Project
PRJEB1986: zebrafish developmental stages transcriptome
Navigation
Downloads
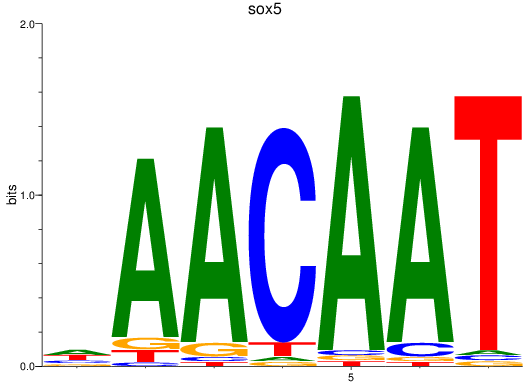
Results for sox5
Z-value: 1.02
Transcription factors associated with sox5
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
sox5
|
ENSDARG00000011582 | SRY-box transcription factor 5 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| sox5 | dr11_v1_chr4_-_17055950_17055950 | -0.44 | 5.9e-02 | Click! |
Activity profile of sox5 motif
Sorted Z-values of sox5 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr9_-_22339582 | 3.61 |
ENSDART00000134805
|
crygm2d1
|
crystallin, gamma M2d1 |
| chr9_-_22135420 | 1.90 |
ENSDART00000184959
|
crygm2d8
|
crystallin, gamma M2d8 |
| chr20_-_47704973 | 1.82 |
ENSDART00000174808
|
tfap2b
|
transcription factor AP-2 beta |
| chr9_-_22272181 | 1.64 |
ENSDART00000113174
|
crygm2d7
|
crystallin, gamma M2d7 |
| chr13_+_25449681 | 1.46 |
ENSDART00000101328
|
atoh7
|
atonal bHLH transcription factor 7 |
| chr22_-_10487490 | 1.41 |
ENSDART00000064798
|
aspn
|
asporin (LRR class 1) |
| chr1_-_30689004 | 1.40 |
ENSDART00000018827
|
dachc
|
dachshund c |
| chr3_-_30158395 | 1.34 |
ENSDART00000103502
|
si:ch211-152f23.5
|
si:ch211-152f23.5 |
| chr24_+_24461558 | 1.29 |
ENSDART00000182424
|
bhlhe22
|
basic helix-loop-helix family, member e22 |
| chr24_+_24461341 | 1.27 |
ENSDART00000147658
|
bhlhe22
|
basic helix-loop-helix family, member e22 |
| chr7_+_39444843 | 1.24 |
ENSDART00000143999
ENSDART00000173554 ENSDART00000173698 ENSDART00000173754 ENSDART00000144075 ENSDART00000138192 ENSDART00000145457 ENSDART00000141750 ENSDART00000103056 ENSDART00000142946 ENSDART00000173748 |
tnnt3b
|
troponin T type 3b (skeletal, fast) |
| chr19_-_43542568 | 1.20 |
ENSDART00000051720
|
BX649384.1
|
|
| chr6_+_43426599 | 1.15 |
ENSDART00000056457
|
mitfa
|
microphthalmia-associated transcription factor a |
| chr8_+_23093155 | 1.14 |
ENSDART00000063075
|
zgc:100920
|
zgc:100920 |
| chr20_+_54383838 | 1.14 |
ENSDART00000157737
|
lrfn5b
|
leucine rich repeat and fibronectin type III domain containing 5b |
| chr18_+_16246806 | 1.13 |
ENSDART00000142584
|
alx1
|
ALX homeobox 1 |
| chr4_+_16323970 | 1.06 |
ENSDART00000190651
|
BX322608.1
|
|
| chr3_-_35602233 | 1.05 |
ENSDART00000055269
|
gng13b
|
guanine nucleotide binding protein (G protein), gamma 13b |
| chr2_-_33455164 | 1.03 |
ENSDART00000134024
ENSDART00000132221 |
ccdc24
|
coiled-coil domain containing 24 |
| chr8_+_31941650 | 1.00 |
ENSDART00000138217
|
htr1aa
|
5-hydroxytryptamine (serotonin) receptor 1A a |
| chr18_+_19972853 | 0.96 |
ENSDART00000180071
|
skor1b
|
SKI family transcriptional corepressor 1b |
| chr23_-_32157865 | 0.96 |
ENSDART00000000876
|
nr4a1
|
nuclear receptor subfamily 4, group A, member 1 |
| chr16_+_10318893 | 0.94 |
ENSDART00000055380
|
tubb5
|
tubulin, beta 5 |
| chr4_-_16412084 | 0.93 |
ENSDART00000188460
|
dcn
|
decorin |
| chr23_+_8797143 | 0.92 |
ENSDART00000132992
|
sox18
|
SRY (sex determining region Y)-box 18 |
| chr16_-_33095161 | 0.90 |
ENSDART00000187648
|
dopey1
|
dopey family member 1 |
| chr2_+_50626476 | 0.89 |
ENSDART00000018150
|
neurod6b
|
neuronal differentiation 6b |
| chr11_-_6048490 | 0.89 |
ENSDART00000066164
|
plvapb
|
plasmalemma vesicle associated protein b |
| chr20_-_14054083 | 0.88 |
ENSDART00000009549
|
rhag
|
Rh associated glycoprotein |
| chr20_+_39283849 | 0.87 |
ENSDART00000002481
ENSDART00000146683 |
scara3
|
scavenger receptor class A, member 3 |
| chr9_+_32978302 | 0.85 |
ENSDART00000007630
|
nhlh2
|
nescient helix loop helix 2 |
| chr1_-_38709551 | 0.85 |
ENSDART00000128794
|
gpm6ab
|
glycoprotein M6Ab |
| chr12_-_35054354 | 0.84 |
ENSDART00000075351
|
zgc:112285
|
zgc:112285 |
| chr9_+_24008879 | 0.82 |
ENSDART00000190419
ENSDART00000191843 ENSDART00000148226 |
mlphb
|
melanophilin b |
| chr18_+_27300700 | 0.82 |
ENSDART00000140444
|
plekha7a
|
pleckstrin homology domain containing, family A member 7a |
| chr8_-_34051548 | 0.80 |
ENSDART00000105204
|
pbx3b
|
pre-B-cell leukemia homeobox 3b |
| chr18_-_10995410 | 0.79 |
ENSDART00000136751
|
tspan33b
|
tetraspanin 33b |
| chr22_-_10541372 | 0.79 |
ENSDART00000179708
|
si:dkey-42i9.4
|
si:dkey-42i9.4 |
| chr8_+_18830759 | 0.78 |
ENSDART00000089079
|
mpnd
|
MPN domain containing |
| chr11_+_25257022 | 0.75 |
ENSDART00000156052
|
tp53inp2
|
tumor protein p53 inducible nuclear protein 2 |
| chr23_+_32335871 | 0.74 |
ENSDART00000149698
|
slc39a5
|
solute carrier family 39 (zinc transporter), member 5 |
| chr2_+_32846602 | 0.73 |
ENSDART00000056649
|
tmem53
|
transmembrane protein 53 |
| chr11_+_39672874 | 0.73 |
ENSDART00000046663
ENSDART00000157659 |
camta1b
|
calmodulin binding transcription activator 1b |
| chr22_+_661711 | 0.72 |
ENSDART00000113795
|
elf3
|
E74-like factor 3 (ets domain transcription factor, epithelial-specific ) |
| chr25_+_14507567 | 0.71 |
ENSDART00000015681
|
dbx1b
|
developing brain homeobox 1b |
| chr7_-_35515931 | 0.70 |
ENSDART00000193324
|
irx6a
|
iroquois homeobox 6a |
| chr13_+_38520927 | 0.70 |
