Project
PRJNA195909:zebrafish embryo and larva development
Navigation
Downloads
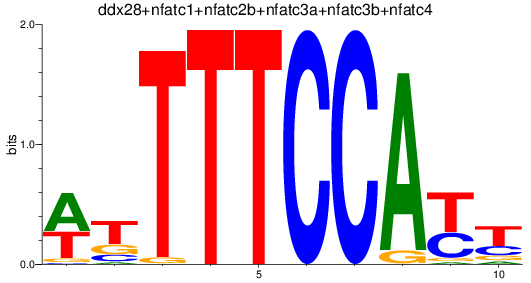
Results for ddx28+nfatc1+nfatc2b+nfatc3a+nfatc3b+nfatc4
Z-value: 2.28
Transcription factors associated with ddx28+nfatc1+nfatc2b+nfatc3a+nfatc3b+nfatc4
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
nfatc1
|
ENSDARG00000036168 | nuclear factor of activated T cells 1 |
|
nfatc3b
|
ENSDARG00000051729 | nuclear factor of activated T cells 3b |
|
nfatc4
|
ENSDARG00000054162 | nuclear factor of activated T cells 4 |
|
nfatc3a
|
ENSDARG00000076297 | nuclear factor of activated T cells 3a |
|
nfatc2b
|
ENSDARG00000079972 | nuclear factor of activated T cells 2b |
|
ddx28
|
ENSDARG00000112133 | nuclear factor of activated T cells 3a |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| nfatc1 | dr11_v1_chr19_+_22216778_22216778 | 0.91 | 5.6e-04 | Click! |
| nfatc3b | dr11_v1_chr25_-_36248053_36248053 | 0.76 | 1.7e-02 | Click! |
| nfatc3a | dr11_v1_chr7_-_34927961_34927961 | -0.63 | 7.0e-02 | Click! |
| nfatc2b | dr11_v1_chr6_-_55254786_55254786 | -0.56 | 1.2e-01 | Click! |
| nfatc4 | dr11_v1_chr2_-_37797577_37797577 | 0.34 | 3.7e-01 | Click! |
Activity profile of ddx28+nfatc1+nfatc2b+nfatc3a+nfatc3b+nfatc4 motif
Sorted Z-values of ddx28+nfatc1+nfatc2b+nfatc3a+nfatc3b+nfatc4 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr2_+_42191592 | 4.90 |
ENSDART00000144716
|
cavin4a
|
caveolae associated protein 4a |
| chr3_-_32817274 | 4.62 |
ENSDART00000142582
|
mylpfa
|
myosin light chain, phosphorylatable, fast skeletal muscle a |
| chr6_-_39313027 | 4.36 |
ENSDART00000012644
|
krt4
|
keratin 4 |
| chr11_-_24191928 | 4.13 |
ENSDART00000136827
|
sox12
|
SRY (sex determining region Y)-box 12 |
| chr3_-_20091964 | 4.04 |
ENSDART00000029386
ENSDART00000020253 ENSDART00000124326 |
slc4a1a
|
solute carrier family 4 (anion exchanger), member 1a (Diego blood group) |
| chr12_-_16636627 | 3.89 |
ENSDART00000128811
|
si:dkey-239j18.3
|
si:dkey-239j18.3 |
| chr1_+_19332837 | 3.85 |
ENSDART00000078594
|
tyrp1b
|
tyrosinase-related protein 1b |
| chr16_+_23397785 | 3.71 |
ENSDART00000148961
|
s100a10b
|
S100 calcium binding protein A10b |
| chr16_+_23398369 | 3.70 |
ENSDART00000037694
|
s100a10b
|
S100 calcium binding protein A10b |
| chr25_-_15045338 | 3.42 |
ENSDART00000161165
ENSDART00000165774 ENSDART00000172538 |
pax6a
|
paired box 6a |
| chr12_-_26064480 | 3.41 |
ENSDART00000158215
ENSDART00000171206 ENSDART00000171212 ENSDART00000182956 ENSDART00000186779 |
ldb3b
|
LIM domain binding 3b |
| chr7_-_52558495 | 3.29 |
ENSDART00000138263
ENSDART00000009938 ENSDART00000174292 ENSDART00000174218 ENSDART00000174335 |
tcf12
|
transcription factor 12 |
| chr12_-_16595406 | 3.23 |
ENSDART00000166798
|
si:dkey-239j18.2
|
si:dkey-239j18.2 |
| chr23_+_36130883 | 3.23 |
ENSDART00000103132
|
hoxc4a
|
homeobox C4a |
| chr3_-_61181018 | 3.23 |
ENSDART00000187970
|
pvalb4
|
parvalbumin 4 |
| chr12_-_16595177 | 3.03 |
ENSDART00000133962
|
si:dkey-239j18.2
|
si:dkey-239j18.2 |
| chr2_-_41518340 | 2.83 |
ENSDART00000130830
|
otomp
|
otolith matrix protein |
| chr4_+_17417111 | 2.81 |
ENSDART00000056005
|
ascl1a
|
achaete-scute family bHLH transcription factor 1a |
| chr12_-_31103187 | 2.72 |
ENSDART00000005562
ENSDART00000031408 ENSDART00000125046 ENSDART00000009237 ENSDART00000122972 ENSDART00000153068 |
tcf7l2
|
transcription factor 7 like 2 |
| chr13_+_23157053 | 2.72 |
ENSDART00000162359
|
sorbs1
|
sorbin and SH3 domain containing 1 |
| chr9_+_3388099 | 2.72 |
ENSDART00000019910
|
dlx1a
|
distal-less homeobox 1a |
| chr20_-_10120442 | 2.71 |
ENSDART00000144970
|
meis2b
|
Meis homeobox 2b |
| chr9_+_53276356 | 2.69 |
ENSDART00000003310
|
sox21b
|
SRY (sex determining region Y)-box 21b |
| chr6_-_49063085 | 2.64 |
ENSDART00000156124
|
si:ch211-105j21.9
|
si:ch211-105j21.9 |
| chr4_+_7677318 | 2.58 |
ENSDART00000149218
|
elk3
|
ELK3, ETS-domain protein |
| chr19_-_41069573 | 2.55 |
ENSDART00000111982
ENSDART00000193142 |
sgce
|
sarcoglycan, epsilon |
| chr12_+_42436920 | 2.53 |
ENSDART00000177303
|
ebf3a
|
early B cell factor 3a |
| chr1_-_12278522 | 2.51 |
ENSDART00000142122
ENSDART00000003825 |
cplx2l
|
complexin 2, like |
| chr3_-_41791178 | 2.46 |
ENSDART00000049687
|
grifin
|
galectin-related inter-fiber protein |
| chr22_-_37349967 | 2.41 |
ENSDART00000104493
|
sox2
|
SRY (sex determining region Y)-box 2 |
| chr5_+_32345187 | 2.39 |
ENSDART00000147132
|
c9
|
complement component 9 |
| chr12_-_16898140 | 2.30 |
ENSDART00000152656
|
MGC174155
|
Cathepsin L1-like |
| chr22_-_26353916 | 2.29 |
ENSDART00000077958
|
capn2b
|
calpain 2, (m/II) large subunit b |
| chr12_-_16923162 | 2.24 |
ENSDART00000106072
|
si:dkey-26g8.5
|
si:dkey-26g8.5 |
| chr7_-_38658411 | 2.23 |
ENSDART00000109463
ENSDART00000017155 |
npsn
|
nephrosin |
| chr12_-_16694092 | 2.22 |
ENSDART00000047916
|
ctslb
|
cathepsin Lb |
| chr12_-_16558106 | 2.20 |
ENSDART00000109033
|
si:dkey-269i1.4
|
si:dkey-269i1.4 |
| chr8_-_40464935 | 2.19 |
ENSDART00000040013
|
myl7
|
myosin, light chain 7, regulatory |
| chr13_+_25449681 | 2.19 |
ENSDART00000101328
|
atoh7
|
atonal bHLH transcription factor 7 |
| chr12_-_16877136 | 2.19 |
ENSDART00000152593
|
si:dkey-269i1.4
|
si:dkey-269i1.4 |
| chr22_-_15587360 | 2.17 |
ENSDART00000142717
ENSDART00000138978 |
tpm4a
|
tropomyosin 4a |
| chr6_+_6491013 | 2.16 |
ENSDART00000140827
|
bcl11ab
|
B cell CLL/lymphoma 11Ab |
| chr6_+_49095646 | 2.14 |
ENSDART00000103385
|
slc25a55a
|
solute carrier family 25, member 55a |
| chr12_-_16720196 | 2.13 |
ENSDART00000187639
|
si:dkey-26g8.4
|
si:dkey-26g8.4 |
| chr4_-_15420452 | 2.09 |
ENSDART00000016230
|
plxna4
|
plexin A4 |
| chr10_-_29903165 | 2.07 |
ENSDART00000078800
|
lim2.1
|
lens intrinsic membrane protein 2.1 |
| chr1_-_38195012 | 2.07 |
ENSDART00000020409
|
hand2
|
heart and neural crest derivatives expressed 2 |
| chr16_+_20910186 | 2.02 |
ENSDART00000046766
|
hoxa10b
|
homeobox A10b |
| chr12_-_16941319 | 2.01 |
ENSDART00000109968
|
zgc:174855
|
zgc:174855 |
| chr2_+_38161318 | 2.01 |
ENSDART00000044264
|
mmp14b
|
matrix metallopeptidase 14b (membrane-inserted) |
| chr22_-_17606575 | 1.99 |
ENSDART00000183951
|
gpx4a
|
glutathione peroxidase 4a |
| chr7_-_52531252 | 1.98 |
ENSDART00000174369
|
tcf12
|
transcription factor 12 |
| chr2_-_30182353 | 1.97 |
ENSDART00000019149
|
rpl7
|
ribosomal protein L7 |
| chr23_+_23119008 | 1.96 |
ENSDART00000132418
|
samd11
|
sterile alpha motif domain containing 11 |
| chr12_+_16949391 | 1.95 |
ENSDART00000152635
|
zgc:174153
|
zgc:174153 |
| chr13_+_17672527 | 1.92 |
ENSDART00000148269
ENSDART00000137776 |
comtd1
|
catechol-O-methyltransferase domain containing 1 |
| chr24_+_11334733 | 1.90 |
ENSDART00000147552
ENSDART00000143171 |
si:dkey-12l12.1
|
si:dkey-12l12.1 |
| chr9_+_26103814 | 1.89 |
ENSDART00000026011
|
efnb2a
|
ephrin-B2a |
| chr20_+_31287356 | 1.85 |
ENSDART00000007688
|
slc22a16
|
solute carrier family 22 (organic cation/carnitine transporter), member 16 |
| chr7_-_40993456 | 1.84 |
ENSDART00000031700
|
en2a
|
engrailed homeobox 2a |
| chr3_+_23721808 | 1.83 |
ENSDART00000012470
|
hoxb4a
|
homeobox B4a |
| chr2_-_48196092 | 1.82 |
ENSDART00000139944
|
snorc
|
secondary ossification center associated regulator of chondrocyte maturation |
| chr20_+_26690036 | 1.82 |
ENSDART00000103232
|
foxf2b
|
forkhead box F2b |
| chr2_-_20923864 | 1.81 |
ENSDART00000006870
|
ptgs2a
|
prostaglandin-endoperoxide synthase 2a |
| chr9_+_31752628 | 1.80 |
ENSDART00000060054
|
itgbl1
|
integrin, beta-like 1 |
| chr25_+_4837915 | 1.80 |
ENSDART00000168016
|
gnb5a
|
guanine nucleotide binding protein (G protein), beta 5a |
| chr13_-_46429220 | 1.80 |
ENSDART00000149125
ENSDART00000098269 ENSDART00000150061 ENSDART00000080916 |
fgfr2
|
fibroblast growth factor receptor 2 |
| chr7_-_30367650 | 1.76 |
ENSDART00000075519
|
aldh1a2
|
aldehyde dehydrogenase 1 family, member A2 |
| chr9_-_1949915 | 1.76 |
ENSDART00000190712
|
hoxd3a
|
homeobox D3a |
| chr16_-_22713152 | 1.73 |
ENSDART00000140953
ENSDART00000143836 |
si:ch211-105c13.3
|
si:ch211-105c13.3 |
| chr2_-_39017838 | 1.72 |
ENSDART00000048838
|
rbp2b
|
retinol binding protein 2b, cellular |
| chr11_+_13630107 | 1.72 |
ENSDART00000172220
|
si:ch211-1a19.3
|
si:ch211-1a19.3 |
| chr9_-_1959917 | 1.72 |
ENSDART00000082359
|
hoxd3a
|
homeobox D3a |
| chr7_-_24022340 | 1.72 |
ENSDART00000149133
|
cideb
|
cell death-inducing DFFA-like effector b |
| chr8_+_17078692 | 1.72 |
ENSDART00000023206
|
plk2b
|
polo-like kinase 2b (Drosophila) |
| chr4_+_19534833 | 1.71 |
ENSDART00000140028
|
lrrc4.1
|
leucine rich repeat containing 4.1 |
| chr23_-_27633730 | 1.67 |
ENSDART00000103639
|
arf3a
|
ADP-ribosylation factor 3a |
| chr23_+_36101185 | 1.66 |
ENSDART00000103139
|
hoxc8a
|
homeobox C8a |
| chr7_-_13882988 | 1.64 |
ENSDART00000169828
|
rlbp1a
|
retinaldehyde binding protein 1a |
| chr14_+_24277556 | 1.63 |
ENSDART00000122660
|
hnrnpa0a
|
heterogeneous nuclear ribonucleoprotein A0a |
| chr2_-_37956768 | 1.61 |
ENSDART00000034595
|
cbln10
|
cerebellin 10 |
| chr20_-_49681850 | 1.59 |
ENSDART00000025926
|
col12a1b
|
collagen, type XII, alpha 1b |
| chr1_-_58664854 | 1.55 |
ENSDART00000109528
|
adgre5b.3
|
adhesion G protein-coupled receptor E5b, duplicate 3 |
| chr12_-_16720432 | 1.55 |
ENSDART00000152261
ENSDART00000152154 |
si:dkey-26g8.4
|
si:dkey-26g8.4 |
| chr1_-_56223913 | 1.55 |
ENSDART00000019573
|
zgc:65894
|
zgc:65894 |
| chr8_+_25616946 | 1.53 |
ENSDART00000133983
|
slc38a5a
|
solute carrier family 38, member 5a |
| chr8_+_37489495 | 1.51 |
ENSDART00000141516
|
fmodb
|
fibromodulin b |
| chr13_+_13681681 | 1.51 |
ENSDART00000057825
|
cfd
|
complement factor D (adipsin) |
| chr14_-_32744464 | 1.50 |
ENSDART00000075617
|
sox3
|
SRY (sex determining region Y)-box 3 |
| chr7_+_15871156 | 1.47 |
ENSDART00000145946
|
pax6b
|
paired box 6b |
| chr11_-_32723851 | 1.47 |
ENSDART00000155592
|
pcdh17
|
protocadherin 17 |
| chr4_+_19535946 | 1.46 |
ENSDART00000192342
ENSDART00000183740 ENSDART00000180812 ENSDART00000180017 |
lrrc4.1
|
leucine rich repeat containing 4.1 |
| chr1_-_38816685 | 1.46 |
ENSDART00000075230
|
asb5b
|
ankyrin repeat and SOCS box containing 5b |
| chr4_+_12013642 | 1.45 |
ENSDART00000067281
|
cry1aa
|
cryptochrome circadian clock 1aa |
| chr10_-_28761454 | 1.44 |
ENSDART00000129400
|
alcama
|
activated leukocyte cell adhesion molecule a |
| chr9_+_33145522 | 1.43 |
ENSDART00000005879
|
atp5po
|
ATP synthase peripheral stalk subunit OSCP |
| chr13_-_31470439 | 1.42 |
ENSDART00000076574
|
rtn1a
|
reticulon 1a |
| chr13_+_1155536 | 1.41 |
ENSDART00000148356
|
perp
|
PERP, TP53 apoptosis effector |
| chr5_+_30179010 | 1.41 |
ENSDART00000134624
|
adamts15a
|
ADAM metallopeptidase with thrombospondin type 1 motif, 15a |
| chr4_+_61995745 | 1.38 |
ENSDART00000171539
|
CT990567.1
|
|
| chr8_+_22931427 | 1.37 |
ENSDART00000063096
|
sypa
|
synaptophysin a |
| chr18_-_39583601 | 1.37 |
ENSDART00000125116
|
tnfaip8l3
|
tumor necrosis factor, alpha-induced protein 8-like 3 |
| chr14_-_448182 | 1.37 |
ENSDART00000180018
|
FAT4
|
FAT atypical cadherin 4 |
| chr23_-_20309505 | 1.36 |
ENSDART00000130856
|
lamb2l
|
laminin, beta 2-like |
| chr10_-_43771447 | 1.35 |
ENSDART00000052307
|
arrdc3b
|
arrestin domain containing 3b |
| chr14_+_49296052 | 1.34 |
ENSDART00000006073
ENSDART00000105346 |
anxa6
|
annexin A6 |
| chr23_-_24682244 | 1.34 |
ENSDART00000104035
|
ctdsp2
|
CTD (carboxy-terminal domain, RNA polymerase II, polypeptide A) small phosphatase 2 |
| chr16_-_35975254 | 1.32 |
ENSDART00000167537
|
eva1ba
|
eva-1 homolog Ba (C. elegans) |
| chr4_-_22311610 | 1.31 |
ENSDART00000137814
|
hcls1
|
hematopoietic cell-specific Lyn substrate 1 |
| chr5_+_37091626 | 1.31 |
ENSDART00000161054
|
tagln2
|
transgelin 2 |
| chr6_-_8311044 | 1.29 |
ENSDART00000129674
|
slc44a2
|
solute carrier family 44 (choline transporter), member 2 |
| chr1_-_28629471 | 1.28 |
ENSDART00000121758
|
ednrba
|
endothelin receptor Ba |
| chr12_+_18234557 | 1.28 |
ENSDART00000130741
|
fam20cb
|
family with sequence similarity 20, member Cb |
| chr11_-_18705303 | 1.28 |
ENSDART00000059732
|
id1
|
inhibitor of DNA binding 1 |
| chr22_+_28446557 | 1.27 |
ENSDART00000089546
|
abi3bpb
|
ABI family, member 3 (NESH) binding protein b |
| chr1_-_22834824 | 1.25 |
ENSDART00000043556
|
ldb2b
|
LIM domain binding 2b |
| chr14_-_33454595 | 1.22 |
ENSDART00000109615
ENSDART00000173267 ENSDART00000185737 ENSDART00000190989 |
tmem255a
|
transmembrane protein 255A |
| chr22_+_28446365 | 1.22 |
ENSDART00000189359
|
abi3bpb
|
ABI family, member 3 (NESH) binding protein b |
| chr20_+_15552657 | 1.22 |
ENSDART00000063912
|
jun
|
Jun proto-oncogene, AP-1 transcription factor subunit |
| chr3_-_39152478 | 1.22 |
ENSDART00000154550
|
si:dkeyp-57f11.2
|
si:dkeyp-57f11.2 |
| chr25_+_27493444 | 1.22 |
ENSDART00000112299
|
gpr37a
|
G protein-coupled receptor 37a |
| chr1_-_21599219 | 1.22 |
ENSDART00000148327
|
adamtsl7
|
ADAMTS-like 7 |
| chr11_-_26666501 | 1.22 |
ENSDART00000188067
ENSDART00000111539 |
efcc1
|
EF-hand and coiled-coil domain containing 1 |
| chr7_-_68373495 | 1.22 |
ENSDART00000167440
|
zfhx3
|
zinc finger homeobox 3 |
| chr19_-_31522625 | 1.20 |
ENSDART00000158438
ENSDART00000035049 |
necab1
|
N-terminal EF-hand calcium binding protein 1 |
| chr19_+_5674907 | 1.20 |
ENSDART00000042189
|
pdk2b
|
pyruvate dehydrogenase kinase, isozyme 2b |
| chr1_+_44911405 | 1.19 |
ENSDART00000182465
|
wu:fc21g02
|
wu:fc21g02 |
| chr9_-_12424791 | 1.18 |
ENSDART00000135447
ENSDART00000088199 |
zgc:162707
|
zgc:162707 |
| chr13_-_39947335 | 1.18 |
ENSDART00000056996
|
sfrp5
|
secreted frizzled-related protein 5 |
| chr6_-_50204262 | 1.16 |
ENSDART00000163648
|
raly
|
RALY heterogeneous nuclear ribonucleoprotein |
| chr11_+_24290130 | 1.16 |
ENSDART00000029695
|
pnp4a
|
purine nucleoside phosphorylase 4a |
| chr4_-_22310956 | 1.16 |
ENSDART00000162585
|
hcls1
|
hematopoietic cell-specific Lyn substrate 1 |
| chr9_-_18716 | 1.15 |
ENSDART00000164763
|
CABZ01078737.1
|
|
| chr2_-_32558795 | 1.14 |
ENSDART00000140026
|
smarcd3a
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 3a |
| chr21_+_2091895 | 1.14 |
ENSDART00000161828
|
si:rp71-1h20.9
|
si:rp71-1h20.9 |
| chr7_-_52417777 | 1.12 |
ENSDART00000110265
|
myzap
|
myocardial zonula adherens protein |
| chr10_+_8767541 | 1.10 |
ENSDART00000170272
|
itga1
|
integrin, alpha 1 |
| chr10_-_42685512 | 1.09 |
ENSDART00000081347
|
stc1l
|
stanniocalcin 1, like |
| chr15_-_20933574 | 1.09 |
ENSDART00000152648
ENSDART00000152448 ENSDART00000152244 |
usp2a
|
ubiquitin specific peptidase 2a |
| chr4_-_9722568 | 1.09 |
ENSDART00000067190
|
tspan9b
|
tetraspanin 9b |
| chr13_-_50139916 | 1.07 |
ENSDART00000099475
|
nid1a
|
nidogen 1a |
| chr18_-_44610992 | 1.07 |
ENSDART00000125968
ENSDART00000185836 |
spred3
|
sprouty-related, EVH1 domain containing 3 |
| chr19_+_18797623 | 1.07 |
ENSDART00000166172
|
ddah2
|
dimethylarginine dimethylaminohydrolase 2 |
| chr6_-_48094342 | 1.06 |
ENSDART00000137458
|
slc2a1b
|
solute carrier family 2 (facilitated glucose transporter), member 1b |
| chr6_-_19023468 | 1.06 |
ENSDART00000184729
|
sept9b
|
septin 9b |
| chr1_+_37391141 | 1.04 |
ENSDART00000083593
ENSDART00000168647 |
sparcl1
|
SPARC-like 1 |
| chr25_+_8955530 | 1.03 |
ENSDART00000156444
|
si:ch211-256a21.4
|
si:ch211-256a21.4 |
| chr12_+_31713239 | 1.03 |
ENSDART00000122379
|
habp2
|
hyaluronan binding protein 2 |
| chr1_+_19303241 | 1.03 |
ENSDART00000129970
|
si:dkeyp-118a3.2
|
si:dkeyp-118a3.2 |
| chr3_-_28209001 | 1.03 |
ENSDART00000151178
|
rbfox1
|
RNA binding fox-1 homolog 1 |
| chr11_-_36350421 | 1.03 |
ENSDART00000141477
|
psma5
|
proteasome subunit alpha 5 |
| chr8_-_34051548 | 1.02 |
ENSDART00000105204
|
pbx3b
|
pre-B-cell leukemia homeobox 3b |
| chr13_+_1575276 | 1.02 |
ENSDART00000165987
|
DST
|
dystonin |
| chr1_-_41192059 | 1.02 |
ENSDART00000084665
ENSDART00000135369 |
dok7
|
docking protein 7 |
| chr3_+_45687266 | 1.02 |
ENSDART00000131652
|
gpr146
|
G protein-coupled receptor 146 |
| chr11_-_11471857 | 1.01 |
ENSDART00000030103
|
krt94
|
keratin 94 |
| chr9_-_8454060 | 1.01 |
ENSDART00000110158
|
irs2b
|
insulin receptor substrate 2b |
| chr13_-_42749916 | 1.01 |
ENSDART00000140019
|
capn2a
|
calpain 2, (m/II) large subunit a |
| chr21_+_29077509 | 1.00 |
ENSDART00000128561
|
ecscr
|
endothelial cell surface expressed chemotaxis and apoptosis regulator |
| chr13_+_42544009 | 1.00 |
ENSDART00000145409
|
si:dkey-221j11.3
|
si:dkey-221j11.3 |
| chr8_-_19904124 | 0.99 |
ENSDART00000129193
|
trabd2b
|
TraB domain containing 2B |
| chr21_-_43131752 | 0.99 |
ENSDART00000024137
|
p4ha2
|
procollagen-proline, 2-oxoglutarate 4-dioxygenase (proline 4-hydroxylase), alpha polypeptide 2 |
| chr9_-_42989297 | 0.99 |
ENSDART00000126871
|
ttn.2
|
titin, tandem duplicate 2 |
| chr2_+_4207209 | 0.98 |
ENSDART00000157903
ENSDART00000166476 |
gata6
|
GATA binding protein 6 |
| chr13_+_32740509 | 0.97 |
ENSDART00000076423
ENSDART00000160138 |
sobpa
|
sine oculis binding protein homolog (Drosophila) a |
| chr17_+_51499789 | 0.96 |
ENSDART00000187701
|
CABZ01067581.1
|
|
| chr11_+_10541258 | 0.96 |
ENSDART00000132365
|
b3gnt5a
|
UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 5a |
| chr24_-_17029374 | 0.96 |
ENSDART00000039267
|
ptgdsb.1
|
prostaglandin D2 synthase b, tandem duplicate 1 |
| chr1_-_51474974 | 0.96 |
ENSDART00000152719
|
spred2a
|
sprouty-related, EVH1 domain containing 2a |
| chr7_-_33961551 | 0.95 |
ENSDART00000100104
|
skor1a
|
SKI family transcriptional corepressor 1a |
| chr6_+_53349966 | 0.94 |
ENSDART00000167079
|
si:ch211-161c3.5
|
si:ch211-161c3.5 |
| chr5_-_34997630 | 0.94 |
ENSDART00000170684
|
btf3
|
basic transcription factor 3 |
| chr14_+_790166 | 0.94 |
ENSDART00000123912
|
adra2da
|
adrenergic, alpha-2D-, receptor a |
| chr7_+_30787903 | 0.94 |
ENSDART00000174000
|
apba2b
|
amyloid beta (A4) precursor protein-binding, family A, member 2b |
| chr6_-_49673476 | 0.94 |
ENSDART00000112226
|
apcdd1l
|
adenomatosis polyposis coli down-regulated 1-like |
| chr11_-_31039533 | 0.93 |
ENSDART00000127355
|
ier2b
|
immediate early response 2b |
| chr12_+_17106117 | 0.93 |
ENSDART00000149990
|
acta2
|
actin, alpha 2, smooth muscle, aorta |
| chr8_+_7144066 | 0.93 |
ENSDART00000146306
|
slc6a6a
|
solute carrier family 6 (neurotransmitter transporter), member 6a |
| chr20_+_1996202 | 0.92 |
ENSDART00000184143
|
CABZ01092781.1
|
|
| chr17_+_33340675 | 0.92 |
ENSDART00000184396
ENSDART00000077553 |
xdh
|
xanthine dehydrogenase |
| chr8_+_40628926 | 0.92 |
ENSDART00000163598
|
dusp2
|
dual specificity phosphatase 2 |
| chr17_+_24848976 | 0.91 |
ENSDART00000062917
|
cx35.4
|
connexin 35.4 |
| chr13_+_4205724 | 0.91 |
ENSDART00000134105
|
dlk2
|
delta-like 2 homolog (Drosophila) |
| chr8_-_21268303 | 0.91 |
ENSDART00000067211
|
gpr37l1b
|
G protein-coupled receptor 37 like 1b |
| chr13_-_40401870 | 0.90 |
ENSDART00000128951
|
nkx3.3
|
NK3 homeobox 3 |
| chr5_-_48307804 | 0.88 |
ENSDART00000182831
ENSDART00000186920 ENSDART00000183585 |
mef2cb
|
myocyte enhancer factor 2cb |
| chr24_+_15670013 | 0.88 |
ENSDART00000185826
|
CU929414.1
|
|
| chr19_-_46775562 | 0.88 |
ENSDART00000187144
|
abrab
|
actin binding Rho activating protein b |
| chr15_-_1066269 | 0.88 |
ENSDART00000093133
|
aadac
|
arylacetamide deacetylase |
| chr19_-_38611814 | 0.87 |
ENSDART00000151958
|
col16a1
|
collagen, type XVI, alpha 1 |
| chr16_+_40301056 | 0.87 |
ENSDART00000058578
|
rspo3
|
R-spondin 3 |
| chr9_+_6587056 | 0.86 |
ENSDART00000193421
|
fhl2a
|
four and a half LIM domains 2a |
| chr13_-_27038407 | 0.85 |
ENSDART00000146712
|
ccdc85a
|
coiled-coil domain containing 85A |
| chr9_+_4429593 | 0.83 |
ENSDART00000184855
|
FP015810.1
|
|
| chr2_-_13216269 | 0.83 |
ENSDART00000149947
|
bcl2b
|
BCL2, apoptosis regulator b |
| chr19_-_10243148 | 0.83 |
ENSDART00000148073
|
shisa7b
|
shisa family member 7 |
| chr13_+_49175947 | 0.83 |
ENSDART00000056927
|
egln1a
|
egl-9 family hypoxia-inducible factor 1a |
| chr10_-_35236949 | 0.82 |
ENSDART00000145804
|
ypel2a
|
yippee-like 2a |
| chr25_-_2081371 | 0.82 |
ENSDART00000104915
ENSDART00000156925 |
wnt7bb
|
wingless-type MMTV integration site family, member 7Bb |
| chr22_+_13886821 | 0.81 |
ENSDART00000130585
ENSDART00000105711 |
sh3bp4a
|
SH3-domain binding protein 4a |
Network of associatons between targets according to the STRING database.
First level regulatory network of ddx28+nfatc1+nfatc2b+nfatc3a+nfatc3b+nfatc4
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.3 | 3.9 | GO:0006957 | complement activation, alternative pathway(GO:0006957) |
| 0.8 | 3.9 | GO:0097066 | response to thyroid hormone(GO:0097066) |
| 0.7 | 2.8 | GO:0061549 | sympathetic ganglion development(GO:0061549) |
| 0.7 | 2.1 | GO:0007414 | axonal defasciculation(GO:0007414) |
| 0.7 | 2.7 | GO:0010226 | response to lithium ion(GO:0010226) |
| 0.6 | 1.8 | GO:0015695 | organic cation transport(GO:0015695) |
| 0.6 | 1.2 | GO:0061031 | endodermal digestive tract morphogenesis(GO:0061031) |
| 0.5 | 2.2 | GO:2000171 | negative regulation of dendrite development(GO:2000171) |
| 0.5 | 2.1 | GO:0048485 | sympathetic nervous system development(GO:0048485) |
| 0.5 | 3.4 | GO:0060221 | retinal rod cell differentiation(GO:0060221) |
| 0.5 | 2.7 | GO:0035881 | amacrine cell differentiation(GO:0035881) |
| 0.5 | 1.8 | GO:0019371 | cyclooxygenase pathway(GO:0019371) |
| 0.4 | 1.8 | GO:0061113 | pancreas morphogenesis(GO:0061113) |
| 0.4 | 2.6 | GO:0010799 | regulation of peptidyl-threonine phosphorylation(GO:0010799) negative regulation of peptidyl-threonine phosphorylation(GO:0010801) |
| 0.4 | 2.4 | GO:0021982 | pineal gland development(GO:0021982) |
| 0.4 | 2.3 | GO:0055014 | atrial cardiac muscle cell development(GO:0055014) |
| 0.4 | 1.5 | GO:1902024 | L-histidine transmembrane transport(GO:0089709) L-histidine transport(GO:1902024) |
| 0.4 | 1.9 | GO:0007412 | axon target recognition(GO:0007412) |
| 0.4 | 1.4 | GO:0042776 | mitochondrial ATP synthesis coupled proton transport(GO:0042776) |
| 0.3 | 1.3 | GO:0001778 | plasma membrane repair(GO:0001778) chondrocyte differentiation involved in endochondral bone morphogenesis(GO:0003413) growth plate cartilage chondrocyte differentiation(GO:0003418) |
| 0.3 | 1.3 | GO:0015871 | choline transport(GO:0015871) |
| 0.3 | 1.3 | GO:0070166 | enamel mineralization(GO:0070166) |
| 0.3 | 1.6 | GO:0051145 | smooth muscle cell differentiation(GO:0051145) |
| 0.3 | 0.9 | GO:0090131 | mesenchyme migration(GO:0090131) |
| 0.3 | 0.9 | GO:0006145 | purine nucleobase catabolic process(GO:0006145) |
| 0.3 | 1.1 | GO:1903428 | regulation of nitric oxide biosynthetic process(GO:0045428) positive regulation of nitric oxide biosynthetic process(GO:0045429) positive regulation of reactive oxygen species biosynthetic process(GO:1903428) positive regulation of nitric oxide metabolic process(GO:1904407) |
| 0.3 | 0.8 | GO:0070914 | UV-damage excision repair(GO:0070914) |
| 0.3 | 1.8 | GO:0033278 | cell proliferation in midbrain(GO:0033278) |
| 0.3 | 0.8 | GO:0061341 | non-canonical Wnt signaling pathway involved in heart development(GO:0061341) |
| 0.3 | 3.3 | GO:0060561 | apoptotic process involved in morphogenesis(GO:0060561) |
| 0.2 | 2.7 | GO:0003209 | cardiac atrium morphogenesis(GO:0003209) |
| 0.2 | 2.0 | GO:0001958 | endochondral ossification(GO:0001958) replacement ossification(GO:0036075) |
| 0.2 | 0.9 | GO:0060829 | negative regulation of canonical Wnt signaling pathway involved in neural plate anterior/posterior pattern formation(GO:0060829) |
| 0.2 | 2.1 | GO:0043490 | acidic amino acid transport(GO:0015800) aspartate transport(GO:0015810) L-glutamate transport(GO:0015813) malate-aspartate shuttle(GO:0043490) |
| 0.2 | 1.5 | GO:0090104 | pancreatic D cell differentiation(GO:0003311) pancreatic epsilon cell differentiation(GO:0090104) |
| 0.2 | 0.8 | GO:0034260 | negative regulation of GTPase activity(GO:0034260) |
| 0.2 | 0.6 | GO:1904088 | regulation of convergent extension involved in axis elongation(GO:1901232) positive regulation of epiboly involved in gastrulation with mouth forming second(GO:1904088) |
| 0.2 | 1.0 | GO:0001773 | myeloid dendritic cell activation(GO:0001773) myeloid dendritic cell differentiation(GO:0043011) |
| 0.2 | 0.8 | GO:0009826 | unidimensional cell growth(GO:0009826) |
| 0.2 | 0.4 | GO:0060827 | regulation of canonical Wnt signaling pathway involved in neural plate anterior/posterior pattern formation(GO:0060827) |
| 0.2 | 0.8 | GO:0071675 | regulation of macrophage chemotaxis(GO:0010758) positive regulation of macrophage chemotaxis(GO:0010759) regulation of odontogenesis(GO:0042481) mononuclear cell migration(GO:0071674) regulation of mononuclear cell migration(GO:0071675) |
| 0.2 | 0.8 | GO:0061010 | gall bladder development(GO:0061010) |
| 0.2 | 0.9 | GO:0071881 | adenylate cyclase-inhibiting adrenergic receptor signaling pathway(GO:0071881) |
| 0.2 | 0.7 | GO:0060406 | copulation(GO:0007620) regulation of epinephrine secretion(GO:0014060) positive regulation of epinephrine secretion(GO:0032812) positive regulation of catecholamine secretion(GO:0033605) penile erection(GO:0043084) epinephrine transport(GO:0048241) epinephrine secretion(GO:0048242) regulation of penile erection(GO:0060405) positive regulation of penile erection(GO:0060406) prolactin secretion(GO:0070459) |
| 0.2 | 0.5 | GO:0007509 | mesoderm migration involved in gastrulation(GO:0007509) |
| 0.2 | 4.0 | GO:0015701 | bicarbonate transport(GO:0015701) |
| 0.2 | 1.2 | GO:0042983 | amyloid precursor protein biosynthetic process(GO:0042983) regulation of amyloid precursor protein biosynthetic process(GO:0042984) |
| 0.2 | 1.1 | GO:0016539 | intein-mediated protein splicing(GO:0016539) protein splicing(GO:0030908) |
| 0.1 | 0.6 | GO:0014909 | smooth muscle cell migration(GO:0014909) |
| 0.1 | 0.4 | GO:0035142 | hindbrain structural organization(GO:0021577) dorsal fin morphogenesis(GO:0035142) axial mesoderm structural organization(GO:0048331) mesoderm structural organization(GO:0048338) cell motility involved in somitogenic axis elongation(GO:0090247) cell migration involved in somitogenic axis elongation(GO:0090248) |
| 0.1 | 2.5 | GO:0018401 | peptidyl-proline hydroxylation to 4-hydroxy-L-proline(GO:0018401) 4-hydroxyproline metabolic process(GO:0019471) |
| 0.1 | 1.1 | GO:0032986 | nucleosome disassembly(GO:0006337) chromatin disassembly(GO:0031498) protein-DNA complex disassembly(GO:0032986) |
| 0.1 | 3.0 | GO:0007634 | optokinetic behavior(GO:0007634) |
| 0.1 | 1.0 | GO:0035479 | angioblast cell migration from lateral mesoderm to midline(GO:0035479) |
| 0.1 | 1.2 | GO:0051597 | response to methylmercury(GO:0051597) |
| 0.1 | 0.5 | GO:1903251 | multi-ciliated epithelial cell differentiation(GO:1903251) |
| 0.1 | 0.4 | GO:0045903 | positive regulation of translational fidelity(GO:0045903) |
| 0.1 | 0.7 | GO:0090243 | fibroblast growth factor receptor signaling pathway involved in somitogenesis(GO:0090243) |
| 0.1 | 0.8 | GO:0048172 | regulation of short-term neuronal synaptic plasticity(GO:0048172) |
| 0.1 | 0.4 | GO:0038003 | opioid receptor signaling pathway(GO:0038003) |
| 0.1 | 1.0 | GO:0048769 | sarcomerogenesis(GO:0048769) |
| 0.1 | 0.6 | GO:0009312 | oligosaccharide biosynthetic process(GO:0009312) |
| 0.1 | 0.4 | GO:0050938 | regulation of xanthophore differentiation(GO:0050938) |
| 0.1 | 1.8 | GO:0007622 | rhythmic behavior(GO:0007622) circadian behavior(GO:0048512) |
| 0.1 | 1.6 | GO:0048679 | regulation of axon regeneration(GO:0048679) regulation of neuron projection regeneration(GO:0070570) |
| 0.1 | 2.4 | GO:0006826 | iron ion transport(GO:0006826) |
| 0.1 | 0.2 | GO:0045579 | positive regulation of B cell differentiation(GO:0045579) |
| 0.1 | 0.6 | GO:0030240 | skeletal muscle thin filament assembly(GO:0030240) pointed-end actin filament capping(GO:0051694) |
| 0.1 | 0.6 | GO:0051013 | microtubule severing(GO:0051013) |
| 0.1 | 1.3 | GO:0048070 | regulation of developmental pigmentation(GO:0048070) |
| 0.1 | 0.3 | GO:0072337 | modified amino acid transport(GO:0072337) |
| 0.1 | 0.8 | GO:0061026 | cardiac muscle tissue regeneration(GO:0061026) |
| 0.1 | 0.7 | GO:0038007 | netrin-activated signaling pathway(GO:0038007) |
| 0.1 | 0.9 | GO:0035970 | peptidyl-threonine dephosphorylation(GO:0035970) |
| 0.1 | 0.8 | GO:0021520 | spinal cord motor neuron cell fate specification(GO:0021520) |
| 0.1 | 0.5 | GO:0048790 | maintenance of presynaptic active zone structure(GO:0048790) maintenance of synapse structure(GO:0099558) |
| 0.1 | 0.5 | GO:0021794 | thalamus development(GO:0021794) |
| 0.1 | 5.9 | GO:0035023 | regulation of Rho protein signal transduction(GO:0035023) |
| 0.1 | 1.5 | GO:0060219 | camera-type eye photoreceptor cell differentiation(GO:0060219) |
| 0.1 | 1.3 | GO:0055003 | cardiac myofibril assembly(GO:0055003) |
| 0.1 | 2.5 | GO:0010675 | regulation of cellular carbohydrate metabolic process(GO:0010675) regulation of glucose metabolic process(GO:0010906) |
| 0.1 | 1.4 | GO:0035845 | photoreceptor cell outer segment organization(GO:0035845) |
| 0.1 | 1.7 | GO:0035118 | embryonic pectoral fin morphogenesis(GO:0035118) |
| 0.1 | 1.6 | GO:0033627 | cell adhesion mediated by integrin(GO:0033627) |
| 0.1 | 0.2 | GO:0036363 | transforming growth factor beta activation(GO:0036363) regulation of transforming growth factor beta production(GO:0071634) negative regulation of transforming growth factor beta production(GO:0071635) |
| 0.1 | 1.4 | GO:0070831 | basement membrane assembly(GO:0070831) |
| 0.0 | 3.0 | GO:0010842 | retina layer formation(GO:0010842) |
| 0.0 | 0.9 | GO:0043049 | otic placode formation(GO:0043049) |
| 0.0 | 2.5 | GO:0046928 | regulation of neurotransmitter secretion(GO:0046928) |
| 0.0 | 0.3 | GO:0097476 | spinal cord motor neuron migration(GO:0097476) lateral motor column neuron migration(GO:0097477) |
| 0.0 | 0.2 | GO:0038083 | peptidyl-tyrosine autophosphorylation(GO:0038083) |
| 0.0 | 0.2 | GO:0010735 | positive regulation of transcription via serum response element binding(GO:0010735) |
| 0.0 | 0.4 | GO:0048208 | vesicle targeting, rough ER to cis-Golgi(GO:0048207) COPII vesicle coating(GO:0048208) |
| 0.0 | 0.5 | GO:0048672 | positive regulation of collateral sprouting(GO:0048672) |
| 0.0 | 0.6 | GO:0016556 | mRNA modification(GO:0016556) |
| 0.0 | 0.7 | GO:0050921 | positive regulation of chemotaxis(GO:0050921) regulation of positive chemotaxis(GO:0050926) positive regulation of positive chemotaxis(GO:0050927) induction of positive chemotaxis(GO:0050930) |
| 0.0 | 1.7 | GO:0042742 | defense response to bacterium(GO:0042742) |
| 0.0 | 7.8 | GO:0048704 | embryonic skeletal system morphogenesis(GO:0048704) |
| 0.0 | 0.3 | GO:0048635 | negative regulation of muscle organ development(GO:0048635) |
| 0.0 | 0.8 | GO:0097192 | signal transduction in absence of ligand(GO:0038034) extrinsic apoptotic signaling pathway in absence of ligand(GO:0097192) |
| 0.0 | 1.0 | GO:0008286 | insulin receptor signaling pathway(GO:0008286) |
| 0.0 | 0.2 | GO:0070293 | renal absorption(GO:0070293) |
| 0.0 | 0.9 | GO:0072348 | sulfur compound transport(GO:0072348) |
| 0.0 | 0.5 | GO:0045807 | positive regulation of endocytosis(GO:0045807) |
| 0.0 | 1.0 | GO:0021854 | hypothalamus development(GO:0021854) |
| 0.0 | 0.1 | GO:0019323 | pentose catabolic process(GO:0019323) |
| 0.0 | 0.5 | GO:0071594 | B cell differentiation(GO:0030183) T cell differentiation in thymus(GO:0033077) thymocyte aggregation(GO:0071594) |
| 0.0 | 0.9 | GO:0045746 | negative regulation of Notch signaling pathway(GO:0045746) |
| 0.0 | 1.1 | GO:0032922 | circadian regulation of gene expression(GO:0032922) |
| 0.0 | 0.3 | GO:0036230 | granulocyte activation(GO:0036230) neutrophil activation(GO:0042119) |
| 0.0 | 16.9 | GO:0006955 | immune response(GO:0006955) |
| 0.0 | 0.7 | GO:0006884 | cell volume homeostasis(GO:0006884) |
| 0.0 | 0.6 | GO:0071910 | determination of liver left/right asymmetry(GO:0071910) |
| 0.0 | 1.6 | GO:0007193 | adenylate cyclase-inhibiting G-protein coupled receptor signaling pathway(GO:0007193) |
| 0.0 | 0.6 | GO:0043171 | peptide catabolic process(GO:0043171) |
| 0.0 | 0.9 | GO:0060914 | heart formation(GO:0060914) |
| 0.0 | 0.1 | GO:0003322 | pancreatic A cell development(GO:0003322) |
| 0.0 | 0.1 | GO:0006699 | bile acid biosynthetic process(GO:0006699) bile acid metabolic process(GO:0008206) |
| 0.0 | 0.2 | GO:0043153 | entrainment of circadian clock by photoperiod(GO:0043153) |
| 0.0 | 0.1 | GO:2000275 | regulation of oxidative phosphorylation uncoupler activity(GO:2000275) |
| 0.0 | 0.4 | GO:0000463 | maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000463) |
| 0.0 | 1.2 | GO:0030835 | regulation of actin filament depolymerization(GO:0030834) negative regulation of actin filament depolymerization(GO:0030835) |
| 0.0 | 0.2 | GO:0030816 | activation of adenylate cyclase activity(GO:0007190) positive regulation of cAMP metabolic process(GO:0030816) positive regulation of cAMP biosynthetic process(GO:0030819) positive regulation of adenylate cyclase activity(GO:0045762) |
| 0.0 | 1.3 | GO:0051592 | response to calcium ion(GO:0051592) |
| 0.0 | 0.6 | GO:0043280 | positive regulation of cysteine-type endopeptidase activity involved in apoptotic process(GO:0043280) positive regulation of cysteine-type endopeptidase activity(GO:2001056) |
| 0.0 | 0.7 | GO:0007548 | sex differentiation(GO:0007548) |
| 0.0 | 1.1 | GO:0007229 | integrin-mediated signaling pathway(GO:0007229) |
| 0.0 | 0.2 | GO:0030042 | actin filament depolymerization(GO:0030042) |
| 0.0 | 0.3 | GO:0022010 | central nervous system myelination(GO:0022010) axon ensheathment in central nervous system(GO:0032291) |
| 0.0 | 2.0 | GO:0006979 | response to oxidative stress(GO:0006979) |
| 0.0 | 1.0 | GO:0007596 | blood coagulation(GO:0007596) |
| 0.0 | 0.4 | GO:0032506 | cytokinetic process(GO:0032506) |
| 0.0 | 0.2 | GO:0030217 | T cell differentiation(GO:0030217) |
| 0.0 | 0.5 | GO:0051923 | sulfation(GO:0051923) |
| 0.0 | 0.3 | GO:0048168 | regulation of neuronal synaptic plasticity(GO:0048168) |
| 0.0 | 2.3 | GO:0030833 | regulation of actin filament polymerization(GO:0030833) |
| 0.0 | 0.4 | GO:0031114 | regulation of microtubule depolymerization(GO:0031114) |
| 0.0 | 0.4 | GO:0006891 | intra-Golgi vesicle-mediated transport(GO:0006891) |
| 0.0 | 0.1 | GO:0035912 | establishment of cell polarity involved in ameboidal cell migration(GO:0003365) aorta morphogenesis(GO:0035909) dorsal aorta morphogenesis(GO:0035912) |
| 0.0 | 0.2 | GO:0071456 | cellular response to hypoxia(GO:0071456) |
| 0.0 | 0.3 | GO:0048268 | clathrin coat assembly(GO:0048268) |
| 0.0 | 0.2 | GO:0030518 | intracellular steroid hormone receptor signaling pathway(GO:0030518) |
| 0.0 | 1.0 | GO:0007160 | cell-matrix adhesion(GO:0007160) |
| 0.0 | 0.3 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.0 | 0.7 | GO:0061640 | cytoskeleton-dependent cytokinesis(GO:0061640) |
| 0.0 | 0.2 | GO:0045116 | protein neddylation(GO:0045116) positive regulation of ubiquitin-protein transferase activity(GO:0051443) |
| 0.0 | 0.5 | GO:0043113 | receptor clustering(GO:0043113) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.8 | 2.4 | GO:0044279 | other organism(GO:0044215) other organism cell(GO:0044216) other organism part(GO:0044217) other organism cell membrane(GO:0044218) other organism membrane(GO:0044279) |
| 0.7 | 2.7 | GO:0071664 | catenin-TCF7L2 complex(GO:0071664) |
| 0.4 | 4.4 | GO:0045095 | keratin filament(GO:0045095) |
| 0.4 | 2.5 | GO:0016012 | sarcoglycan complex(GO:0016012) |
| 0.3 | 3.9 | GO:0033162 | melanosome membrane(GO:0033162) chitosome(GO:0045009) |
| 0.2 | 1.4 | GO:0045261 | proton-transporting ATP synthase complex, catalytic core F(1)(GO:0045261) |
| 0.2 | 0.8 | GO:0000110 | nucleotide-excision repair factor 1 complex(GO:0000110) |
| 0.2 | 1.4 | GO:0071914 | prominosome(GO:0071914) |
| 0.2 | 0.9 | GO:0005854 | nascent polypeptide-associated complex(GO:0005854) |
| 0.1 | 4.9 | GO:0005901 | caveola(GO:0005901) plasma membrane raft(GO:0044853) |
| 0.1 | 2.5 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.1 | 1.8 | GO:0002116 | semaphorin receptor complex(GO:0002116) |
| 0.1 | 0.4 | GO:0005850 | eukaryotic translation initiation factor 2 complex(GO:0005850) |
| 0.1 | 0.9 | GO:0019773 | proteasome core complex, alpha-subunit complex(GO:0019773) |
| 0.1 | 3.3 | GO:0031941 | filamentous actin(GO:0031941) |
| 0.1 | 1.1 | GO:0032591 | dendritic spine membrane(GO:0032591) |
| 0.1 | 2.9 | GO:0008305 | integrin complex(GO:0008305) |
| 0.1 | 1.4 | GO:0005605 | basal lamina(GO:0005605) laminin complex(GO:0043256) |
| 0.1 | 4.3 | GO:0005884 | actin filament(GO:0005884) |
| 0.1 | 0.9 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.1 | 16.4 | GO:0005764 | lysosome(GO:0005764) |
| 0.1 | 1.3 | GO:0042470 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.0 | 0.7 | GO:0090665 | dystrophin-associated glycoprotein complex(GO:0016010) glycoprotein complex(GO:0090665) |
| 0.0 | 0.7 | GO:0071564 | npBAF complex(GO:0071564) |
| 0.0 | 4.1 | GO:0016323 | basolateral plasma membrane(GO:0016323) |
| 0.0 | 0.8 | GO:0031430 | M band(GO:0031430) |
| 0.0 | 0.4 | GO:0014704 | intercalated disc(GO:0014704) |
| 0.0 | 0.6 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.0 | 2.0 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.0 | 0.3 | GO:0005665 | DNA-directed RNA polymerase II, core complex(GO:0005665) |
| 0.0 | 2.7 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.0 | 0.4 | GO:0036038 | MKS complex(GO:0036038) |
| 0.0 | 1.8 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.0 | 1.7 | GO:0030018 | Z disc(GO:0030018) |
| 0.0 | 1.1 | GO:0005604 | basement membrane(GO:0005604) |
| 0.0 | 0.2 | GO:0005688 | U6 snRNP(GO:0005688) |
| 0.0 | 0.2 | GO:0033165 | interphotoreceptor matrix(GO:0033165) |
| 0.0 | 0.7 | GO:0043204 | perikaryon(GO:0043204) |
| 0.0 | 0.7 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.0 | 1.2 | GO:0099501 | synaptic vesicle membrane(GO:0030672) exocytic vesicle membrane(GO:0099501) |
| 0.0 | 0.4 | GO:0030127 | COPII vesicle coat(GO:0030127) |
| 0.0 | 1.7 | GO:0000922 | spindle pole(GO:0000922) |
| 0.0 | 0.3 | GO:0043209 | myelin sheath(GO:0043209) |
| 0.0 | 0.5 | GO:0098831 | cytoskeleton of presynaptic active zone(GO:0048788) presynaptic active zone cytoplasmic component(GO:0098831) |
| 0.0 | 0.2 | GO:1990589 | ATF4-CREB1 transcription factor complex(GO:1990589) |
| 0.0 | 0.8 | GO:0016342 | catenin complex(GO:0016342) |
| 0.0 | 0.7 | GO:0000315 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.0 | 1.0 | GO:0005814 | centriole(GO:0005814) |
| 0.0 | 3.6 | GO:0031012 | extracellular matrix(GO:0031012) |
| 0.0 | 4.0 | GO:0043235 | receptor complex(GO:0043235) |
| 0.0 | 0.9 | GO:0030017 | sarcomere(GO:0030017) |
| 0.0 | 12.3 | GO:0005615 | extracellular space(GO:0005615) |
| 0.0 | 1.1 | GO:0005882 | intermediate filament(GO:0005882) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.8 | 2.5 | GO:0048030 | disaccharide binding(GO:0048030) |
| 0.5 | 1.8 | GO:0004666 | prostaglandin-endoperoxide synthase activity(GO:0004666) |
| 0.4 | 2.0 | GO:0047066 | phospholipid-hydroperoxide glutathione peroxidase activity(GO:0047066) |
| 0.3 | 9.1 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.3 | 1.3 | GO:0015220 | choline transmembrane transporter activity(GO:0015220) |
| 0.3 | 1.0 | GO:0047256 | beta-galactosyl-N-acetylglucosaminylgalactosylglucosyl-ceramide beta-1,3-acetylglucosaminyltransferase activity(GO:0008457) lactosylceramide 1,3-N-acetyl-beta-D-glucosaminyltransferase activity(GO:0047256) |
| 0.2 | 4.0 | GO:0005452 | inorganic anion exchanger activity(GO:0005452) |
| 0.2 | 1.3 | GO:0004962 | endothelin receptor activity(GO:0004962) |
| 0.2 | 1.3 | GO:0030274 | LIM domain binding(GO:0030274) |
| 0.2 | 2.1 | GO:0015183 | L-aspartate transmembrane transporter activity(GO:0015183) |
| 0.2 | 3.4 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) alpha-actinin binding(GO:0051393) |
| 0.2 | 2.4 | GO:0031682 | G-protein gamma-subunit binding(GO:0031682) |
| 0.2 | 0.9 | GO:0004938 | alpha2-adrenergic receptor activity(GO:0004938) |
| 0.2 | 0.8 | GO:0016519 | gastric inhibitory peptide receptor activity(GO:0016519) |
| 0.2 | 1.5 | GO:0015186 | L-glutamine transmembrane transporter activity(GO:0015186) |
| 0.2 | 1.1 | GO:0055056 | D-glucose transmembrane transporter activity(GO:0055056) |
| 0.2 | 1.2 | GO:0004740 | pyruvate dehydrogenase (acetyl-transferring) kinase activity(GO:0004740) |
| 0.1 | 1.3 | GO:0008420 | CTD phosphatase activity(GO:0008420) |
| 0.1 | 0.7 | GO:0042609 | CD4 receptor binding(GO:0042609) |
| 0.1 | 0.6 | GO:0005017 | platelet-derived growth factor-activated receptor activity(GO:0005017) platelet-derived growth factor binding(GO:0048407) |
| 0.1 | 21.7 | GO:0004197 | cysteine-type endopeptidase activity(GO:0004197) |
| 0.1 | 1.8 | GO:0005007 | fibroblast growth factor-activated receptor activity(GO:0005007) |
| 0.1 | 1.4 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 0.1 | 0.4 | GO:0005183 | gonadotropin hormone-releasing hormone activity(GO:0005183) gonadotropin-releasing hormone receptor binding(GO:0031530) |
| 0.1 | 1.2 | GO:0004731 | purine-nucleoside phosphorylase activity(GO:0004731) |
| 0.1 | 1.8 | GO:0004029 | aldehyde dehydrogenase (NAD) activity(GO:0004029) |
| 0.1 | 2.5 | GO:0031545 | peptidyl-proline 4-dioxygenase activity(GO:0031545) |
| 0.1 | 0.4 | GO:0038046 | enkephalin receptor activity(GO:0038046) |
| 0.1 | 1.0 | GO:0001972 | retinoic acid binding(GO:0001972) |
| 0.1 | 1.1 | GO:0005113 | patched binding(GO:0005113) |
| 0.1 | 1.1 | GO:0016813 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amidines(GO:0016813) |
| 0.1 | 0.5 | GO:0031697 | beta-1 adrenergic receptor binding(GO:0031697) type II activin receptor binding(GO:0070699) |
| 0.1 | 2.1 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.1 | 0.7 | GO:0070180 | large ribosomal subunit rRNA binding(GO:0070180) |
| 0.1 | 0.5 | GO:0046592 | polyamine oxidase activity(GO:0046592) |
| 0.1 | 0.9 | GO:0016725 | oxidoreductase activity, acting on CH or CH2 groups(GO:0016725) |
| 0.1 | 3.9 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.1 | 0.9 | GO:0099583 | neurotransmitter receptor activity involved in regulation of postsynaptic cytosolic calcium ion concentration(GO:0099583) |
| 0.1 | 0.7 | GO:0038036 | sphingosine-1-phosphate receptor activity(GO:0038036) |
| 0.1 | 1.9 | GO:0046875 | ephrin receptor binding(GO:0046875) |
| 0.1 | 0.8 | GO:0004499 | N,N-dimethylaniline monooxygenase activity(GO:0004499) |
| 0.1 | 0.3 | GO:0005153 | interleukin-8 receptor binding(GO:0005153) |
| 0.1 | 1.0 | GO:0005158 | insulin receptor binding(GO:0005158) |
| 0.1 | 0.5 | GO:0034714 | type III transforming growth factor beta receptor binding(GO:0034714) |
| 0.1 | 0.7 | GO:0008273 | calcium, potassium:sodium antiporter activity(GO:0008273) |
| 0.1 | 0.7 | GO:0008511 | sodium:potassium:chloride symporter activity(GO:0008511) |
| 0.1 | 3.9 | GO:0008013 | beta-catenin binding(GO:0008013) |
| 0.1 | 1.5 | GO:0051019 | mitogen-activated protein kinase binding(GO:0051019) |
| 0.1 | 0.4 | GO:0008158 | hedgehog receptor activity(GO:0008158) |
| 0.1 | 0.4 | GO:1990756 | protein binding, bridging involved in substrate recognition for ubiquitination(GO:1990756) |
| 0.1 | 0.7 | GO:0005042 | netrin receptor activity(GO:0005042) |
| 0.1 | 1.8 | GO:0008171 | O-methyltransferase activity(GO:0008171) |
| 0.1 | 1.0 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.1 | 0.3 | GO:0070324 | thyroid hormone binding(GO:0070324) |
| 0.1 | 0.4 | GO:0016316 | phosphatidylinositol-3,4-bisphosphate 4-phosphatase activity(GO:0016316) |
| 0.1 | 0.7 | GO:0015101 | organic cation transmembrane transporter activity(GO:0015101) |
| 0.1 | 8.5 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.1 | 0.3 | GO:0004960 | thromboxane receptor activity(GO:0004960) |
| 0.1 | 1.6 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.1 | 0.5 | GO:0004062 | aryl sulfotransferase activity(GO:0004062) |
| 0.0 | 8.2 | GO:0001228 | transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001228) |
| 0.0 | 0.8 | GO:0005092 | GDP-dissociation inhibitor activity(GO:0005092) |
| 0.0 | 2.4 | GO:0015485 | cholesterol binding(GO:0015485) |
| 0.0 | 0.2 | GO:0043914 | NADPH:sulfur oxidoreductase activity(GO:0043914) |
| 0.0 | 2.4 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.0 | 1.6 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.0 | 0.6 | GO:2001069 | glycogen binding(GO:2001069) |
| 0.0 | 1.8 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 0.5 | GO:0005165 | nerve growth factor receptor binding(GO:0005163) neurotrophin receptor binding(GO:0005165) |
| 0.0 | 0.9 | GO:0005112 | Notch binding(GO:0005112) |
| 0.0 | 1.0 | GO:0005518 | collagen binding(GO:0005518) |
| 0.0 | 0.6 | GO:0003785 | actin monomer binding(GO:0003785) tropomyosin binding(GO:0005523) |
| 0.0 | 0.5 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.0 | 0.2 | GO:1903924 | estradiol binding(GO:1903924) |
| 0.0 | 0.5 | GO:0098882 | structural constituent of presynaptic active zone(GO:0098882) structural constituent of synapse(GO:0098918) |
| 0.0 | 0.9 | GO:0005343 | organic acid:sodium symporter activity(GO:0005343) |
| 0.0 | 0.3 | GO:0030507 | spectrin binding(GO:0030507) |
| 0.0 | 1.9 | GO:1902936 | phosphatidylinositol bisphosphate binding(GO:1902936) |
| 0.0 | 20.9 | GO:0005509 | calcium ion binding(GO:0005509) |
| 0.0 | 1.5 | GO:0008201 | heparin binding(GO:0008201) |
| 0.0 | 0.3 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 0.0 | 0.6 | GO:0070006 | metalloaminopeptidase activity(GO:0070006) |
| 0.0 | 0.8 | GO:0003684 | damaged DNA binding(GO:0003684) |
| 0.0 | 0.2 | GO:0016500 | protein-hormone receptor activity(GO:0016500) |
| 0.0 | 1.5 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.0 | 0.7 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.0 | 0.9 | GO:0042826 | histone deacetylase binding(GO:0042826) |
| 0.0 | 0.3 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.0 | 0.3 | GO:0030546 | receptor activator activity(GO:0030546) receptor agonist activity(GO:0048018) |
| 0.0 | 0.4 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.0 | 0.1 | GO:0015057 | thrombin receptor activity(GO:0015057) |
| 0.0 | 22.1 | GO:0000978 | RNA polymerase II core promoter proximal region sequence-specific DNA binding(GO:0000978) |
| 0.0 | 0.6 | GO:0003727 | single-stranded RNA binding(GO:0003727) |
| 0.0 | 1.3 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.0 | 1.1 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 0.3 | GO:0003899 | DNA-directed RNA polymerase activity(GO:0003899) RNA polymerase activity(GO:0034062) |
| 0.0 | 1.2 | GO:0003730 | mRNA 3'-UTR binding(GO:0003730) |
| 0.0 | 0.2 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.0 | 0.1 | GO:0015280 | ligand-gated sodium channel activity(GO:0015280) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.0 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
| 0.1 | 1.1 | PID LYMPH ANGIOGENESIS PATHWAY | VEGFR3 signaling in lymphatic endothelium |
| 0.1 | 1.3 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.1 | 0.8 | PID S1P S1P2 PATHWAY | S1P2 pathway |
| 0.1 | 1.3 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.1 | 1.3 | PID IL3 PATHWAY | IL3-mediated signaling events |
| 0.1 | 2.5 | PID FCER1 PATHWAY | Fc-epsilon receptor I signaling in mast cells |
| 0.1 | 0.6 | PID S1P S1P1 PATHWAY | S1P1 pathway |
| 0.1 | 2.5 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.1 | 2.7 | SIG INSULIN RECEPTOR PATHWAY IN CARDIAC MYOCYTES | Genes related to the insulin receptor pathway |
| 0.1 | 4.7 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.1 | 0.5 | PID VEGFR1 PATHWAY | VEGFR1 specific signals |
| 0.1 | 2.4 | PID TAP63 PATHWAY | Validated transcriptional targets of TAp63 isoforms |
| 0.1 | 2.0 | PID FGF PATHWAY | FGF signaling pathway |
| 0.1 | 0.6 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.0 | 0.6 | PID IL12 STAT4 PATHWAY | IL12 signaling mediated by STAT4 |
| 0.0 | 0.7 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
| 0.0 | 1.9 | PID AP1 PATHWAY | AP-1 transcription factor network |
| 0.0 | 0.5 | PID MAPK TRK PATHWAY | Trk receptor signaling mediated by the MAPK pathway |
| 0.0 | 1.0 | PID HES HEY PATHWAY | Notch-mediated HES/HEY network |
| 0.0 | 0.2 | ST TYPE I INTERFERON PATHWAY | Type I Interferon (alpha/beta IFN) Pathway |
| 0.0 | 0.2 | PID PI3K PLC TRK PATHWAY | Trk receptor signaling mediated by PI3K and PLC-gamma |
| 0.0 | 0.2 | PID VEGFR1 2 PATHWAY | Signaling events mediated by VEGFR1 and VEGFR2 |
| 0.0 | 0.5 | PID AJDISS 2PATHWAY | Posttranslational regulation of adherens junction stability and dissassembly |
| 0.0 | 1.0 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.0 | 0.2 | PID ARF6 PATHWAY | Arf6 signaling events |
| 0.0 | 1.9 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 1.8 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.8 | REACTOME ORGANIC CATION ANION ZWITTERION TRANSPORT | Genes involved in Organic cation/anion/zwitterion transport |
| 0.3 | 1.8 | REACTOME FGFR2C LIGAND BINDING AND ACTIVATION | Genes involved in FGFR2c ligand binding and activation |
| 0.2 | 5.1 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.2 | 0.9 | REACTOME PURINE CATABOLISM | Genes involved in Purine catabolism |
| 0.2 | 5.7 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.1 | 3.9 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 0.1 | 1.2 | REACTOME ACTIVATION OF THE AP1 FAMILY OF TRANSCRIPTION FACTORS | Genes involved in Activation of the AP-1 family of transcription factors |
| 0.1 | 0.7 | REACTOME ROLE OF DCC IN REGULATING APOPTOSIS | Genes involved in Role of DCC in regulating apoptosis |
| 0.1 | 0.6 | REACTOME P38MAPK EVENTS | Genes involved in p38MAPK events |
| 0.1 | 1.1 | REACTOME SYNTHESIS OF PC | Genes involved in Synthesis of PC |
| 0.1 | 2.5 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.0 | 0.8 | REACTOME ADHERENS JUNCTIONS INTERACTIONS | Genes involved in Adherens junctions interactions |
| 0.0 | 0.5 | REACTOME SHC MEDIATED SIGNALLING | Genes involved in SHC-mediated signalling |
| 0.0 | 1.3 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |
| 0.0 | 0.2 | REACTOME THE NLRP3 INFLAMMASOME | Genes involved in The NLRP3 inflammasome |
| 0.0 | 0.8 | REACTOME FORMATION OF INCISION COMPLEX IN GG NER | Genes involved in Formation of incision complex in GG-NER |
| 0.0 | 0.1 | REACTOME SYNTHESIS OF BILE ACIDS AND BILE SALTS VIA 24 HYDROXYCHOLESTEROL | Genes involved in Synthesis of bile acids and bile salts via 24-hydroxycholesterol |
| 0.0 | 0.5 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION | Genes involved in TRAF6 mediated IRF7 activation |
| 0.0 | 0.1 | REACTOME CD28 DEPENDENT VAV1 PATHWAY | Genes involved in CD28 dependent Vav1 pathway |
| 0.0 | 0.9 | REACTOME CROSS PRESENTATION OF SOLUBLE EXOGENOUS ANTIGENS ENDOSOMES | Genes involved in Cross-presentation of soluble exogenous antigens (endosomes) |
| 0.0 | 2.4 | REACTOME 3 UTR MEDIATED TRANSLATIONAL REGULATION | Genes involved in 3' -UTR-mediated translational regulation |
| 0.0 | 0.2 | REACTOME REGULATION OF IFNA SIGNALING | Genes involved in Regulation of IFNA signaling |
| 0.0 | 1.0 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.0 | 0.5 | REACTOME PHASE1 FUNCTIONALIZATION OF COMPOUNDS | Genes involved in Phase 1 - Functionalization of compounds |
| 0.0 | 1.3 | REACTOME TRANSPORT OF INORGANIC CATIONS ANIONS AND AMINO ACIDS OLIGOPEPTIDES | Genes involved in Transport of inorganic cations/anions and amino acids/oligopeptides |
| 0.0 | 0.3 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.0 | 0.1 | REACTOME ADP SIGNALLING THROUGH P2RY12 | Genes involved in ADP signalling through P2Y purinoceptor 12 |