Project
PRJNA195909:zebrafish embryo and larva development
Navigation
Downloads
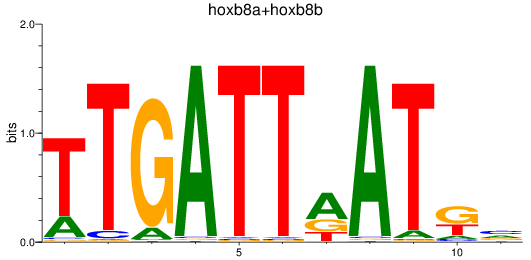
Results for hoxb8a+hoxb8b
Z-value: 1.41
Transcription factors associated with hoxb8a+hoxb8b
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
hoxb8b
|
ENSDARG00000054025 | homeobox B8b |
|
hoxb8a
|
ENSDARG00000056027 | homeobox B8a |
|
hoxb8b
|
ENSDARG00000111056 | homeobox B8b |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| hoxb8b | dr11_v1_chr12_+_27117609_27117609 | 0.47 | 2.1e-01 | Click! |
| hoxb8a | dr11_v1_chr3_+_23687909_23687909 | 0.45 | 2.2e-01 | Click! |
Activity profile of hoxb8a+hoxb8b motif
Sorted Z-values of hoxb8a+hoxb8b motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr24_+_38301080 | 3.48 |
ENSDART00000105672
|
mybpc2b
|
myosin binding protein C, fast type b |
| chr2_+_33368414 | 2.09 |
ENSDART00000077462
|
slc6a9
|
solute carrier family 6 (neurotransmitter transporter, glycine), member 9 |
| chr3_+_33300522 | 1.88 |
ENSDART00000114023
|
hspb9
|
heat shock protein, alpha-crystallin-related, 9 |
| chr10_-_39011514 | 1.82 |
ENSDART00000075123
|
pcp4a
|
Purkinje cell protein 4a |
| chr1_+_17676745 | 1.68 |
ENSDART00000030665
|
slc25a4
|
solute carrier family 25 (mitochondrial carrier; adenine nucleotide translocator), member 4 |
| chr12_-_26406323 | 1.56 |
ENSDART00000131896
|
myoz1b
|
myozenin 1b |
| chr6_+_52804267 | 1.55 |
ENSDART00000065681
|
matn4
|
matrilin 4 |
| chr19_-_3123963 | 1.48 |
ENSDART00000122816
|
si:ch211-80h18.1
|
si:ch211-80h18.1 |
| chr4_-_4387012 | 1.28 |
ENSDART00000191836
|
CU468826.3
|
Danio rerio U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit-related protein 1-like (LOC100331497), mRNA. |
| chr7_+_31879649 | 1.19 |
ENSDART00000099789
|
mybpc3
|
myosin binding protein C, cardiac |
| chr13_+_23282549 | 1.18 |
ENSDART00000101134
|
khdrbs2
|
KH domain containing, RNA binding, signal transduction associated 2 |
| chr1_+_16144615 | 1.17 |
ENSDART00000054707
|
tusc3
|
tumor suppressor candidate 3 |
| chr18_-_38088099 | 1.15 |
ENSDART00000146120
|
luzp2
|
leucine zipper protein 2 |
| chr15_+_4632782 | 1.13 |
ENSDART00000156012
|
si:dkey-35i13.1
|
si:dkey-35i13.1 |
| chr18_+_21408794 | 1.03 |
ENSDART00000140161
|
necab2
|
N-terminal EF-hand calcium binding protein 2 |
| chr6_-_43449013 | 1.03 |
ENSDART00000122423
|
eevs
|
2-epi-5-epi-valiolone synthase |
| chr7_+_31879986 | 1.03 |
ENSDART00000138491
|
mybpc3
|
myosin binding protein C, cardiac |
| chr9_+_42066030 | 1.01 |
ENSDART00000185311
ENSDART00000015267 |
pcbp3
|
poly(rC) binding protein 3 |
| chr23_-_11870962 | 1.01 |
ENSDART00000143481
|
si:dkey-178k16.1
|
si:dkey-178k16.1 |
| chr20_+_20672163 | 1.00 |
ENSDART00000027758
|
rtn1b
|
reticulon 1b |
| chr3_+_23752150 | 0.98 |
ENSDART00000146636
|
hoxb2a
|
homeobox B2a |
| chr25_+_15647750 | 0.98 |
ENSDART00000137375
|
spon1b
|
spondin 1b |
| chr15_+_28202170 | 0.96 |
ENSDART00000077736
|
vtna
|
vitronectin a |
| chr2_+_2223837 | 0.96 |
ENSDART00000101038
ENSDART00000129354 |
tmie
|
transmembrane inner ear |
| chr13_-_30027730 | 0.93 |
ENSDART00000044009
|
scdb
|
stearoyl-CoA desaturase b |
| chr16_+_10776688 | 0.91 |
ENSDART00000161969
ENSDART00000172657 |
atp1a3b
|
ATPase Na+/K+ transporting subunit alpha 3b |
| chr9_-_23217196 | 0.91 |
ENSDART00000083567
|
kif5c
|
kinesin family member 5C |
| chr2_-_9646857 | 0.91 |
ENSDART00000056901
|
zgc:153615
|
zgc:153615 |
| chr14_+_25465346 | 0.87 |
ENSDART00000173436
|
si:dkey-280e21.3
|
si:dkey-280e21.3 |
| chr12_+_39685485 | 0.87 |
ENSDART00000163403
|
LO017650.1
|
|
| chr10_-_7785930 | 0.87 |
ENSDART00000043961
ENSDART00000111058 |
mpx
|
myeloid-specific peroxidase |
| chr8_-_14484599 | 0.82 |
ENSDART00000057644
|
lhx4
|
LIM homeobox 4 |
