Project
PRJNA195909:zebrafish embryo and larva development
Navigation
Downloads
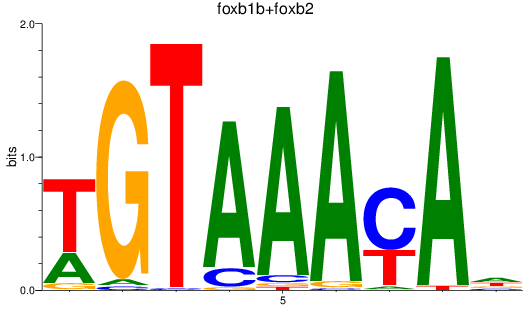
Results for foxb1b+foxb2
Z-value: 2.44
Transcription factors associated with foxb1b+foxb2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
foxb2
|
ENSDARG00000037475 | forkhead box B2 |
|
foxb1b
|
ENSDARG00000053650 | forkhead box B1b |
|
foxb1b
|
ENSDARG00000110408 | forkhead box B1b |
|
foxb1b
|
ENSDARG00000113373 | forkhead box B1b |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| foxb1b | dr11_v1_chr7_+_29461060_29461060 | -0.61 | 8.3e-02 | Click! |
| foxb2 | dr11_v1_chr8_-_38506339_38506339 | -0.32 | 4.0e-01 | Click! |
Activity profile of foxb1b+foxb2 motif
Sorted Z-values of foxb1b+foxb2 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr8_+_37749263 | 4.88 |
ENSDART00000108556
ENSDART00000147942 |
npm2a
|
nucleophosmin/nucleoplasmin, 2a |
| chr11_-_6452444 | 4.59 |
ENSDART00000137879
ENSDART00000134957 ENSDART00000004483 |
larp6b
|
La ribonucleoprotein domain family, member 6b |
| chr9_-_8314028 | 4.58 |
ENSDART00000102739
|
si:ch211-145c1.1
|
si:ch211-145c1.1 |
| chr5_-_33236637 | 4.46 |
ENSDART00000085512
ENSDART00000144694 |
kank1b
|
KN motif and ankyrin repeat domains 1b |
| chr23_+_44236281 | 3.72 |
ENSDART00000149842
|
MEPCE
|
si:ch1073-157b13.1 |
| chr14_+_15155684 | 3.63 |
ENSDART00000167966
|
zgc:158852
|
zgc:158852 |
| chr5_-_17876709 | 3.43 |
ENSDART00000141978
|
si:dkey-112e17.1
|
si:dkey-112e17.1 |
| chr16_-_47381519 | 3.36 |
ENSDART00000032188
ENSDART00000150136 |
si:dkey-256h2.1
|
si:dkey-256h2.1 |
| chr12_+_2648043 | 3.19 |
ENSDART00000082220
|
gdf2
|
growth differentiation factor 2 |
| chr8_-_25771474 | 3.12 |
ENSDART00000193883
|
suv39h1b
|
suppressor of variegation 3-9 homolog 1b |
| chr7_-_37555208 | 3.10 |
ENSDART00000148905
ENSDART00000150229 |
cylda
|
cylindromatosis (turban tumor syndrome), a |
| chr14_-_7409364 | 3.03 |
ENSDART00000036463
|
dnd1
|
DND microRNA-mediated repression inhibitor 1 |
| chr11_+_11120532 | 3.01 |
ENSDART00000026135
ENSDART00000189872 |
ly75
|
lymphocyte antigen 75 |
| chr13_-_33207367 | 2.95 |
ENSDART00000146138
ENSDART00000109667 ENSDART00000182741 |
trip11
|
thyroid hormone receptor interactor 11 |
| chr5_-_68782641 | 2.86 |
ENSDART00000141699
|
mepce
|
methylphosphate capping enzyme |
| chr17_+_24036791 | 2.73 |
ENSDART00000140767
|
b3gnt2b
|
UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 2b |
| chr7_+_46019780 | 2.69 |
ENSDART00000163991
|
ccne1
|
cyclin E1 |
| chr22_-_38819603 | 2.68 |
ENSDART00000104437
|
si:ch211-262h13.5
|
si:ch211-262h13.5 |
| chr1_-_8917902 | 2.67 |
ENSDART00000137900
|
grin2ab
|
glutamate receptor, ionotropic, N-methyl D-aspartate 2A, b |
| chr8_+_23726708 | 2.59 |
ENSDART00000142395
|
mkrn4
|
makorin, ring finger protein, 4 |
| chr20_-_52928541 | 2.51 |
ENSDART00000162812
|
fdft1
|
farnesyl-diphosphate farnesyltransferase 1 |
| chr20_+_42918755 | 2.49 |
ENSDART00000134855
|
efr3bb
|
EFR3 homolog Bb (S. cerevisiae) |
| chr22_-_5171829 | 2.49 |
ENSDART00000140313
|
tnfaip8l1
|
tumor necrosis factor, alpha-induced protein 8-like 1 |
| chr15_+_19991280 | 2.38 |
ENSDART00000186677
|
zgc:112083
|
zgc:112083 |
| chr17_+_44030692 | 2.37 |
ENSDART00000049503
|
peli2
|
pellino E3 ubiquitin protein ligase family member 2 |
| chr25_-_13408760 | 2.33 |
ENSDART00000154445
|
gins3
|
GINS complex subunit 3 |
| chr9_-_27391908 | 2.29 |
ENSDART00000135221
|
nepro
|
nucleolus and neural progenitor protein |
| chr4_-_12781182 | 2.28 |
ENSDART00000058020
|
helb
|
helicase (DNA) B |
| chr9_-_46072805 | 2.27 |
ENSDART00000169682
|
hdac4
|
histone deacetylase 4 |
| chr3_+_14512670 | 2.26 |
ENSDART00000161403
|
rab3db
|
RAB3D, member RAS oncogene family, b |
| chr12_+_38807604 | 2.23 |
ENSDART00000155563
|
abca5
|
ATP-binding cassette, sub-family A (ABC1), member 5 |
| chr22_-_17631675 | 2.22 |
ENSDART00000132565
|
hmha1b
|
histocompatibility (minor) HA-1 b |
| chr3_-_48603471 | 2.22 |
ENSDART00000189027
|
ndel1b
|
nudE neurodevelopment protein 1-like 1b |
| chr1_+_21937201 | 2.20 |
ENSDART00000087729
|
kdm4c
|
lysine (K)-specific demethylase 4C |
| chr4_-_14207471 | 2.17 |
ENSDART00000015134
|
twf1b
|
twinfilin actin-binding protein 1b |
| chr16_-_25233515 | 2.16 |
ENSDART00000058943
|
zgc:110182
|
zgc:110182 |
| chr6_-_41138854 | 2.14 |
ENSDART00000128723
ENSDART00000151055 ENSDART00000132484 |
slc6a22.1
|
solute carrier family 6 member 22, tandem duplicate 1 |
| chr25_+_15994100 | 2.14 |
ENSDART00000144723
|
ppfibp2b
|
PTPRF interacting protein, binding protein 2b (liprin beta 2) |
| chr15_-_8856391 | 2.13 |
ENSDART00000008273
|
rab4b
|
RAB4B, member RAS oncogene family |
| chr7_+_59677273 | 2.11 |
ENSDART00000039535
ENSDART00000132044 |
trmt44
|
tRNA methyltransferase 44 homolog |
| chr20_+_46741074 | 2.11 |
ENSDART00000145294
|
si:ch211-57i17.1
|
si:ch211-57i17.1 |
| chr22_-_5171362 | 2.10 |
ENSDART00000124889
|
tnfaip8l1
|
tumor necrosis factor, alpha-induced protein 8-like 1 |
| chr10_-_6587066 | 2.09 |
ENSDART00000171833
|
chd1
|
chromodomain helicase DNA binding protein 1 |
| chr14_+_30285613 | 2.09 |
ENSDART00000173090
|
mtus1a
|
microtubule associated tumor suppressor 1a |
| chr17_-_1705013 | 2.04 |
ENSDART00000182864
|
CABZ01086293.1
|
|
| chr8_+_23726244 | 1.99 |
ENSDART00000132734
|
mkrn4
|
makorin, ring finger protein, 4 |
| chr23_-_1571682 | 1.97 |
ENSDART00000013635
|
fbxo30b
|
F-box protein 30b |
| chr2_-_4787566 | 1.96 |
ENSDART00000160663
ENSDART00000157808 |
tnk2b
|
tyrosine kinase, non-receptor, 2b |
| chr5_-_50084310 | 1.95 |
ENSDART00000074599
ENSDART00000189970 |
fam172a
|
family with sequence similarity 172, member A |
| chr17_-_30652738 | 1.95 |
ENSDART00000154960
|
sh3yl1
|
SH3 and SYLF domain containing 1 |
| chr20_-_35512932 | 1.95 |
ENSDART00000137690
|
adgrf3b
|
adhesion G protein-coupled receptor F3b |
| chr12_+_18916285 | 1.94 |
ENSDART00000127536
|
cbx7b
|
chromobox homolog 7b |
| chr14_+_50918769 | 1.94 |
ENSDART00000146918
|
rnf44
|
ring finger protein 44 |
| chr21_+_3928947 | 1.93 |
ENSDART00000149777
|
setx
|
senataxin |
| chr22_+_38276024 | 1.91 |
ENSDART00000143792
|
rcor3
|
REST corepressor 3 |
| chr6_-_8466717 | 1.90 |
ENSDART00000151577
ENSDART00000151800 ENSDART00000151227 |
si:dkey-217d24.6
|
si:dkey-217d24.6 |
| chr13_+_43050562 | 1.90 |
ENSDART00000016602
|
cdh23
|
cadherin-related 23 |
| chr25_-_18435481 | 1.90 |
ENSDART00000004771
|
poc1b
|
POC1 centriolar protein B |
| chr19_-_10915898 | 1.89 |
ENSDART00000163179
|
pip5k1aa
|
phosphatidylinositol-4-phosphate 5-kinase, type I, alpha, a |
| chr11_-_25257045 | 1.88 |
ENSDART00000130477
|
snai1a
|
snail family zinc finger 1a |
| chr4_-_14624481 | 1.87 |
ENSDART00000137847
|
plxnb2a
|
plexin b2a |
| chr16_-_26820634 | 1.86 |
ENSDART00000111156
|
pdp1
|
pyruvate dehyrogenase phosphatase catalytic subunit 1 |
| chr17_+_25213229 | 1.86 |
ENSDART00000110451
|
arhgef10
|
Rho guanine nucleotide exchange factor (GEF) 10 |
| chr11_+_2600612 | 1.82 |
ENSDART00000173442
|
dnajc14
|
DnaJ (Hsp40) homolog, subfamily C, member 14 |
| chr2_-_57941037 | 1.82 |
ENSDART00000131420
|
si:dkeyp-68b7.5
|
si:dkeyp-68b7.5 |
| chr2_+_34112100 | 1.81 |
ENSDART00000056666
ENSDART00000146624 |
klhl20
|
kelch-like family member 20 |
| chr13_-_15799391 | 1.80 |
ENSDART00000124688
|
bag5
|
BCL2 associated athanogene 5 |
| chr21_-_22871901 | 1.79 |
ENSDART00000065556
|
polq
|
polymerase (DNA directed), theta |
| chr7_+_33372680 | 1.77 |
ENSDART00000193436
ENSDART00000099988 |
glceb
|
glucuronic acid epimerase b |
| chr19_-_46091497 | 1.77 |
ENSDART00000178772
ENSDART00000167255 |
ptdss1b
si:dkey-108k24.2
|
phosphatidylserine synthase 1b si:dkey-108k24.2 |
| chr8_-_35960987 | 1.76 |
ENSDART00000160503
|
slc15a4
|
solute carrier family 15 (oligopeptide transporter), member 4 |
| chr2_-_389867 | 1.76 |
ENSDART00000004848
|
wrnip1
|
Werner helicase interacting protein 1 |
| chr6_-_39700965 | 1.76 |
ENSDART00000156645
|
espl1
|
extra spindle pole bodies like 1, separase |
| chr9_-_3519717 | 1.74 |
ENSDART00000145043
|
dcaf17
|
ddb1 and cul4 associated factor 17 |
| chr23_+_21978816 | 1.74 |
ENSDART00000087110
|
eif4g3b
|
eukaryotic translation initiation factor 4 gamma, 3b |
| chr25_-_12412704 | 1.73 |
ENSDART00000168275
|
det1
|
DET1, COP1 ubiquitin ligase partner |
| chr5_-_65121747 | 1.72 |
ENSDART00000165556
|
tor2a
|
torsin family 2, member A |
| chr12_-_28537615 | 1.72 |
ENSDART00000067762
|
MYO1D
|
si:ch211-94l19.4 |
| chr22_+_9003090 | 1.71 |
ENSDART00000106414
|
rnh1
|
ribonuclease/angiogenin inhibitor 1 |
| chr9_+_54237100 | 1.70 |
ENSDART00000148928
|
rbm27
|
RNA binding motif protein 27 |
| chr23_+_21978584 | 1.70 |
ENSDART00000145172
|
eif4g3b
|
eukaryotic translation initiation factor 4 gamma, 3b |
| chr20_-_34388324 | 1.69 |
ENSDART00000133593
ENSDART00000136591 |
swt1
|
SWT1 RNA endoribonuclease homolog |
| chr7_-_58178807 | 1.69 |
ENSDART00000188531
|
nsmaf
|
neutral sphingomyelinase (N-SMase) activation associated factor |
| chr9_+_2343096 | 1.68 |
ENSDART00000062292
ENSDART00000191722 ENSDART00000135180 |
atf2
|
activating transcription factor 2 |
| chr4_-_13613148 | 1.67 |
ENSDART00000067164
ENSDART00000111247 |
irf5
|
interferon regulatory factor 5 |
| chr19_-_3821678 | 1.67 |
ENSDART00000169639
|
si:dkey-206d17.12
|
si:dkey-206d17.12 |
| chr16_+_40954481 | 1.66 |
ENSDART00000058587
|
gbp
|
glycogen synthase kinase binding protein |
| chr5_-_41709234 | 1.65 |
ENSDART00000083656
|
atxn2
|
ataxin 2 |
| chr15_-_37589600 | 1.65 |