ENSDART00000137299
ENSDART00000111080 |
adgrb3
|
adhesion G protein-coupled receptor B3 |
| chr17_-_30666037 | 0.70 |
ENSDART00000156509
|
alkal2b
|
ALK and LTK ligand 2b |
| chr3_-_28828242 | 0.68 |
ENSDART00000151445
|
si:ch211-76l23.4
|
si:ch211-76l23.4 |
| chr14_-_26704829 | 0.67 |
ENSDART00000078563
|
neurog1
|
neurogenin 1 |
| chr13_-_40411908 | 0.67 |
ENSDART00000057094
ENSDART00000150091 |
nkx2.3
|
NK2 homeobox 3 |
| chr19_-_103289 | 0.66 |
ENSDART00000143118
|
adgrb1b
|
adhesion G protein-coupled receptor B1b |
| chr18_-_20466061 | 0.66 |
ENSDART00000060311
|
paqr5a
|
progestin and adipoQ receptor family member Va |
| chr14_-_46442459 | 0.66 |
ENSDART00000158770
|
fgfbp1b
|
fibroblast growth factor binding protein 1b |
| chr14_+_38786298 | 0.66 |
ENSDART00000164440
|
si:ch211-195b11.3
|
si:ch211-195b11.3 |
| chr8_+_21195420 | 0.66 |
ENSDART00000100234
ENSDART00000091307 |
col2a1a
|
collagen, type II, alpha 1a |
| chr12_+_6195191 | 0.64 |
ENSDART00000043236
ENSDART00000186420 |
prkg1b
|
protein kinase, cGMP-dependent, type Ib |
| chr19_+_19777437 | 0.63 |
ENSDART00000170662
|
hoxa3a
|
homeobox A3a |
| chr15_+_28685892 | 0.63 |
ENSDART00000155815
ENSDART00000060244 |
nova2
|
neuro-oncological ventral antigen 2 |
| chr16_+_28754403 | 0.62 |
ENSDART00000103340
|
s100v1
|
S100 calcium binding protein V1 |
| chr3_+_23742868 | 0.62 |
ENSDART00000153512
|
hoxb3a
|
homeobox B3a |
| chr13_+_29773153 | 0.61 |
ENSDART00000144443
ENSDART00000133796 ENSDART00000141310 ENSDART00000139782 |
pax2a
|
paired box 2a |
| chr5_-_67911111 | 0.61 |
ENSDART00000051833
|
gsx1
|
GS homeobox 1 |
| chr3_+_29714775 | 0.60 |
ENSDART00000041388
|
cacng2a
|
calcium channel, voltage-dependent, gamma subunit 2a |
| chr16_-_28856112 | 0.60 |
ENSDART00000078543
|
syt11b
|
synaptotagmin XIb |
| chr19_+_37848830 | 0.59 |
ENSDART00000042276
ENSDART00000180872 |
nxph1
|
neurexophilin 1 |
| chr19_+_15571290 | 0.59 |
ENSDART00000131134
|
foxo6b
|
forkhead box O6 b |
| chr15_-_41689981 | 0.59 |
ENSDART00000059327
|
spsb4b
|
splA/ryanodine receptor domain and SOCS box containing 4b |
| chr2_+_56463167 | 0.59 |
ENSDART00000123392
|
rab11bb
|
RAB11B, member RAS oncogene family, b |
| chr3_+_23697997 | 0.59 |
ENSDART00000184299
ENSDART00000078466 |
hoxb3a
|
homeobox B3a |
| chr13_+_12045758 | 0.58 |
ENSDART00000079398
ENSDART00000165467 ENSDART00000165880 |
gng2
|
guanine nucleotide binding protein (G protein), gamma 2 |
| chr13_+_12045475 | 0.56 |
ENSDART00000163053
ENSDART00000160812 ENSDART00000158244 |
gng2
|
guanine nucleotide binding protein (G protein), gamma 2 |
| chr25_+_26441054 | 0.56 |
ENSDART00000158607
|
dapk2b
|
death-associated protein kinase 2b |
| chr8_-_31938364 | 0.56 |
ENSDART00000137647
|
RNF180
|
si:dkey-250l23.4 |
| chr3_+_52999962 | 0.56 |
ENSDART00000104683
|
pbx4
|
pre-B-cell leukemia transcription factor 4 |
| chr10_+_23022263 | 0.55 |
ENSDART00000138955
|
si:dkey-175g6.2
|
si:dkey-175g6.2 |
| chr7_-_18601206 | 0.54 |
ENSDART00000111636
|
DTX4
|
si:ch211-119e14.2 |
| chr19_-_38872650 | 0.54 |
ENSDART00000146641
|
adgrb2
|
adhesion G protein-coupled receptor B2 |
| chr18_-_14836600 | 0.53 |
ENSDART00000045232
|
mtss1la
|
metastasis suppressor 1-like a |
| chr12_+_45677293 | 0.53 |
ENSDART00000152850
ENSDART00000153047 |
si:ch73-111m19.2
|
si:ch73-111m19.2 |
| chr23_-_3759692 | 0.53 |
ENSDART00000028885
|
hmga1a
|
high mobility group AT-hook 1a |
| chr19_-_868187 | 0.53 |
ENSDART00000186626
|
eomesa
|
eomesodermin homolog a |
| chr25_-_12923482 | 0.52 |
ENSDART00000161754
|
CR450808.1
|
|
| chr15_+_29728377 | 0.52 |
ENSDART00000099958
|
zgc:153372
|
zgc:153372 |
| chr4_-_6567355 | 0.52 |
ENSDART00000134820
ENSDART00000142087 |
foxp2
|
forkhead box P2 |
| chr5_+_56026031 | 0.51 |
ENSDART00000050970
|
fzd9a
|
frizzled class receptor 9a |
| chr23_+_36118738 | 0.51 |
ENSDART00000139319
|
hoxc5a
|
homeobox C5a |
| chr15_-_20233105 | 0.51 |
ENSDART00000123910
|
ppp1r14ab
|
protein phosphatase 1, regulatory (inhibitor) subunit 14Ab |
| chr18_+_30421528 | 0.50 |
ENSDART00000140908
|
gse1
|
Gse1 coiled-coil protein |
| chr12_-_33357655 | 0.50 |
ENSDART00000066233
ENSDART00000148165 |
slc16a3
|
solute carrier family 16 (monocarboxylate transporter), member 3 |
| chr2_-_21170517 | 0.50 |
ENSDART00000135417
|
bmi1b
|
bmi1 polycomb ring finger oncogene 1b |
| chr14_-_24761132 | 0.50 |
ENSDART00000146299
|
slit3
|
slit homolog 3 (Drosophila) |
| chr16_+_23811554 | 0.50 |
ENSDART00000114336
|
si:dkey-7f3.9
|
si:dkey-7f3.9 |
| chr11_-_3865472 | 0.49 |
ENSDART00000161426
|
gata2a
|
GATA binding protein 2a |
| chr1_+_45958904 | 0.49 |
ENSDART00000108528
|
arhgef7b
|
Rho guanine nucleotide exchange factor (GEF) 7b |
| chr13_+_29771463 | 0.48 |
ENSDART00000134424
ENSDART00000138332 ENSDART00000134330 ENSDART00000160944 ENSDART00000076992 ENSDART00000160921 |
pax2a
|
paired box 2a |
| chr23_+_30730121 | 0.48 |
ENSDART00000134141
|
asxl1
|
additional sex combs like transcriptional regulator 1 |
| chr6_+_42819337 | 0.47 |
ENSDART00000046498
|
sema3fa
|
sema domain, immunoglobulin domain (Ig), short basic domain, secreted, (semaphorin) 3Fa |
| chr14_+_21754521 | 0.46 |
ENSDART00000111839
|
kdm2ab
|
lysine (K)-specific demethylase 2Ab |
| chr19_-_2318391 | 0.46 |
ENSDART00000012791
|
sp8a
|
sp8 transcription factor a |
| chr6_+_18520859 | 0.46 |
ENSDART00000158263
|
si:dkey-10p5.10
|
si:dkey-10p5.10 |
| chr7_-_31938938 | 0.46 |
ENSDART00000132353
|
bdnf
|
brain-derived neurotrophic factor |
| chr1_+_14372680 | 0.46 |
ENSDART00000046874
|
BX530018.1
|
|
| chr10_-_28761454 | 0.46 |
ENSDART00000129400
|
alcama
|
activated leukocyte cell adhesion molecule a |
| chr19_+_20793388 | 0.46 |
ENSDART00000142463
|
txnl4a
|
thioredoxin-like 4A |
| chr3_+_52487695 | 0.45 |
ENSDART00000155451
|
si:ch211-241f5.3
|
si:ch211-241f5.3 |
| chr16_+_29492937 | 0.45 |
ENSDART00000011497
|
ctsk
|
cathepsin K |
| chr15_+_19797918 | 0.45 |
ENSDART00000113314
|
si:ch211-229d2.5
|
si:ch211-229d2.5 |
| chr10_+_15255198 | 0.45 |
ENSDART00000139047
ENSDART00000172107 ENSDART00000183413 ENSDART00000185314 |
vldlr
|
very low density lipoprotein receptor |
| chr16_-_7443388 | 0.44 |
ENSDART00000017445
|
prdm1a
|
PR domain containing 1a, with ZNF domain |
| chr19_+_19762183 | 0.44 |
ENSDART00000163611
ENSDART00000187604 |
hoxa3a
|
homeobox A3a |
| chr17_-_30975978 | 0.44 |
ENSDART00000051697
|
evla
|
Enah/Vasp-like a |
| chr19_+_43604643 | 0.44 |
ENSDART00000151168
|
si:ch211-199g17.9
|
si:ch211-199g17.9 |
| chr16_+_40560622 | 0.43 |
ENSDART00000038294
|
tp53inp1
|
tumor protein p53 inducible nuclear protein 1 |
| chr3_-_31804481 | 0.43 |
ENSDART00000028270
|
gfap
|
glial fibrillary acidic protein |
| chr8_+_13106760 | 0.43 |
ENSDART00000029308
|
itgb4
|
integrin, beta 4 |
| chr14_-_25949951 | 0.43 |
ENSDART00000141304
|
sparc
|
secreted protein, acidic, cysteine-rich (osteonectin) |
| chr21_-_18991165 | 0.43 |
ENSDART00000012574
|
si:ch211-222n4.6
|
si:ch211-222n4.6 |
| chr3_+_51684963 | 0.42 |
ENSDART00000091180
ENSDART00000183711 ENSDART00000159493 |
baiap2a
|
BAI1-associated protein 2a |
| chr19_+_43604256 | 0.42 |
ENSDART00000151080
ENSDART00000110305 |
si:ch211-199g17.9
|
si:ch211-199g17.9 |
| chr16_+_19732543 | 0.42 |
ENSDART00000149901
ENSDART00000052927 |
twist1b
|
twist family bHLH transcription factor 1b |
| chr13_+_28854438 | 0.42 |
ENSDART00000193407
ENSDART00000189554 |
CU639469.1
|
|
| chr15_-_41689684 | 0.42 |
ENSDART00000143447
|
spsb4b
|
splA/ryanodine receptor domain and SOCS box containing 4b |
| chr5_+_17624463 | 0.42 |
ENSDART00000183869
ENSDART00000081064 |
fbrsl1
|
fibrosin-like 1 |
| chr25_-_19139259 | 0.41 |
ENSDART00000067327
|
abhd2b
|
abhydrolase domain containing 2b |
| chr6_-_21873266 | 0.41 |
ENSDART00000151658
ENSDART00000151152 ENSDART00000151179 |
si:dkey-19e4.5
|
si:dkey-19e4.5 |
| chr6_+_32382743 | 0.41 |
ENSDART00000190009
|
dock7
|
dedicator of cytokinesis 7 |
| chr5_-_44843738 | 0.41 |
ENSDART00000003926
|
fbp1a
|
fructose-1,6-bisphosphatase 1a |
| chr17_-_30975707 | 0.41 |
ENSDART00000138346
|
evla
|
Enah/Vasp-like a |
| chr14_+_31529958 | 0.41 |
ENSDART00000053026
|
fam122b
|
family with sequence similarity 122B |
| chr7_+_20471315 | 0.40 |
ENSDART00000173714
|
si:dkey-19b23.13
|
si:dkey-19b23.13 |
| chr11_+_39135050 | 0.40 |
ENSDART00000180571
ENSDART00000189685 |
cdc42
|
cell division cycle 42 |
| chr20_+_34915945 | 0.40 |
ENSDART00000153064
|
snap25a
|
synaptosomal-associated protein, 25a |
| chr24_+_32411753 | 0.40 |
ENSDART00000058530
|
neurod6a
|
neuronal differentiation 6a |
| chr25_+_7321675 | 0.39 |
ENSDART00000104712
ENSDART00000142934 |
hmg20a
|
high mobility group 20A |
| chr9_+_11293830 | 0.39 |
ENSDART00000144440
|
wnt6b
|
wingless-type MMTV integration site family, member 6b |
| chr9_+_41040294 | 0.39 |
ENSDART00000000693
|
osgepl1
|
O-sialoglycoprotein endopeptidase-like 1 |
| chr17_+_43595692 | 0.39 |
ENSDART00000156271
|
cfap99
|
cilia and flagella associated protein 99 |
| chr9_+_42607138 | 0.39 |
ENSDART00000138133
ENSDART00000002027 |
gulp1a
|
GULP, engulfment adaptor PTB domain containing 1a |
| chr23_-_33654889 | 0.38 |
ENSDART00000146180
|
csrnp2
|
cysteine-serine-rich nuclear protein 2 |
| chr10_-_31104983 | 0.38 |
ENSDART00000186941
|
pknox2
|
pbx/knotted 1 homeobox 2 |
| chr18_-_14836862 | 0.38 |
ENSDART00000124843
|
mtss1la
|
metastasis suppressor 1-like a |
| chr21_+_7605803 | 0.38 |
ENSDART00000121813
|
wdr41
|
WD repeat domain 41 |
| chr14_-_36397768 | 0.37 |
ENSDART00000185199
ENSDART00000052562 |
spata4
|
spermatogenesis associated 4 |
| chr10_+_15255012 | 0.37 |
ENSDART00000023766
|
vldlr
|
very low density lipoprotein receptor |
| chr3_+_54047342 | 0.37 |
ENSDART00000178486
|
olfm2a
|
olfactomedin 2a |
| chr7_+_25003851 | 0.37 |
ENSDART00000144526
|
naa40
|
N(alpha)-acetyltransferase 40, NatD catalytic subunit, homolog (S. cerevisiae) |
| chr1_+_16466771 | 0.37 |
ENSDART00000162389
|
vps37a
|
vacuolar protein sorting 37A |
| chr6_-_28943056 | 0.36 |
ENSDART00000065138
|
tbc1d23
|
TBC1 domain family, member 23 |
| chr8_+_7311478 | 0.36 |
ENSDART00000133358
|
ssuh2rs1
|
ssu-2 homolog, related sequence 1 |
| chr3_-_26244256 | 0.36 |
ENSDART00000103741
|
ppp4ca
|
protein phosphatase 4, catalytic subunit a |
| chr6_-_32093830 | 0.36 |
ENSDART00000017695
|
foxd3
|
forkhead box D3 |
| chr4_-_19016396 | 0.36 |
ENSDART00000166160
|
si:dkey-31f5.11
|
si:dkey-31f5.11 |
| chr7_+_47287307 | 0.36 |
ENSDART00000114669
|
dpy19l3
|
dpy-19 like C-mannosyltransferase 3 |
| chr23_-_33640959 | 0.36 |
ENSDART00000187599
ENSDART00000189475 |
csrnp2
|
cysteine-serine-rich nuclear protein 2 |
| chr16_+_29492749 | 0.36 |
ENSDART00000179680
|
ctsk
|
cathepsin K |
| chr19_-_45960191 | 0.35 |
ENSDART00000052434
ENSDART00000172732 |
eif3hb
|
eukaryotic translation initiation factor 3, subunit H, b |
| chr19_+_19761966 | 0.35 |
ENSDART00000163697
|
hoxa3a
|
homeobox A3a |
| chr20_-_25369767 | 0.35 |
ENSDART00000180278
|
itsn2a
|
intersectin 2a |
| chr14_-_2588216 | 0.35 |
ENSDART00000188082
ENSDART00000167293 |
fbxw11a
|
F-box and WD repeat domain containing 11a |
| chr5_-_28606916 | 0.35 |
ENSDART00000026107
ENSDART00000137717 |
tnc
|
tenascin C |
| chr2_-_30246781 | 0.34 |
ENSDART00000175071
|
elocb
|
elongin C paralog b |
| chr17_+_29276544 | 0.34 |
ENSDART00000049399
|
ankrd9
|
ankyrin repeat domain 9 |
| chr16_-_21047872 | 0.34 |
ENSDART00000131582
|
cbx3b
|
chromobox homolog 3b |
| chr6_-_38418862 | 0.34 |
ENSDART00000104135
|
gabra5
|
gamma-aminobutyric acid (GABA) A receptor, alpha 5 |
| chr23_+_20523617 | 0.34 |
ENSDART00000176404
|
adnpb
|
activity-dependent neuroprotector homeobox b |
| chr23_-_3759345 | 0.33 |
ENSDART00000132205
ENSDART00000137707 ENSDART00000189382 |
hmga1a
|
high mobility group AT-hook 1a |
| chr5_+_41793001 | 0.33 |
ENSDART00000136439
ENSDART00000190477 ENSDART00000192289 |
bcl7a
|
BCL tumor suppressor 7A |
| chr25_+_28253844 | 0.33 |
ENSDART00000151891
|
cadps2
|
Ca++-dependent secretion activator 2 |
| chr22_-_14128716 | 0.33 |
ENSDART00000140323
|
si:ch211-246m6.4
|
si:ch211-246m6.4 |
| chr18_-_19484919 | 0.33 |
ENSDART00000177690
ENSDART00000184949 |
lctlb
|
lactase-like b |
| chr5_-_20893993 | 0.33 |
ENSDART00000137603
|
si:ch211-225b11.4
|
si:ch211-225b11.4 |
| chr19_+_19759577 | 0.32 |
ENSDART00000169480
|
hoxa5a
|
homeobox A5a |
| chr23_-_33944597 | 0.32 |
ENSDART00000133223
|
si:dkey-190g6.2
|
si:dkey-190g6.2 |
| chr24_-_3426620 | 0.32 |
ENSDART00000184346
|
nck1b
|
NCK adaptor protein 1b |
| chr16_+_19033554 | 0.32 |
ENSDART00000182430
|
rapgef5b
|
Rap guanine nucleotide exchange factor (GEF) 5b |
| chr14_-_41478265 | 0.32 |
ENSDART00000149886
ENSDART00000016002 |
tspan7
|
tetraspanin 7 |
| chr13_+_3938526 | 0.32 |
ENSDART00000012759
|
yipf3
|
Yip1 domain family, member 3 |
| chr21_-_27201495 | 0.32 |
ENSDART00000145013
|
zgc:77262
|
zgc:77262 |
| chr2_-_12079718 | 0.31 |
ENSDART00000155821
|
kcnt2
|
potassium channel, subfamily T, member 2 |
| chr7_+_21180747 | 0.31 |
ENSDART00000185543
|
serpinh2
|
serine (or cysteine) peptidase inhibitor, clade H, member 2 |
| chr8_-_39978767 | 0.31 |
ENSDART00000083066
|
asphd2
|
aspartate beta-hydroxylase domain containing 2 |
| chr18_+_48428713 | 0.30 |
ENSDART00000076861
|
fli1a
|
Fli-1 proto-oncogene, ETS transcription factor a |
| chr6_-_46742455 | 0.30 |
ENSDART00000011970
|
zgc:66479
|
zgc:66479 |
| chr18_+_6039141 | 0.30 |
ENSDART00000138972
|
si:ch73-386h18.1
|
si:ch73-386h18.1 |
| chr13_+_11440389 | 0.30 |
ENSDART00000186463
|
zbtb18
|
zinc finger and BTB domain containing 18 |
| chr23_-_30787932 | 0.30 |
ENSDART00000135771
|
myt1a
|
myelin transcription factor 1a |
| chr11_-_16152400 | 0.29 |
ENSDART00000123665
|
arpc4l
|
actin related protein 2/3 complex, subunit 4, like |
| chr6_+_9872261 | 0.29 |
ENSDART00000045070
|
MPP4 (1 of many)
|
si:ch211-222n4.6 |
| chr15_-_19705707 | 0.29 |
ENSDART00000047643
|
sytl2b
|
synaptotagmin-like 2b |
| chr11_+_16153207 | 0.29 |
ENSDART00000192356
|
tada3l
|
transcriptional adaptor 3 (NGG1 homolog, yeast)-like |
| chr8_+_20488322 | 0.29 |
ENSDART00000036630
|
zgc:101100
|
zgc:101100 |
| chr17_+_30545895 | 0.29 |
ENSDART00000076739
|
nhsl1a
|
NHS-like 1a |
| chr11_+_16152316 | 0.29 |
ENSDART00000081054
|
tada3l
|
transcriptional adaptor 3 (NGG1 homolog, yeast)-like |
| chr25_-_29134000 | 0.28 |
ENSDART00000172027
ENSDART00000190447 |
parp6b
|
poly (ADP-ribose) polymerase family, member 6b |
| chr7_-_38861741 | 0.28 |
ENSDART00000173629
ENSDART00000037361 ENSDART00000173953 |
phf21aa
|
PHD finger protein 21Aa |
| chr15_-_5799170 | 0.28 |
ENSDART00000142334
ENSDART00000171528 |
hnrnpl2
|
heterogeneous nuclear ribonucleoprotein L2 |
| chr17_+_23968214 | 0.27 |
ENSDART00000183053
|
xpo1b
|
exportin 1 (CRM1 homolog, yeast) b |
| chr21_+_28502340 | 0.27 |
ENSDART00000077897
ENSDART00000140229 |
otub1a
|
OTU deubiquitinase, ubiquitin aldehyde binding 1a |
| chr11_+_37612214 | 0.27 |
ENSDART00000172899
ENSDART00000077496 |
hp1bp3
|
heterochromatin protein 1, binding protein 3 |
Network of associatons between targets according to the STRING database.
First level regulatory network of sox5
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.8 | GO:0045218 | zonula adherens maintenance(GO:0045218) |
| 0.2 | 1.1 | GO:0010718 | positive regulation of epithelial to mesenchymal transition(GO:0010718) |
| 0.2 | 0.9 | GO:0015840 | urea transport(GO:0015840) |
| 0.2 | 1.4 | GO:0060017 | parathyroid gland development(GO:0060017) |
| 0.2 | 0.5 | GO:0060571 | invagination involved in gastrulation with mouth forming second(GO:0055109) morphogenesis of an epithelial fold(GO:0060571) |
| 0.2 | 0.7 | GO:0090386 | phagosome maturation involved in apoptotic cell clearance(GO:0090386) phagolysosome assembly involved in apoptotic cell clearance(GO:0090387) |
| 0.2 | 0.5 | GO:1902895 | pri-miRNA transcription from RNA polymerase II promoter(GO:0061614) regulation of pri-miRNA transcription from RNA polymerase II promoter(GO:1902893) positive regulation of pri-miRNA transcription from RNA polymerase II promoter(GO:1902895) |
| 0.2 | 0.5 | GO:0071896 | protein localization to adherens junction(GO:0071896) |
| 0.2 | 0.5 | GO:0018872 | arsonoacetate metabolic process(GO:0018872) |
| 0.1 | 0.7 | GO:0016322 | neuron remodeling(GO:0016322) |
| 0.1 | 1.1 | GO:0021588 | cerebellum formation(GO:0021588) |
| 0.1 | 0.7 | GO:0021516 | dorsal spinal cord development(GO:0021516) anterior lateral line nerve development(GO:0048909) |
| 0.1 | 0.4 | GO:1990403 | embryonic brain development(GO:1990403) |
| 0.1 | 0.4 | GO:0018317 | protein C-linked glycosylation(GO:0018103) peptidyl-tryptophan modification(GO:0018211) protein C-linked glycosylation via tryptophan(GO:0018317) protein C-linked glycosylation via 2'-alpha-mannosyl-L-tryptophan(GO:0018406) |
| 0.1 | 0.2 | GO:0032776 | DNA methylation on cytosine(GO:0032776) |
| 0.1 | 0.7 | GO:0070376 | ERK5 cascade(GO:0070375) regulation of ERK5 cascade(GO:0070376) positive regulation of ERK5 cascade(GO:0070378) |
| 0.1 | 0.3 | GO:0045887 | regulation of synaptic growth at neuromuscular junction(GO:0008582) axon regeneration at neuromuscular junction(GO:0014814) positive regulation of synaptic growth at neuromuscular junction(GO:0045887) |
| 0.1 | 1.0 | GO:0071376 | response to corticotropin-releasing hormone(GO:0043435) cellular response to corticotropin-releasing hormone stimulus(GO:0071376) |
| 0.1 | 0.6 | GO:0003232 | bulbus arteriosus development(GO:0003232) |
| 0.1 | 1.0 | GO:0050909 | sensory perception of taste(GO:0050909) |
| 0.1 | 0.3 | GO:0001954 | positive regulation of cell-matrix adhesion(GO:0001954) |
| 0.1 | 0.4 | GO:0002949 | tRNA threonylcarbamoyladenosine modification(GO:0002949) |
| 0.1 | 0.4 | GO:0051792 | medium-chain fatty acid biosynthetic process(GO:0051792) |
| 0.1 | 0.7 | GO:0071578 | zinc II ion transmembrane import(GO:0071578) |
| 0.1 | 0.2 | GO:0090131 | mesenchyme migration(GO:0090131) |
| 0.1 | 0.4 | GO:0097066 | response to thyroid hormone(GO:0097066) |
| 0.1 | 0.5 | GO:0033687 | osteoblast proliferation(GO:0033687) |
| 0.1 | 0.2 | GO:0032241 | regulation of nucleobase-containing compound transport(GO:0032239) positive regulation of nucleobase-containing compound transport(GO:0032241) positive regulation of nucleocytoplasmic transport(GO:0046824) regulation of RNA export from nucleus(GO:0046831) positive regulation of RNA export from nucleus(GO:0046833) messenger ribonucleoprotein complex assembly(GO:1990120) |
| 0.1 | 1.7 | GO:0010508 | positive regulation of autophagy(GO:0010508) |
| 0.1 | 0.2 | GO:0030238 | male sex determination(GO:0030238) |
| 0.1 | 0.2 | GO:0043504 | mitochondrial DNA repair(GO:0043504) |
| 0.1 | 0.2 | GO:0071548 | cellular response to cortisol stimulus(GO:0071387) response to dexamethasone(GO:0071548) |
| 0.1 | 1.2 | GO:0051489 | regulation of filopodium assembly(GO:0051489) |
| 0.0 | 1.2 | GO:0006942 | regulation of striated muscle contraction(GO:0006942) |
| 0.0 | 0.9 | GO:0007130 | synaptonemal complex assembly(GO:0007130) synaptonemal complex organization(GO:0070193) |
| 0.0 | 1.0 | GO:0007198 | adenylate cyclase-inhibiting serotonin receptor signaling pathway(GO:0007198) |
| 0.0 | 0.9 | GO:0097178 | ruffle assembly(GO:0097178) |
| 0.0 | 0.2 | GO:0010032 | meiotic chromosome condensation(GO:0010032) |
| 0.0 | 0.3 | GO:1900045 | negative regulation of histone ubiquitination(GO:0033183) regulation of protein K63-linked ubiquitination(GO:1900044) negative regulation of protein K63-linked ubiquitination(GO:1900045) regulation of histone H2A K63-linked ubiquitination(GO:1901314) negative regulation of histone H2A K63-linked ubiquitination(GO:1901315) regulation of protein polyubiquitination(GO:1902914) negative regulation of protein polyubiquitination(GO:1902915) |
| 0.0 | 0.4 | GO:0001709 | cell fate determination(GO:0001709) |
| 0.0 | 0.3 | GO:0061298 | retina vasculature development in camera-type eye(GO:0061298) |
| 0.0 | 5.7 | GO:0002088 | lens development in camera-type eye(GO:0002088) |
| 0.0 | 0.2 | GO:0006844 | acyl carnitine transport(GO:0006844) |
| 0.0 | 0.2 | GO:0003322 | pancreatic A cell development(GO:0003322) |
| 0.0 | 0.1 | GO:0097264 | self proteolysis(GO:0097264) |
| 0.0 | 0.2 | GO:0010269 | response to selenium ion(GO:0010269) |
| 0.0 | 0.2 | GO:0033499 | galactose catabolic process(GO:0019388) galactose catabolic process via UDP-galactose(GO:0033499) |
| 0.0 | 0.2 | GO:0060760 | positive regulation of cytokine-mediated signaling pathway(GO:0001961) positive regulation of response to cytokine stimulus(GO:0060760) |
| 0.0 | 0.2 | GO:0005513 | detection of calcium ion(GO:0005513) |
| 0.0 | 0.2 | GO:0030326 | embryonic limb morphogenesis(GO:0030326) |
| 0.0 | 0.5 | GO:0048672 | positive regulation of collateral sprouting(GO:0048672) |
| 0.0 | 0.6 | GO:0098943 | neurotransmitter receptor transport, postsynaptic endosome to lysosome(GO:0098943) |
| 0.0 | 0.2 | GO:0061056 | sclerotome development(GO:0061056) |
| 0.0 | 0.6 | GO:0043114 | regulation of vascular permeability(GO:0043114) |
| 0.0 | 0.9 | GO:0021952 | central nervous system projection neuron axonogenesis(GO:0021952) |
| 0.0 | 0.5 | GO:0045471 | response to ethanol(GO:0045471) |
| 0.0 | 0.2 | GO:0032656 | interleukin-13 production(GO:0032616) regulation of interleukin-13 production(GO:0032656) |
| 0.0 | 0.2 | GO:0038065 | collagen-activated tyrosine kinase receptor signaling pathway(GO:0038063) collagen-activated signaling pathway(GO:0038065) |
| 0.0 | 0.3 | GO:0000056 | ribosomal small subunit export from nucleus(GO:0000056) |
| 0.0 | 0.4 | GO:0036353 | histone H2A-K119 monoubiquitination(GO:0036353) |
| 0.0 | 0.5 | GO:0043433 | negative regulation of sequence-specific DNA binding transcription factor activity(GO:0043433) |
| 0.0 | 1.1 | GO:0048010 | vascular endothelial growth factor receptor signaling pathway(GO:0048010) |
| 0.0 | 0.3 | GO:0003160 | endocardium morphogenesis(GO:0003160) |
| 0.0 | 0.4 | GO:0006474 | N-terminal protein amino acid acetylation(GO:0006474) |
| 0.0 | 0.1 | GO:0046952 | ketone body catabolic process(GO:0046952) |
| 0.0 | 0.1 | GO:0099612 | protein localization to juxtaparanode region of axon(GO:0071205) protein localization to axon(GO:0099612) |
| 0.0 | 0.1 | GO:2000726 | negative regulation of striated muscle cell differentiation(GO:0051154) negative regulation of cardiac muscle tissue development(GO:0055026) cardiac muscle cell fate commitment(GO:0060923) canonical Wnt signaling pathway involved in cardiac muscle cell fate commitment(GO:0061317) regulation of canonical Wnt signaling pathway involved in cardiac muscle cell fate commitment(GO:1901295) negative regulation of canonical Wnt signaling pathway involved in cardiac muscle cell fate commitment(GO:1901296) negative regulation of cardiocyte differentiation(GO:1905208) negative regulation of cardiac muscle cell differentiation(GO:2000726) |
| 0.0 | 0.4 | GO:0031581 | hemidesmosome assembly(GO:0031581) |
| 0.0 | 0.2 | GO:0031639 | plasminogen activation(GO:0031639) |
| 0.0 | 0.2 | GO:0000098 | sulfur amino acid catabolic process(GO:0000098) |
| 0.0 | 1.1 | GO:0036269 | swimming behavior(GO:0036269) |
| 0.0 | 0.7 | GO:0060037 | pharyngeal system development(GO:0060037) |
| 0.0 | 0.2 | GO:0006177 | GMP biosynthetic process(GO:0006177) GMP metabolic process(GO:0046037) |
| 0.0 | 0.4 | GO:0043162 | ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway(GO:0043162) |
| 0.0 | 0.2 | GO:1990845 | adaptive thermogenesis(GO:1990845) |
| 0.0 | 0.1 | GO:0007252 | I-kappaB phosphorylation(GO:0007252) |
| 0.0 | 0.2 | GO:0007525 | somatic muscle development(GO:0007525) |
| 0.0 | 0.6 | GO:0043967 | histone H4 acetylation(GO:0043967) |
| 0.0 | 0.1 | GO:0060623 | regulation of chromosome condensation(GO:0060623) |
| 0.0 | 1.0 | GO:0030514 | negative regulation of BMP signaling pathway(GO:0030514) |
| 0.0 | 1.0 | GO:0001946 | lymphangiogenesis(GO:0001946) |
| 0.0 | 0.7 | GO:0030199 | collagen fibril organization(GO:0030199) |
| 0.0 | 0.2 | GO:0098887 | neurotransmitter receptor transport, endosome to postsynaptic membrane(GO:0098887) endosome to plasma membrane protein transport(GO:0099638) neurotransmitter receptor transport, endosome to plasma membrane(GO:0099639) |
| 0.0 | 0.5 | GO:0016525 | negative regulation of angiogenesis(GO:0016525) negative regulation of vasculature development(GO:1901343) negative regulation of blood vessel morphogenesis(GO:2000181) |
| 0.0 | 0.2 | GO:0045741 | positive regulation of epidermal growth factor-activated receptor activity(GO:0045741) positive regulation of receptor activity(GO:2000273) |
| 0.0 | 0.3 | GO:0008078 | mesodermal cell migration(GO:0008078) |
| 0.0 | 0.1 | GO:1904036 | negative regulation of epithelial cell apoptotic process(GO:1904036) |
| 0.0 | 0.2 | GO:0043551 | regulation of lipid kinase activity(GO:0043550) regulation of phosphatidylinositol 3-kinase activity(GO:0043551) |
| 0.0 | 0.3 | GO:1901663 | ubiquinone metabolic process(GO:0006743) ubiquinone biosynthetic process(GO:0006744) quinone biosynthetic process(GO:1901663) |
| 0.0 | 0.4 | GO:0043249 | erythrocyte maturation(GO:0043249) |
| 0.0 | 0.2 | GO:0048096 | chromatin-mediated maintenance of transcription(GO:0048096) |
| 0.0 | 0.1 | GO:0034063 | stress granule assembly(GO:0034063) |
| 0.0 | 0.0 | GO:0019240 | citrulline biosynthetic process(GO:0019240) |
| 0.0 | 0.9 | GO:0030318 | melanocyte differentiation(GO:0030318) |
| 0.0 | 0.7 | GO:0014046 | dopamine secretion(GO:0014046) regulation of dopamine secretion(GO:0014059) |
| 0.0 | 0.3 | GO:0051932 | synaptic transmission, GABAergic(GO:0051932) |
| 0.0 | 0.5 | GO:0006471 | protein ADP-ribosylation(GO:0006471) |
| 0.0 | 0.2 | GO:0030816 | activation of adenylate cyclase activity(GO:0007190) positive regulation of cAMP metabolic process(GO:0030816) positive regulation of cAMP biosynthetic process(GO:0030819) positive regulation of adenylate cyclase activity(GO:0045762) |
| 0.0 | 0.4 | GO:0060536 | cartilage morphogenesis(GO:0060536) |
| 0.0 | 0.4 | GO:0042476 | odontogenesis(GO:0042476) |
| 0.0 | 0.5 | GO:0006482 | protein demethylation(GO:0006482) protein dealkylation(GO:0008214) |
| 0.0 | 0.4 | GO:0001732 | formation of cytoplasmic translation initiation complex(GO:0001732) |
| 0.0 | 0.1 | GO:0007023 | post-chaperonin tubulin folding pathway(GO:0007023) |
| 0.0 | 0.0 | GO:0060231 | mesenchymal to epithelial transition(GO:0060231) |
| 0.0 | 0.1 | GO:0045743 | positive regulation of fibroblast growth factor receptor signaling pathway(GO:0045743) |
| 0.0 | 0.1 | GO:0001682 | tRNA 5'-leader removal(GO:0001682) |
| 0.0 | 1.0 | GO:0030838 | positive regulation of actin filament polymerization(GO:0030838) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.8 | GO:0005915 | cell-cell adherens junction(GO:0005913) zonula adherens(GO:0005915) |
| 0.1 | 0.7 | GO:0043083 | synaptic cleft(GO:0043083) |
| 0.1 | 0.4 | GO:0097524 | sperm plasma membrane(GO:0097524) |
| 0.1 | 0.4 | GO:0071203 | WASH complex(GO:0071203) |
| 0.1 | 0.8 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.1 | 0.5 | GO:0035517 | PR-DUB complex(GO:0035517) |
| 0.1 | 0.4 | GO:0070032 | synaptobrevin 2-SNAP-25-syntaxin-1a-complexin I complex(GO:0070032) |
| 0.1 | 0.2 | GO:0042382 | paraspeckles(GO:0042382) |
| 0.1 | 0.6 | GO:0005671 | Ada2/Gcn5/Ada3 transcription activator complex(GO:0005671) |
| 0.1 | 0.4 | GO:0000408 | EKC/KEOPS complex(GO:0000408) |
| 0.1 | 1.0 | GO:0031680 | G-protein beta/gamma-subunit complex(GO:0031680) |
| 0.1 | 0.8 | GO:0034361 | very-low-density lipoprotein particle(GO:0034361) |
| 0.1 | 0.2 | GO:0005828 | kinetochore microtubule(GO:0005828) |
| 0.0 | 0.7 | GO:0005682 | U5 snRNP(GO:0005682) |
| 0.0 | 0.2 | GO:0098837 | postsynaptic recycling endosome(GO:0098837) |
| 0.0 | 0.4 | GO:0042788 | polysomal ribosome(GO:0042788) |
| 0.0 | 0.2 | GO:0016012 | sarcoglycan complex(GO:0016012) |
| 0.0 | 0.9 | GO:0000795 | synaptonemal complex(GO:0000795) |
| 0.0 | 0.1 | GO:0061673 | mitotic spindle astral microtubule(GO:0061673) |
| 0.0 | 0.2 | GO:0098651 | collagen type IV trimer(GO:0005587) network-forming collagen trimer(GO:0098642) collagen network(GO:0098645) basement membrane collagen trimer(GO:0098651) |
| 0.0 | 1.0 | GO:0016605 | PML body(GO:0016605) |
| 0.0 | 1.0 | GO:0005861 | troponin complex(GO:0005861) |
| 0.0 | 0.4 | GO:0000813 | ESCRT I complex(GO:0000813) |
| 0.0 | 1.0 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.0 | 0.8 | GO:0030864 | cortical actin cytoskeleton(GO:0030864) |
| 0.0 | 0.1 | GO:0008385 | IkappaB kinase complex(GO:0008385) |
| 0.0 | 0.4 | GO:0035102 | PRC1 complex(GO:0035102) |
| 0.0 | 0.3 | GO:0033180 | proton-transporting V-type ATPase, V1 domain(GO:0033180) |
| 0.0 | 0.1 | GO:0033010 | paranodal junction(GO:0033010) |
| 0.0 | 0.2 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.0 | 0.3 | GO:0048188 | Set1C/COMPASS complex(GO:0048188) |
| 0.0 | 0.6 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.0 | 0.1 | GO:0000172 | ribonuclease MRP complex(GO:0000172) |
| 0.0 | 0.3 | GO:0030175 | filopodium(GO:0030175) |
| 0.0 | 0.1 | GO:0000811 | GINS complex(GO:0000811) |
| 0.0 | 1.0 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.0 | 1.2 | GO:0030027 | lamellipodium(GO:0030027) |
| 0.0 | 3.9 | GO:0005667 | transcription factor complex(GO:0005667) |
| 0.0 | 0.3 | GO:0032590 | neuron projection membrane(GO:0032589) dendrite membrane(GO:0032590) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.8 | GO:0070097 | delta-catenin binding(GO:0070097) |
| 0.2 | 0.8 | GO:0030228 | lipoprotein particle receptor activity(GO:0030228) |
| 0.2 | 0.5 | GO:0030792 | arsenite methyltransferase activity(GO:0030791) methylarsonite methyltransferase activity(GO:0030792) |
| 0.2 | 0.6 | GO:0051059 | NF-kappaB binding(GO:0051059) |
| 0.1 | 0.4 | GO:0061711 | N(6)-L-threonylcarbamoyladenine synthase(GO:0061711) |
| 0.1 | 0.7 | GO:0030298 | receptor signaling protein tyrosine kinase activator activity(GO:0030298) |
| 0.1 | 0.6 | GO:0004692 | cGMP-dependent protein kinase activity(GO:0004692) |
| 0.1 | 2.1 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.1 | 0.7 | GO:0070888 | E-box binding(GO:0070888) |
| 0.1 | 0.5 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.1 | 1.0 | GO:0031013 | troponin C binding(GO:0030172) troponin I binding(GO:0031013) |
| 0.1 | 0.4 | GO:0043998 | H2A histone acetyltransferase activity(GO:0043998) |
| 0.1 | 1.0 | GO:0035259 | glucocorticoid receptor binding(GO:0035259) |
| 0.1 | 0.9 | GO:0005522 | profilin binding(GO:0005522) |
| 0.1 | 2.8 | GO:0005518 | collagen binding(GO:0005518) |
| 0.1 | 0.4 | GO:0008126 | acetylesterase activity(GO:0008126) |
| 0.1 | 0.3 | GO:0019784 | NEDD8-specific protease activity(GO:0019784) |
| 0.1 | 0.2 | GO:0003978 | UDP-glucose 4-epimerase activity(GO:0003978) |
| 0.1 | 5.5 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.1 | 1.1 | GO:0070122 | isopeptidase activity(GO:0070122) |
| 0.1 | 0.4 | GO:0004045 | aminoacyl-tRNA hydrolase activity(GO:0004045) |
| 0.1 | 0.5 | GO:0004865 | protein serine/threonine phosphatase inhibitor activity(GO:0004865) |
| 0.0 | 0.7 | GO:0038191 | neuropilin binding(GO:0038191) |
| 0.0 | 1.0 | GO:0043176 | amine binding(GO:0043176) serotonin binding(GO:0051378) |
| 0.0 | 0.2 | GO:0015227 | acyl carnitine transmembrane transporter activity(GO:0015227) |
| 0.0 | 0.5 | GO:0015129 | lactate transmembrane transporter activity(GO:0015129) |
| 0.0 | 0.2 | GO:0017077 | oxidative phosphorylation uncoupler activity(GO:0017077) |
| 0.0 | 0.9 | GO:0008519 | ammonium transmembrane transporter activity(GO:0008519) |
| 0.0 | 0.3 | GO:0048039 | ubiquinone binding(GO:0048039) |
| 0.0 | 0.5 | GO:0005163 | nerve growth factor receptor binding(GO:0005163) neurotrophin receptor binding(GO:0005165) |
| 0.0 | 0.2 | GO:0003938 | IMP dehydrogenase activity(GO:0003938) |
| 0.0 | 0.5 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.0 | 0.5 | GO:0032041 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.0 | 0.5 | GO:0048495 | Roundabout binding(GO:0048495) |
| 0.0 | 0.1 | GO:0017064 | fatty acid amide hydrolase activity(GO:0017064) |
| 0.0 | 0.4 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 0.0 | 0.3 | GO:0008422 | beta-glucosidase activity(GO:0008422) |
| 0.0 | 0.3 | GO:0005049 | nuclear export signal receptor activity(GO:0005049) |
| 0.0 | 0.7 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.0 | 0.1 | GO:0008260 | 3-oxoacid CoA-transferase activity(GO:0008260) |
| 0.0 | 0.1 | GO:0008384 | IkappaB kinase activity(GO:0008384) |
| 0.0 | 0.2 | GO:0008515 | sugar:proton symporter activity(GO:0005351) cation:sugar symporter activity(GO:0005402) sucrose:proton symporter activity(GO:0008506) sucrose transmembrane transporter activity(GO:0008515) disaccharide transmembrane transporter activity(GO:0015154) oligosaccharide transmembrane transporter activity(GO:0015157) |
| 0.0 | 0.3 | GO:0022851 | benzodiazepine receptor activity(GO:0008503) GABA-gated chloride ion channel activity(GO:0022851) |
| 0.0 | 0.2 | GO:0070579 | methylcytosine dioxygenase activity(GO:0070579) |
| 0.0 | 0.6 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.0 | 0.5 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.0 | 0.2 | GO:0008568 | microtubule-severing ATPase activity(GO:0008568) |
| 0.0 | 1.0 | GO:0001786 | phosphatidylserine binding(GO:0001786) |
| 0.0 | 0.1 | GO:0070698 | type I activin receptor binding(GO:0070698) |
| 0.0 | 0.2 | GO:0043515 | kinetochore binding(GO:0043515) |
| 0.0 | 0.7 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.0 | 0.9 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.0 | 8.2 | GO:0046983 | protein dimerization activity(GO:0046983) |
| 0.0 | 0.2 | GO:0015269 | calcium-activated potassium channel activity(GO:0015269) |
| 0.0 | 0.5 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.0 | 0.8 | GO:0046332 | SMAD binding(GO:0046332) |
| 0.0 | 0.1 | GO:0043014 | alpha-tubulin binding(GO:0043014) |
| 0.0 | 0.6 | GO:0004402 | histone acetyltransferase activity(GO:0004402) |
| 0.0 | 0.2 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 0.0 | 0.4 | GO:0000030 | mannosyltransferase activity(GO:0000030) |
| 0.0 | 1.9 | GO:0001228 | transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001228) |
| 0.0 | 0.1 | GO:0004985 | opioid receptor activity(GO:0004985) |
| 0.0 | 0.1 | GO:0034450 | ubiquitin-ubiquitin ligase activity(GO:0034450) |
| 0.0 | 0.3 | GO:0003899 | DNA-directed RNA polymerase activity(GO:0003899) RNA polymerase activity(GO:0034062) |
| 0.0 | 0.4 | GO:0098631 | protein binding involved in cell adhesion(GO:0098631) protein binding involved in cell-cell adhesion(GO:0098632) |
| 0.0 | 0.5 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.0 | 0.5 | GO:0017022 | myosin binding(GO:0017022) |
| 0.0 | 0.0 | GO:0015105 | arsenite transmembrane transporter activity(GO:0015105) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.1 | PID THROMBIN PAR4 PATHWAY | PAR4-mediated thrombin signaling events |
| 0.1 | 2.3 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 1.0 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.0 | 0.8 | PID REELIN PATHWAY | Reelin signaling pathway |
| 0.0 | 0.4 | PID S1P S1P4 PATHWAY | S1P4 pathway |
| 0.0 | 0.6 | PID RANBP2 PATHWAY | Sumoylation by RanBP2 regulates transcriptional repression |
| 0.0 | 0.2 | PID ARF6 DOWNSTREAM PATHWAY | Arf6 downstream pathway |
| 0.0 | 0.3 | PID INTEGRIN A9B1 PATHWAY | Alpha9 beta1 integrin signaling events |
| 0.0 | 0.4 | PID INTEGRIN CS PATHWAY | Integrin family cell surface interactions |
| 0.0 | 0.1 | PID TCR JNK PATHWAY | JNK signaling in the CD4+ TCR pathway |
| 0.0 | 0.2 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.9 | REACTOME CS DS DEGRADATION | Genes involved in CS/DS degradation |
| 0.1 | 0.8 | REACTOME TRAFFICKING AND PROCESSING OF ENDOSOMAL TLR | Genes involved in Trafficking and processing of endosomal TLR |
| 0.1 | 1.1 | REACTOME INHIBITION OF INSULIN SECRETION BY ADRENALINE NORADRENALINE | Genes involved in Inhibition of Insulin Secretion by Adrenaline/Noradrenaline |
| 0.1 | 0.4 | REACTOME CD28 DEPENDENT VAV1 PATHWAY | Genes involved in CD28 dependent Vav1 pathway |
| 0.1 | 0.7 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.0 | 0.9 | REACTOME AMINE COMPOUND SLC TRANSPORTERS | Genes involved in Amine compound SLC transporters |
| 0.0 | 1.1 | REACTOME PIP3 ACTIVATES AKT SIGNALING | Genes involved in PIP3 activates AKT signaling |
| 0.0 | 0.5 | REACTOME BILE SALT AND ORGANIC ANION SLC TRANSPORTERS | Genes involved in Bile salt and organic anion SLC transporters |
| 0.0 | 0.4 | REACTOME MEMBRANE BINDING AND TARGETTING OF GAG PROTEINS | Genes involved in Membrane binding and targetting of GAG proteins |
| 0.0 | 0.5 | REACTOME ACTIVATED NOTCH1 TRANSMITS SIGNAL TO THE NUCLEUS | Genes involved in Activated NOTCH1 Transmits Signal to the Nucleus |
| 0.0 | 0.3 | REACTOME GABA A RECEPTOR ACTIVATION | Genes involved in GABA A receptor activation |
| 0.0 | 0.2 | REACTOME ARMS MEDIATED ACTIVATION | Genes involved in ARMS-mediated activation |
| 0.0 | 0.3 | REACTOME INTEGRATION OF ENERGY METABOLISM | Genes involved in Integration of energy metabolism |
| 0.0 | 0.5 | REACTOME G0 AND EARLY G1 | Genes involved in G0 and Early G1 |
| 0.0 | 0.2 | REACTOME PI3K EVENTS IN ERBB4 SIGNALING | Genes involved in PI3K events in ERBB4 signaling |
| 0.0 | 0.5 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.0 | 0.5 | REACTOME MRNA SPLICING MINOR PATHWAY | Genes involved in mRNA Splicing - Minor Pathway |