| chr19_+_12444943 | 0.82 |
ENSDART00000135706
|
ldlrad4a
|
low density lipoprotein receptor class A domain containing 4a |
| chr5_-_825920 | 0.82 |
ENSDART00000126982
|
zgc:158463
|
zgc:158463 |
| chr13_-_5569562 | 0.81 |
ENSDART00000102576
|
meis1b
|
Meis homeobox 1 b |
| chr6_+_14949950 | 0.79 |
ENSDART00000149202
ENSDART00000149949 |
pou3f3b
|
POU class 3 homeobox 3b |
| chr24_-_28893251 | 0.77 |
ENSDART00000042065
ENSDART00000003503 |
col11a1a
|
collagen, type XI, alpha 1a |
| chr24_+_24461341 | 0.76 |
ENSDART00000147658
|
bhlhe22
|
basic helix-loop-helix family, member e22 |
| chr22_+_18389271 | 0.75 |
ENSDART00000088270
|
yjefn3
|
YjeF N-terminal domain containing 3 |
| chr8_-_34052019 | 0.75 |
ENSDART00000040126
ENSDART00000159208 ENSDART00000048994 ENSDART00000098822 |
pbx3b
|
pre-B-cell leukemia homeobox 3b |
| chr3_-_34337969 | 0.74 |
ENSDART00000151634
|
tnrc6c1
|
trinucleotide repeat containing 6C1 |
| chr25_+_15647993 | 0.73 |
ENSDART00000186578
ENSDART00000031828 |
spon1b
|
spondin 1b |
| chr24_+_24461558 | 0.73 |
ENSDART00000182424
|
bhlhe22
|
basic helix-loop-helix family, member e22 |
| chr4_-_9891874 | 0.72 |
ENSDART00000067193
|
adm2a
|
adrenomedullin 2a |
| chr13_-_5568928 | 0.71 |
ENSDART00000192679
|
meis1b
|
Meis homeobox 1 b |
| chr18_+_783936 | 0.67 |
ENSDART00000193357
|
rpp25b
|
ribonuclease P and MRP subunit p25, b |
| chr15_-_34567370 | 0.60 |
ENSDART00000099793
|
sostdc1a
|
sclerostin domain containing 1a |
| chr4_+_13810811 | 0.59 |
ENSDART00000067168
|
pdzrn4
|
PDZ domain containing ring finger 4 |
| chr17_+_39242437 | 0.59 |
ENSDART00000156138
ENSDART00000128863 |
zgc:174356
|
zgc:174356 |
| chr21_-_27338639 | 0.59 |
ENSDART00000130632
|
hif1al2
|
hypoxia-inducible factor 1, alpha subunit, like 2 |
| chr9_-_1951144 | 0.59 |
ENSDART00000082355
|
hoxd4a
|
homeobox D4a |
| chr6_-_28980756 | 0.58 |
ENSDART00000014661
|
glmnb
|
glomulin, FKBP associated protein b |
| chr7_-_35126374 | 0.57 |
ENSDART00000141211
|
hsd11b2
|
hydroxysteroid (11-beta) dehydrogenase 2 |
| chr2_+_7818368 | 0.57 |
ENSDART00000007068
|
kcnmb2
|
potassium large conductance calcium-activated channel, subfamily M, beta member 2 |
| chr8_-_34051548 | 0.55 |
ENSDART00000105204
|
pbx3b
|
pre-B-cell leukemia homeobox 3b |
| chr3_+_59784632 | 0.55 |
ENSDART00000084729
|
pecam1
|
platelet/endothelial cell adhesion molecule 1 |
| chr20_-_29420713 | 0.55 |
ENSDART00000147464
|
ryr3
|
ryanodine receptor 3 |
| chr6_+_37894914 | 0.54 |
ENSDART00000148817
|
oca2
|
oculocutaneous albinism II |
| chr20_-_19590378 | 0.53 |
ENSDART00000152588
|
baalcb
|
brain and acute leukemia, cytoplasmic b |
| chr1_-_19215336 | 0.52 |
ENSDART00000162949
ENSDART00000170680 |
ptprdb
|
protein tyrosine phosphatase, receptor type, D, b |
| chr16_+_17389116 | 0.50 |
ENSDART00000103750
ENSDART00000173448 |
fam131bb
|
family with sequence similarity 131, member Bb |
| chr12_-_17201028 | 0.50 |
ENSDART00000020541
|
lipf
|
lipase, gastric |
| chr24_-_26328721 | 0.48 |
ENSDART00000125468
|
apodb
|
apolipoprotein Db |
| chr18_+_13792490 | 0.48 |
ENSDART00000136754
|
cdh13
|
cadherin 13, H-cadherin (heart) |
| chr17_-_30666037 | 0.48 |
ENSDART00000156509
|
alkal2b
|
ALK and LTK ligand 2b |
| chr6_-_18960105 | 0.48 |
ENSDART00000185278
ENSDART00000162968 |
sept9b
|
septin 9b |
| chr11_+_23265157 | 0.46 |
ENSDART00000110152
|
csf1a
|
colony stimulating factor 1a (macrophage) |
| chr8_-_52413032 | 0.45 |
ENSDART00000183039
|
CABZ01070469.1
|
|
| chr12_-_11457625 | 0.45 |
ENSDART00000012318
|
htra1b
|
HtrA serine peptidase 1b |
| chr1_-_23157583 | 0.44 |
ENSDART00000144208
|
adgrl3.1
|
adhesion G protein-coupled receptor L3.1 |
| chr21_+_43669943 | 0.44 |
ENSDART00000136025
|
tmlhe
|
trimethyllysine hydroxylase, epsilon |