ENSDART00000154641
|
proser3
|
proline and serine rich 3 |
| chr9_-_32158288 | 1.65 |
ENSDART00000037182
|
ankrd44
|
ankyrin repeat domain 44 |
| chr4_+_6735053 | 1.65 |
ENSDART00000150389
|
tmem168a
|
transmembrane protein 168a |
| chr22_-_34979139 | 1.65 |
ENSDART00000116455
ENSDART00000133537 |
arhgap19
|
Rho GTPase activating protein 19 |
| chr17_+_30591287 | 1.65 |
ENSDART00000154243
|
si:dkey-190l8.2
|
si:dkey-190l8.2 |
| chr16_+_25296389 | 1.65 |
ENSDART00000114528
|
tbc1d31
|
TBC1 domain family, member 31 |
| chr12_-_7234915 | 1.63 |
ENSDART00000048866
|
ipmkb
|
inositol polyphosphate multikinase b |
| chr21_-_13661631 | 1.63 |
ENSDART00000184408
|
pnpla7a
|
patatin-like phospholipase domain containing 7a |
| chr14_-_15990361 | 1.63 |
ENSDART00000168075
|
trim105
|
tripartite motif containing 105 |
| chr23_-_31400792 | 1.63 |
ENSDART00000132736
|
lca5
|
Leber congenital amaurosis 5 |
| chr11_-_25257595 | 1.62 |
ENSDART00000123567
|
snai1a
|
snail family zinc finger 1a |
| chr15_+_19990068 | 1.62 |
ENSDART00000154033
ENSDART00000054428 |
zgc:112083
|
zgc:112083 |
| chr16_-_13612650 | 1.61 |
ENSDART00000080372
|
dbpb
|
D site albumin promoter binding protein b |
| chr16_+_48714048 | 1.61 |
ENSDART00000148709
ENSDART00000150121 |
brd2b
|
bromodomain containing 2b |
| chr20_-_40360571 | 1.61 |
ENSDART00000144768
|
smpdl3a
|
sphingomyelin phosphodiesterase, acid-like 3A |
| chr13_+_48617974 | 1.60 |
ENSDART00000168596
|
si:ch1073-268j14.1
|
si:ch1073-268j14.1 |
| chr8_-_20862443 | 1.58 |
ENSDART00000147267
|
si:ch211-133l5.8
|
si:ch211-133l5.8 |
| chr18_+_3579829 | 1.58 |
ENSDART00000158763
ENSDART00000182850 ENSDART00000162754 ENSDART00000178789 ENSDART00000172656 |
lrch3
|
leucine-rich repeats and calponin homology (CH) domain containing 3 |
| chr20_+_36629173 | 1.58 |
ENSDART00000161241
|
ephx1
|
epoxide hydrolase 1, microsomal (xenobiotic) |
| chr21_+_19547806 | 1.58 |
ENSDART00000159707
ENSDART00000184869 ENSDART00000181321 ENSDART00000058487 ENSDART00000058485 |
rai14
|
retinoic acid induced 14 |
| chr11_+_6295370 | 1.57 |
ENSDART00000139882
|
ranbp3a
|
RAN binding protein 3a |
| chr6_-_25384526 | 1.57 |
ENSDART00000160544
|
PKN2 (1 of many)
|
zgc:153916 |
| chr19_+_30450125 | 1.56 |
ENSDART00000073704
|
si:ch211-215a10.4
|
si:ch211-215a10.4 |
| chr3_-_29910547 | 1.56 |
ENSDART00000151501
|
RUNDC1
|
si:dkey-151m15.5 |
| chr17_-_27200634 | 1.56 |
ENSDART00000185332
|
asap3
|
ArfGAP with SH3 domain, ankyrin repeat and PH domain 3 |
| chr21_+_20901505 | 1.55 |
ENSDART00000132741
|
c7b
|
complement component 7b |
| chr19_+_34230108 | 1.55 |
ENSDART00000141950
|
galnt12
|
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 12 |
| chr16_+_30117798 | 1.55 |
ENSDART00000135723
ENSDART00000000198 |
sema6e
|
sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6E |
| chr20_+_27713210 | 1.54 |
ENSDART00000132222
|
zbtb1
|
zinc finger and BTB domain containing 1 |
| chr8_-_29822527 | 1.54 |
ENSDART00000167487
|
slc20a2
|
solute carrier family 20 (phosphate transporter), member 2 |
| chr25_-_29087925 | 1.54 |
ENSDART00000171758
|
rpp25a
|
ribonuclease P and MRP subunit p25, a |
| chr21_-_13662237 | 1.54 |
ENSDART00000091647
ENSDART00000151547 |
pnpla7a
|
patatin-like phospholipase domain containing 7a |
| chr3_+_19685873 | 1.53 |
ENSDART00000006490
|
tlk2
|
tousled-like kinase 2 |
| chr15_-_25365570 | 1.53 |
ENSDART00000152754
|
cluha
|
clustered mitochondria (cluA/CLU1) homolog a |
| chr11_-_30508843 | 1.52 |
ENSDART00000101667
ENSDART00000179930 |
map4k3a
|
mitogen-activated protein kinase kinase kinase kinase 3a |
| chr14_-_34633960 | 1.52 |
ENSDART00000128869
ENSDART00000179977 |
afap1l1a
|
actin filament associated protein 1-like 1a |
| chr25_-_18948816 | 1.51 |
ENSDART00000091549
|
nt5dc3
|
5'-nucleotidase domain containing 3 |
| chr20_-_44090624 | 1.51 |
ENSDART00000048978
ENSDART00000082283 ENSDART00000082276 |
runx2b
|
runt-related transcription factor 2b |
| chr23_-_24542952 | 1.51 |
ENSDART00000088777
|
atp13a2
|
ATPase 13A2 |
| chr7_+_52712807 | 1.50 |
ENSDART00000174095
ENSDART00000174377 ENSDART00000174061 ENSDART00000174094 ENSDART00000110906 ENSDART00000174071 ENSDART00000174238 |
znf280d
|
zinc finger protein 280D |
| chr8_+_7778770 | 1.49 |
ENSDART00000171325
|
tfe3a
|
transcription factor binding to IGHM enhancer 3a |
| chr3_+_25907266 | 1.49 |
ENSDART00000170324
ENSDART00000192633 |
tom1
|
target of myb1 membrane trafficking protein |
| chr9_-_41025062 | 1.48 |
ENSDART00000002053
|
pms1
|
PMS1 homolog 1, mismatch repair system component |
| chr9_-_6991650 | 1.48 |
ENSDART00000081718
|
slc9a2
|
solute carrier family 9, subfamily A (NHE2, cation proton antiporter 2), member 2 |
| chr8_+_26034623 | 1.48 |
ENSDART00000004521
ENSDART00000142555 |
arih2
|
ariadne homolog 2 (Drosophila) |
| chr1_-_55238610 | 1.47 |
ENSDART00000110818
|
FO704915.1
|
|
| chr16_-_7828838 | 1.47 |
ENSDART00000191434
ENSDART00000108653 |
tcaim
|
T cell activation inhibitor, mitochondrial |
| chr12_+_27061720 | 1.47 |
ENSDART00000153426
|
srcap
|
Snf2-related CREBBP activator protein |
| chr5_+_62340799 | 1.47 |
ENSDART00000074117
|
aspa
|
aspartoacylase |
| chr19_+_40122160 | 1.47 |
ENSDART00000143966
|
si:ch211-173p18.3
|
si:ch211-173p18.3 |
| chr21_-_43398122 | 1.47 |
ENSDART00000050533
|
ccni2
|
cyclin I family, member 2 |
| chr22_+_10713713 | 1.46 |
ENSDART00000122349
|
hiat1b
|
hippocampus abundant transcript 1b |
| chr18_-_13121983 | 1.46 |
ENSDART00000092648
|
rxylt1
|
ribitol xylosyltransferase 1 |
| chr11_+_15878343 | 1.46 |
ENSDART00000167191
ENSDART00000171862 ENSDART00000163992 ENSDART00000170065 |
pank4
|
pantothenate kinase 4 |
| chr16_+_42667560 | 1.46 |
ENSDART00000023452
|
dpy19l1l
|
dpy-19-like 1, like (H. sapiens) |
| chr9_-_41024438 | 1.46 |
ENSDART00000157398
|
pms1
|
PMS1 homolog 1, mismatch repair system component |
| chr1_+_29766725 | 1.45 |
ENSDART00000054064
|
zc3h13
|
zinc finger CCCH-type containing 13 |
| chr5_+_44944778 | 1.45 |
ENSDART00000130428
ENSDART00000044361 ENSDART00000128825 ENSDART00000124637 ENSDART00000126066 ENSDART00000177635 |
dmrt1
|
doublesex and mab-3 related transcription factor 1 |
| chr5_+_61944453 | 1.45 |
ENSDART00000134344
|
si:dkeyp-117b8.4
|
si:dkeyp-117b8.4 |
| chr20_-_211920 | 1.43 |
ENSDART00000104790
|
znf292b
|
zinc finger protein 292b |
| chr5_-_66749535 | 1.43 |
ENSDART00000132183
|
kat5b
|
K(lysine) acetyltransferase 5b |
| chr5_-_18961694 | 1.43 |
ENSDART00000142531
ENSDART00000090521 |
ankle2
|
ankyrin repeat and LEM domain containing 2 |
| chr25_+_33046060 | 1.42 |
ENSDART00000165345
|
tln2b
|
talin 2b |
| chr15_-_1485086 | 1.42 |
ENSDART00000191651
|
si:dkeyp-97b10.3
|
si:dkeyp-97b10.3 |
| chr14_-_32824380 | 1.42 |
ENSDART00000172791
ENSDART00000105745 |
inppl1b
|
inositol polyphosphate phosphatase-like 1b |
| chr19_+_42227400 | 1.42 |
ENSDART00000131574
ENSDART00000135436 |
jtb
|
jumping translocation breakpoint |
| chr11_+_42765963 | 1.42 |
ENSDART00000156080
ENSDART00000179888 |
tdrd3
|
tudor domain containing 3 |
| chr16_-_32975951 | 1.42 |
ENSDART00000101969
ENSDART00000175149 |
me1
|
malic enzyme 1, NADP(+)-dependent, cytosolic |
| chr7_+_40094081 | 1.41 |
ENSDART00000186054
|
si:ch73-174h16.4
|
si:ch73-174h16.4 |
| chr16_+_19029297 | 1.41 |
ENSDART00000115263
ENSDART00000114954 |
rapgef5b
|
Rap guanine nucleotide exchange factor (GEF) 5b |
| chr23_+_19594608 | 1.41 |
ENSDART00000134865
|
slmapb
|
sarcolemma associated protein b |
| chr15_-_1590858 | 1.41 |
ENSDART00000081875
|
nnr
|
nanor |
| chr5_-_68058168 | 1.40 |
ENSDART00000177026
|
rnf167
|
ring finger protein 167 |
| chr16_-_30847192 | 1.40 |
ENSDART00000191106
ENSDART00000128158 |
ptk2ab
|
protein tyrosine kinase 2ab |
| chr9_-_30494727 | 1.40 |
ENSDART00000148729
|
si:dkey-229b18.3
|
si:dkey-229b18.3 |
| chr21_-_43457554 | 1.40 |
ENSDART00000085039
|
stk26
|
serine/threonine protein kinase 26 |
| chr11_-_14102131 | 1.40 |
ENSDART00000085158
ENSDART00000191962 |
tmem259
|
transmembrane protein 259 |
| chr7_+_15446229 | 1.40 |
ENSDART00000046542
|
igf1rb
|
insulin-like growth factor 1b receptor |
| chr1_-_49947290 | 1.40 |
ENSDART00000141476
|
sgms2
|
sphingomyelin synthase 2 |
| chr7_-_58178980 | 1.40 |
ENSDART00000073635
|
nsmaf
|
neutral sphingomyelinase (N-SMase) activation associated factor |
| chr3_+_13929860 | 1.40 |
ENSDART00000164179
|
syce2
|
synaptonemal complex central element protein 2 |
| chr20_-_9428021 | 1.39 |
ENSDART00000025330
|
rdh14b
|
retinol dehydrogenase 14b |
| chr7_-_46019756 | 1.39 |
ENSDART00000162583
|
zgc:162297
|
zgc:162297 |
| chr1_-_23370395 | 1.38 |
ENSDART00000143014
ENSDART00000126785 ENSDART00000159138 |
pds5a
|
PDS5 cohesin associated factor A |
| chr2_-_37462462 | 1.38 |
ENSDART00000145896
|
si:dkey-57k2.7
|
si:dkey-57k2.7 |
| chr23_+_25172976 | 1.38 |
ENSDART00000140789
|
si:dkey-151g10.3
|
si:dkey-151g10.3 |
| chr20_-_7582936 | 1.38 |
ENSDART00000083890
|
usp24
|
ubiquitin specific peptidase 24 |
| chr13_+_29925397 | 1.38 |
ENSDART00000123482
|
cuedc2
|
CUE domain containing 2 |
| chr10_-_39283883 | 1.37 |
ENSDART00000023831
|
cry5
|
cryptochrome circadian clock 5 |
| chr13_+_40815012 | 1.37 |
ENSDART00000016960
|
prkg1a
|
protein kinase, cGMP-dependent, type Ia |
| chr17_-_9962578 | 1.37 |
ENSDART00000021942
|
eapp
|
e2f-associated phosphoprotein |
| chr3_+_29476085 | 1.36 |
ENSDART00000184495
ENSDART00000181058 |
fam83fa
|
family with sequence similarity 83, member Fa |
| chr8_-_4031121 | 1.36 |
ENSDART00000169474
ENSDART00000163754 |
mtmr3
|
myotubularin related protein 3 |
| chr10_+_43406796 | 1.35 |
ENSDART00000184337
ENSDART00000183034 ENSDART00000180623 ENSDART00000132486 |
rasa1b
|
RAS p21 protein activator (GTPase activating protein) 1b |
| chr21_+_13387965 | 1.34 |
ENSDART00000134347
|
zgc:113162
|
zgc:113162 |
| chr9_+_33154841 | 1.34 |
ENSDART00000132465
|
dopey2
|
dopey family member 2 |
| chr20_+_27712714 | 1.33 |
ENSDART00000008306
|
zbtb1
|
zinc finger and BTB domain containing 1 |
| chr22_-_11054244 | 1.33 |
ENSDART00000105823
|
insrb
|
insulin receptor b |
| chr1_-_23268013 | 1.32 |
ENSDART00000146575
|
rfc1
|
replication factor C (activator 1) 1 |
| chr13_+_36958086 | 1.32 |
ENSDART00000024386
|
frmd6
|
FERM domain containing 6 |
| chr20_+_30378803 | 1.32 |
ENSDART00000148242
ENSDART00000169140 ENSDART00000062441 |
rnaseh1
|
ribonuclease H1 |
| chr9_+_28140089 | 1.31 |
ENSDART00000046880
|
plekhm3
|
pleckstrin homology domain containing, family M, member 3 |
| chr6_-_54348568 | 1.30 |
ENSDART00000156501
|
zgc:101577
|
zgc:101577 |
| chr10_+_41939963 | 1.30 |
ENSDART00000126248
|
tmem120b
|
transmembrane protein 120B |
| chr23_-_17003533 | 1.30 |
ENSDART00000080545
|
dnmt3bb.2
|
DNA (cytosine-5-)-methyltransferase 3 beta, duplicate b.2 |
| chr19_+_43669122 | 1.30 |
ENSDART00000139151
|
si:ch211-193k19.1
|
si:ch211-193k19.1 |
| chr5_-_25236340 | 1.30 |
ENSDART00000162774
|
abca2
|
ATP-binding cassette, sub-family A (ABC1), member 2 |
| chr3_-_26191960 | 1.29 |
ENSDART00000113843
|
ypel3
|
yippee-like 3 |
| chr19_-_23249822 | 1.29 |
ENSDART00000140665
|
grb10a
|
growth factor receptor-bound protein 10a |
| chr5_+_1965296 | 1.29 |
ENSDART00000156224
|
dhx33
|
DEAH (Asp-Glu-Ala-His) box polypeptide 33 |
| chr12_+_23991639 | 1.29 |
ENSDART00000003143
|
psme4b
|
proteasome activator subunit 4b |
| chr9_+_37366973 | 1.27 |
ENSDART00000016370
|
dirc2
|
disrupted in renal carcinoma 2 |
Network of associatons between targets according to the STRING database.
First level regulatory network of foxb1b+foxb2
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 3.1 | GO:0002522 | leukocyte migration involved in immune response(GO:0002522) |
| 1.0 | 2.9 | GO:1904869 | positive regulation of protein localization to nucleus(GO:1900182) regulation of protein localization to Cajal body(GO:1904869) positive regulation of protein localization to Cajal body(GO:1904871) |
| 0.7 | 3.4 | GO:1903722 | regulation of centriole elongation(GO:1903722) |
| 0.7 | 3.3 | GO:0008592 | regulation of Toll signaling pathway(GO:0008592) |
| 0.6 | 3.5 | GO:0035092 | spermatid nucleus differentiation(GO:0007289) sperm chromatin condensation(GO:0035092) spermatogenesis, exchange of chromosomal proteins(GO:0035093) |
| 0.6 | 1.7 | GO:0048917 | posterior lateral line ganglion development(GO:0048917) |
| 0.6 | 2.3 | GO:0006269 | DNA replication, synthesis of RNA primer(GO:0006269) |
| 0.6 | 2.3 | GO:0090299 | regulation of neural crest formation(GO:0090299) |
| 0.5 | 3.2 | GO:1990108 | protein linear deubiquitination(GO:1990108) |
| 0.5 | 3.0 | GO:0060965 | negative regulation of posttranscriptional gene silencing(GO:0060149) negative regulation of gene silencing by miRNA(GO:0060965) negative regulation of gene silencing by RNA(GO:0060967) |
| 0.5 | 1.5 | GO:0018103 | protein C-linked glycosylation(GO:0018103) peptidyl-tryptophan modification(GO:0018211) protein C-linked glycosylation via tryptophan(GO:0018317) protein C-linked glycosylation via 2'-alpha-mannosyl-L-tryptophan(GO:0018406) |
| 0.5 | 1.4 | GO:0030238 | male sex determination(GO:0030238) |
| 0.5 | 1.9 | GO:1904184 | regulation of pyruvate dehydrogenase activity(GO:1904182) positive regulation of pyruvate dehydrogenase activity(GO:1904184) |
| 0.4 | 2.2 | GO:0010591 | regulation of lamellipodium assembly(GO:0010591) |
| 0.4 | 1.7 | GO:0061158 | 3'-UTR-mediated mRNA destabilization(GO:0061158) |
| 0.4 | 1.2 | GO:0002940 | tRNA N2-guanine methylation(GO:0002940) |
| 0.4 | 1.9 | GO:0035678 | neuromast hair cell morphogenesis(GO:0035678) |
| 0.4 | 2.2 | GO:0060052 | neurofilament cytoskeleton organization(GO:0060052) |
| 0.4 | 1.1 | GO:1901223 | negative regulation of NIK/NF-kappaB signaling(GO:1901223) |
| 0.4 | 1.4 | GO:1900120 | regulation of cytokine activity(GO:0060300) regulation of receptor binding(GO:1900120) regulation of chemokine activity(GO:1900136) |
| 0.3 | 1.0 | GO:0032780 | negative regulation of ATPase activity(GO:0032780) |
| 0.3 | 2.3 | GO:1902975 | cell cycle DNA replication initiation(GO:1902292) nuclear cell cycle DNA replication initiation(GO:1902315) mitotic DNA replication initiation(GO:1902975) |
| 0.3 | 2.5 | GO:0045338 | farnesyl diphosphate metabolic process(GO:0045338) |
| 0.3 | 0.9 | GO:0070278 | extracellular matrix constituent secretion(GO:0070278) |
| 0.3 | 0.9 | GO:0018008 | N-terminal protein myristoylation(GO:0006499) N-terminal peptidyl-glycine N-myristoylation(GO:0018008) protein myristoylation(GO:0018377) |
| 0.3 | 1.2 | GO:2000677 | histone H3-K79 methylation(GO:0034729) regulation of transcription regulatory region DNA binding(GO:2000677) |
| 0.3 | 0.9 | GO:0035773 | insulin secretion involved in cellular response to glucose stimulus(GO:0035773) regulation of insulin secretion involved in cellular response to glucose stimulus(GO:0061178) |
| 0.3 | 0.9 | GO:0015760 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 0.3 | 1.1 | GO:0044210 | 'de novo' CTP biosynthetic process(GO:0044210) |
| 0.3 | 2.0 | GO:0031048 | chromatin silencing by small RNA(GO:0031048) |
| 0.3 | 0.8 | GO:0042823 | pyridoxal phosphate metabolic process(GO:0042822) pyridoxal phosphate biosynthetic process(GO:0042823) |
| 0.3 | 1.7 | GO:0010603 | regulation of cytoplasmic mRNA processing body assembly(GO:0010603) |
| 0.3 | 1.9 | GO:0006369 | termination of RNA polymerase II transcription(GO:0006369) |
| 0.3 | 1.3 | GO:0032776 | DNA methylation on cytosine(GO:0032776) |
| 0.3 | 0.8 | GO:0001514 | selenocysteine incorporation(GO:0001514) translational readthrough(GO:0006451) |
| 0.2 | 1.7 | GO:0045943 | positive regulation of transcription from RNA polymerase I promoter(GO:0045943) |
| 0.2 | 1.7 | GO:0048069 | eye pigmentation(GO:0048069) |
| 0.2 | 3.9 | GO:0035268 | protein mannosylation(GO:0035268) mannosylation(GO:0097502) |
| 0.2 | 1.5 | GO:0080009 | mRNA methylation(GO:0080009) |
| 0.2 | 2.4 | GO:0006108 | malate metabolic process(GO:0006108) |
| 0.2 | 0.7 | GO:0045191 | histone H4-K20 methylation(GO:0034770) histone H4-K20 trimethylation(GO:0034773) regulation of isotype switching(GO:0045191) positive regulation of isotype switching(GO:0045830) |
| 0.2 | 0.9 | GO:0046168 | NADH oxidation(GO:0006116) glycerol-3-phosphate catabolic process(GO:0046168) |
| 0.2 | 0.7 | GO:1903646 | regulation of protein folding(GO:1903332) positive regulation of protein folding(GO:1903334) regulation of chaperone-mediated protein folding(GO:1903644) positive regulation of chaperone-mediated protein folding(GO:1903646) |
| 0.2 | 2.3 | GO:0006685 | sphingomyelin catabolic process(GO:0006685) |
| 0.2 | 1.6 | GO:0032287 | peripheral nervous system myelin maintenance(GO:0032287) |
| 0.2 | 1.3 | GO:0043137 | DNA replication, removal of RNA primer(GO:0043137) |
| 0.2 | 0.9 | GO:0006750 | glutathione biosynthetic process(GO:0006750) |
| 0.2 | 1.9 | GO:0007084 | mitotic nuclear envelope reassembly(GO:0007084) |
| 0.2 | 1.0 | GO:0071596 | ubiquitin-dependent protein catabolic process via the N-end rule pathway(GO:0071596) |
| 0.2 | 0.8 | GO:0051876 | pigment granule dispersal(GO:0051876) |
| 0.2 | 0.6 | GO:0070589 | cell wall biogenesis(GO:0042546) cell wall macromolecule biosynthetic process(GO:0044038) cellular component macromolecule biosynthetic process(GO:0070589) |
| 0.2 | 1.0 | GO:0010867 | regulation of triglyceride biosynthetic process(GO:0010866) positive regulation of triglyceride biosynthetic process(GO:0010867) |
| 0.2 | 1.2 | GO:1900028 | wound healing, spreading of epidermal cells(GO:0035313) negative regulation of ruffle assembly(GO:1900028) |
| 0.2 | 1.8 | GO:0006857 | oligopeptide transport(GO:0006857) |
| 0.2 | 1.0 | GO:0051972 | regulation of telomerase activity(GO:0051972) |
| 0.2 | 1.2 | GO:0010735 | positive regulation of transcription via serum response element binding(GO:0010735) |
| 0.2 | 1.5 | GO:0001672 | regulation of chromatin assembly or disassembly(GO:0001672) |
| 0.2 | 0.7 | GO:1990167 | protein K27-linked deubiquitination(GO:1990167) |
| 0.2 | 1.8 | GO:0006659 | phosphatidylserine biosynthetic process(GO:0006659) |
| 0.2 | 0.2 | GO:0071421 | manganese ion transmembrane transport(GO:0071421) |
| 0.2 | 0.9 | GO:0051659 | maintenance of mitochondrion location(GO:0051659) |
| 0.2 | 1.5 | GO:0001682 | tRNA 5'-leader removal(GO:0001682) |
| 0.2 | 3.5 | GO:0008078 | mesodermal cell migration(GO:0008078) |
| 0.2 | 1.5 | GO:0006971 | hypotonic response(GO:0006971) hypotonic salinity response(GO:0042539) |
| 0.2 | 2.7 | GO:0051897 | positive regulation of protein kinase B signaling(GO:0051897) |
| 0.2 | 0.6 | GO:0010807 | regulation of synaptic vesicle priming(GO:0010807) |
| 0.2 | 1.1 | GO:0030728 | ovulation(GO:0030728) |
| 0.2 | 0.6 | GO:0019532 | oxalate transport(GO:0019532) |
| 0.2 | 1.1 | GO:0070584 | mitochondrion morphogenesis(GO:0070584) |
| 0.2 | 3.9 | GO:0006884 | cell volume homeostasis(GO:0006884) |
| 0.2 | 0.6 | GO:0019287 | isopentenyl diphosphate biosynthetic process, mevalonate pathway(GO:0019287) |
| 0.2 | 1.1 | GO:0006556 | S-adenosylmethionine biosynthetic process(GO:0006556) |
| 0.1 | 1.2 | GO:0031113 | regulation of microtubule polymerization(GO:0031113) |
| 0.1 | 1.8 | GO:0030202 | heparin metabolic process(GO:0030202) heparin biosynthetic process(GO:0030210) |
| 0.1 | 0.4 | GO:0090134 | mesendoderm migration(GO:0090133) cell migration involved in mesendoderm migration(GO:0090134) |
| 0.1 | 1.0 | GO:0000290 | deadenylation-dependent decapping of nuclear-transcribed mRNA(GO:0000290) |
| 0.1 | 1.2 | GO:0070816 | phosphorylation of RNA polymerase II C-terminal domain(GO:0070816) |
| 0.1 | 0.7 | GO:0034427 | nuclear-transcribed mRNA catabolic process, exonucleolytic, 3'-5'(GO:0034427) |
| 0.1 | 0.4 | GO:0048785 | hatching gland development(GO:0048785) |
| 0.1 | 1.0 | GO:0046716 | muscle cell cellular homeostasis(GO:0046716) |
| 0.1 | 2.5 | GO:0060079 | excitatory postsynaptic potential(GO:0060079) |
| 0.1 | 1.0 | GO:0046958 | nonassociative learning(GO:0046958) habituation(GO:0046959) |
| 0.1 | 0.6 | GO:1904292 | regulation of ERAD pathway(GO:1904292) |
| 0.1 | 2.9 | GO:0006298 | mismatch repair(GO:0006298) |
| 0.1 | 1.4 | GO:0070307 | lens fiber cell development(GO:0070307) lens fiber cell morphogenesis(GO:0070309) |
| 0.1 | 1.5 | GO:0048312 | intracellular distribution of mitochondria(GO:0048312) |
| 0.1 | 2.3 | GO:0045747 | positive regulation of Notch signaling pathway(GO:0045747) |
| 0.1 | 0.6 | GO:0030242 | pexophagy(GO:0030242) |
| 0.1 | 0.5 | GO:0061010 | gall bladder development(GO:0061010) |
| 0.1 | 0.5 | GO:1903059 | regulation of lipoprotein metabolic process(GO:0050746) regulation of protein lipidation(GO:1903059) |
| 0.1 | 1.7 | GO:0046627 | negative regulation of insulin receptor signaling pathway(GO:0046627) negative regulation of cellular response to insulin stimulus(GO:1900077) |
| 0.1 | 0.7 | GO:0048755 | branching morphogenesis of a nerve(GO:0048755) |
| 0.1 | 1.5 | GO:0030311 | poly-N-acetyllactosamine metabolic process(GO:0030309) poly-N-acetyllactosamine biosynthetic process(GO:0030311) |
| 0.1 | 1.5 | GO:0045670 | regulation of osteoclast differentiation(GO:0045670) |
| 0.1 | 0.3 | GO:1990168 | protein K29-linked deubiquitination(GO:0035523) protein K33-linked deubiquitination(GO:1990168) |
| 0.1 | 1.1 | GO:0061709 | reticulophagy(GO:0061709) |
| 0.1 | 0.4 | GO:0033345 | asparagine catabolic process(GO:0006530) asparagine catabolic process via L-aspartate(GO:0033345) |
| 0.1 | 0.7 | GO:0070650 | actin filament bundle distribution(GO:0070650) |
| 0.1 | 2.0 | GO:0016558 | protein import into peroxisome matrix(GO:0016558) |
| 0.1 | 1.2 | GO:0070301 | cellular response to hydrogen peroxide(GO:0070301) |
| 0.1 | 0.6 | GO:0003272 | endocardial cushion formation(GO:0003272) |
| 0.1 | 0.4 | GO:0046951 | ketone body biosynthetic process(GO:0046951) |
| 0.1 | 4.8 | GO:0032784 | regulation of DNA-templated transcription, elongation(GO:0032784) |
| 0.1 | 0.4 | GO:1902373 | negative regulation of RNA catabolic process(GO:1902369) negative regulation of mRNA catabolic process(GO:1902373) regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000622) negative regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000623) |
| 0.1 | 1.5 | GO:0015937 | coenzyme A biosynthetic process(GO:0015937) |
| 0.1 | 1.9 | GO:0022011 | myelination in peripheral nervous system(GO:0022011) |
| 0.1 | 0.8 | GO:0033206 | meiotic cytokinesis(GO:0033206) polar body extrusion after meiotic divisions(GO:0040038) |
| 0.1 | 1.2 | GO:0003139 | secondary heart field specification(GO:0003139) |
| 0.1 | 2.2 | GO:0060287 | epithelial cilium movement involved in determination of left/right asymmetry(GO:0060287) |
| 0.1 | 0.7 | GO:0035627 | ceramide transport(GO:0035627) |
| 0.1 | 3.1 | GO:0030488 | tRNA methylation(GO:0030488) |
| 0.1 | 5.3 | GO:0000281 | mitotic cytokinesis(GO:0000281) |
| 0.1 | 2.0 | GO:0021551 | central nervous system morphogenesis(GO:0021551) |
| 0.1 | 1.2 | GO:0019730 | antimicrobial humoral response(GO:0019730) |
| 0.1 | 0.4 | GO:0002495 | antigen processing and presentation of peptide antigen via MHC class II(GO:0002495) |
| 0.1 | 0.6 | GO:0006621 | protein retention in ER lumen(GO:0006621) |
| 0.1 | 0.4 | GO:0009794 | regulation of mitotic cell cycle, embryonic(GO:0009794) mitotic cell cycle, embryonic(GO:0045448) |
| 0.1 | 0.3 | GO:0099553 | trans-synaptic signaling by lipid, modulating synaptic transmission(GO:0099552) trans-synaptic signaling by endocannabinoid, modulating synaptic transmission(GO:0099553) |
| 0.1 | 1.0 | GO:0071479 | cellular response to ionizing radiation(GO:0071479) |
| 0.1 | 3.5 | GO:0032007 | negative regulation of TOR signaling(GO:0032007) |
| 0.1 | 0.7 | GO:0031063 | regulation of histone deacetylation(GO:0031063) |
| 0.1 | 1.4 | GO:1902307 | positive regulation of sodium ion transport(GO:0010765) positive regulation of sodium ion transmembrane transport(GO:1902307) regulation of potassium ion import(GO:1903286) positive regulation of potassium ion import(GO:1903288) positive regulation of sodium ion transmembrane transporter activity(GO:2000651) |
| 0.1 | 1.6 | GO:0032958 | inositol phosphate biosynthetic process(GO:0032958) |
| 0.1 | 0.6 | GO:0097340 | inhibition of cysteine-type endopeptidase activity(GO:0097340) zymogen inhibition(GO:0097341) |
| 0.1 | 1.1 | GO:0007130 | synaptonemal complex assembly(GO:0007130) synaptonemal complex organization(GO:0070193) |
| 0.1 | 1.5 | GO:0035435 | phosphate ion transmembrane transport(GO:0035435) |
| 0.1 | 0.8 | GO:0003160 | endocardium morphogenesis(GO:0003160) |
| 0.1 | 0.3 | GO:0030970 | retrograde protein transport, ER to cytosol(GO:0030970) |
| 0.1 | 1.8 | GO:0051307 | meiotic chromosome separation(GO:0051307) |
| 0.1 | 0.6 | GO:0071578 | zinc II ion transmembrane import(GO:0071578) |
| 0.1 | 9.2 | GO:0006338 | chromatin remodeling(GO:0006338) |
| 0.1 | 4.2 | GO:0034968 | histone lysine methylation(GO:0034968) |
| 0.1 | 1.1 | GO:0018216 | peptidyl-arginine methylation(GO:0018216) |
| 0.1 | 0.4 | GO:0000395 | mRNA 5'-splice site recognition(GO:0000395) |
| 0.1 | 1.2 | GO:0086010 | membrane depolarization during action potential(GO:0086010) |
| 0.1 | 1.9 | GO:0034472 | snRNA 3'-end processing(GO:0034472) |
| 0.1 | 1.4 | GO:0008156 | negative regulation of DNA replication(GO:0008156) |
| 0.1 | 1.1 | GO:0006003 | fructose 2,6-bisphosphate metabolic process(GO:0006003) |
| 0.1 | 0.7 | GO:0071941 | nitrogen cycle metabolic process(GO:0071941) |
| 0.1 | 3.4 | GO:1904029 | regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0000079) regulation of cyclin-dependent protein kinase activity(GO:1904029) |
| 0.1 | 0.3 | GO:0070166 | enamel mineralization(GO:0070166) |
| 0.1 | 0.7 | GO:0006622 | protein targeting to lysosome(GO:0006622) |
| 0.1 | 1.4 | GO:0043153 | entrainment of circadian clock by photoperiod(GO:0043153) |
| 0.1 | 0.4 | GO:2001256 | regulation of store-operated calcium entry(GO:2001256) |
| 0.1 | 1.7 | GO:0007131 | reciprocal meiotic recombination(GO:0007131) |
| 0.1 | 0.2 | GO:0032957 | inositol trisphosphate metabolic process(GO:0032957) inositol phosphorylation(GO:0052746) |
| 0.1 | 0.8 | GO:0033198 | response to ATP(GO:0033198) |
| 0.1 | 1.0 | GO:0007031 | peroxisome organization(GO:0007031) |
| 0.1 | 1.0 | GO:0006368 | transcription elongation from RNA polymerase II promoter(GO:0006368) |
| 0.1 | 1.6 | GO:0019835 | cytolysis(GO:0019835) |
| 0.1 | 0.8 | GO:0099560 | synaptic membrane adhesion(GO:0099560) |
| 0.1 | 0.8 | GO:0032508 | DNA geometric change(GO:0032392) DNA duplex unwinding(GO:0032508) |
| 0.1 | 1.8 | GO:0050821 | protein stabilization(GO:0050821) |
| 0.1 | 0.9 | GO:0035675 | neuromast hair cell development(GO:0035675) |
| 0.1 | 0.8 | GO:0045022 | early endosome to late endosome transport(GO:0045022) vesicle-mediated transport between endosomal compartments(GO:0098927) |
| 0.1 | 1.0 | GO:0002098 | tRNA wobble uridine modification(GO:0002098) |
| 0.1 | 2.9 | GO:1901800 | positive regulation of proteasomal ubiquitin-dependent protein catabolic process(GO:0032436) positive regulation of proteasomal protein catabolic process(GO:1901800) |
| 0.1 | 1.6 | GO:0042073 | intraciliary transport(GO:0042073) |
| 0.1 | 0.7 | GO:0000479 | endonucleolytic cleavage involved in rRNA processing(GO:0000478) endonucleolytic cleavage of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000479) |
| 0.1 | 1.7 | GO:0051085 | chaperone mediated protein folding requiring cofactor(GO:0051085) |
| 0.0 | 0.9 | GO:0043949 | regulation of cAMP-mediated signaling(GO:0043949) |
| 0.0 | 1.6 | GO:0009636 | response to toxic substance(GO:0009636) |
| 0.0 | 0.2 | GO:0032790 | ribosome disassembly(GO:0032790) |
| 0.0 | 0.3 | GO:0017187 | peptidyl-glutamic acid carboxylation(GO:0017187) protein carboxylation(GO:0018214) |
| 0.0 | 1.2 | GO:0071456 | cellular response to hypoxia(GO:0071456) |
| 0.0 | 0.8 | GO:0016082 | synaptic vesicle priming(GO:0016082) |
| 0.0 | 0.8 | GO:0006998 | nuclear envelope organization(GO:0006998) |
| 0.0 | 1.2 | GO:0045494 | photoreceptor cell maintenance(GO:0045494) |
| 0.0 | 0.6 | GO:0098734 | protein depalmitoylation(GO:0002084) macromolecule depalmitoylation(GO:0098734) |
| 0.0 | 0.6 | GO:0016199 | axon midline choice point recognition(GO:0016199) |
| 0.0 | 1.8 | GO:0007029 | endoplasmic reticulum organization(GO:0007029) |
| 0.0 | 3.8 | GO:0032272 | negative regulation of actin filament polymerization(GO:0030837) negative regulation of protein polymerization(GO:0032272) |
| 0.0 | 0.6 | GO:0036353 | histone H2A monoubiquitination(GO:0035518) histone H2A-K119 monoubiquitination(GO:0036353) negative regulation of intrinsic apoptotic signaling pathway by p53 class mediator(GO:1902254) |
| 0.0 | 1.3 | GO:0032482 | Rab protein signal transduction(GO:0032482) |
| 0.0 | 2.1 | GO:0046856 | phosphatidylinositol dephosphorylation(GO:0046856) |
| 0.0 | 0.4 | GO:0006658 | phosphatidylserine metabolic process(GO:0006658) |
| 0.0 | 0.4 | GO:0048172 | regulation of short-term neuronal synaptic plasticity(GO:0048172) |
| 0.0 | 0.3 | GO:0045905 | positive regulation of translational termination(GO:0045905) |
| 0.0 | 1.3 | GO:0060037 | pharyngeal system development(GO:0060037) |
| 0.0 | 1.0 | GO:0035329 | hippo signaling(GO:0035329) |
| 0.0 | 1.0 | GO:0048264 | determination of ventral identity(GO:0048264) |
| 0.0 | 1.0 | GO:0043249 | erythrocyte maturation(GO:0043249) |
| 0.0 | 3.5 | GO:0007030 | Golgi organization(GO:0007030) |
| 0.0 | 1.0 | GO:0006879 | cellular iron ion homeostasis(GO:0006879) |
| 0.0 | 0.7 | GO:0048026 | positive regulation of mRNA splicing, via spliceosome(GO:0048026) |
| 0.0 | 1.1 | GO:0006400 | tRNA modification(GO:0006400) |
| 0.0 | 0.9 | GO:0043631 | mRNA polyadenylation(GO:0006378) RNA polyadenylation(GO:0043631) |
| 0.0 | 0.7 | GO:0000462 | maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000462) |
| 0.0 | 0.7 | GO:0002574 | thrombocyte differentiation(GO:0002574) |
| 0.0 | 0.2 | GO:0031179 | peptide modification(GO:0031179) |
| 0.0 | 2.1 | GO:0051607 | defense response to virus(GO:0051607) |
| 0.0 | 0.4 | GO:0006611 | protein export from nucleus(GO:0006611) |
| 0.0 | 0.9 | GO:0000154 | rRNA modification(GO:0000154) |
| 0.0 | 2.3 | GO:0031047 | gene silencing by RNA(GO:0031047) |
| 0.0 | 0.2 | GO:0010996 | response to auditory stimulus(GO:0010996) |
| 0.0 | 1.9 | GO:0031146 | SCF-dependent proteasomal ubiquitin-dependent protein catabolic process(GO:0031146) |
| 0.0 | 0.8 | GO:0050870 | positive regulation of homotypic cell-cell adhesion(GO:0034112) positive regulation of T cell activation(GO:0050870) positive regulation of leukocyte cell-cell adhesion(GO:1903039) |
| 0.0 | 3.3 | GO:0031098 | stress-activated protein kinase signaling cascade(GO:0031098) |
| 0.0 | 1.0 | GO:0030261 | chromosome condensation(GO:0030261) |
| 0.0 | 1.2 | GO:0030433 | ER-associated ubiquitin-dependent protein catabolic process(GO:0030433) |
| 0.0 | 0.6 | GO:0035469 | determination of pancreatic left/right asymmetry(GO:0035469) |
| 0.0 | 0.1 | GO:1901492 | positive regulation of lymphangiogenesis(GO:1901492) |
| 0.0 | 0.6 | GO:0006289 | nucleotide-excision repair(GO:0006289) |
| 0.0 | 2.7 | GO:0006413 | translational initiation(GO:0006413) |
| 0.0 | 0.2 | GO:0010824 | regulation of centrosome duplication(GO:0010824) |
| 0.0 | 0.7 | GO:0006891 | intra-Golgi vesicle-mediated transport(GO:0006891) |
| 0.0 | 0.0 | GO:0071514 | genetic imprinting(GO:0071514) |
| 0.0 | 0.4 | GO:0060078 | regulation of postsynaptic membrane potential(GO:0060078) |
| 0.0 | 2.9 | GO:0046488 | phosphatidylinositol metabolic process(GO:0046488) |
| 0.0 | 0.6 | GO:0032402 | melanosome transport(GO:0032402) pigment granule transport(GO:0051904) |
| 0.0 | 0.9 | GO:0006904 | vesicle docking involved in exocytosis(GO:0006904) |
| 0.0 | 2.3 | GO:0090504 | epiboly(GO:0090504) |
| 0.0 | 0.5 | GO:0040001 | establishment of mitotic spindle localization(GO:0040001) |
| 0.0 | 1.8 | GO:0006261 | DNA-dependent DNA replication(GO:0006261) |
| 0.0 | 0.1 | GO:0006043 | glucosamine metabolic process(GO:0006041) glucosamine catabolic process(GO:0006043) |
| 0.0 | 0.5 | GO:0016925 | protein sumoylation(GO:0016925) |
| 0.0 | 1.8 | GO:0016573 | histone acetylation(GO:0016573) |
| 0.0 | 0.3 | GO:0022010 | central nervous system myelination(GO:0022010) axon ensheathment in central nervous system(GO:0032291) |
| 0.0 | 0.5 | GO:0048821 | erythrocyte development(GO:0048821) |
| 0.0 | 0.7 | GO:0018230 | peptidyl-L-cysteine S-palmitoylation(GO:0018230) peptidyl-S-diacylglycerol-L-cysteine biosynthetic process from peptidyl-cysteine(GO:0018231) |
| 0.0 | 0.7 | GO:0035088 | establishment or maintenance of apical/basal cell polarity(GO:0035088) establishment or maintenance of epithelial cell apical/basal polarity(GO:0045197) establishment or maintenance of bipolar cell polarity(GO:0061245) |
| 0.0 | 0.3 | GO:0032465 | regulation of cytokinesis(GO:0032465) |
| 0.0 | 0.3 | GO:0006491 | N-glycan processing(GO:0006491) |
| 0.0 | 1.6 | GO:0016579 | protein deubiquitination(GO:0016579) |
| 0.0 | 0.8 | GO:0048843 | negative regulation of axon extension involved in axon guidance(GO:0048843) negative regulation of axon guidance(GO:1902668) |
| 0.0 | 0.1 | GO:0007032 | endosome organization(GO:0007032) |
| 0.0 | 1.8 | GO:0000209 | protein polyubiquitination(GO:0000209) |
| 0.0 | 1.7 | GO:0090630 | activation of GTPase activity(GO:0090630) |
| 0.0 | 0.1 | GO:0006895 | Golgi to endosome transport(GO:0006895) |
| 0.0 | 3.3 | GO:0006869 | lipid transport(GO:0006869) |
| 0.0 | 0.3 | GO:0007198 | adenylate cyclase-inhibiting serotonin receptor signaling pathway(GO:0007198) |
| 0.0 | 1.4 | GO:0043547 | positive regulation of GTPase activity(GO:0043547) |
| 0.0 | 0.2 | GO:0040019 | positive regulation of embryonic development(GO:0040019) |
| 0.0 | 0.5 | GO:0007098 | centrosome cycle(GO:0007098) |
| 0.0 | 1.0 | GO:0007281 | germ cell development(GO:0007281) |
| 0.0 | 0.8 | GO:0010951 | negative regulation of endopeptidase activity(GO:0010951) |
| 0.0 | 0.1 | GO:0036058 | filtration diaphragm assembly(GO:0036058) slit diaphragm assembly(GO:0036060) |
| 0.0 | 0.4 | GO:0034314 | Arp2/3 complex-mediated actin nucleation(GO:0034314) |
| 0.0 | 0.1 | GO:0009651 | response to salt stress(GO:0009651) |
| 0.0 | 0.4 | GO:0051781 | positive regulation of cell division(GO:0051781) |
| 0.0 | 0.3 | GO:0017121 | phospholipid scrambling(GO:0017121) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 3.5 | GO:1990111 | spermatoproteasome complex(GO:1990111) |
| 0.7 | 2.7 | GO:0097134 | cyclin E1-CDK2 complex(GO:0097134) |
| 0.6 | 2.4 | GO:0043073 | germ cell nucleus(GO:0043073) |
| 0.5 | 2.7 | GO:0005899 | insulin receptor complex(GO:0005899) |
| 0.5 | 3.3 | GO:0000347 | THO complex(GO:0000347) THO complex part of transcription export complex(GO:0000445) |
| 0.4 | 1.5 | GO:0036396 | MIS complex(GO:0036396) mRNA editing complex(GO:0045293) |
| 0.3 | 1.4 | GO:0000801 | central element(GO:0000801) |
| 0.3 | 1.6 | GO:0000811 | GINS complex(GO:0000811) |
| 0.3 | 2.9 | GO:0032300 | mismatch repair complex(GO:0032300) |
| 0.3 | 1.1 | GO:0097268 | cytoophidium(GO:0097268) |
| 0.3 | 1.1 | GO:0048269 | methionine adenosyltransferase complex(GO:0048269) |
| 0.2 | 0.7 | GO:0072380 | TRC complex(GO:0072380) |
| 0.2 | 1.5 | GO:0000172 | ribonuclease MRP complex(GO:0000172) |
| 0.2 | 2.1 | GO:0032593 | insulin-responsive compartment(GO:0032593) |
| 0.2 | 1.0 | GO:0071595 | Nem1-Spo7 phosphatase complex(GO:0071595) |
| 0.2 | 0.9 | GO:0034657 | GID complex(GO:0034657) |
| 0.2 | 1.2 | GO:0000439 | core TFIIH complex(GO:0000439) |
| 0.2 | 2.7 | GO:0017146 | NMDA selective glutamate receptor complex(GO:0017146) |
| 0.2 | 0.7 | GO:0044326 | dendritic spine neck(GO:0044326) dendritic filopodium(GO:1902737) |
| 0.2 | 3.4 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.2 | 0.7 | GO:0030896 | checkpoint clamp complex(GO:0030896) |
| 0.2 | 0.6 | GO:0070044 | synaptobrevin 2-SNAP-25-syntaxin-1a complex(GO:0070044) |
| 0.2 | 2.3 | GO:0035267 | NuA4 histone acetyltransferase complex(GO:0035267) H4/H2A histone acetyltransferase complex(GO:0043189) H4 histone acetyltransferase complex(GO:1902562) |
| 0.2 | 0.9 | GO:0009331 | glycerol-3-phosphate dehydrogenase complex(GO:0009331) |
| 0.2 | 0.5 | GO:0005828 | kinetochore microtubule(GO:0005828) |
| 0.2 | 0.5 | GO:0031213 | RSF complex(GO:0031213) |
| 0.1 | 1.5 | GO:0000812 | Swr1 complex(GO:0000812) |
| 0.1 | 0.4 | GO:0034457 | Mpp10 complex(GO:0034457) |
| 0.1 | 1.4 | GO:0035145 | exon-exon junction complex(GO:0035145) |
| 0.1 | 0.7 | GO:0055087 | Ski complex(GO:0055087) |
| 0.1 | 0.9 | GO:0030897 | HOPS complex(GO:0030897) |
| 0.1 | 1.0 | GO:0000796 | condensin complex(GO:0000796) |
| 0.1 | 0.8 | GO:0070695 | FHF complex(GO:0070695) |
| 0.1 | 0.9 | GO:0030121 | AP-1 adaptor complex(GO:0030121) |
| 0.1 | 0.6 | GO:0071203 | WASH complex(GO:0071203) |
| 0.1 | 1.0 | GO:0042645 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.1 | 2.5 | GO:0035102 | PRC1 complex(GO:0035102) |
| 0.1 | 1.0 | GO:0033588 | Elongator holoenzyme complex(GO:0033588) |
| 0.1 | 1.6 | GO:0005579 | membrane attack complex(GO:0005579) |
| 0.1 | 0.7 | GO:0034388 | Pwp2p-containing subcomplex of 90S preribosome(GO:0034388) |
| 0.1 | 1.0 | GO:0070449 | elongin complex(GO:0070449) |
| 0.1 | 0.6 | GO:0030914 | STAGA complex(GO:0030914) |
| 0.1 | 1.9 | GO:0032420 | stereocilium(GO:0032420) |
| 0.1 | 1.9 | GO:0032039 | integrator complex(GO:0032039) |
| 0.1 | 0.6 | GO:0070847 | core mediator complex(GO:0070847) |
| 0.1 | 2.3 | GO:0000407 | pre-autophagosomal structure(GO:0000407) |
| 0.1 | 0.5 | GO:0033557 | Slx1-Slx4 complex(GO:0033557) |
| 0.1 | 0.8 | GO:0001669 | acrosomal vesicle(GO:0001669) |
| 0.1 | 1.8 | GO:0030014 | CCR4-NOT complex(GO:0030014) |
| 0.1 | 1.5 | GO:0030686 | 90S preribosome(GO:0030686) |
| 0.1 | 1.5 | GO:0031229 | integral component of nuclear inner membrane(GO:0005639) intrinsic component of nuclear inner membrane(GO:0031229) nuclear membrane part(GO:0044453) |
| 0.1 | 0.7 | GO:0032021 | NELF complex(GO:0032021) |
| 0.1 | 0.3 | GO:0017177 | glucosidase II complex(GO:0017177) |
| 0.1 | 2.2 | GO:0080008 | Cul4-RING E3 ubiquitin ligase complex(GO:0080008) |
| 0.1 | 0.3 | GO:0000818 | nuclear MIS12/MIND complex(GO:0000818) |
| 0.1 | 0.9 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.1 | 0.6 | GO:0005952 | cAMP-dependent protein kinase complex(GO:0005952) |
| 0.1 | 3.0 | GO:0005778 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) |
| 0.1 | 0.5 | GO:0016586 | RSC complex(GO:0016586) |
| 0.1 | 1.2 | GO:0030992 | intraciliary transport particle B(GO:0030992) |
| 0.1 | 0.3 | GO:0035032 | phosphatidylinositol 3-kinase complex, class III(GO:0035032) |
| 0.1 | 0.3 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.1 | 0.7 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.1 | 1.0 | GO:0048188 | Set1C/COMPASS complex(GO:0048188) |
| 0.1 | 2.9 | GO:0030496 | midbody(GO:0030496) |
| 0.1 | 0.7 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.1 | 1.7 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
| 0.0 | 0.3 | GO:0031907 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.0 | 0.8 | GO:0032154 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.0 | 3.9 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.0 | 3.5 | GO:0005814 | centriole(GO:0005814) |
| 0.0 | 0.4 | GO:0016272 | prefoldin complex(GO:0016272) |
| 0.0 | 1.0 | GO:0000783 | telomere cap complex(GO:0000782) nuclear telomere cap complex(GO:0000783) nuclear chromosome, telomeric region(GO:0000784) |
| 0.0 | 5.6 | GO:0005925 | focal adhesion(GO:0005925) |
| 0.0 | 2.0 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.0 | 0.3 | GO:0031931 | TORC1 complex(GO:0031931) |
| 0.0 | 1.8 | GO:0072686 | mitotic spindle(GO:0072686) |
| 0.0 | 3.6 | GO:0036464 | cytoplasmic ribonucleoprotein granule(GO:0036464) |
| 0.0 | 1.1 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.0 | 0.5 | GO:0071782 | endoplasmic reticulum tubular network(GO:0071782) endoplasmic reticulum subcompartment(GO:0098827) |
| 0.0 | 10.0 | GO:0005730 | nucleolus(GO:0005730) |
| 0.0 | 4.4 | GO:0031234 | extrinsic component of cytoplasmic side of plasma membrane(GO:0031234) |
| 0.0 | 1.4 | GO:0030175 | filopodium(GO:0030175) |
| 0.0 | 0.4 | GO:0005868 | cytoplasmic dynein complex(GO:0005868) |
| 0.0 | 2.8 | GO:0005819 | spindle(GO:0005819) |
| 0.0 | 0.6 | GO:0042555 | MCM complex(GO:0042555) |
| 0.0 | 0.6 | GO:0070822 | Sin3-type complex(GO:0070822) |
| 0.0 | 1.7 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.0 | 2.7 | GO:0000775 | chromosome, centromeric region(GO:0000775) |
| 0.0 | 2.8 | GO:0043197 | dendritic spine(GO:0043197) neuron spine(GO:0044309) |
| 0.0 | 0.4 | GO:0005956 | protein kinase CK2 complex(GO:0005956) |
| 0.0 | 0.5 | GO:0005885 | Arp2/3 protein complex(GO:0005885) |
| 0.0 | 1.2 | GO:0030864 | cortical actin cytoskeleton(GO:0030864) |
| 0.0 | 0.4 | GO:0005682 | U5 snRNP(GO:0005682) |
| 0.0 | 0.4 | GO:0044545 | NSL complex(GO:0044545) |
| 0.0 | 6.8 | GO:0000151 | ubiquitin ligase complex(GO:0000151) |
| 0.0 | 0.9 | GO:0016514 | SWI/SNF complex(GO:0016514) |
| 0.0 | 0.7 | GO:0098878 | ionotropic glutamate receptor complex(GO:0008328) neurotransmitter receptor complex(GO:0098878) |
| 0.0 | 0.2 | GO:0031080 | nuclear pore outer ring(GO:0031080) |
| 0.0 | 1.2 | GO:0005811 | lipid particle(GO:0005811) |
| 0.0 | 0.3 | GO:0098839 | postsynaptic density membrane(GO:0098839) postsynaptic specialization membrane(GO:0099634) |
| 0.0 | 0.4 | GO:0038201 | TORC2 complex(GO:0031932) TOR complex(GO:0038201) |
| 0.0 | 0.6 | GO:0008305 | integrin complex(GO:0008305) |
| 0.0 | 0.8 | GO:0017053 | transcriptional repressor complex(GO:0017053) |
| 0.0 | 0.7 | GO:0000118 | histone deacetylase complex(GO:0000118) |
| 0.0 | 0.1 | GO:0043514 | interleukin-12 complex(GO:0043514) interleukin-23 complex(GO:0070743) |
| 0.0 | 0.7 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.0 | 1.9 | GO:0005813 | centrosome(GO:0005813) |
| 0.0 | 1.2 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.0 | 0.5 | GO:0030125 | clathrin vesicle coat(GO:0030125) |
| 0.0 | 0.2 | GO:0017119 | Golgi transport complex(GO:0017119) |
| 0.0 | 0.9 | GO:0005930 | axoneme(GO:0005930) ciliary plasm(GO:0097014) |
| 0.0 | 2.0 | GO:0005681 | spliceosomal complex(GO:0005681) |
| 0.0 | 1.3 | GO:0005884 | actin filament(GO:0005884) |
| 0.0 | 0.4 | GO:0032589 | neuron projection membrane(GO:0032589) dendrite membrane(GO:0032590) |
| 0.0 | 0.3 | GO:0009898 | cytoplasmic side of plasma membrane(GO:0009898) |
| 0.0 | 0.8 | GO:0030134 | ER to Golgi transport vesicle(GO:0030134) |
| 0.0 | 1.0 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.0 | 3.9 | GO:1990904 | ribonucleoprotein complex(GO:1990904) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 3.5 | GO:0051059 | NF-kappaB binding(GO:0051059) |
| 0.8 | 2.5 | GO:0004311 | farnesyltranstransferase activity(GO:0004311) |
| 0.7 | 2.1 | GO:0016300 | tRNA (uracil) methyltransferase activity(GO:0016300) |
| 0.6 | 1.9 | GO:0001160 | transcription termination site sequence-specific DNA binding(GO:0001147) transcription termination site DNA binding(GO:0001160) |
| 0.6 | 2.4 | GO:0004471 | malic enzyme activity(GO:0004470) malate dehydrogenase (decarboxylating) (NAD+) activity(GO:0004471) malate dehydrogenase (decarboxylating) (NADP+) activity(GO:0004473) |
| 0.6 | 3.4 | GO:0016715 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced ascorbate as one donor, and incorporation of one atom of oxygen(GO:0016715) |
| 0.5 | 2.7 | GO:0043560 | insulin-activated receptor activity(GO:0005009) insulin receptor substrate binding(GO:0043560) |
| 0.5 | 1.6 | GO:0008440 | inositol-1,4,5-trisphosphate 3-kinase activity(GO:0008440) |
| 0.5 | 1.6 | GO:0033961 | cis-stilbene-oxide hydrolase activity(GO:0033961) |
| 0.5 | 1.9 | GO:0016308 | 1-phosphatidylinositol-4-phosphate 5-kinase activity(GO:0016308) |
| 0.5 | 1.4 | GO:0031701 | angiotensin receptor binding(GO:0031701) |
| 0.5 | 1.9 | GO:0004741 | [pyruvate dehydrogenase (lipoamide)] phosphatase activity(GO:0004741) |
| 0.5 | 1.4 | GO:0003913 | DNA photolyase activity(GO:0003913) |
| 0.4 | 1.8 | GO:0047464 | heparosan-N-sulfate-glucuronate 5-epimerase activity(GO:0047464) |
| 0.4 | 1.2 | GO:0042806 | fucose binding(GO:0042806) |
| 0.4 | 2.0 | GO:0035197 | siRNA binding(GO:0035197) |
| 0.4 | 3.3 | GO:0016922 | ligand-dependent nuclear receptor binding(GO:0016922) |
| 0.4 | 3.5 | GO:0016504 | peptidase activator activity(GO:0016504) |
| 0.3 | 3.8 | GO:0043142 | single-stranded DNA-dependent ATP-dependent DNA helicase activity(GO:0017116) single-stranded DNA-dependent ATPase activity(GO:0043142) |
| 0.3 | 1.7 | GO:1990756 | protein binding, bridging involved in substrate recognition for ubiquitination(GO:1990756) |
| 0.3 | 3.1 | GO:0046974 | histone methyltransferase activity (H3-K9 specific)(GO:0046974) |
| 0.3 | 1.0 | GO:0004736 | pyruvate carboxylase activity(GO:0004736) |
| 0.3 | 1.0 | GO:0032050 | clathrin heavy chain binding(GO:0032050) |
| 0.3 | 1.2 | GO:0004809 | tRNA (guanine-N2-)-methyltransferase activity(GO:0004809) |
| 0.3 | 0.9 | GO:0004379 | glycylpeptide N-tetradecanoyltransferase activity(GO:0004379) |
| 0.3 | 1.2 | GO:0031151 | histone methyltransferase activity (H3-K79 specific)(GO:0031151) |
| 0.3 | 1.5 | GO:0042809 | vitamin D receptor binding(GO:0042809) |
| 0.3 | 0.9 | GO:0061513 | hexose phosphate transmembrane transporter activity(GO:0015119) organophosphate:inorganic phosphate antiporter activity(GO:0015315) hexose-phosphate:inorganic phosphate antiporter activity(GO:0015526) glucose 6-phosphate:inorganic phosphate antiporter activity(GO:0061513) |
| 0.3 | 1.1 | GO:0003883 | CTP synthase activity(GO:0003883) |
| 0.3 | 1.6 | GO:0008427 | calcium-dependent protein kinase inhibitor activity(GO:0008427) calcium-dependent protein kinase regulator activity(GO:0010858) |
| 0.3 | 3.7 | GO:0051864 | histone demethylase activity (H3-K36 specific)(GO:0051864) |
| 0.3 | 2.6 | GO:0052629 | phosphatidylinositol-3,5-bisphosphate 3-phosphatase activity(GO:0052629) |
| 0.3 | 1.5 | GO:0008349 | MAP kinase kinase kinase kinase activity(GO:0008349) |
| 0.2 | 1.0 | GO:0010521 | telomerase inhibitor activity(GO:0010521) |
| 0.2 | 1.5 | GO:0004594 | pantothenate kinase activity(GO:0004594) |
| 0.2 | 0.7 | GO:0042799 | histone methyltransferase activity (H4-K20 specific)(GO:0042799) |
| 0.2 | 0.9 | GO:0005463 | UDP-glucuronic acid transmembrane transporter activity(GO:0005461) UDP-N-acetylgalactosamine transmembrane transporter activity(GO:0005463) |
| 0.2 | 0.9 | GO:0004367 | glycerol-3-phosphate dehydrogenase [NAD+] activity(GO:0004367) |
| 0.2 | 1.4 | GO:0004692 | cGMP-dependent protein kinase activity(GO:0004692) |
| 0.2 | 0.9 | GO:0004652 | polynucleotide adenylyltransferase activity(GO:0004652) |
| 0.2 | 1.5 | GO:0034450 | ubiquitin-ubiquitin ligase activity(GO:0034450) |
| 0.2 | 3.1 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.2 | 1.2 | GO:0043914 | NADPH:sulfur oxidoreductase activity(GO:0043914) |
| 0.2 | 1.1 | GO:0015616 | DNA translocase activity(GO:0015616) |
| 0.2 | 1.3 | GO:0004523 | RNA-DNA hybrid ribonuclease activity(GO:0004523) |
| 0.2 | 0.9 | GO:0000179 | rRNA (adenine-N6,N6-)-dimethyltransferase activity(GO:0000179) |
| 0.2 | 1.5 | GO:0046912 | transferase activity, transferring acyl groups, acyl groups converted into alkyl on transfer(GO:0046912) |
| 0.2 | 1.8 | GO:0000774 | adenyl-nucleotide exchange factor activity(GO:0000774) |
| 0.2 | 1.1 | GO:0004022 | alcohol dehydrogenase (NAD) activity(GO:0004022) |
| 0.2 | 1.1 | GO:0070568 | guanylyltransferase activity(GO:0070568) |
| 0.2 | 0.9 | GO:0046592 | polyamine oxidase activity(GO:0046592) |
| 0.2 | 0.9 | GO:0043295 | glutathione binding(GO:0043295) |
| 0.2 | 2.4 | GO:0004169 | dolichyl-phosphate-mannose-protein mannosyltransferase activity(GO:0004169) |
| 0.2 | 1.2 | GO:0008332 | low voltage-gated calcium channel activity(GO:0008332) |
| 0.2 | 0.7 | GO:0034511 | U3 snoRNA binding(GO:0034511) |
| 0.2 | 2.1 | GO:0005332 | gamma-aminobutyric acid:sodium symporter activity(GO:0005332) |
| 0.2 | 1.0 | GO:0022889 | serine transmembrane transporter activity(GO:0022889) |
| 0.2 | 2.9 | GO:0030983 | mismatched DNA binding(GO:0030983) |
| 0.2 | 2.3 | GO:0032041 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.2 | 2.3 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.2 | 0.5 | GO:0032574 | 5'-3' RNA helicase activity(GO:0032574) |
| 0.2 | 3.9 | GO:0015377 | cation:chloride symporter activity(GO:0015377) potassium:chloride symporter activity(GO:0015379) |
| 0.2 | 1.9 | GO:0005158 | insulin receptor binding(GO:0005158) |
| 0.1 | 0.9 | GO:0004719 | protein-L-isoaspartate (D-aspartate) O-methyltransferase activity(GO:0004719) |
| 0.1 | 2.9 | GO:0008171 | O-methyltransferase activity(GO:0008171) |
| 0.1 | 1.1 | GO:0035241 | protein-arginine omega-N monomethyltransferase activity(GO:0035241) |
| 0.1 | 0.7 | GO:0035620 | ceramide transporter activity(GO:0035620) ceramide 1-phosphate binding(GO:1902387) ceramide 1-phosphate transporter activity(GO:1902388) |
| 0.1 | 3.3 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.1 | 2.4 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.1 | 0.9 | GO:0008603 | cAMP-dependent protein kinase regulator activity(GO:0008603) |
| 0.1 | 1.5 | GO:0015385 | monovalent cation:proton antiporter activity(GO:0005451) sodium:proton antiporter activity(GO:0015385) potassium:proton antiporter activity(GO:0015386) |
| 0.1 | 3.2 | GO:0031267 | small GTPase binding(GO:0031267) |
| 0.1 | 2.7 | GO:0004972 | NMDA glutamate receptor activity(GO:0004972) |
| 0.1 | 1.0 | GO:0050072 | m7G(5')pppN diphosphatase activity(GO:0050072) |
| 0.1 | 1.0 | GO:0004740 | pyruvate dehydrogenase (acetyl-transferring) kinase activity(GO:0004740) |
| 0.1 | 0.8 | GO:0004839 | ubiquitin activating enzyme activity(GO:0004839) |
| 0.1 | 2.2 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.1 | 1.5 | GO:0008532 | N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase activity(GO:0008532) |
| 0.1 | 0.6 | GO:0005483 | soluble NSF attachment protein activity(GO:0005483) |
| 0.1 | 0.4 | GO:0004419 | hydroxymethylglutaryl-CoA lyase activity(GO:0004419) |
| 0.1 | 1.5 | GO:0005315 | inorganic phosphate transmembrane transporter activity(GO:0005315) |
| 0.1 | 1.3 | GO:0003886 | DNA (cytosine-5-)-methyltransferase activity(GO:0003886) |
| 0.1 | 0.7 | GO:0098809 | nitrite reductase activity(GO:0098809) |
| 0.1 | 2.0 | GO:0050321 | tau-protein kinase activity(GO:0050321) |
| 0.1 | 1.4 | GO:0051721 | protein phosphatase 2A binding(GO:0051721) |
| 0.1 | 0.6 | GO:0070628 | proteasome binding(GO:0070628) |
| 0.1 | 1.5 | GO:0000030 | mannosyltransferase activity(GO:0000030) |
| 0.1 | 0.4 | GO:0008798 | beta-aspartyl-peptidase activity(GO:0008798) |
| 0.1 | 0.9 | GO:0030628 | pre-mRNA 3'-splice site binding(GO:0030628) |
| 0.1 | 1.4 | GO:0019869 | chloride channel inhibitor activity(GO:0019869) potassium channel inhibitor activity(GO:0019870) |
| 0.1 | 0.3 | GO:0030626 | U12 snRNA binding(GO:0030626) |
| 0.1 | 0.6 | GO:0010859 | calcium-dependent cysteine-type endopeptidase inhibitor activity(GO:0010859) |
| 0.1 | 1.1 | GO:0003873 | 6-phosphofructo-2-kinase activity(GO:0003873) |
| 0.1 | 1.8 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.1 | 0.7 | GO:0051920 | peroxiredoxin activity(GO:0051920) |
| 0.1 | 5.8 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.1 | 0.3 | GO:0008488 | gamma-glutamyl carboxylase activity(GO:0008488) |
| 0.1 | 2.3 | GO:0070491 | repressing transcription factor binding(GO:0070491) |
| 0.1 | 1.1 | GO:0002039 | p53 binding(GO:0002039) |
| 0.1 | 6.4 | GO:0001227 | transcriptional repressor activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001227) |
| 0.1 | 0.4 | GO:0016312 | inositol bisphosphate phosphatase activity(GO:0016312) |
| 0.1 | 0.5 | GO:0016863 | intramolecular oxidoreductase activity, transposing C=C bonds(GO:0016863) |
| 0.1 | 2.1 | GO:0032266 | phosphatidylinositol-3-phosphate binding(GO:0032266) |
| 0.1 | 1.6 | GO:0000339 | RNA cap binding(GO:0000339) |
| 0.1 | 0.8 | GO:0043495 | protein anchor(GO:0043495) |
| 0.1 | 0.2 | GO:0051766 | inositol tetrakisphosphate 1-kinase activity(GO:0047325) inositol tetrakisphosphate kinase activity(GO:0051765) inositol trisphosphate kinase activity(GO:0051766) inositol-1,3,4-trisphosphate 6-kinase activity(GO:0052725) inositol-1,3,4-trisphosphate 5-kinase activity(GO:0052726) |
| 0.1 | 0.8 | GO:0035381 | extracellular ATP-gated cation channel activity(GO:0004931) ATP-gated ion channel activity(GO:0035381) |
| 0.1 | 0.4 | GO:0019136 | deoxynucleoside kinase activity(GO:0019136) |
| 0.1 | 4.4 | GO:0003678 | DNA helicase activity(GO:0003678) |
| 0.1 | 2.1 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.1 | 0.7 | GO:0005537 | mannose binding(GO:0005537) |
| 0.1 | 0.2 | GO:0051117 | ATPase binding(GO:0051117) |
| 0.1 | 0.4 | GO:0035035 | histone acetyltransferase binding(GO:0035035) |
| 0.1 | 0.2 | GO:0045545 | syndecan binding(GO:0045545) |
| 0.1 | 0.3 | GO:0008502 | melatonin receptor activity(GO:0008502) |
| 0.1 | 0.4 | GO:0008515 | sugar:proton symporter activity(GO:0005351) cation:sugar symporter activity(GO:0005402) sucrose:proton symporter activity(GO:0008506) sucrose transmembrane transporter activity(GO:0008515) disaccharide transmembrane transporter activity(GO:0015154) oligosaccharide transmembrane transporter activity(GO:0015157) |
| 0.1 | 1.7 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.1 | 0.9 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.1 | 0.5 | GO:0043515 | kinetochore binding(GO:0043515) |
| 0.0 | 0.6 | GO:0043014 | alpha-tubulin binding(GO:0043014) |
| 0.0 | 0.2 | GO:0005384 | manganese ion transmembrane transporter activity(GO:0005384) |
| 0.0 | 0.1 | GO:0070016 | armadillo repeat domain binding(GO:0070016) |
| 0.0 | 2.3 | GO:0004521 | endoribonuclease activity(GO:0004521) |
| 0.0 | 1.0 | GO:0031210 | phosphatidylcholine binding(GO:0031210) |
| 0.0 | 0.5 | GO:0061665 | SUMO ligase activity(GO:0061665) |
| 0.0 | 4.8 | GO:0101005 | thiol-dependent ubiquitin-specific protease activity(GO:0004843) thiol-dependent ubiquitinyl hydrolase activity(GO:0036459) ubiquitinyl hydrolase activity(GO:0101005) |
| 0.0 | 1.6 | GO:0030674 | protein binding, bridging(GO:0030674) |
| 0.0 | 3.8 | GO:0030276 | clathrin binding(GO:0030276) |
| 0.0 | 6.6 | GO:0042393 | histone binding(GO:0042393) |
| 0.0 | 1.1 | GO:0004016 | adenylate cyclase activity(GO:0004016) |
| 0.0 | 0.6 | GO:0016861 | intramolecular oxidoreductase activity, interconverting aldoses and ketoses(GO:0016861) |
| 0.0 | 0.1 | GO:1990381 | ubiquitin-specific protease binding(GO:1990381) |
| 0.0 | 0.3 | GO:0099530 | G-protein coupled receptor activity involved in regulation of postsynaptic membrane potential(GO:0099530) |
| 0.0 | 1.5 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
| 0.0 | 0.1 | GO:1902945 | metalloendopeptidase activity involved in amyloid precursor protein catabolic process(GO:1902945) |
| 0.0 | 0.4 | GO:0008568 | microtubule-severing ATPase activity(GO:0008568) |
| 0.0 | 3.7 | GO:0003730 | mRNA 3'-UTR binding(GO:0003730) |
| 0.0 | 1.8 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.0 | 0.1 | GO:0070182 | DNA polymerase binding(GO:0070182) |
| 0.0 | 0.4 | GO:0022851 | benzodiazepine receptor activity(GO:0008503) GABA-gated chloride ion channel activity(GO:0022851) |
| 0.0 | 3.5 | GO:0005319 | lipid transporter activity(GO:0005319) |
| 0.0 | 0.1 | GO:0004362 | glutathione-disulfide reductase activity(GO:0004362) |
| 0.0 | 5.6 | GO:0035091 | phosphatidylinositol binding(GO:0035091) |
| 0.0 | 0.2 | GO:0000048 | peptidyltransferase activity(GO:0000048) glutathione hydrolase activity(GO:0036374) |
| 0.0 | 0.6 | GO:0008474 | palmitoyl-(protein) hydrolase activity(GO:0008474) palmitoyl hydrolase activity(GO:0098599) |
| 0.0 | 0.7 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.0 | 0.3 | GO:0031994 | insulin-like growth factor I binding(GO:0031994) insulin-like growth factor II binding(GO:0031995) |
| 0.0 | 0.7 | GO:0004869 | cysteine-type endopeptidase inhibitor activity(GO:0004869) |
| 0.0 | 0.6 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.0 | 0.4 | GO:0045504 | dynein heavy chain binding(GO:0045504) |
| 0.0 | 1.5 | GO:0016811 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amides(GO:0016811) |
| 0.0 | 0.1 | GO:0032052 | bile acid binding(GO:0032052) |
| 0.0 | 0.6 | GO:0000062 | fatty-acyl-CoA binding(GO:0000062) |
| 0.0 | 0.4 | GO:0047617 | acyl-CoA hydrolase activity(GO:0047617) |
| 0.0 | 8.2 | GO:0005096 | GTPase activator activity(GO:0005096) |
| 0.0 | 0.5 | GO:0008253 | 5'-nucleotidase activity(GO:0008253) |
| 0.0 | 0.1 | GO:0042164 | interleukin-12 binding(GO:0019972) interleukin-12 alpha subunit binding(GO:0042164) |
| 0.0 | 0.3 | GO:0030515 | snoRNA binding(GO:0030515) |
| 0.0 | 1.7 | GO:0004386 | helicase activity(GO:0004386) |
| 0.0 | 0.2 | GO:0015299 | solute:proton antiporter activity(GO:0015299) |
| 0.0 | 0.7 | GO:0051959 | dynein light intermediate chain binding(GO:0051959) |
| 0.0 | 0.7 | GO:0019707 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.0 | 1.1 | GO:0019905 | syntaxin binding(GO:0019905) |
| 0.0 | 5.3 | GO:0016887 | ATPase activity(GO:0016887) |
| 0.0 | 0.6 | GO:0000049 | tRNA binding(GO:0000049) |
| 0.0 | 0.4 | GO:0042826 | histone deacetylase binding(GO:0042826) |
| 0.0 | 1.4 | GO:0015171 | amino acid transmembrane transporter activity(GO:0015171) |
| 0.0 | 0.2 | GO:0004322 | ferroxidase activity(GO:0004322) oxidoreductase activity, oxidizing metal ions, oxygen as acceptor(GO:0016724) |
| 0.0 | 0.5 | GO:0003727 | single-stranded RNA binding(GO:0003727) |
| 0.0 | 0.5 | GO:0001085 | RNA polymerase II transcription factor binding(GO:0001085) |
| 0.0 | 1.0 | GO:0003774 | motor activity(GO:0003774) |
| 0.0 | 0.5 | GO:0009931 | calcium-dependent protein serine/threonine kinase activity(GO:0009931) |
| 0.0 | 0.5 | GO:0018024 | histone-lysine N-methyltransferase activity(GO:0018024) |
| 0.0 | 4.5 | GO:0061630 | ubiquitin protein ligase activity(GO:0061630) |
| 0.0 | 0.7 | GO:0008408 | 3'-5' exonuclease activity(GO:0008408) |
| 0.0 | 0.3 | GO:0043176 | amine binding(GO:0043176) serotonin binding(GO:0051378) |
| 0.0 | 0.4 | GO:0004707 | MAP kinase activity(GO:0004707) |
| 0.0 | 0.2 | GO:0030165 | PDZ domain binding(GO:0030165) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 3.3 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.2 | 2.8 | SA REG CASCADE OF CYCLIN EXPR | Expression of cyclins regulates progression through the cell cycle by activating cyclin-dependent kinases. |
| 0.2 | 2.8 | PID RANBP2 PATHWAY | Sumoylation by RanBP2 regulates transcriptional repression |
| 0.1 | 4.3 | PID CERAMIDE PATHWAY | Ceramide signaling pathway |
| 0.1 | 2.2 | PID ARF6 PATHWAY | Arf6 signaling events |
| 0.1 | 2.2 | PID ALK1 PATHWAY | ALK1 signaling events |
| 0.1 | 1.2 | PID S1P S1P2 PATHWAY | S1P2 pathway |
| 0.1 | 2.2 | PID RHOA PATHWAY | RhoA signaling pathway |
| 0.1 | 1.1 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 0.1 | 0.3 | ST GAQ PATHWAY | G alpha q Pathway |
| 0.1 | 0.3 | PID ARF6 DOWNSTREAM PATHWAY | Arf6 downstream pathway |
| 0.1 | 2.4 | PID LKB1 PATHWAY | LKB1 signaling events |
| 0.1 | 2.6 | ST JNK MAPK PATHWAY | JNK MAPK Pathway |
| 0.1 | 0.6 | PID MYC PATHWAY | C-MYC pathway |
| 0.0 | 1.0 | PID LIS1 PATHWAY | Lissencephaly gene (LIS1) in neuronal migration and development |
| 0.0 | 2.4 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.0 | 1.9 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
| 0.0 | 1.2 | PID ANGIOPOIETIN RECEPTOR PATHWAY | Angiopoietin receptor Tie2-mediated signaling |
| 0.0 | 0.9 | PID PRL SIGNALING EVENTS PATHWAY | Signaling events mediated by PRL |
| 0.0 | 0.4 | PID AR NONGENOMIC PATHWAY | Nongenotropic Androgen signaling |
| 0.0 | 0.6 | PID P38 ALPHA BETA DOWNSTREAM PATHWAY | Signaling mediated by p38-alpha and p38-beta |
| 0.0 | 1.9 | PID AR PATHWAY | Coregulation of Androgen receptor activity |
| 0.0 | 0.6 | PID ERBB4 PATHWAY | ErbB4 signaling events |
| 0.0 | 0.4 | PID CXCR3 PATHWAY | CXCR3-mediated signaling events |
| 0.0 | 0.7 | PID RAS PATHWAY | Regulation of Ras family activation |
| 0.0 | 1.0 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
| 0.0 | 2.2 | PID P53 DOWNSTREAM PATHWAY | Direct p53 effectors |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 2.8 | REACTOME REGULATION OF PYRUVATE DEHYDROGENASE PDH COMPLEX | Genes involved in Regulation of pyruvate dehydrogenase (PDH) complex |
| 0.2 | 2.4 | REACTOME IRAK1 RECRUITS IKK COMPLEX | Genes involved in IRAK1 recruits IKK complex |
| 0.2 | 2.1 | REACTOME REGULATION OF AMPK ACTIVITY VIA LKB1 | Genes involved in Regulation of AMPK activity via LKB1 |
| 0.2 | 1.7 | REACTOME ACTIVATION OF THE AP1 FAMILY OF TRANSCRIPTION FACTORS | Genes involved in Activation of the AP-1 family of transcription factors |
| 0.2 | 0.8 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 0.1 | 2.5 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.1 | 3.1 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.1 | 1.5 | REACTOME VITAMIN B5 PANTOTHENATE METABOLISM | Genes involved in Vitamin B5 (pantothenate) metabolism |
| 0.1 | 1.8 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.1 | 1.3 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.1 | 0.9 | REACTOME SOS MEDIATED SIGNALLING | Genes involved in SOS-mediated signalling |
| 0.1 | 2.7 | REACTOME G1 S SPECIFIC TRANSCRIPTION | Genes involved in G1/S-Specific Transcription |
| 0.1 | 0.9 | REACTOME MEMBRANE BINDING AND TARGETTING OF GAG PROTEINS | Genes involved in Membrane binding and targetting of GAG proteins |
| 0.1 | 0.5 | REACTOME TGF BETA RECEPTOR SIGNALING IN EMT EPITHELIAL TO MESENCHYMAL TRANSITION | Genes involved in TGF-beta receptor signaling in EMT (epithelial to mesenchymal transition) |
| 0.1 | 1.1 | REACTOME ADENYLATE CYCLASE ACTIVATING PATHWAY | Genes involved in Adenylate cyclase activating pathway |
| 0.1 | 1.0 | REACTOME MRNA DECAY BY 3 TO 5 EXORIBONUCLEASE | Genes involved in mRNA Decay by 3' to 5' Exoribonuclease |
| 0.1 | 2.1 | REACTOME SYNTHESIS OF PIPS AT THE PLASMA MEMBRANE | Genes involved in Synthesis of PIPs at the plasma membrane |
| 0.1 | 1.2 | REACTOME SIGNALING BY CONSTITUTIVELY ACTIVE EGFR | Genes involved in Signaling by constitutively active EGFR |
| 0.1 | 2.2 | REACTOME PYRUVATE METABOLISM AND CITRIC ACID TCA CYCLE | Genes involved in Pyruvate metabolism and Citric Acid (TCA) cycle |
| 0.1 | 1.0 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |
| 0.1 | 4.8 | REACTOME TRANSPORT OF INORGANIC CATIONS ANIONS AND AMINO ACIDS OLIGOPEPTIDES | Genes involved in Transport of inorganic cations/anions and amino acids/oligopeptides |
| 0.1 | 0.9 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.1 | 1.1 | REACTOME SULFUR AMINO ACID METABOLISM | Genes involved in Sulfur amino acid metabolism |
| 0.1 | 1.0 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.1 | 0.4 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 7ALPHA HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 7alpha-hydroxycholesterol |
| 0.1 | 0.3 | REACTOME INTEGRATION OF PROVIRUS | Genes involved in Integration of provirus |
| 0.1 | 1.4 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.0 | 1.6 | REACTOME NOTCH1 INTRACELLULAR DOMAIN REGULATES TRANSCRIPTION | Genes involved in NOTCH1 Intracellular Domain Regulates Transcription |
| 0.0 | 0.7 | REACTOME IONOTROPIC ACTIVITY OF KAINATE RECEPTORS | Genes involved in Ionotropic activity of Kainate Receptors |
| 0.0 | 0.6 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.0 | 1.2 | REACTOME FORMATION OF INCISION COMPLEX IN GG NER | Genes involved in Formation of incision complex in GG-NER |
| 0.0 | 0.8 | REACTOME DOWNREGULATION OF SMAD2 3 SMAD4 TRANSCRIPTIONAL ACTIVITY | Genes involved in Downregulation of SMAD2/3:SMAD4 transcriptional activity |
| 0.0 | 0.4 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.0 | 0.4 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.0 | 0.3 | REACTOME ADVANCED GLYCOSYLATION ENDPRODUCT RECEPTOR SIGNALING | Genes involved in Advanced glycosylation endproduct receptor signaling |
| 0.0 | 0.9 | REACTOME SYNTHESIS OF PA | Genes involved in Synthesis of PA |
| 0.0 | 1.2 | REACTOME INTERFERON GAMMA SIGNALING | Genes involved in Interferon gamma signaling |
| 0.0 | 0.3 | REACTOME MTORC1 MEDIATED SIGNALLING | Genes involved in mTORC1-mediated signalling |
| 0.0 | 1.5 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |
| 0.0 | 7.3 | REACTOME ANTIGEN PROCESSING UBIQUITINATION PROTEASOME DEGRADATION | Genes involved in Antigen processing: Ubiquitination & Proteasome degradation |
| 0.0 | 0.3 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.0 | 0.3 | REACTOME GAMMA CARBOXYLATION TRANSPORT AND AMINO TERMINAL CLEAVAGE OF PROTEINS | Genes involved in Gamma-carboxylation, transport, and amino-terminal cleavage of proteins |
| 0.0 | 2.3 | REACTOME PPARA ACTIVATES GENE EXPRESSION | Genes involved in PPARA Activates Gene Expression |
| 0.0 | 3.5 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 0.4 | REACTOME GABA A RECEPTOR ACTIVATION | Genes involved in GABA A receptor activation |
| 0.0 | 1.0 | REACTOME TRANSPORT TO THE GOLGI AND SUBSEQUENT MODIFICATION | Genes involved in Transport to the Golgi and subsequent modification |
| 0.0 | 0.4 | REACTOME SYNTHESIS OF SUBSTRATES IN N GLYCAN BIOSYTHESIS | Genes involved in Synthesis of substrates in N-glycan biosythesis |
| 0.0 | 0.6 | REACTOME LYSOSOME VESICLE BIOGENESIS | Genes involved in Lysosome Vesicle Biogenesis |
| 0.0 | 0.4 | REACTOME SIGNAL TRANSDUCTION BY L1 | Genes involved in Signal transduction by L1 |
| 0.0 | 0.4 | REACTOME PIP3 ACTIVATES AKT SIGNALING | Genes involved in PIP3 activates AKT signaling |
| 0.0 | 0.7 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.0 | 0.6 | REACTOME NCAM1 INTERACTIONS | Genes involved in NCAM1 interactions |
| 0.0 | 0.1 | REACTOME REGULATION OF THE FANCONI ANEMIA PATHWAY | Genes involved in Regulation of the Fanconi anemia pathway |
| 0.0 | 0.6 | REACTOME SIGNALING BY EGFR IN CANCER | Genes involved in Signaling by EGFR in Cancer |
| 0.0 | 0.4 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.0 | 0.1 | REACTOME ASSOCIATION OF LICENSING FACTORS WITH THE PRE REPLICATIVE COMPLEX | Genes involved in Association of licensing factors with the pre-replicative complex |
| 0.0 | 0.0 | REACTOME SIGNALLING TO P38 VIA RIT AND RIN | Genes involved in Signalling to p38 via RIT and RIN |