Project
PRJEB1986: zebrafish developmental stages transcriptome
Navigation
Downloads
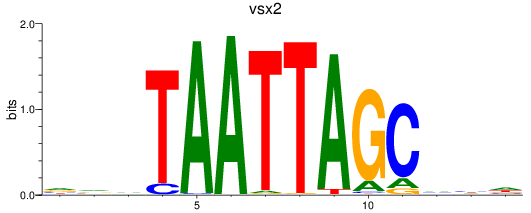
Results for vsx2
Z-value: 0.29
Transcription factors associated with vsx2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
vsx2
|
ENSDARG00000005574 | visual system homeobox 2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| vsx2 | dr11_v1_chr17_-_31659670_31659670 | -0.30 | 2.1e-01 | Click! |
Activity profile of vsx2 motif
Sorted Z-values of vsx2 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr21_-_13085242 | 0.66 |
ENSDART00000044504
|
zgc:109965
|
zgc:109965 |
| chr16_+_31804590 | 0.56 |
ENSDART00000167321
|
wnt4b
|
wingless-type MMTV integration site family, member 4b |
| chr3_-_43356082 | 0.49 |
ENSDART00000171213
|
uncx
|
UNC homeobox |
| chr20_-_9436521 | 0.46 |
ENSDART00000133000
|
zgc:101840
|
zgc:101840 |
| chr16_+_46111849 | 0.34 |
ENSDART00000172232
|
sv2a
|
synaptic vesicle glycoprotein 2A |
| chr6_+_28208973 | 0.33 |
ENSDART00000171216
ENSDART00000171377 ENSDART00000167389 ENSDART00000166988 |
LSM2 (1 of many)
|
si:ch73-14h10.2 |
| chr4_+_21129752 | 0.31 |
ENSDART00000169764
|
syt1a
|
synaptotagmin Ia |
| chr21_+_6751760 | 0.30 |
ENSDART00000135914
|
olfm1b
|
olfactomedin 1b |
| chr22_-_20011476 | 0.30 |
ENSDART00000093312
ENSDART00000093310 |
celf5a
|
cugbp, Elav-like family member 5a |
| chr24_+_35881124 | 0.29 |
ENSDART00000143015
|
klhl14
|
kelch-like family member 14 |
| chr21_+_6751405 | 0.29 |
ENSDART00000037265
ENSDART00000146371 |
olfm1b
|
olfactomedin 1b |
| chr2_-_35566938 | 0.28 |
ENSDART00000029006
ENSDART00000077178 ENSDART00000125298 |
tnn
|
tenascin N |
| chr23_+_28731379 | 0.27 |
ENSDART00000047378
|
cort
|
cortistatin |
| chr5_-_16996482 | 0.26 |
ENSDART00000144501
|
galnt9
|
polypeptide N-acetylgalactosaminyltransferase 9 |
| chr2_-_16216568 | 0.24 |
ENSDART00000173758
|
arhgef4
|
Rho guanine nucleotide exchange factor (GEF) 4 |
| chr12_+_20352400 | 0.22 |
ENSDART00000066383
|
hbae5
|
hemoglobin, alpha embryonic 5 |
| chr20_+_37294112 | 0.22 |
ENSDART00000076293
ENSDART00000140450 |
cx23
|
connexin 23 |
| chr15_-_9272328 | 0.22 |
ENSDART00000172114
|
calm2a
|
calmodulin 2a (phosphorylase kinase, delta) |
| chr13_+_19322686 | 0.20 |
ENSDART00000058036
|
emx2
|
empty spiracles homeobox 2 |
| chr21_+_28958471 | 0.19 |
ENSDART00000144331
ENSDART00000005929 |
ppp3ca
|
protein phosphatase 3, catalytic subunit, alpha isozyme |
| chr7_+_36898850 | 0.19 |
ENSDART00000113342
|
tox3
|
TOX high mobility group box family member 3 |
| chr11_-_3865472 | 0.18 |
ENSDART00000161426
|
gata2a
|
GATA binding protein 2a |
| chr2_-_15324837 | 0.18 |
ENSDART00000015655
|
tecrl2b
|
trans-2,3-enoyl-CoA reductase-like 2b |
| chr1_+_9290103 | 0.18 |
ENSDART00000055009
|
uncx4.1
|
Unc4.1 homeobox (C. elegans) |
| chr9_+_50001746 | 0.18 |
ENSDART00000058892
|
slc38a11
|
solute carrier family 38, member 11 |
| chr24_-_21923930 | 0.17 |
ENSDART00000131944
|
tagln3b
|
transgelin 3b |
| chr21_-_37790727 | 0.17 |
ENSDART00000162907
|
gabrb4
|
gamma-aminobutyric acid (GABA) A receptor, beta 4 |
| chr20_+_35382482 | 0.15 |
ENSDART00000135284
|
vsnl1a
|
visinin-like 1a |
| chr14_+_23717165 | 0.15 |
ENSDART00000006373
|
ndfip1
|
Nedd4 family interacting protein 1 |
| chr17_+_31185276 | 0.14 |
ENSDART00000062887
|
disp2
|
dispatched homolog 2 (Drosophila) |
| chr21_+_25231160 | 0.14 |
ENSDART00000063089
ENSDART00000139127 |
gng8
|
guanine nucleotide binding protein (G protein), gamma 8 |
| chr12_-_19862912 | 0.14 |
ENSDART00000145788
|
shisa9a
|
shisa family member 9a |
| chr3_-_46817499 | 0.14 |
ENSDART00000013717
|
elavl3
|
ELAV like neuron-specific RNA binding protein 3 |
| chr13_-_42560662 | 0.13 |
ENSDART00000124898
|
CR792417.1
|
|
| chr5_-_64831207 | 0.12 |
ENSDART00000144816
|
lix1
|
limb and CNS expressed 1 |
| chr18_-_1185772 | 0.12 |
ENSDART00000143245
|
nptnb
|
neuroplastin b |
| chr14_-_7207961 | 0.11 |
ENSDART00000167994
ENSDART00000166532 |
stox2b
|
storkhead box 2b |
| chr15_-_16098531 | 0.11 |
ENSDART00000080377
|
aldoca
|
aldolase C, fructose-bisphosphate, a |
| chr21_+_33172526 | 0.11 |
ENSDART00000183532
|
arl3l1
|
ADP-ribosylation factor-like 3, like 1 |
| chr11_-_29768054 | 0.11 |
ENSDART00000079117
|
plekha3
|
pleckstrin homology domain containing, family A (phosphoinositide binding specific) member 3 |
| chr20_-_37813863 | 0.10 |
ENSDART00000147529
|
batf3
|
basic leucine zipper transcription factor, ATF-like 3 |
| chr1_+_55755304 | 0.10 |
ENSDART00000144983
|
tecrb
|
trans-2,3-enoyl-CoA reductase b |
| chr24_-_25004553 | 0.10 |
ENSDART00000080997
ENSDART00000136860 |
zdhhc20b
|
zinc finger, DHHC-type containing 20b |
| chr20_+_29209615 | 0.10 |
ENSDART00000062350
|
katnbl1
|
katanin p80 subunit B-like 1 |
| chr14_+_25817628 | 0.10 |
ENSDART00000047680
|
glra1
|
glycine receptor, alpha 1 |
| chr20_+_29209767 | 0.09 |
ENSDART00000141252
|
katnbl1
|
katanin p80 subunit B-like 1 |
| chr18_+_14529005 | 0.09 |
ENSDART00000186379
|
kcng4a
|
potassium voltage-gated channel, subfamily G, member 4a |
| chr6_-_32726848 | 0.09 |
ENSDART00000155294
|
zc3h3
|
zinc finger CCCH-type containing 3 |
| chr23_-_19230627 | 0.08 |
ENSDART00000007122
|
guca1b
|
guanylate cyclase activator 1B |
| chr9_+_11532025 | 0.08 |
ENSDART00000109037
|
cdk5r2b
|
cyclin-dependent kinase 5, regulatory subunit 2b (p39) |
| chr24_-_23320223 | 0.07 |
ENSDART00000135846
|
zfhx4
|
zinc finger homeobox 4 |
| chr17_+_3379673 | 0.07 |
ENSDART00000176354
|
sntg2
|
syntrophin, gamma 2 |
| chr11_+_18873619 | 0.07 |
ENSDART00000176141
|
magi1b
|
membrane associated guanylate kinase, WW and PDZ domain containing 1b |
| chr14_+_23874062 | 0.06 |
ENSDART00000172149
|
sh3rf2
|
SH3 domain containing ring finger 2 |
| chr21_+_32820175 | 0.06 |
ENSDART00000076903
|
adra2db
|
adrenergic, alpha-2D-, receptor b |
| chr20_+_29209926 | 0.06 |
ENSDART00000152949
ENSDART00000153016 |
katnbl1
|
katanin p80 subunit B-like 1 |
| chr25_+_11456696 | 0.06 |
ENSDART00000171408
|
AGBL1
|
si:ch73-141f14.1 |
| chr23_+_28582865 | 0.05 |
ENSDART00000020296
|
l1cama
|
L1 cell adhesion molecule, paralog a |
| chr24_+_1023839 | 0.05 |
ENSDART00000082526
|
zgc:111976
|
zgc:111976 |
| chr12_-_35095414 | 0.05 |
ENSDART00000153229
|
si:dkey-21e13.3
|
si:dkey-21e13.3 |
| chr19_+_22062202 | 0.05 |
ENSDART00000100181
|
sall3b
|
spalt-like transcription factor 3b |
| chr9_-_5263947 | 0.04 |
ENSDART00000088342
|
cytip
|
cytohesin 1 interacting protein |
| chr20_-_20270191 | 0.04 |
ENSDART00000009356
|
ppp2r5ea
|
protein phosphatase 2, regulatory subunit B', epsilon isoform a |
| chr19_-_5265155 | 0.03 |
ENSDART00000145003
|
prf1.3
|
perforin 1.3 |
| chr3_-_46811611 | 0.03 |
ENSDART00000134092
|
elavl3
|
ELAV like neuron-specific RNA binding protein 3 |
| chr24_+_39518774 | 0.03 |
ENSDART00000132939
|
dcun1d3
|
defective in cullin neddylation 1 domain containing 3 |
| chr3_+_17537352 | 0.03 |
ENSDART00000104549
|
hcrt
|
hypocretin (orexin) neuropeptide precursor |
| chr23_-_24195519 | 0.03 |
ENSDART00000112370
ENSDART00000180377 |
ano11
|
anoctamin 11 |
| chr5_-_27438812 | 0.03 |
ENSDART00000078755
|
drd7
|
dopamine receptor D7 |
| chr5_-_48268049 | 0.03 |
ENSDART00000187454
|
mef2cb
|
myocyte enhancer factor 2cb |
| chr11_+_18873113 | 0.02 |
ENSDART00000103969
ENSDART00000103968 |
magi1b
|
membrane associated guanylate kinase, WW and PDZ domain containing 1b |
| chr6_-_39198912 | 0.02 |
ENSDART00000077938
|
c1galt1b
|
core 1 synthase, glycoprotein-N-acetylgalactosamine 3-beta-galactosyltransferase, 1b |
| chr6_-_39199070 | 0.02 |
ENSDART00000131793
|
c1galt1b
|
core 1 synthase, glycoprotein-N-acetylgalactosamine 3-beta-galactosyltransferase, 1b |
| chr8_+_18588551 | 0.02 |
ENSDART00000177476
ENSDART00000063539 |
prrg1
|
proline rich Gla (G-carboxyglutamic acid) 1 |
| chr1_-_17715493 | 0.02 |
ENSDART00000133027
|
si:dkey-256e7.8
|
si:dkey-256e7.8 |
| chr14_+_21820034 | 0.02 |
ENSDART00000122739
|
ctbp1
|
C-terminal binding protein 1 |
| chr8_-_23776399 | 0.01 |
ENSDART00000114800
|
INAVA
|
si:ch211-163l21.4 |
| chr20_-_46362606 | 0.01 |
ENSDART00000153087
|
bmf2
|
BCL2 modifying factor 2 |
| chr22_+_9003090 | 0.01 |
ENSDART00000106414
|
rnh1
|
ribonuclease/angiogenin inhibitor 1 |
| chr6_-_8264751 | 0.01 |
ENSDART00000091628
|
ccdc151
|
coiled-coil domain containing 151 |
| chr5_-_28109416 | 0.01 |
ENSDART00000042002
|
si:ch211-48m9.1
|
si:ch211-48m9.1 |
| chr4_+_11464255 | 0.01 |
ENSDART00000008584
|
gdi2
|
GDP dissociation inhibitor 2 |
Network of associatons between targets according to the STRING database.
First level regulatory network of vsx2
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.2 | GO:0061614 | pri-miRNA transcription from RNA polymerase II promoter(GO:0061614) regulation of pri-miRNA transcription from RNA polymerase II promoter(GO:1902893) positive regulation of pri-miRNA transcription from RNA polymerase II promoter(GO:1902895) |
| 0.0 | 0.1 | GO:0071361 | cellular response to ethanol(GO:0071361) |
| 0.0 | 0.2 | GO:0060118 | vestibular receptor cell differentiation(GO:0060114) vestibular receptor cell development(GO:0060118) |
| 0.0 | 0.6 | GO:0045839 | negative regulation of mitotic nuclear division(GO:0045839) |
| 0.0 | 0.1 | GO:0097067 | cellular response to thyroid hormone stimulus(GO:0097067) |
| 0.0 | 0.1 | GO:0035627 | ceramide transport(GO:0035627) |
| 0.0 | 0.1 | GO:0048172 | regulation of short-term neuronal synaptic plasticity(GO:0048172) |
| 0.0 | 0.3 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
| 0.0 | 0.2 | GO:0033173 | calcineurin-NFAT signaling cascade(GO:0033173) |
| 0.0 | 0.1 | GO:0071881 | adenylate cyclase-inhibiting adrenergic receptor signaling pathway(GO:0071881) |
| 0.0 | 0.2 | GO:0015671 | oxygen transport(GO:0015671) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.3 | GO:0042584 | chromaffin granule membrane(GO:0042584) |
| 0.0 | 0.2 | GO:0005955 | calcineurin complex(GO:0005955) |
| 0.0 | 0.2 | GO:0031838 | haptoglobin-hemoglobin complex(GO:0031838) |
| 0.0 | 0.1 | GO:0016533 | cyclin-dependent protein kinase 5 holoenzyme complex(GO:0016533) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | GO:0004723 | calcium-dependent protein serine/threonine phosphatase activity(GO:0004723) calmodulin-dependent protein phosphatase activity(GO:0033192) |
| 0.0 | 0.2 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.0 | 0.1 | GO:1902387 | ceramide transporter activity(GO:0035620) ceramide 1-phosphate binding(GO:1902387) ceramide 1-phosphate transporter activity(GO:1902388) |
| 0.0 | 0.2 | GO:0031720 | haptoglobin binding(GO:0031720) |
| 0.0 | 0.1 | GO:0004332 | fructose-bisphosphate aldolase activity(GO:0004332) |
| 0.0 | 0.6 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.0 | 0.1 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 0.0 | 0.1 | GO:0016933 | extracellular-glycine-gated ion channel activity(GO:0016933) extracellular-glycine-gated chloride channel activity(GO:0016934) |
| 0.0 | 0.1 | GO:0042809 | vitamin D receptor binding(GO:0042809) |
| 0.0 | 0.1 | GO:0008048 | calcium sensitive guanylate cyclase activator activity(GO:0008048) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |