Project
PRJNA195909:zebrafish embryo and larva development
Navigation
Downloads
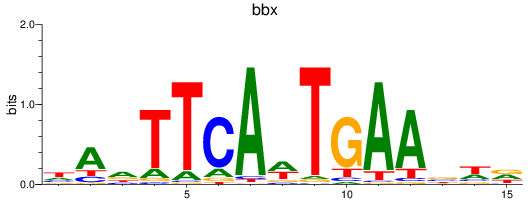
Results for bbx
Z-value: 0.51
Transcription factors associated with bbx
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
bbx
|
ENSDARG00000012699 | BBX high mobility group box domain containing |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| bbx | dr11_v1_chr10_-_28519505_28519505 | 0.38 | 3.1e-01 | Click! |
Activity profile of bbx motif
Sorted Z-values of bbx motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr13_-_7233811 | 0.45 |
ENSDART00000162026
|
ninl
|
ninein-like |
| chr9_+_48755638 | 0.33 |
ENSDART00000047401
|
abcb11a
|
ATP-binding cassette, sub-family B (MDR/TAP), member 11a |
| chr21_+_21263988 | 0.31 |
ENSDART00000089651
ENSDART00000108978 |
ccdc61
|
coiled-coil domain containing 61 |
| chr12_+_23812530 | 0.27 |
ENSDART00000066331
|
svila
|
supervillin a |
| chr17_-_51893123 | 0.22 |
ENSDART00000103350
ENSDART00000017329 |
numb
|
numb homolog (Drosophila) |
| chr18_-_26510545 | 0.21 |
ENSDART00000135133
|
FRMD5
|
si:ch211-69m14.1 |
| chr21_-_22678195 | 0.21 |
ENSDART00000171231
|
gig2g
|
grass carp reovirus (GCRV)-induced gene 2g |
| chr23_+_40139765 | 0.20 |
ENSDART00000185376
|
gpsm2l
|
G protein signaling modulator 2, like |
| chr7_+_24729558 | 0.20 |
ENSDART00000111542
ENSDART00000170100 |
shroom4
|
shroom family member 4 |
| chr12_-_9700605 | 0.20 |
ENSDART00000161063
|
heatr1
|
HEAT repeat containing 1 |
| chr15_+_44201056 | 0.18 |
ENSDART00000162433
ENSDART00000148336 |
CU655961.4
|
|
| chr14_-_4177311 | 0.17 |
ENSDART00000128129
|
si:dkey-185e18.7
|
si:dkey-185e18.7 |
| chr8_-_40205712 | 0.16 |
ENSDART00000158927
|
anapc5
|
anaphase promoting complex subunit 5 |
| chr6_+_30430591 | 0.16 |
ENSDART00000108943
|
shroom2a
|
shroom family member 2a |
| chr17_+_43867889 | 0.16 |
ENSDART00000132673
ENSDART00000167214 |
zgc:66313
|
zgc:66313 |
| chr8_+_28593707 | 0.16 |
ENSDART00000097213
|
tcf15
|
transcription factor 15 |
| chr15_-_15983428 | 0.16 |
ENSDART00000115129
|
synrg
|
synergin, gamma |
| chr13_+_23095228 | 0.16 |
ENSDART00000189068
ENSDART00000188624 |
pik3ap1
|
phosphoinositide-3-kinase adaptor protein 1 |
| chr1_+_35956435 | 0.16 |
ENSDART00000085021
ENSDART00000148505 |
mmaa
|
methylmalonic aciduria (cobalamin deficiency) cblA type |
| chr23_+_2703044 | 0.16 |
ENSDART00000182512
ENSDART00000105286 |
ncoa6
|
nuclear receptor coactivator 6 |
| chr24_+_41989108 | 0.16 |
ENSDART00000169725
|
zbtb14
|
zinc finger and BTB domain containing 14 |
| chr8_+_3431671 | 0.15 |
ENSDART00000017850
|
ctu1
|
cytosolic thiouridylase subunit 1 homolog (S. pombe) |
| chr19_-_438337 | 0.15 |
ENSDART00000127985
|
MTERF1
|
mitochondrial transcription termination factor 1 |
| chr24_-_9002038 | 0.15 |
ENSDART00000066783
ENSDART00000150185 |
mppe1
|
metallophosphoesterase 1 |
| chr8_+_31016180 | 0.15 |
ENSDART00000130870
ENSDART00000143604 |
odf2b
|
outer dense fiber of sperm tails 2b |
| chr12_-_18568834 | 0.15 |
ENSDART00000039693
|
pgp
|
phosphoglycolate phosphatase |
| chr11_-_18323059 | 0.15 |
ENSDART00000182590
|
SFMBT1
|
Scm like with four mbt domains 1 |
| chr16_+_12740790 | 0.15 |
ENSDART00000132957
|
epn1
|
epsin 1 |
| chr23_-_30781875 | 0.15 |
ENSDART00000114628
ENSDART00000180949 ENSDART00000191313 |
myt1a
|
myelin transcription factor 1a |
| chr16_-_30826712 | 0.15 |
ENSDART00000122474
|
ptk2ab
|
protein tyrosine kinase 2ab |
| chr13_-_24669258 | 0.14 |
ENSDART00000000831
|
znf511
|
zinc finger protein 511 |
| chr2_-_32237916 | 0.14 |
ENSDART00000141418
|
fam49ba
|
family with sequence similarity 49, member Ba |
| chr9_-_30555725 | 0.14 |
ENSDART00000079222
|
chaf1b
|
chromatin assembly factor 1, subunit B |
| chr15_-_15983183 | 0.14 |
ENSDART00000154841
|
synrg
|
synergin, gamma |
| chr3_+_32789605 | 0.14 |
ENSDART00000171895
|
tbc1d10b
|
TBC1 domain family, member 10b |
| chr15_-_31109760 | 0.13 |
ENSDART00000154254
|
ksr1b
|
kinase suppressor of ras 1b |
| chr16_-_27566552 | 0.13 |
ENSDART00000142102
|
zgc:153215
|
zgc:153215 |
| chr8_+_21114338 | 0.13 |
ENSDART00000002186
|
uck2a
|
uridine-cytidine kinase 2a |
| chr9_+_19623363 | 0.13 |
ENSDART00000142471
ENSDART00000147662 ENSDART00000136053 |
pdxka
|
pyridoxal (pyridoxine, vitamin B6) kinase a |
| chr17_-_8656155 | 0.13 |
ENSDART00000148990
|
ctbp2a
|
C-terminal binding protein 2a |
| chr18_+_39060687 | 0.13 |
ENSDART00000098729
ENSDART00000136367 |
zgc:171509
|
zgc:171509 |
| chr15_+_23528310 | 0.12 |
ENSDART00000152523
|
si:dkey-182i3.8
|
si:dkey-182i3.8 |
| chr10_-_26218354 | 0.12 |
ENSDART00000180764
|
arfip2b
|
ADP-ribosylation factor interacting protein 2b |
| chr11_-_7410537 | 0.12 |
ENSDART00000009859
|
adgrl4
|
adhesion G protein-coupled receptor L4 |
| chr17_+_21546993 | 0.12 |
ENSDART00000182387
|
chst15
|
carbohydrate (N-acetylgalactosamine 4-sulfate 6-O) sulfotransferase 15 |
| chr8_-_33154677 | 0.12 |
ENSDART00000133300
|
zbtb34
|
zinc finger and BTB domain containing 34 |
| chr13_+_6086730 | 0.12 |
ENSDART00000049328
|
fam120b
|
family with sequence similarity 120B |
| chr12_+_4222854 | 0.12 |
ENSDART00000144881
|
mapk7
|
mitogen-activated protein kinase 7 |
| chr17_+_8799451 | 0.12 |
ENSDART00000189814
ENSDART00000191577 |
tonsl
|
tonsoku-like, DNA repair protein |
| chr2_+_49246099 | 0.12 |
ENSDART00000179089
|
sema6ba
|
sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6Ba |
| chr4_-_14531687 | 0.11 |
ENSDART00000182093
ENSDART00000159447 |
plxnb2a
|
plexin b2a |
| chr25_+_16646113 | 0.11 |
ENSDART00000110426
|
cecr2
|
cat eye syndrome chromosome region, candidate 2 |
| chr23_-_30781510 | 0.11 |
ENSDART00000183364
|
myt1a
|
myelin transcription factor 1a |
| chr19_-_3773905 | 0.11 |
ENSDART00000168433
|
btr20
|
bloodthirsty-related gene family, member 20 |
| chr19_+_33093577 | 0.11 |
ENSDART00000180317
|
fam91a1
|
family with sequence similarity 91, member A1 |
| chr16_+_32014552 | 0.11 |
ENSDART00000047570
|
mboat7
|
membrane bound O-acyltransferase domain containing 7 |
| chr17_+_8799661 | 0.11 |
ENSDART00000105326
|
tonsl
|
tonsoku-like, DNA repair protein |
| chr12_-_33770299 | 0.11 |
ENSDART00000189849
|
llgl2
|
lethal giant larvae homolog 2 (Drosophila) |
| chr20_-_20248408 | 0.11 |
ENSDART00000183234
|
ppp2r5ea
|
protein phosphatase 2, regulatory subunit B', epsilon isoform a |
| chr8_+_23034718 | 0.11 |
ENSDART00000184512
|
ythdf1
|
YTH N(6)-methyladenosine RNA binding protein 1 |
| chr13_+_31205439 | 0.11 |
ENSDART00000132326
|
ptpn20
|
protein tyrosine phosphatase, non-receptor type 20 |
| chr7_+_6941583 | 0.10 |
ENSDART00000160709
ENSDART00000157634 |
rbm14b
|
RNA binding motif protein 14b |
| chr19_+_43341424 | 0.10 |
ENSDART00000134815
|
sesn2
|
sestrin 2 |
| chr9_-_7640692 | 0.10 |
ENSDART00000135616
|
dnajb2
|
DnaJ (Hsp40) homolog, subfamily B, member 2 |
| chr8_+_20438884 | 0.10 |
ENSDART00000016422
ENSDART00000133794 |
mknk2b
|
MAP kinase interacting serine/threonine kinase 2b |
| chr16_+_14707960 | 0.10 |
ENSDART00000137912
|
col14a1a
|
collagen, type XIV, alpha 1a |
| chr23_-_24343363 | 0.10 |
ENSDART00000166392
|
fam131c
|
family with sequence similarity 131, member C |
| chr6_-_40921412 | 0.10 |
ENSDART00000076097
ENSDART00000188364 |
sfi1
|
SFI1 centrin binding protein |
| chr17_+_39834947 | 0.10 |
ENSDART00000172260
|
LO018474.1
|
|
| chr13_+_18371208 | 0.10 |
ENSDART00000138172
|
ccar1
|
cell division cycle and apoptosis regulator 1 |
| chr17_+_22381215 | 0.10 |
ENSDART00000162670
ENSDART00000128875 |
slc8a1b
|
solute carrier family 8 (sodium/calcium exchanger), member 1b |
| chr7_-_1163829 | 0.09 |
ENSDART00000181937
|
CT030017.1
|
|
| chr18_-_6856380 | 0.09 |
ENSDART00000175747
|
ppp6r2b
|
protein phosphatase 6, regulatory subunit 2b |
| chr25_-_12730260 | 0.09 |
ENSDART00000171834
|
klhdc4
|
kelch domain containing 4 |
| chr1_+_55643198 | 0.09 |
ENSDART00000060693
|
adgre7
|
adhesion G protein-coupled receptor E7 |
| chr9_+_40546677 | 0.09 |
ENSDART00000132911
|
vwc2l
|
von Willebrand factor C domain containing protein 2-like |
| chr21_+_38732945 | 0.09 |
ENSDART00000076157
|
rab24
|
RAB24, member RAS oncogene family |
| chr19_+_33093395 | 0.08 |
ENSDART00000019459
|
fam91a1
|
family with sequence similarity 91, member A1 |
| chr1_+_2431956 | 0.08 |
ENSDART00000183832
|
farp1
|
FERM, RhoGEF (ARHGEF) and pleckstrin domain protein 1 (chondrocyte-derived) |
| chr18_+_17493859 | 0.08 |
ENSDART00000090754
|
si:dkey-102f14.5
|
si:dkey-102f14.5 |
| chr7_+_10701938 | 0.08 |
ENSDART00000158162
|
arnt2
|
aryl-hydrocarbon receptor nuclear translocator 2 |
| chr17_+_23549117 | 0.08 |
ENSDART00000063953
|
zgc:65997
|
zgc:65997 |
| chr15_-_4596623 | 0.08 |
ENSDART00000132227
|
eif4h
|
eukaryotic translation initiation factor 4h |
| chr9_-_48407408 | 0.08 |
ENSDART00000058248
|
zgc:172182
|
zgc:172182 |
| chr21_-_25416391 | 0.08 |
ENSDART00000114081
|
sms
|
spermine synthase |
| chr7_+_5910467 | 0.08 |
ENSDART00000173232
|
si:ch211-113a14.22
|
si:ch211-113a14.22 |
| chr13_+_18507592 | 0.08 |
ENSDART00000142622
|
si:ch211-198a12.6
|
si:ch211-198a12.6 |
| chr12_+_27156943 | 0.08 |
ENSDART00000153030
ENSDART00000001737 |
skap1
|
src kinase associated phosphoprotein 1 |
| chr17_+_44697604 | 0.07 |
ENSDART00000156625
|
pgfb
|
placental growth factor b |
| chr5_+_61556172 | 0.07 |
ENSDART00000131937
|
orai2
|
ORAI calcium release-activated calcium modulator 2 |
| chr21_-_22648007 | 0.07 |
ENSDART00000121788
|
gig2l
|
grass carp reovirus (GCRV)-induced gene 2l |
| chr7_-_50604626 | 0.07 |
ENSDART00000073903
ENSDART00000174031 |
crtc3
|
CREB regulated transcription coactivator 3 |
| chr1_-_28831848 | 0.07 |
ENSDART00000148536
|
GK3P
|
zgc:172295 |
| chr24_-_28229618 | 0.07 |
ENSDART00000145290
|
bcl2a
|
BCL2, apoptosis regulator a |
| chr17_+_8212477 | 0.07 |
ENSDART00000064665
|
slc18b1
|
solute carrier family 18, subfamily B, member 1 |
| chr13_-_23095006 | 0.07 |
ENSDART00000089242
|
kif1bp
|
kif1 binding protein |
| chr15_-_35212462 | 0.07 |
ENSDART00000043960
|
agfg1a
|
ArfGAP with FG repeats 1a |
| chr6_+_36839509 | 0.07 |
ENSDART00000190605
ENSDART00000104160 |
zgc:110788
|
zgc:110788 |
| chr1_+_29858032 | 0.07 |
ENSDART00000054066
|
zic2b
|
zic family member 2 (odd-paired homolog, Drosophila) b |
| chr12_+_41348969 | 0.06 |
ENSDART00000171352
|
si:ch211-27e6.1
|
si:ch211-27e6.1 |
| chr1_-_524433 | 0.06 |
ENSDART00000147610
|
si:ch73-41e3.7
|
si:ch73-41e3.7 |
| chr19_+_10860485 | 0.06 |
ENSDART00000169962
|
si:ch73-347e22.4
|
si:ch73-347e22.4 |
| chr3_-_34816893 | 0.06 |
ENSDART00000084448
ENSDART00000154696 |
psmd11a
|
proteasome 26S subunit, non-ATPase 11a |
| chr11_-_23322182 | 0.06 |
ENSDART00000111289
|
kiss1
|
KiSS-1 metastasis-suppressor |
| chr23_+_35504824 | 0.06 |
ENSDART00000082647
ENSDART00000159218 |
acp1
|
acid phosphatase 1 |
| chr12_+_34770531 | 0.06 |
ENSDART00000153320
|
slc38a10
|
solute carrier family 38, member 10 |
| chr10_-_34107919 | 0.06 |
ENSDART00000167428
ENSDART00000145279 |
pimr144
pimr146
|
Pim proto-oncogene, serine/threonine kinase, related 144 Pim proto-oncogene, serine/threonine kinase, related 146 |
| chr7_+_72003301 | 0.06 |
ENSDART00000012918
ENSDART00000182268 ENSDART00000185750 |
psmd9
|
proteasome 26S subunit, non-ATPase 9 |
| chr5_+_36650096 | 0.05 |
ENSDART00000111414
|
alkbh6
|
alkB homolog 6 |
| chr6_+_6780873 | 0.05 |
ENSDART00000011865
|
sec23b
|
Sec23 homolog B, COPII coat complex component |
| chr6_-_43922813 | 0.05 |
ENSDART00000123341
|
prok2
|
prokineticin 2 |
| chr16_+_25535993 | 0.05 |
ENSDART00000077436
|
mylipb
|
myosin regulatory light chain interacting protein b |
| chr2_+_22659787 | 0.05 |
ENSDART00000043956
|
zgc:161973
|
zgc:161973 |
| chr5_+_37744625 | 0.05 |
ENSDART00000014031
|
dpf2
|
D4, zinc and double PHD fingers family 2 |
| chr1_+_19433004 | 0.04 |
ENSDART00000133959
|
clockb
|
clock circadian regulator b |
| chr12_-_17686404 | 0.04 |
ENSDART00000079065
|
ccz1
|
CCZ1 homolog, vacuolar protein trafficking and biogenesis associated |
| chr6_-_57539141 | 0.04 |
ENSDART00000156967
|
itcha
|
itchy E3 ubiquitin protein ligase a |
| chr23_+_3721042 | 0.04 |
ENSDART00000143323
|
smim29
|
small integral membrane protein 29 |
| chr6_+_37301341 | 0.04 |
ENSDART00000104180
|
zranb2
|
zinc finger, RAN-binding domain containing 2 |
| chr4_+_34121902 | 0.04 |
ENSDART00000170225
|
si:ch211-223g7.6
|
si:ch211-223g7.6 |
| chr15_-_5222924 | 0.04 |
ENSDART00000128924
|
or128-4
|
odorant receptor, family E, subfamily 128, member 4 |
| chr18_+_26899316 | 0.04 |
ENSDART00000050230
|
tspan3a
|
tetraspanin 3a |
| chr3_+_54761569 | 0.04 |
ENSDART00000135913
ENSDART00000180983 |
si:ch211-74m13.1
|
si:ch211-74m13.1 |
| chr18_-_17724295 | 0.04 |
ENSDART00000121553
|
fam192a
|
family with sequence similarity 192, member A |
| chr5_-_42883761 | 0.04 |
ENSDART00000167374
|
BX323596.2
|
|
| chr1_-_46244523 | 0.04 |
ENSDART00000143908
|
si:ch211-138g9.3
|
si:ch211-138g9.3 |
| chr1_+_52481332 | 0.04 |
ENSDART00000074231
|
cldnd1b
|
claudin domain containing 1b |
| chr20_-_32752457 | 0.03 |
ENSDART00000153287
|
si:dkey-6f10.3
|
si:dkey-6f10.3 |
| chr7_+_10701770 | 0.03 |
ENSDART00000167323
|
arnt2
|
aryl-hydrocarbon receptor nuclear translocator 2 |
| chr6_+_21395051 | 0.03 |
ENSDART00000017774
|
cacng5a
|
calcium channel, voltage-dependent, gamma subunit 5a |
| chr1_+_55535827 | 0.03 |
ENSDART00000152784
|
adgre16
|
adhesion G protein-coupled receptor E16 |
| chr24_-_20926812 | 0.03 |
ENSDART00000130958
|
ccdc58
|
coiled-coil domain containing 58 |
| chr15_-_19051152 | 0.03 |
ENSDART00000186453
|
arhgap32a
|
Rho GTPase activating protein 32a |
| chr6_+_14980761 | 0.03 |
ENSDART00000087782
|
mrps9
|
mitochondrial ribosomal protein S9 |
| chr15_-_20190052 | 0.03 |
ENSDART00000157149
|
exoc3l2b
|
exocyst complex component 3-like 2b |
| chr23_-_18381361 | 0.03 |
ENSDART00000016891
|
hsd17b10
|
hydroxysteroid (17-beta) dehydrogenase 10 |
| chr23_-_3721444 | 0.03 |
ENSDART00000141682
|
nudt3a
|
nudix (nucleoside diphosphate linked moiety X)-type motif 3a |
| chr2_-_22659450 | 0.03 |
ENSDART00000115025
|
thap4
|
THAP domain containing 4 |
| chr10_-_42751641 | 0.02 |
ENSDART00000182734
ENSDART00000113926 |
zgc:100918
|
zgc:100918 |
| chr17_-_24439672 | 0.02 |
ENSDART00000155020
|
wdpcp
|
WD repeat containing planar cell polarity effector |
| chr6_-_35052388 | 0.02 |
ENSDART00000181000
ENSDART00000170116 |
uap1
|
UDP-N-acetylglucosamine pyrophosphorylase 1 |
| chr4_+_60296027 | 0.02 |
ENSDART00000172479
|
si:dkey-248e17.5
|
si:dkey-248e17.5 |
| chr8_-_22157301 | 0.02 |
ENSDART00000158383
|
nphp4
|
nephronophthisis 4 |
| chr1_-_45039726 | 0.02 |
ENSDART00000186188
|
smu1b
|
SMU1, DNA replication regulator and spliceosomal factor b |
| chr5_-_55933420 | 0.02 |
ENSDART00000050966
|
slc25a46
|
solute carrier family 25, member 46 |
| chr5_+_69733096 | 0.02 |
ENSDART00000169013
|
arl6ip4
|
ADP-ribosylation factor-like 6 interacting protein 4 |
| chr18_-_4957022 | 0.02 |
ENSDART00000101686
|
zbbx
|
zinc finger, B-box domain containing |
| chr8_-_36327328 | 0.02 |
ENSDART00000183333
|
zgc:103700
|
zgc:103700 |
| chr16_-_47427016 | 0.01 |
ENSDART00000074575
|
sept7b
|
septin 7b |
| chr3_-_12026741 | 0.01 |
ENSDART00000132238
|
cfap70
|
cilia and flagella associated protein 70 |
| chr1_+_55563532 | 0.01 |
ENSDART00000152549
|
adgre15
|
adhesion G protein-coupled receptor E15 |
| chr8_+_23784471 | 0.01 |
ENSDART00000189457
|
si:ch211-163l21.8
|
si:ch211-163l21.8 |
| chr15_+_36310442 | 0.01 |
ENSDART00000154678
|
gmnc
|
geminin coiled-coil domain containing |
| chr20_+_25626479 | 0.01 |
ENSDART00000143883
|
ppat
|
phosphoribosyl pyrophosphate amidotransferase |
| chr4_-_33071267 | 0.01 |
ENSDART00000186314
|
si:dkeyp-4f2.1
|
si:dkeyp-4f2.1 |
| chr7_+_36898850 | 0.01 |
ENSDART00000113342
|
tox3
|
TOX high mobility group box family member 3 |
| chr3_+_13862753 | 0.01 |
ENSDART00000168315
|
ilf3b
|
interleukin enhancer binding factor 3b |
| chr25_-_16076257 | 0.00 |
ENSDART00000140780
|
ovch2
|
ovochymase 2 |
| chr17_-_14523722 | 0.00 |
ENSDART00000024726
|
daam1a
|
dishevelled associated activator of morphogenesis 1a |
| chr1_+_55662491 | 0.00 |
ENSDART00000152386
|
adgre8
|
adhesion G protein-coupled receptor E8 |
| chr15_+_16886196 | 0.00 |
ENSDART00000139296
ENSDART00000049196 |
gdpd1
|
glycerophosphodiester phosphodiesterase domain containing 1 |
| chr23_+_17512037 | 0.00 |
ENSDART00000054738
|
gid8b
|
GID complex subunit 8 homolog b (S. cerevisiae) |
| chr4_+_47636303 | 0.00 |
ENSDART00000167272
ENSDART00000166961 |
BX324142.1
|
|
| chr4_+_76442674 | 0.00 |
ENSDART00000164825
|
si:dkey-16p6.1
|
si:dkey-16p6.1 |
Network of associatons between targets according to the STRING database.
First level regulatory network of bbx
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | GO:0034454 | microtubule anchoring at centrosome(GO:0034454) |
| 0.1 | 0.3 | GO:0032782 | canalicular bile acid transport(GO:0015722) bile acid secretion(GO:0032782) |
| 0.0 | 0.1 | GO:0042823 | pyridoxal phosphate metabolic process(GO:0042822) pyridoxal phosphate biosynthetic process(GO:0042823) |
| 0.0 | 0.1 | GO:0006114 | glycerol biosynthetic process(GO:0006114) alditol biosynthetic process(GO:0019401) |
| 0.0 | 0.2 | GO:0045943 | positive regulation of transcription from RNA polymerase I promoter(GO:0045943) |
| 0.0 | 0.1 | GO:0036149 | phosphatidylinositol acyl-chain remodeling(GO:0036149) |
| 0.0 | 0.2 | GO:0002143 | tRNA wobble position uridine thiolation(GO:0002143) protein urmylation(GO:0032447) |
| 0.0 | 0.1 | GO:0010719 | photoreceptor cell morphogenesis(GO:0008594) negative regulation of epithelial to mesenchymal transition(GO:0010719) |
| 0.0 | 0.1 | GO:0070073 | clustering of voltage-gated calcium channels(GO:0070073) |
| 0.0 | 0.1 | GO:0044211 | CTP salvage(GO:0044211) |
| 0.0 | 0.2 | GO:0070979 | protein K11-linked ubiquitination(GO:0070979) |
| 0.0 | 0.1 | GO:1901031 | response to leucine(GO:0043201) cellular response to leucine(GO:0071233) regulation of response to reactive oxygen species(GO:1901031) |
| 0.0 | 0.2 | GO:0045176 | asymmetric protein localization(GO:0008105) apical protein localization(GO:0045176) |
| 0.0 | 0.1 | GO:0070309 | lens fiber cell development(GO:0070307) lens fiber cell morphogenesis(GO:0070309) |
| 0.0 | 0.1 | GO:0097010 | eukaryotic translation initiation factor 4F complex assembly(GO:0097010) |
| 0.0 | 0.2 | GO:0031297 | replication fork processing(GO:0031297) |
| 0.0 | 0.1 | GO:0032793 | positive regulation of CREB transcription factor activity(GO:0032793) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | GO:0033186 | CAF-1 complex(GO:0033186) |
| 0.0 | 0.2 | GO:0005662 | DNA replication factor A complex(GO:0005662) |
| 0.0 | 0.3 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.0 | 0.1 | GO:0044233 | ER-mitochondrion membrane contact site(GO:0044233) |
| 0.0 | 0.3 | GO:0012510 | trans-Golgi network transport vesicle membrane(GO:0012510) clathrin coat of trans-Golgi network vesicle(GO:0030130) |
| 0.0 | 0.1 | GO:0061700 | GATOR2 complex(GO:0061700) |
| 0.0 | 0.1 | GO:0005614 | interstitial matrix(GO:0005614) |
| 0.0 | 0.5 | GO:0030864 | cortical actin cytoskeleton(GO:0030864) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0015432 | canalicular bile acid transmembrane transporter activity(GO:0015126) bile acid-exporting ATPase activity(GO:0015432) |
| 0.0 | 0.3 | GO:0036312 | phosphatidylinositol 3-kinase regulatory subunit binding(GO:0036312) |
| 0.0 | 0.1 | GO:1990247 | N6-methyladenosine-containing RNA binding(GO:1990247) |
| 0.0 | 0.1 | GO:0004849 | uridine kinase activity(GO:0004849) |
| 0.0 | 0.1 | GO:0070728 | leucine binding(GO:0070728) |
| 0.0 | 0.0 | GO:0030882 | lipid antigen binding(GO:0030882) |
| 0.0 | 0.1 | GO:0035197 | siRNA binding(GO:0035197) |
| 0.0 | 0.1 | GO:0004726 | non-membrane spanning protein tyrosine phosphatase activity(GO:0004726) |
| 0.0 | 0.1 | GO:0033592 | RNA strand annealing activity(GO:0033592) RNA strand-exchange activity(GO:0034057) |
| 0.0 | 0.1 | GO:0008140 | cAMP response element binding protein binding(GO:0008140) |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.1 | REACTOME ACYL CHAIN REMODELLING OF PI | Genes involved in Acyl chain remodelling of PI |