| chr1_+_26071869 | 0.44 |
ENSDART00000059264
|
mxd4
|
MAX dimerization protein 4 |
| chr11_+_6819050 | 0.42 |
ENSDART00000104289
|
rab3ab
|
RAB3A, member RAS oncogene family, b |
| chr24_+_25259154 | 0.41 |
ENSDART00000171125
|
gabrr3b
|
gamma-aminobutyric acid (GABA) A receptor, rho 3b |
| chr12_+_18458502 | 0.41 |
ENSDART00000108745
|
rnf151
|
ring finger protein 151 |
| chr12_-_3756405 | 0.41 |
ENSDART00000150839
|
fam57bb
|
family with sequence similarity 57, member Bb |
| chr11_-_3959889 | 0.40 |
ENSDART00000159683
|
pbrm1
|
polybromo 1 |
| chr19_+_37925616 | 0.40 |
ENSDART00000148348
|
nxph1
|
neurexophilin 1 |
| chr2_+_47623202 | 0.40 |
ENSDART00000154465
|
si:ch211-165b10.3
|
si:ch211-165b10.3 |
| chr11_+_43419809 | 0.39 |
ENSDART00000172982
|
slc29a1b
|
solute carrier family 29 (equilibrative nucleoside transporter), member 1b |
| chr12_+_17201522 | 0.37 |
ENSDART00000014536
ENSDART00000152452 |
rnls
|
renalase, FAD-dependent amine oxidase |
| chr2_+_3809226 | 0.37 |
ENSDART00000147261
|
egfra
|
epidermal growth factor receptor a (erythroblastic leukemia viral (v-erb-b) oncogene homolog, avian) |
| chr10_+_31244619 | 0.35 |
ENSDART00000145562
ENSDART00000184412 |
robo4
|
roundabout, axon guidance receptor, homolog 4 (Drosophila) |
| chr14_+_36231126 | 0.34 |
ENSDART00000141766
|
elovl6
|
ELOVL fatty acid elongase 6 |
| chr5_+_9382301 | 0.34 |
ENSDART00000124017
|
ugt2a7
|
UDP glucuronosyltransferase 2 family, polypeptide A7 |
| chr23_-_32304650 | 0.33 |
ENSDART00000143772
ENSDART00000085642 ENSDART00000188989 |
dgkaa
|
diacylglycerol kinase, alpha a |
| chr21_-_26918901 | 0.33 |
ENSDART00000100685
|
lrfn4a
|
leucine rich repeat and fibronectin type III domain containing 4a |
| chr5_+_9377005 | 0.33 |
ENSDART00000124924
|
ugt2a7
|
UDP glucuronosyltransferase 2 family, polypeptide A7 |
| chr10_+_26571174 | 0.32 |
ENSDART00000148617
ENSDART00000112956 |
slc9a6b
|
solute carrier family 9, subfamily A (NHE6, cation proton antiporter 6), member 6b |
| chr16_-_35952789 | 0.32 |
ENSDART00000180118
|
eva1ba
|
eva-1 homolog Ba (C. elegans) |
| chr10_+_36026576 | 0.32 |
ENSDART00000193786
|
hmgb1a
|
high mobility group box 1a |
| chr7_+_13382852 | 0.32 |
ENSDART00000166318
|
dagla
|
diacylglycerol lipase, alpha |
| chr20_+_32152355 | 0.31 |
ENSDART00000152904
ENSDART00000139507 |
sesn1
|
sestrin 1 |
| chr2_-_10188598 | 0.30 |
ENSDART00000189122
|
dmbx1a
|
diencephalon/mesencephalon homeobox 1a |
| chr24_+_5840258 | 0.30 |
ENSDART00000087034
|
trpc1
|
transient receptor potential cation channel, subfamily C, member 1 |
| chr25_-_31702970 | 0.30 |
ENSDART00000153586
|
lamb4
|
laminin, beta 4 |
| chr14_+_35237613 | 0.30 |
ENSDART00000163465
|
ebf3a
|
early B cell factor 3a |
| chr1_-_46984142 | 0.29 |
ENSDART00000125032
|
pknox1.2
|
pbx/knotted 1 homeobox 1.2 |
| chr4_+_10754802 | 0.29 |
ENSDART00000181470
|
stab2
|
stabilin 2 |
| chr2_-_54039293 | 0.29 |
ENSDART00000166013
|
abhd8a
|
abhydrolase domain containing 8a |
| chr20_-_2949028 | 0.28 |
ENSDART00000104667
ENSDART00000193151 ENSDART00000131946 |
cdk19
|
cyclin-dependent kinase 19 |
| chr18_+_27218737 | 0.28 |
ENSDART00000111450
|
plekha7a
|
pleckstrin homology domain containing, family A member 7a |
| chr11_+_36243774 | 0.28 |
ENSDART00000023323
|
zgc:172270
|
zgc:172270 |
| chr1_-_8980665 | 0.28 |
ENSDART00000148182
|
si:ch73-191k20.3
|
si:ch73-191k20.3 |
| chr12_-_38548299 | 0.28 |
ENSDART00000153374
|
si:dkey-1f1.3
|
si:dkey-1f1.3 |
| chr11_-_3959477 | 0.27 |
ENSDART00000045971
|
pbrm1
|
polybromo 1 |
| chr14_+_6954579 | 0.27 |
ENSDART00000060998
|
nme5
|
NME/NM23 family member 5 |
| chr13_-_28272299 | 0.27 |
ENSDART00000006393
|
tlx1
|
T cell leukemia homeobox 1 |
| chr21_+_22423286 | 0.27 |
ENSDART00000133190
|
capslb
|
calcyphosine-like b |
| chr20_-_8443425 | 0.27 |
ENSDART00000083908
|
dab1a
|
Dab, reelin signal transducer, homolog 1a (Drosophila) |
| chr4_+_7391110 | 0.27 |
ENSDART00000160708
ENSDART00000187823 |
tnni4a
|
troponin I4a |
| chr23_+_21479958 | 0.27 |
ENSDART00000188302
ENSDART00000144320 |
si:dkey-1c11.1
|
si:dkey-1c11.1 |
| chr19_+_28256076 | 0.27 |
ENSDART00000133354
|
irx4b
|
iroquois homeobox 4b |
| chr10_-_27741793 | 0.26 |
ENSDART00000129369
ENSDART00000192440 ENSDART00000189808 ENSDART00000138149 |
auts2a
si:dkey-33o22.1
|
autism susceptibility candidate 2a si:dkey-33o22.1 |
| chr15_-_21132480 | 0.26 |
ENSDART00000078734
ENSDART00000157481 |
a2ml
|
alpha-2-macroglobulin-like |
| chr24_+_18948665 | 0.26 |
ENSDART00000106186
|
prex2
|
phosphatidylinositol-3,4,5-trisphosphate-dependent Rac exchange factor 2 |
| chr13_+_30912385 | 0.26 |
ENSDART00000182642
|
drgx
|
dorsal root ganglia homeobox |
| chr5_-_201600 | 0.25 |
ENSDART00000158495
|
CABZ01088906.1
|
|
| chr8_+_18588551 | 0.25 |
ENSDART00000177476
ENSDART00000063539 |
prrg1
|
proline rich Gla (G-carboxyglutamic acid) 1 |
| chr14_-_6666854 | 0.25 |
ENSDART00000133031
|
si:dkeyp-44a8.4
|
si:dkeyp-44a8.4 |
| chr17_-_19626357 | 0.25 |
ENSDART00000011432
|
reep3a
|
receptor accessory protein 3a |
| chr2_+_31665836 | 0.24 |
ENSDART00000135411
ENSDART00000143914 |
si:ch211-106h4.12
|
si:ch211-106h4.12 |
| chr20_-_26491567 | 0.24 |
ENSDART00000147154
|
mthfd1l
|
methylenetetrahydrofolate dehydrogenase (NADP+ dependent) 1 like |
| chr8_-_20380750 | 0.24 |
ENSDART00000113910
|
adamts10
|
ADAM metallopeptidase with thrombospondin type 1 motif, 10 |
| chr12_-_43428542 | 0.23 |
ENSDART00000192266
|
ptprea
|
protein tyrosine phosphatase, receptor type, E, a |
| chr24_-_6898302 | 0.23 |
ENSDART00000158646
|
dpp6a
|
dipeptidyl-peptidase 6a |
| chr7_+_53755054 | 0.22 |
ENSDART00000181629
|
neo1a
|
neogenin 1a |
| chr23_+_5490854 | 0.22 |
ENSDART00000175403
|
tulp1a
|
tubby like protein 1a |
| chr16_+_46401576 | 0.22 |
ENSDART00000130264
|
rpz
|
rapunzel |
| chr18_-_14836862 | 0.22 |
ENSDART00000124843
|
mtss1la
|
metastasis suppressor 1-like a |
| chr15_-_5467477 | 0.22 |
ENSDART00000123839
|
arrb1
|
arrestin, beta 1 |
| chr12_+_22404108 | 0.22 |
ENSDART00000153055
|
hdlbpb
|
high density lipoprotein binding protein b |
| chr7_-_46777876 | 0.21 |
ENSDART00000193954
|
tshz3b
|
teashirt zinc finger homeobox 3b |
| chr18_+_34225520 | 0.20 |
ENSDART00000126115
|
v2rl1
|
vomeronasal 2 receptor, l1 |
| chr24_+_13735616 | 0.20 |
ENSDART00000184267
|
msc
|
musculin (activated B-cell factor-1) |
| chr5_+_19261391 | 0.20 |
ENSDART00000089173
|
atp8b5a
|
ATPase phospholipid transporting 8B5a |
| chr14_+_35024521 | 0.20 |
ENSDART00000158634
ENSDART00000170631 |
ebf3a
|
early B cell factor 3a |
| chr17_-_38778826 | 0.20 |
ENSDART00000168182
ENSDART00000124041 ENSDART00000136921 |
dglucy
|
D-glutamate cyclase |
| chr5_-_35743741 | 0.19 |
ENSDART00000046157
ENSDART00000123562 |
ophn1
|
oligophrenin 1 |
| chr10_-_33156789 | 0.19 |
ENSDART00000192268
ENSDART00000182065 ENSDART00000081170 |
cux1a
|
cut-like homeobox 1a |
| chr1_-_681116 | 0.19 |
ENSDART00000165894
|
adamts1
|
ADAM metallopeptidase with thrombospondin type 1 motif, 1 |
| chr9_+_20519846 | 0.19 |
ENSDART00000109680
ENSDART00000142220 |
vtcn1
|
V-set domain containing T cell activation inhibitor 1 |
| chr5_+_19261687 | 0.18 |
ENSDART00000139401
|
atp8b5a
|
ATPase phospholipid transporting 8B5a |
| chr7_-_33248773 | 0.18 |
ENSDART00000052389
|
itga11a
|
integrin, alpha 11a |
| chr7_-_18598661 | 0.18 |
ENSDART00000182109
|
DTX4
|
si:ch211-119e14.2 |
| chr1_-_21901589 | 0.17 |
ENSDART00000140553
|
frmpd1a
|
FERM and PDZ domain containing 1a |
| chr6_+_23931236 | 0.16 |
ENSDART00000166079
|
gadd45ab
|
growth arrest and DNA-damage-inducible, alpha, b |
| chr9_+_38075851 | 0.15 |
ENSDART00000135314
|
cacnb4a
|
calcium channel, voltage-dependent, beta 4a subunit |
| chr15_-_21155641 | 0.15 |
ENSDART00000061098
ENSDART00000046443 |
A2ML1 (1 of many)
|
si:dkey-105h12.2 |
| chr24_+_41915878 | 0.14 |
ENSDART00000171523
|
TMEM200C
|
transmembrane protein 200C |
| chr1_+_23784905 | 0.14 |
ENSDART00000171951
ENSDART00000188521 ENSDART00000183029 ENSDART00000187183 |
slit2
|
slit homolog 2 (Drosophila) |
| chr14_+_46274611 | 0.13 |
ENSDART00000134363
|
cabp2b
|
calcium binding protein 2b |
| chr1_+_57151926 | 0.13 |
ENSDART00000152628
|
si:ch73-94k4.5
|
si:ch73-94k4.5 |
| chr13_-_8776474 | 0.13 |
ENSDART00000122371
|
STPG4
|
si:ch211-93n23.7 |
| chr11_-_2838699 | 0.12 |
ENSDART00000066189
|
lhfpl5a
|
LHFPL tetraspan subfamily member 5a |
| chr10_+_43039947 | 0.11 |
ENSDART00000193434
|
atg10
|
ATG10 autophagy related 10 homolog (S. cerevisiae) |
| chr5_-_34185115 | 0.10 |
ENSDART00000192771
|
fibcd1
|
fibrinogen C domain containing 1 |
| chr1_+_56900580 | 0.10 |
ENSDART00000152568
|
si:ch211-152f2.3
|
si:ch211-152f2.3 |
| chr3_-_29508959 | 0.10 |
ENSDART00000055408
|
cyth4a
|
cytohesin 4a |
| chr7_-_69983948 | 0.09 |
ENSDART00000185827
|
KCNIP4
|
potassium voltage-gated channel interacting protein 4 |
| chr5_+_45007962 | 0.08 |
ENSDART00000010786
|
dmrt2a
|
doublesex and mab-3 related transcription factor 2a |
| chr20_-_54924593 | 0.08 |
ENSDART00000151522
|
si:dkey-15f23.1
|
si:dkey-15f23.1 |
| chr1_+_56873359 | 0.08 |
ENSDART00000152713
|
si:ch211-152f2.2
|
si:ch211-152f2.2 |
| chr4_+_12292274 | 0.07 |
ENSDART00000061070
ENSDART00000150786 |
mkrn1
|
makorin, ring finger protein, 1 |
| chr21_-_22812417 | 0.06 |
ENSDART00000079151
ENSDART00000112175 |
trpc6a
|
transient receptor potential cation channel, subfamily C, member 6a |
| chr21_+_40092301 | 0.06 |
ENSDART00000145150
|
serpinf2a
|
serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), member 2a |
| chr1_+_57208005 | 0.06 |
ENSDART00000152410
|
si:dkey-27j5.10
|
si:dkey-27j5.10 |
| chr11_-_44030962 | 0.06 |
ENSDART00000171910
|
FP016005.1
|
|
| chr15_+_28116129 | 0.06 |
ENSDART00000185157
|
unc119a
|
unc-119 homolog a (C. elegans) |
| chr13_-_36621926 | 0.05 |
ENSDART00000057155
|
cdkn3
|
cyclin-dependent kinase inhibitor 3 |
| chr18_+_10840071 | 0.05 |
ENSDART00000014496
|
mical3a
|
microtubule associated monooxygenase, calponin and LIM domain containing 3a |
| chr9_-_20372977 | 0.05 |
ENSDART00000113418
|
igsf3
|
immunoglobulin superfamily, member 3 |
| chr1_+_57218301 | 0.04 |
ENSDART00000152476
ENSDART00000166419 |
si:dkey-27j5.8
|
si:dkey-27j5.8 |
| chr14_+_4796168 | 0.04 |
ENSDART00000041468
|
ap1ar
|
adaptor-related protein complex 1 associated regulatory protein |
| chr9_-_33081978 | 0.04 |
ENSDART00000100918
|
zgc:172053
|
zgc:172053 |
| chr15_-_21239416 | 0.04 |
ENSDART00000155787
|
A2ML1 (1 of many)
|
si:dkey-105h12.2 |
| chr13_+_36622100 | 0.04 |
ENSDART00000133198
|
si:ch211-67f24.7
|
si:ch211-67f24.7 |
| chr5_+_9405283 | 0.03 |
ENSDART00000127306
|
ugt2a7
|
UDP glucuronosyltransferase 2 family, polypeptide A7 |
| chr23_-_30960506 | 0.03 |
ENSDART00000142661
|
osbpl2a
|
oxysterol binding protein-like 2a |
| chr18_+_50890749 | 0.02 |
ENSDART00000174109
|
si:ch1073-450f2.1
|
si:ch1073-450f2.1 |
| chr6_-_23931442 | 0.02 |
ENSDART00000160547
|
sec16b
|
SEC16 homolog B, endoplasmic reticulum export factor |
| chr9_-_33062891 | 0.02 |
ENSDART00000161182
|
si:ch211-125e6.5
|
si:ch211-125e6.5 |
| chr1_-_43862638 | 0.02 |
ENSDART00000145044
|
tacr3a
|
tachykinin receptor 3a |
| chr4_+_30363474 | 0.01 |
ENSDART00000168421
|
si:dkey-199m13.7
|
si:dkey-199m13.7 |
| chr2_+_37480669 | 0.01 |
ENSDART00000029801
|
sppl2
|
signal peptide peptidase-like 2 |
| chr10_+_37083654 | 0.01 |
ENSDART00000132162
ENSDART00000029843 |
vezf1a
|
vascular endothelial zinc finger 1a |
| chr2_+_45068366 | 0.01 |
ENSDART00000142175
|
si:dkey-76d14.2
|
si:dkey-76d14.2 |
| chr12_-_46145909 | 0.01 |
ENSDART00000139094
|
zgc:153932
|
zgc:153932 |
| chr1_+_57050899 | 0.01 |
ENSDART00000152601
|
si:ch211-1f22.14
|
si:ch211-1f22.14 |
Network of associatons between targets according to the STRING database.
First level regulatory network of hoxb8a+hoxb8b
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 2.1 | GO:0060092 | regulation of synaptic transmission, glycinergic(GO:0060092) |
| 0.4 | 2.2 | GO:0003210 | cardiac atrium formation(GO:0003210) |
| 0.4 | 1.7 | GO:0015867 | intracellular nucleoside transport(GO:0015859) purine nucleoside transmembrane transport(GO:0015860) purine nucleotide transport(GO:0015865) ATP transport(GO:0015867) purine ribonucleotide transport(GO:0015868) adenine nucleotide transport(GO:0051503) mitochondrial ATP transmembrane transport(GO:1990544) |
| 0.2 | 0.9 | GO:1903964 | monounsaturated fatty acid metabolic process(GO:1903964) monounsaturated fatty acid biosynthetic process(GO:1903966) |
| 0.2 | 0.8 | GO:0010874 | regulation of cholesterol efflux(GO:0010874) |
| 0.2 | 0.9 | GO:0035332 | positive regulation of hippo signaling(GO:0035332) |
| 0.1 | 1.0 | GO:0042983 | amyloid precursor protein biosynthetic process(GO:0042983) regulation of amyloid precursor protein biosynthetic process(GO:0042984) |
| 0.1 | 0.4 | GO:0031630 | regulation of synaptic vesicle fusion to presynaptic membrane(GO:0031630) |
| 0.1 | 0.9 | GO:0098971 | dendritic transport(GO:0098935) anterograde dendritic transport(GO:0098937) anterograde dendritic transport of neurotransmitter receptor complex(GO:0098971) |
| 0.1 | 1.9 | GO:0072554 | endothelial tube morphogenesis(GO:0061154) blood vessel lumenization(GO:0072554) |
| 0.1 | 0.8 | GO:0060394 | negative regulation of pathway-restricted SMAD protein phosphorylation(GO:0060394) |
| 0.1 | 0.3 | GO:0001954 | positive regulation of cell-matrix adhesion(GO:0001954) |
| 0.1 | 0.5 | GO:0070378 | ERK5 cascade(GO:0070375) regulation of ERK5 cascade(GO:0070376) positive regulation of ERK5 cascade(GO:0070378) |
| 0.1 | 0.6 | GO:0005513 | detection of calcium ion(GO:0005513) |
| 0.1 | 0.3 | GO:0045218 | zonula adherens maintenance(GO:0045218) |
| 0.1 | 0.3 | GO:1902175 | regulation of oxidative stress-induced intrinsic apoptotic signaling pathway(GO:1902175) negative regulation of oxidative stress-induced intrinsic apoptotic signaling pathway(GO:1902176) |
| 0.1 | 0.8 | GO:0021521 | ventral spinal cord interneuron specification(GO:0021521) cell fate specification involved in pattern specification(GO:0060573) |
| 0.1 | 0.6 | GO:0034650 | cortisol metabolic process(GO:0034650) |
| 0.1 | 0.3 | GO:0098920 | endocannabinoid signaling pathway(GO:0071926) retrograde trans-synaptic signaling by lipid(GO:0098920) retrograde trans-synaptic signaling by endocannabinoid(GO:0098921) |
| 0.1 | 0.7 | GO:0001682 | tRNA 5'-leader removal(GO:0001682) |
| 0.1 | 0.3 | GO:1901492 | positive regulation of lymphangiogenesis(GO:1901492) |
| 0.1 | 1.2 | GO:0015693 | magnesium ion transport(GO:0015693) |
| 0.1 | 0.9 | GO:2000377 | regulation of reactive oxygen species metabolic process(GO:2000377) |
| 0.1 | 0.4 | GO:0015862 | uridine transport(GO:0015862) pyrimidine nucleoside transport(GO:0015864) |
| 0.1 | 0.4 | GO:0045329 | carnitine biosynthetic process(GO:0045329) |
| 0.1 | 0.5 | GO:1902262 | apoptotic process involved in patterning of blood vessels(GO:1902262) |
| 0.1 | 0.5 | GO:0042438 | melanin biosynthetic process(GO:0042438) |
| 0.1 | 0.2 | GO:0060300 | microglial cell activation(GO:0001774) regulation of cytokine activity(GO:0060300) regulation of receptor binding(GO:1900120) regulation of chemokine activity(GO:1900136) |
| 0.1 | 1.0 | GO:0060088 | auditory receptor cell stereocilium organization(GO:0060088) |
| 0.0 | 0.9 | GO:0071436 | cellular potassium ion homeostasis(GO:0030007) sodium ion export from cell(GO:0036376) sodium ion export(GO:0071436) |
| 0.0 | 0.7 | GO:0040033 | miRNA mediated inhibition of translation(GO:0035278) negative regulation of translation, ncRNA-mediated(GO:0040033) regulation of translation, ncRNA-mediated(GO:0045974) |
| 0.0 | 0.1 | GO:0007414 | axonal defasciculation(GO:0007414) |
| 0.0 | 0.3 | GO:0097477 | spinal cord motor neuron migration(GO:0097476) lateral motor column neuron migration(GO:0097477) |
| 0.0 | 0.3 | GO:0097369 | sodium ion import(GO:0097369) sodium ion import across plasma membrane(GO:0098719) sodium ion import into cell(GO:1990118) |
| 0.0 | 0.6 | GO:0071456 | cellular response to hypoxia(GO:0071456) |
| 0.0 | 0.3 | GO:0001765 | membrane raft assembly(GO:0001765) plasma membrane raft assembly(GO:0044854) plasma membrane raft organization(GO:0044857) caveola assembly(GO:0070836) |
| 0.0 | 0.2 | GO:0071786 | endoplasmic reticulum tubular network organization(GO:0071786) |
| 0.0 | 0.2 | GO:0000185 | activation of MAPKKK activity(GO:0000185) |
| 0.0 | 0.5 | GO:0007568 | aging(GO:0007568) |
| 0.0 | 0.3 | GO:0034626 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.0 | 0.2 | GO:0035999 | tetrahydrofolate interconversion(GO:0035999) |
| 0.0 | 0.3 | GO:0061074 | regulation of neural retina development(GO:0061074) |
| 0.0 | 1.0 | GO:0030866 | cortical actin cytoskeleton organization(GO:0030866) |
| 0.0 | 0.3 | GO:0009409 | response to cold(GO:0009409) |
| 0.0 | 0.3 | GO:0007205 | protein kinase C-activating G-protein coupled receptor signaling pathway(GO:0007205) |
| 0.0 | 0.4 | GO:0006828 | manganese ion transport(GO:0006828) |
| 0.0 | 0.6 | GO:0030514 | negative regulation of BMP signaling pathway(GO:0030514) |
| 0.0 | 0.7 | GO:0043044 | ATP-dependent chromatin remodeling(GO:0043044) |
| 0.0 | 0.2 | GO:0097178 | ruffle assembly(GO:0097178) |
| 0.0 | 0.0 | GO:0010812 | negative regulation of cell-substrate adhesion(GO:0010812) |
| 0.0 | 0.3 | GO:0070831 | basement membrane assembly(GO:0070831) |
| 0.0 | 0.3 | GO:0021884 | forebrain neuron development(GO:0021884) |
| 0.0 | 1.0 | GO:0000381 | regulation of alternative mRNA splicing, via spliceosome(GO:0000381) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.7 | GO:0000172 | ribonuclease MRP complex(GO:0000172) |
| 0.1 | 0.7 | GO:0016586 | RSC complex(GO:0016586) |
| 0.1 | 1.2 | GO:0008250 | oligosaccharyltransferase complex(GO:0008250) |
| 0.1 | 0.6 | GO:0005790 | smooth endoplasmic reticulum(GO:0005790) |
| 0.0 | 0.3 | GO:0005915 | cell-cell adherens junction(GO:0005913) zonula adherens(GO:0005915) |
| 0.0 | 0.2 | GO:0071782 | endoplasmic reticulum tubular network(GO:0071782) endoplasmic reticulum subcompartment(GO:0098827) |
| 0.0 | 1.6 | GO:0030018 | Z disc(GO:0030018) |
| 0.0 | 0.3 | GO:0005605 | basal lamina(GO:0005605) laminin complex(GO:0043256) |
| 0.0 | 0.5 | GO:0031105 | septin ring(GO:0005940) septin complex(GO:0031105) septin cytoskeleton(GO:0032156) |
| 0.0 | 0.8 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.0 | 0.4 | GO:1902711 | GABA receptor complex(GO:1902710) GABA-A receptor complex(GO:1902711) |
| 0.0 | 0.9 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.0 | 0.8 | GO:0034705 | voltage-gated potassium channel complex(GO:0008076) potassium channel complex(GO:0034705) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.7 | GO:0005471 | ATP:ADP antiporter activity(GO:0005471) |
| 0.2 | 0.9 | GO:0032896 | stearoyl-CoA 9-desaturase activity(GO:0004768) acyl-CoA desaturase activity(GO:0016215) palmitoyl-CoA 9-desaturase activity(GO:0032896) |
| 0.2 | 2.2 | GO:0032036 | myosin heavy chain binding(GO:0032036) |
| 0.2 | 0.8 | GO:0070412 | R-SMAD binding(GO:0070412) |
| 0.2 | 1.6 | GO:0031433 | telethonin binding(GO:0031433) FATZ binding(GO:0051373) |
| 0.1 | 2.1 | GO:0015187 | glycine transmembrane transporter activity(GO:0015187) |
| 0.1 | 0.4 | GO:0005006 | epidermal growth factor-activated receptor activity(GO:0005006) |
| 0.1 | 1.0 | GO:0016838 | carbon-oxygen lyase activity, acting on phosphates(GO:0016838) |
| 0.1 | 0.6 | GO:0005219 | ryanodine-sensitive calcium-release channel activity(GO:0005219) calcium-induced calcium release activity(GO:0048763) |
| 0.1 | 0.5 | GO:0030298 | receptor signaling protein tyrosine kinase activator activity(GO:0030298) |
| 0.1 | 0.3 | GO:0070097 | delta-catenin binding(GO:0070097) |
| 0.1 | 0.9 | GO:0005391 | sodium:potassium-exchanging ATPase activity(GO:0005391) |
| 0.1 | 0.6 | GO:0036122 | BMP binding(GO:0036122) |
| 0.1 | 1.2 | GO:0008143 | poly(A) binding(GO:0008143) |
| 0.0 | 0.1 | GO:0019777 | Atg12 transferase activity(GO:0019777) |
| 0.0 | 1.0 | GO:0030247 | pattern binding(GO:0001871) polysaccharide binding(GO:0030247) |
| 0.0 | 0.2 | GO:0004329 | formate-tetrahydrofolate ligase activity(GO:0004329) |
| 0.0 | 0.6 | GO:0015269 | calcium-activated potassium channel activity(GO:0015269) |
| 0.0 | 0.8 | GO:0016854 | racemase and epimerase activity(GO:0016854) |
| 0.0 | 0.8 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.0 | 0.3 | GO:0005451 | monovalent cation:proton antiporter activity(GO:0005451) sodium:proton antiporter activity(GO:0015385) potassium:proton antiporter activity(GO:0015386) |
| 0.0 | 0.9 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.0 | 0.6 | GO:0033764 | steroid dehydrogenase activity, acting on the CH-OH group of donors, NAD or NADP as acceptor(GO:0033764) |
| 0.0 | 0.3 | GO:0008353 | RNA polymerase II carboxy-terminal domain kinase activity(GO:0008353) |
| 0.0 | 0.3 | GO:0008301 | DNA binding, bending(GO:0008301) |
| 0.0 | 0.3 | GO:0102336 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.0 | 0.4 | GO:0015279 | store-operated calcium channel activity(GO:0015279) |
| 0.0 | 0.4 | GO:0005520 | insulin-like growth factor binding(GO:0005520) |
| 0.0 | 0.9 | GO:0004601 | peroxidase activity(GO:0004601) |
| 0.0 | 0.3 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.0 | 0.2 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.0 | 0.7 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.0 | 0.4 | GO:0004890 | GABA-A receptor activity(GO:0004890) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.6 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
| 0.0 | 2.0 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 0.2 | PID IL8 CXCR1 PATHWAY | IL8- and CXCR1-mediated signaling events |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.1 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.1 | 0.6 | REACTOME PECAM1 INTERACTIONS | Genes involved in PECAM1 interactions |
| 0.0 | 0.3 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.0 | 1.7 | REACTOME INTERACTIONS OF VPR WITH HOST CELLULAR PROTEINS | Genes involved in Interactions of Vpr with host cellular proteins |
| 0.0 | 0.6 | REACTOME CGMP EFFECTS | Genes involved in cGMP effects |
| 0.0 | 0.3 | REACTOME HYALURONAN UPTAKE AND DEGRADATION | Genes involved in Hyaluronan uptake and degradation |
| 0.0 | 0.5 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.0 | 0.4 | REACTOME ACTIVATED NOTCH1 TRANSMITS SIGNAL TO THE NUCLEUS | Genes involved in Activated NOTCH1 Transmits Signal to the Nucleus |