902710) GABA-A receptor complex(GO:1902711) |
| 0.0 | 0.6 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.0 | 0.1 | GO:0005744 | mitochondrial inner membrane presequence translocase complex(GO:0005744) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.7 | GO:0016841 | folic acid binding(GO:0005542) ammonia-lyase activity(GO:0016841) |
| 0.1 | 0.4 | GO:0001635 | adrenomedullin receptor activity(GO:0001605) calcitonin gene-related peptide receptor activity(GO:0001635) |
| 0.1 | 0.9 | GO:0038036 | sphingosine-1-phosphate receptor activity(GO:0038036) |
| 0.1 | 1.2 | GO:0005021 | vascular endothelial growth factor-activated receptor activity(GO:0005021) |
| 0.1 | 0.5 | GO:0019215 | intermediate filament binding(GO:0019215) |
| 0.1 | 0.5 | GO:0055056 | D-glucose transmembrane transporter activity(GO:0055056) |
| 0.1 | 1.5 | GO:0022848 | acetylcholine-gated cation channel activity(GO:0022848) |
| 0.1 | 0.8 | GO:0016413 | O-acetyltransferase activity(GO:0016413) |
| 0.1 | 0.2 | GO:0019777 | Atg12 transferase activity(GO:0019777) |
| 0.0 | 1.4 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.0 | 0.3 | GO:0005436 | sodium:phosphate symporter activity(GO:0005436) |
| 0.0 | 0.2 | GO:0050431 | transforming growth factor beta binding(GO:0050431) |
| 0.0 | 0.1 | GO:0052905 | tRNA (guanine(9)-N(1))-methyltransferase activity(GO:0052905) |
| 0.0 | 0.2 | GO:0001609 | G-protein coupled adenosine receptor activity(GO:0001609) |
| 0.0 | 0.2 | GO:0008379 | thioredoxin peroxidase activity(GO:0008379) |
| 0.0 | 0.4 | GO:0005247 | voltage-gated chloride channel activity(GO:0005247) |
| 0.0 | 0.2 | GO:0016176 | superoxide-generating NADPH oxidase activator activity(GO:0016176) |
| 0.0 | 0.4 | GO:0080025 | phosphatidylinositol-3,5-bisphosphate binding(GO:0080025) |
| 0.0 | 0.2 | GO:0000268 | peroxisome targeting sequence binding(GO:0000268) |
| 0.0 | 0.2 | GO:0005451 | monovalent cation:proton antiporter activity(GO:0005451) sodium:proton antiporter activity(GO:0015385) potassium:proton antiporter activity(GO:0015386) |
| 0.0 | 0.4 | GO:0004890 | GABA-A receptor activity(GO:0004890) |
| 0.0 | 0.4 | GO:0008483 | transaminase activity(GO:0008483) |
| 0.0 | 0.3 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.0 | 0.3 | GO:0017110 | nucleoside-diphosphatase activity(GO:0017110) |
| 0.0 | 0.4 | GO:0098632 | protein binding involved in cell-cell adhesion(GO:0098632) |
| 0.0 | 0.1 | GO:0039706 | co-receptor binding(GO:0039706) |
| 0.0 | 0.0 | GO:0034618 | arginine binding(GO:0034618) |
| 0.0 | 0.2 | GO:0016018 | cyclosporin A binding(GO:0016018) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.9 | PID S1P META PATHWAY | Sphingosine 1-phosphate (S1P) pathway |
| 0.1 | 0.3 | PID VEGF VEGFR PATHWAY | VEGF and VEGFR signaling network |
| 0.0 | 1.0 | PID BMP PATHWAY | BMP receptor signaling |
| 0.0 | 0.2 | PID NFKAPPAB ATYPICAL PATHWAY | Atypical NF-kappaB pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.5 | REACTOME PRESYNAPTIC NICOTINIC ACETYLCHOLINE RECEPTORS | Genes involved in Presynaptic nicotinic acetylcholine receptors |
| 0.1 | 0.5 | REACTOME FACILITATIVE NA INDEPENDENT GLUCOSE TRANSPORTERS | Genes involved in Facilitative Na+-independent glucose transporters |
| 0.1 | 0.5 | REACTOME DCC MEDIATED ATTRACTIVE SIGNALING | Genes involved in DCC mediated attractive signaling |
| 0.0 | 0.3 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.0 | 1.0 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.0 | 0.4 | REACTOME GABA A RECEPTOR ACTIVATION | Genes involved in GABA A receptor activation |
| 0.0 | 0.5 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.0 | 0.2 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.0 | 0.6 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |