Project
PRJNA195909:zebrafish embryo and larva development
Navigation
Downloads
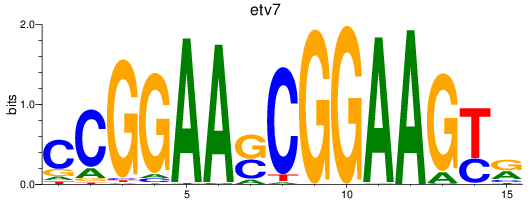
Results for etv7
Z-value: 1.36
Transcription factors associated with etv7
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
etv7
|
ENSDARG00000089434 | ETS variant transcription factor 7 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| etv7 | dr11_v1_chr23_-_5101847_5101847 | -0.56 | 1.2e-01 | Click! |
Activity profile of etv7 motif
Sorted Z-values of etv7 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr23_+_32101202 | 2.30 |
ENSDART00000000992
|
zgc:56699
|
zgc:56699 |
| chr23_+_32101361 | 2.18 |
ENSDART00000138849
|
zgc:56699
|
zgc:56699 |
| chr7_-_58826164 | 1.66 |
ENSDART00000171095
|
sox32
|
SRY (sex determining region Y)-box 32 |
| chr7_+_46019780 | 1.54 |
ENSDART00000163991
|
ccne1
|
cyclin E1 |
| chr13_-_4979029 | 1.26 |
ENSDART00000132931
|
nolc1
|
nucleolar and coiled-body phosphoprotein 1 |
| chr14_-_10617923 | 1.21 |
ENSDART00000133723
ENSDART00000131939 ENSDART00000136649 |
si:dkey-92i17.2
|
si:dkey-92i17.2 |
| chr16_+_13860299 | 1.15 |
ENSDART00000121998
|
grwd1
|
glutamate-rich WD repeat containing 1 |
| chr5_+_28497956 | 1.11 |
ENSDART00000191935
|
nfr
|
notochord formation related |
| chr10_+_17747880 | 0.97 |
ENSDART00000135044
|
pigo
|
phosphatidylinositol glycan anchor biosynthesis, class O |
| chr1_-_354115 | 0.95 |
ENSDART00000141590
ENSDART00000098627 |
pros1
|
protein S |
| chr13_-_33700461 | 0.95 |
ENSDART00000160520
|
mad2l1bp
|
MAD2L1 binding protein |
| chr17_+_43863708 | 0.93 |
ENSDART00000133874
ENSDART00000140316 ENSDART00000142929 ENSDART00000148090 |
zgc:66313
|
zgc:66313 |
| chr16_+_38338721 | 0.85 |
ENSDART00000076528
ENSDART00000142885 |
gabpb2b
|
GA binding protein transcription factor, beta subunit 2b |
| chr5_+_872299 | 0.80 |
ENSDART00000130042
|
fubp3
|
far upstream element (FUSE) binding protein 3 |
| chr1_-_46505310 | 0.79 |
ENSDART00000178072
|
si:busm1-105l16.2
|
si:busm1-105l16.2 |
| chr13_+_9559461 | 0.79 |
ENSDART00000047740
|
wdr32
|
WD repeat domain 32 |
| chr13_+_6188759 | 0.75 |
ENSDART00000161062
|
ppm1g
|
protein phosphatase, Mg2+/Mn2+ dependent, 1G |
| chr15_-_25365319 | 0.74 |
ENSDART00000152651
|
cluha
|
clustered mitochondria (cluA/CLU1) homolog a |
| chr16_-_9453591 | 0.74 |
ENSDART00000126154
|
prpf3
|
PRP3 pre-mRNA processing factor 3 homolog (yeast) |
| chr17_+_34244345 | 0.73 |
ENSDART00000006058
|
eif2s1a
|
eukaryotic translation initiation factor 2, subunit 1 alpha a |
| chr17_-_43863700 | 0.73 |
ENSDART00000157530
|
ahsa1b
|
AHA1, activator of heat shock protein ATPase homolog 1b |
| chr13_-_44819254 | 0.70 |
ENSDART00000131673
|
si:dkeyp-2e4.8
|
si:dkeyp-2e4.8 |
| chr1_+_39865748 | 0.69 |
ENSDART00000131954
ENSDART00000130270 |
irf2a
|
interferon regulatory factor 2a |
| chr16_+_33930864 | 0.67 |
ENSDART00000125945
|
snip1
|
Smad nuclear interacting protein |
| chr7_-_46019756 | 0.66 |
ENSDART00000162583
|
zgc:162297
|
zgc:162297 |
| chr6_+_60125033 | 0.65 |
ENSDART00000148557
ENSDART00000008224 |
aurka
|
aurora kinase A |
| chr20_-_37820939 | 0.64 |
ENSDART00000032978
|
nsl1
|
NSL1, MIS12 kinetochore complex component |
| chr11_+_3959495 | 0.64 |
ENSDART00000122953
|
gnl3
|
guanine nucleotide binding protein-like 3 (nucleolar) |
| chr18_+_50707179 | 0.59 |
ENSDART00000160206
|
rap2b
|
RAP2B, member of RAS oncogene family |
| chr21_-_36453594 | 0.58 |
ENSDART00000193176
|
cnot8
|
CCR4-NOT transcription complex, subunit 8 |
| chr13_+_6189203 | 0.57 |
ENSDART00000109665
|
ppm1g
|
protein phosphatase, Mg2+/Mn2+ dependent, 1G |
| chr8_-_31053872 | 0.56 |
ENSDART00000109885
|
snrnp200
|
small nuclear ribonucleoprotein 200 (U5) |
| chr1_+_494297 | 0.56 |
ENSDART00000108579
ENSDART00000146732 |
blzf1
|
basic leucine zipper nuclear factor 1 |
| chr8_+_4803906 | 0.55 |
ENSDART00000045533
|
tmem127
|
transmembrane protein 127 |
| chr25_+_16880990 | 0.54 |
ENSDART00000020259
|
zgc:77158
|
zgc:77158 |
| chr15_-_26931541 | 0.54 |
ENSDART00000027563
|
ccdc9
|
coiled-coil domain containing 9 |
| chr14_+_49220026 | 0.54 |
ENSDART00000063643
ENSDART00000128744 |
rmnd5b
|
required for meiotic nuclear division 5 homolog B |
| chr5_+_3927989 | 0.53 |
ENSDART00000030125
|
znhit3
|
zinc finger, HIT-type containing 3 |
| chr2_-_37134169 | 0.53 |
ENSDART00000146123
ENSDART00000146533 ENSDART00000040427 |
elavl1a
|
ELAV like RNA binding protein 1a |
| chr22_-_22242884 | 0.52 |
ENSDART00000020937
|
hdgfl2
|
HDGF like 2 |
| chr19_+_9050852 | 0.51 |
ENSDART00000151031
|
ash1l
|
ash1 (absent, small, or homeotic)-like (Drosophila) |
| chr17_-_51121525 | 0.51 |
ENSDART00000130412
ENSDART00000013418 |
aqr
|
aquarius intron-binding spliceosomal factor |
| chr5_-_4297459 | 0.51 |
ENSDART00000018895
|
srrt
|
serrate RNA effector molecule homolog (Arabidopsis) |
| chr3_+_62140077 | 0.50 |
ENSDART00000108945
|
GID4
|
GID complex subunit 4 homolog |
| chr3_-_25492361 | 0.50 |
ENSDART00000147322
ENSDART00000055473 |
grb2b
|
growth factor receptor-bound protein 2b |
| chr15_+_37546391 | 0.49 |
ENSDART00000186625
|
psenen
|
presenilin enhancer gamma secretase subunit |
| chr23_+_39611688 | 0.48 |
ENSDART00000034690
|
otud3
|
OTU deubiquitinase 3 |
| chr3_+_24511959 | 0.48 |
ENSDART00000133898
|
dnal4a
|
dynein, axonemal, light chain 4a |
| chr13_-_330004 | 0.48 |
ENSDART00000093149
|
ddx21
|
DEAD (Asp-Glu-Ala-Asp) box helicase 21 |
| chr19_-_37508571 | 0.47 |
ENSDART00000018255
|
ilf2
|
interleukin enhancer binding factor 2 |
| chr16_+_33931032 | 0.47 |
ENSDART00000167240
|
snip1
|
Smad nuclear interacting protein |
| chr3_-_15444396 | 0.47 |
ENSDART00000104361
|
si:dkey-56d12.4
|
si:dkey-56d12.4 |
| chr15_-_25365570 | 0.46 |
ENSDART00000152754
|
cluha
|
clustered mitochondria (cluA/CLU1) homolog a |
| chr13_+_49727333 | 0.44 |
ENSDART00000168799
ENSDART00000037559 |
ggps1
|
geranylgeranyl diphosphate synthase 1 |
| chr8_+_387622 | 0.44 |
ENSDART00000167361
|
pym1
|
PYM homolog 1, exon junction complex associated factor |
| chr6_+_18367388 | 0.44 |
ENSDART00000163394
|
dgke
|
diacylglycerol kinase, epsilon |
| chr21_-_41617372 | 0.42 |
ENSDART00000187171
|
BX005397.1
|
|
| chr15_+_37545855 | 0.41 |
ENSDART00000099456
|
psenen
|
presenilin enhancer gamma secretase subunit |
| chr12_-_31461412 | 0.41 |
ENSDART00000175929
ENSDART00000186342 |
acsl5
|
acyl-CoA synthetase long chain family member 5 |
| chr6_-_19381993 | 0.40 |
ENSDART00000164114
|
grb2a
|
growth factor receptor-bound protein 2a |
| chr10_-_1697037 | 0.40 |
ENSDART00000125188
ENSDART00000002985 |
srsf9
|
serine/arginine-rich splicing factor 9 |
| chr3_+_1150348 | 0.40 |
ENSDART00000148524
|
nol12
|
nucleolar protein 12 |
| chr8_+_52530889 | 0.39 |
ENSDART00000127729
ENSDART00000170360 ENSDART00000162687 |
stambpb
|
STAM binding protein b |
| chr16_+_38337783 | 0.39 |
ENSDART00000135008
|
gabpb2b
|
GA binding protein transcription factor, beta subunit 2b |
| chr3_-_8388344 | 0.39 |
ENSDART00000146856
|
rbfox3b
|
RNA binding fox-1 homolog 3b |
| chr8_+_7854130 | 0.38 |
ENSDART00000165575
|
cxxc1a
|
CXXC finger protein 1a |
| chr7_+_26545911 | 0.38 |
ENSDART00000135313
|
tnk1
|
tyrosine kinase, non-receptor, 1 |
| chr24_+_29352039 | 0.38 |
ENSDART00000101641
|
prmt6
|
protein arginine methyltransferase 6 |
| chr11_-_18254 | 0.38 |
ENSDART00000167814
|
prr13
|
proline rich 13 |
| chr23_+_45229198 | 0.37 |
ENSDART00000172445
|
ttc39b
|
tetratricopeptide repeat domain 39B |
| chr7_+_29177191 | 0.36 |
ENSDART00000008096
|
aph1b
|
APH1B gamma secretase subunit |
| chr7_+_26545502 | 0.35 |
ENSDART00000140528
|
tnk1
|
tyrosine kinase, non-receptor, 1 |
| chr6_+_49723289 | 0.35 |
ENSDART00000190452
|
stx16
|
syntaxin 16 |
| chr6_-_57426097 | 0.35 |
ENSDART00000171531
|
znfx1
|
zinc finger, NFX1-type containing 1 |
| chr24_+_33462800 | 0.35 |
ENSDART00000166666
ENSDART00000050826 |
rmc1
|
regulator of MON1-CCZ1 |
| chr19_+_348729 | 0.35 |
ENSDART00000114284
|
mcl1a
|
MCL1, BCL2 family apoptosis regulator a |
| chr8_-_33154677 | 0.34 |
ENSDART00000133300
|
zbtb34
|
zinc finger and BTB domain containing 34 |
| chr13_+_6086730 | 0.34 |
ENSDART00000049328
|
fam120b
|
family with sequence similarity 120B |
| chr21_-_18932761 | 0.34 |
ENSDART00000140129
|
med15
|
mediator complex subunit 15 |
| chr20_+_27298783 | 0.33 |
ENSDART00000013861
|
ppp4r4
|
protein phosphatase 4, regulatory subunit 4 |
| chr13_-_35760969 | 0.33 |
ENSDART00000127476
|
erlec1
|
endoplasmic reticulum lectin 1 |
| chr5_-_38777852 | 0.33 |
ENSDART00000131603
|
si:dkey-58f10.4
|
si:dkey-58f10.4 |
| chr21_-_30254185 | 0.32 |
ENSDART00000101054
|
dnajc18
|
DnaJ (Hsp40) homolog, subfamily C, member 18 |
| chr6_+_49722970 | 0.32 |
ENSDART00000155934
ENSDART00000154738 |
stx16
|
syntaxin 16 |
| chr1_+_59314675 | 0.32 |
ENSDART00000161872
ENSDART00000160658 ENSDART00000169792 ENSDART00000160735 |
parn
|
poly(A)-specific ribonuclease (deadenylation nuclease) |
| chr13_+_5978809 | 0.31 |
ENSDART00000102563
ENSDART00000121598 |
phf10
|
PHD finger protein 10 |
| chr11_+_5681762 | 0.31 |
ENSDART00000179139
|
arid3a
|
AT rich interactive domain 3A (BRIGHT-like) |
| chr21_-_36453417 | 0.30 |
ENSDART00000018350
|
cnot8
|
CCR4-NOT transcription complex, subunit 8 |
| chr7_+_29962559 | 0.30 |
ENSDART00000075538
|
fbxo22
|
F-box protein 22 |
| chr22_-_506522 | 0.29 |
ENSDART00000106645
ENSDART00000067637 |
dstyk
|
dual serine/threonine and tyrosine protein kinase |
| chr22_+_94692 | 0.29 |
ENSDART00000114777
|
nckipsd
|
NCK interacting protein with SH3 domain |
| chr14_+_52408619 | 0.29 |
ENSDART00000163856
|
noa1
|
nitric oxide associated 1 |
| chr20_-_4793450 | 0.28 |
ENSDART00000053870
|
galca
|
galactosylceramidase a |
| chr25_-_18948816 | 0.27 |
ENSDART00000091549
|
nt5dc3
|
5'-nucleotidase domain containing 3 |
| chr13_-_35761266 | 0.27 |
ENSDART00000190217
|
erlec1
|
endoplasmic reticulum lectin 1 |
| chr4_-_2637689 | 0.27 |
ENSDART00000192550
ENSDART00000021953 ENSDART00000150344 |
cog5
|
component of oligomeric golgi complex 5 |
| chr12_+_10115964 | 0.26 |
ENSDART00000152369
|
si:dkeyp-118b1.2
|
si:dkeyp-118b1.2 |
| chr14_+_23668730 | 0.26 |
ENSDART00000157741
|
med12
|
mediator complex subunit 12 |
| chr1_+_54889733 | 0.26 |
ENSDART00000140375
|
zfyve27
|
zinc finger, FYVE domain containing 27 |
| chr16_+_21789703 | 0.26 |
ENSDART00000153617
|
trim108
|
tripartite motif containing 108 |
| chr18_+_15778110 | 0.25 |
ENSDART00000014188
|
ube2na
|
ubiquitin-conjugating enzyme E2Na |
| chr20_+_39344889 | 0.25 |
ENSDART00000009164
|
esco2
|
establishment of sister chromatid cohesion N-acetyltransferase 2 |
| chr9_+_44304980 | 0.25 |
ENSDART00000147990
|
ssfa2
|
sperm specific antigen 2 |
| chr8_+_17184602 | 0.25 |
ENSDART00000050228
ENSDART00000140531 |
dimt1l
|
DIM1 dimethyladenosine transferase 1-like (S. cerevisiae) |
| chr4_+_2637947 | 0.24 |
ENSDART00000130623
|
dus4l
|
dihydrouridine synthase 4-like (S. cerevisiae) |
| chr4_+_29773917 | 0.24 |
ENSDART00000170798
|
BX530097.1
|
|
| chr2_+_26479676 | 0.23 |
ENSDART00000056795
ENSDART00000144837 |
hectd3
|
HECT domain containing 3 |
| chr25_-_37338048 | 0.22 |
ENSDART00000073439
|
trim44
|
tripartite motif containing 44 |
| chr10_-_41400049 | 0.22 |
ENSDART00000009838
|
gpat4
|
glycerol-3-phosphate acyltransferase 4 |
| chr4_+_77957611 | 0.22 |
ENSDART00000156692
|
arfgap3
|
ADP-ribosylation factor GTPase activating protein 3 |
| chr21_-_13123176 | 0.21 |
ENSDART00000144866
ENSDART00000024616 |
fam219aa
|
family with sequence similarity 219, member Aa |
| chr14_+_12178915 | 0.21 |
ENSDART00000054626
|
hdac3
|
histone deacetylase 3 |
| chr12_+_46708920 | 0.21 |
ENSDART00000153089
|
exoc7
|
exocyst complex component 7 |
| chr13_-_3324764 | 0.20 |
ENSDART00000102748
ENSDART00000114040 |
ubr2
|
ubiquitin protein ligase E3 component n-recognin 2 |
| chr7_-_33351485 | 0.20 |
ENSDART00000146420
|
anp32a
|
acidic (leucine-rich) nuclear phosphoprotein 32 family, member A |
| chr21_-_33126697 | 0.19 |
ENSDART00000189293
ENSDART00000084559 |
CU019662.1
|
|
| chr25_-_3627130 | 0.18 |
ENSDART00000171863
|
si:ch211-272n13.3
|
si:ch211-272n13.3 |
| chr2_+_2772447 | 0.17 |
ENSDART00000124882
|
thoc1
|
THO complex 1 |
| chr21_-_44556411 | 0.16 |
ENSDART00000163983
|
brcc3
|
BRCA1/BRCA2-containing complex, subunit 3 |
| chr9_-_7287375 | 0.16 |
ENSDART00000128352
|
mitd1
|
MIT, microtubule interacting and transport, domain containing 1 |
| chr22_+_17118225 | 0.16 |
ENSDART00000135604
|
frem1b
|
Fras1 related extracellular matrix 1b |
| chr22_+_9862466 | 0.14 |
ENSDART00000146864
|
si:dkey-253d23.3
|
si:dkey-253d23.3 |
| chr23_-_45318760 | 0.13 |
ENSDART00000166883
|
ccdc171
|
coiled-coil domain containing 171 |
| chr3_+_36972586 | 0.13 |
ENSDART00000102784
|
si:ch211-18i17.2
|
si:ch211-18i17.2 |
| chr21_-_22678195 | 0.13 |
ENSDART00000171231
|
gig2g
|
grass carp reovirus (GCRV)-induced gene 2g |
| chr9_-_29003245 | 0.12 |
ENSDART00000183391
ENSDART00000188836 |
ptpn4a
|
protein tyrosine phosphatase, non-receptor type 4a |
| chr8_+_41453691 | 0.12 |
ENSDART00000061144
|
arpc5la
|
actin related protein 2/3 complex, subunit 5-like, a |
| chr15_-_31409013 | 0.12 |
ENSDART00000140456
|
or111-9
|
odorant receptor, family D, subfamily 111, member 9 |
| chr19_+_42660158 | 0.12 |
ENSDART00000018328
|
fbxl2
|
F-box and leucine-rich repeat protein 2 |
| chr7_+_26762958 | 0.12 |
ENSDART00000167956
ENSDART00000134717 |
tspan18a
|
tetraspanin 18a |
| chr5_-_24124118 | 0.12 |
ENSDART00000051550
|
capga
|
capping protein (actin filament), gelsolin-like a |
| chr22_+_9523479 | 0.12 |
ENSDART00000189473
ENSDART00000143953 |
strip1
|
striatin interacting protein 1 |
| chr11_-_18449 | 0.11 |
ENSDART00000172050
|
prr13
|
proline rich 13 |
| chr2_+_58008980 | 0.11 |
ENSDART00000171264
|
si:ch211-155e24.3
|
si:ch211-155e24.3 |
| chr23_+_20644511 | 0.11 |
ENSDART00000133131
|
uba1
|
ubiquitin-like modifier activating enzyme 1 |
| chr16_+_12730311 | 0.11 |
ENSDART00000162030
ENSDART00000124544 |
epn1
|
epsin 1 |
| chr8_-_6918721 | 0.11 |
ENSDART00000014915
|
asb6
|
ankyrin repeat and SOCS box containing 6 |
| chr20_-_37831849 | 0.10 |
ENSDART00000188483
ENSDART00000153005 ENSDART00000142364 |
si:ch211-147d7.5
|
si:ch211-147d7.5 |
| chr1_+_9199031 | 0.10 |
ENSDART00000092058
ENSDART00000182771 |
chtf18
|
CTF18, chromosome transmission fidelity factor 18 homolog (S. cerevisiae) |
| chr2_-_58142854 | 0.10 |
ENSDART00000169909
|
CABZ01110881.1
|
|
| chr21_-_44556148 | 0.10 |
ENSDART00000163955
|
brcc3
|
BRCA1/BRCA2-containing complex, subunit 3 |
| chr16_-_34477805 | 0.10 |
ENSDART00000136546
|
serinc2l
|
serine incorporator 2, like |
| chr21_+_170038 | 0.09 |
ENSDART00000157614
|
klhl8
|
kelch-like family member 8 |
| chr24_+_15670013 | 0.09 |
ENSDART00000185826
|
CU929414.1
|
|
| chr20_+_27093042 | 0.09 |
ENSDART00000024595
|
ubr7
|
ubiquitin protein ligase E3 component n-recognin 7 |
| chr3_+_28831450 | 0.08 |
ENSDART00000055422
|
flr
|
fleer |
| chr22_+_9862243 | 0.08 |
ENSDART00000105942
|
si:dkey-253d23.3
|
si:dkey-253d23.3 |
| chr1_-_494280 | 0.08 |
ENSDART00000144406
|
ercc5
|
excision repair cross-complementation group 5 |
| chr7_-_26049282 | 0.08 |
ENSDART00000136389
ENSDART00000101124 |
rnaseka
|
ribonuclease, RNase K a |
| chr2_-_24317240 | 0.08 |
ENSDART00000078975
|
trnau1apb
|
tRNA selenocysteine 1 associated protein 1b |
| chr5_-_68495967 | 0.07 |
ENSDART00000188107
|
ephb4a
|
eph receptor B4a |
| chr25_-_7946728 | 0.07 |
ENSDART00000183847
ENSDART00000114653 |
nox5
|
NADPH oxidase, EF-hand calcium binding domain 5 |
| chr8_+_1187928 | 0.07 |
ENSDART00000127252
|
slc35d2
|
solute carrier family 35 (UDP-GlcNAc/UDP-glucose transporter), member D2 |
| chr9_-_33136990 | 0.06 |
ENSDART00000131742
|
slc5a3b
|
solute carrier family 5 (sodium/myo-inositol cotransporter), member 3b |
| chr19_-_32888758 | 0.06 |
ENSDART00000052080
|
laptm4b
|
lysosomal protein transmembrane 4 beta |
| chr14_+_49251331 | 0.05 |
ENSDART00000148882
|
anxa6
|
annexin A6 |
| chr16_+_53489676 | 0.04 |
ENSDART00000074653
|
grinab
|
glutamate receptor, ionotropic, N-methyl D-aspartate-associated protein 1b (glutamate binding) |
| chr19_+_63567 | 0.04 |
ENSDART00000165657
ENSDART00000165183 |
zhx2b
|
zinc fingers and homeoboxes 2b |
| chr8_-_45430817 | 0.04 |
ENSDART00000150067
ENSDART00000112394 |
ywhabb
|
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, beta polypeptide b |
| chr12_+_47280839 | 0.04 |
ENSDART00000187574
|
RYR2
|
ryanodine receptor 2 |
| chr2_+_54389750 | 0.03 |
ENSDART00000189236
|
rab12
|
RAB12, member RAS oncogene family |
| chr5_-_50084310 | 0.03 |
ENSDART00000074599
ENSDART00000189970 |
fam172a
|
family with sequence similarity 172, member A |
| chr25_+_469855 | 0.03 |
ENSDART00000104717
|
rsl24d1
|
ribosomal L24 domain containing 1 |
| chr4_-_5371639 | 0.03 |
ENSDART00000150697
|
si:dkey-14d8.1
|
si:dkey-14d8.1 |
| chr10_-_105100 | 0.02 |
ENSDART00000145716
|
ttc3
|
tetratricopeptide repeat domain 3 |
| chr17_+_27140806 | 0.02 |
ENSDART00000044378
|
rps6ka1
|
ribosomal protein S6 kinase a, polypeptide 1 |
| chr1_+_57757456 | 0.02 |
ENSDART00000152650
|
si:dkey-1c7.1
|
si:dkey-1c7.1 |
| chr1_+_961607 | 0.02 |
ENSDART00000184660
|
n6amt1
|
N-6 adenine-specific DNA methyltransferase 1 |
| chr20_+_37820992 | 0.00 |
ENSDART00000064692
|
tatdn3
|
TatD DNase domain containing 3 |
| chr7_-_50272912 | 0.00 |
ENSDART00000098842
|
hsd17b12b
|
hydroxysteroid (17-beta) dehydrogenase 12b |
Network of associatons between targets according to the STRING database.
First level regulatory network of etv7
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.0 | GO:0042730 | fibrinolysis(GO:0042730) |
| 0.2 | 0.6 | GO:0000212 | meiotic spindle organization(GO:0000212) |
| 0.2 | 1.7 | GO:0003262 | endocardial progenitor cell migration to the midline involved in heart field formation(GO:0003262) |
| 0.1 | 1.3 | GO:0007220 | Notch receptor processing(GO:0007220) |
| 0.1 | 0.4 | GO:0010746 | regulation of plasma membrane long-chain fatty acid transport(GO:0010746) positive regulation of plasma membrane long-chain fatty acid transport(GO:0010747) plasma membrane long-chain fatty acid transport(GO:0015911) fatty acid transmembrane transport(GO:1902001) positive regulation of anion transmembrane transport(GO:1903961) regulation of fatty acid transport(GO:2000191) positive regulation of fatty acid transport(GO:2000193) |
| 0.1 | 0.5 | GO:1990167 | protein K27-linked deubiquitination(GO:1990167) |
| 0.1 | 1.2 | GO:0032781 | positive regulation of ATPase activity(GO:0032781) |
| 0.1 | 0.5 | GO:0031053 | primary miRNA processing(GO:0031053) |
| 0.1 | 1.2 | GO:0048312 | intracellular distribution of mitochondria(GO:0048312) |
| 0.1 | 0.4 | GO:0034969 | histone arginine methylation(GO:0034969) |
| 0.1 | 0.6 | GO:0000393 | spliceosomal conformational changes to generate catalytic conformation(GO:0000393) |
| 0.1 | 1.6 | GO:0007096 | regulation of exit from mitosis(GO:0007096) |
| 0.1 | 0.3 | GO:0070537 | histone H2A K63-linked deubiquitination(GO:0070537) |
| 0.1 | 0.2 | GO:0034421 | post-translational protein acetylation(GO:0034421) |
| 0.1 | 1.1 | GO:0035196 | production of miRNAs involved in gene silencing by miRNA(GO:0035196) |
| 0.1 | 0.9 | GO:0043928 | exonucleolytic nuclear-transcribed mRNA catabolic process involved in deadenylation-dependent decay(GO:0043928) |
| 0.1 | 0.5 | GO:0000492 | box C/D snoRNP assembly(GO:0000492) |
| 0.1 | 0.2 | GO:0048205 | COPI-coated vesicle budding(GO:0035964) Golgi transport vesicle coating(GO:0048200) COPI coating of Golgi vesicle(GO:0048205) |
| 0.1 | 0.6 | GO:0032486 | Rap protein signal transduction(GO:0032486) |
| 0.1 | 0.3 | GO:0090579 | transcriptional activation by promoter-enhancer looping(GO:0071733) gene looping(GO:0090202) dsDNA loop formation(GO:0090579) |
| 0.1 | 0.7 | GO:0039023 | pronephric duct morphogenesis(GO:0039023) |
| 0.1 | 0.4 | GO:0031048 | chromatin silencing by small RNA(GO:0031048) |
| 0.0 | 0.3 | GO:0048011 | neurotrophin TRK receptor signaling pathway(GO:0048011) |
| 0.0 | 0.3 | GO:0042769 | DNA damage response, detection of DNA damage(GO:0042769) |
| 0.0 | 0.3 | GO:0032515 | negative regulation of phosphoprotein phosphatase activity(GO:0032515) |
| 0.0 | 0.2 | GO:0071596 | ubiquitin-dependent protein catabolic process via the N-end rule pathway(GO:0071596) |
| 0.0 | 0.3 | GO:0048742 | regulation of skeletal muscle fiber development(GO:0048742) |
| 0.0 | 4.5 | GO:0007283 | spermatogenesis(GO:0007283) |
| 0.0 | 0.3 | GO:0046477 | glycosylceramide catabolic process(GO:0046477) |
| 0.0 | 1.0 | GO:0006506 | GPI anchor biosynthetic process(GO:0006506) |
| 0.0 | 0.6 | GO:0043001 | Golgi to plasma membrane protein transport(GO:0043001) |
| 0.0 | 0.2 | GO:0070933 | histone H4 deacetylation(GO:0070933) |
| 0.0 | 0.1 | GO:0006295 | nucleotide-excision repair, DNA incision, 3'-to lesion(GO:0006295) |
| 0.0 | 0.1 | GO:0033212 | iron assimilation(GO:0033212) |
| 0.0 | 0.3 | GO:0043154 | negative regulation of cysteine-type endopeptidase activity involved in apoptotic process(GO:0043154) |
| 0.0 | 1.5 | GO:0000082 | G1/S transition of mitotic cell cycle(GO:0000082) |
| 0.0 | 0.2 | GO:0060036 | notochord cell vacuolation(GO:0060036) |
| 0.0 | 0.4 | GO:0007205 | protein kinase C-activating G-protein coupled receptor signaling pathway(GO:0007205) |
| 0.0 | 0.4 | GO:0070536 | protein K63-linked deubiquitination(GO:0070536) |
| 0.0 | 0.4 | GO:0000184 | nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:0000184) |
| 0.0 | 0.3 | GO:0071218 | response to misfolded protein(GO:0051788) cellular response to misfolded protein(GO:0071218) |
| 0.0 | 0.1 | GO:0001778 | plasma membrane repair(GO:0001778) chondrocyte differentiation involved in endochondral bone morphogenesis(GO:0003413) growth plate cartilage chondrocyte differentiation(GO:0003418) |
| 0.0 | 0.2 | GO:0031167 | rRNA methylation(GO:0031167) |
| 0.0 | 0.6 | GO:0032007 | negative regulation of TOR signaling(GO:0032007) |
| 0.0 | 0.1 | GO:0035475 | angioblast cell migration involved in selective angioblast sprouting(GO:0035475) |
| 0.0 | 0.1 | GO:0036372 | opsin transport(GO:0036372) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.5 | GO:0097134 | cyclin E1-CDK2 complex(GO:0097134) |
| 0.2 | 0.7 | GO:0043614 | multi-eIF complex(GO:0043614) |
| 0.2 | 1.3 | GO:0070765 | gamma-secretase complex(GO:0070765) |
| 0.1 | 0.6 | GO:0000444 | MIS12/MIND type complex(GO:0000444) |
| 0.1 | 0.6 | GO:0032133 | chromosome passenger complex(GO:0032133) |
| 0.1 | 0.5 | GO:0034657 | GID complex(GO:0034657) |
| 0.1 | 0.9 | GO:0030015 | CCR4-NOT core complex(GO:0030015) |
| 0.1 | 0.3 | GO:0032584 | growth cone membrane(GO:0032584) |
| 0.0 | 0.4 | GO:0035145 | exon-exon junction complex(GO:0035145) |
| 0.0 | 0.3 | GO:0070552 | BRISC complex(GO:0070552) |
| 0.0 | 0.7 | GO:0046540 | U4/U6 x U5 tri-snRNP complex(GO:0046540) |
| 0.0 | 0.8 | GO:0080008 | Cul4-RING E3 ubiquitin ligase complex(GO:0080008) |
| 0.0 | 0.2 | GO:0000445 | THO complex(GO:0000347) THO complex part of transcription export complex(GO:0000445) |
| 0.0 | 0.4 | GO:0048188 | Set1C/COMPASS complex(GO:0048188) |
| 0.0 | 0.3 | GO:0017119 | Golgi transport complex(GO:0017119) |
| 0.0 | 0.3 | GO:0071564 | npBAF complex(GO:0071564) |
| 0.0 | 0.1 | GO:0005879 | axonemal microtubule(GO:0005879) |
| 0.0 | 0.6 | GO:0055038 | recycling endosome membrane(GO:0055038) |
| 0.0 | 0.5 | GO:0071013 | catalytic step 2 spliceosome(GO:0071013) |
| 0.0 | 0.4 | GO:0032154 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.0 | 3.2 | GO:0005730 | nucleolus(GO:0005730) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.0 | GO:0051377 | mannose-ethanolamine phosphotransferase activity(GO:0051377) |
| 0.1 | 0.6 | GO:0035173 | histone kinase activity(GO:0035173) |
| 0.1 | 0.3 | GO:0043621 | protein self-association(GO:0043621) |
| 0.1 | 1.3 | GO:0004724 | magnesium-dependent protein serine/threonine phosphatase activity(GO:0004724) |
| 0.1 | 1.2 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.1 | 0.4 | GO:0008469 | histone-arginine N-methyltransferase activity(GO:0008469) |
| 0.1 | 0.5 | GO:0008506 | sugar:proton symporter activity(GO:0005351) cation:sugar symporter activity(GO:0005402) sucrose:proton symporter activity(GO:0008506) sucrose transmembrane transporter activity(GO:0008515) disaccharide transmembrane transporter activity(GO:0015154) oligosaccharide transmembrane transporter activity(GO:0015157) |
| 0.0 | 0.2 | GO:0000179 | rRNA (adenine-N6,N6-)-dimethyltransferase activity(GO:0000179) |
| 0.0 | 0.7 | GO:0051879 | Hsp90 protein binding(GO:0051879) |
| 0.0 | 0.2 | GO:0017150 | tRNA dihydrouridine synthase activity(GO:0017150) |
| 0.0 | 0.4 | GO:0047676 | arachidonate-CoA ligase activity(GO:0047676) |
| 0.0 | 0.6 | GO:0070122 | isopeptidase activity(GO:0070122) |
| 0.0 | 0.1 | GO:0070735 | protein-glycine ligase activity(GO:0070735) tubulin-glycine ligase activity(GO:0070738) |
| 0.0 | 0.3 | GO:0030544 | Hsp70 protein binding(GO:0030544) |
| 0.0 | 0.1 | GO:0000293 | ferric-chelate reductase activity(GO:0000293) |
| 0.0 | 1.2 | GO:0043022 | ribosome binding(GO:0043022) |
| 0.0 | 0.4 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.0 | 0.4 | GO:0004659 | prenyltransferase activity(GO:0004659) |
| 0.0 | 0.1 | GO:0005461 | UDP-glucuronic acid transmembrane transporter activity(GO:0005461) UDP-N-acetylgalactosamine transmembrane transporter activity(GO:0005463) |
| 0.0 | 0.6 | GO:0019003 | GDP binding(GO:0019003) |
| 0.0 | 0.1 | GO:0004839 | ubiquitin activating enzyme activity(GO:0004839) |
| 0.0 | 0.2 | GO:0031078 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.0 | 0.3 | GO:0008253 | 5'-nucleotidase activity(GO:0008253) |
| 0.0 | 1.0 | GO:0003724 | RNA helicase activity(GO:0003724) |
| 0.0 | 0.5 | GO:0003725 | double-stranded RNA binding(GO:0003725) |
| 0.0 | 3.5 | GO:0044822 | mRNA binding(GO:0003729) poly(A) RNA binding(GO:0044822) |
| 0.0 | 0.1 | GO:0005412 | glucose:sodium symporter activity(GO:0005412) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.4 | SA REG CASCADE OF CYCLIN EXPR | Expression of cyclins regulates progression through the cell cycle by activating cyclin-dependent kinases. |
| 0.1 | 1.3 | PID SYNDECAN 3 PATHWAY | Syndecan-3-mediated signaling events |
| 0.0 | 0.6 | PID AURORA A PATHWAY | Aurora A signaling |
| 0.0 | 1.3 | SIG PIP3 SIGNALING IN CARDIAC MYOCTES | Genes related to PIP3 signaling in cardiac myocytes |
| 0.0 | 1.5 | PID SMAD2 3NUCLEAR PATHWAY | Regulation of nuclear SMAD2/3 signaling |
| 0.0 | 0.3 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.0 | 0.2 | PID RETINOIC ACID PATHWAY | Retinoic acid receptors-mediated signaling |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.0 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 0.1 | 1.3 | REACTOME SIGNALING BY NOTCH4 | Genes involved in Signaling by NOTCH4 |
| 0.1 | 1.2 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.1 | 1.5 | REACTOME G1 S SPECIFIC TRANSCRIPTION | Genes involved in G1/S-Specific Transcription |
| 0.1 | 1.0 | REACTOME SYNTHESIS OF GLYCOSYLPHOSPHATIDYLINOSITOL GPI | Genes involved in Synthesis of glycosylphosphatidylinositol (GPI) |
| 0.1 | 0.4 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.0 | 0.7 | REACTOME PROTEOLYTIC CLEAVAGE OF SNARE COMPLEX PROTEINS | Genes involved in Proteolytic cleavage of SNARE complex proteins |
| 0.0 | 0.4 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.0 | 0.3 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.0 | 0.5 | REACTOME MRNA SPLICING MINOR PATHWAY | Genes involved in mRNA Splicing - Minor Pathway |
| 0.0 | 0.4 | REACTOME MRNA 3 END PROCESSING | Genes involved in mRNA 3'-end processing |