Project
PRJEB1986: zebrafish developmental stages transcriptome
Navigation
Downloads
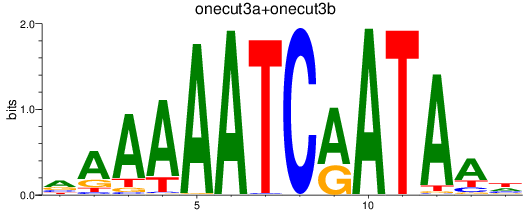
Results for onecut3a+onecut3b
Z-value: 0.67
Transcription factors associated with onecut3a+onecut3b
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
onecut3a
|
ENSDARG00000056395 | one cut homeobox 3a |
|
onecut3b
|
ENSDARG00000091059 | one cut homeobox 3b |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| onecut3a | dr11_v1_chr22_+_20461488_20461488 | 0.73 | 4.4e-04 | Click! |
| onecut3b | dr11_v1_chr2_+_57504073_57504073 | 0.50 | 2.9e-02 | Click! |
Activity profile of onecut3a+onecut3b motif
Sorted Z-values of onecut3a+onecut3b motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr24_-_38113208 | 1.68 |
ENSDART00000079003
|
crp
|
c-reactive protein, pentraxin-related |
| chr18_+_642889 | 1.66 |
ENSDART00000189007
|
CABZ01078320.1
|
|
| chr14_+_35748385 | 1.35 |
ENSDART00000064617
ENSDART00000074671 ENSDART00000172803 |
gria2b
|
glutamate receptor, ionotropic, AMPA 2b |
| chr15_-_34418525 | 1.33 |
ENSDART00000147582
|
agmo
|
alkylglycerol monooxygenase |
| chr1_-_21409877 | 1.29 |
ENSDART00000102782
|
gria2a
|
glutamate receptor, ionotropic, AMPA 2a |
| chr17_-_20897407 | 1.27 |
ENSDART00000149481
|
ank3b
|
ankyrin 3b |
| chr14_+_35748206 | 1.24 |
ENSDART00000177391
|
gria2b
|
glutamate receptor, ionotropic, AMPA 2b |
| chr3_-_60142530 | 1.23 |
ENSDART00000153247
|
si:ch211-120g10.1
|
si:ch211-120g10.1 |
| chr4_-_5291256 | 1.22 |
ENSDART00000150864
|
SNAP91 (1 of many)
|
si:ch211-214j24.9 |
| chr1_+_19332837 | 1.21 |
ENSDART00000078594
|
tyrp1b
|
tyrosinase-related protein 1b |
| chr3_-_28665291 | 1.19 |
ENSDART00000151670
|
fbxl16
|
F-box and leucine-rich repeat protein 16 |
| chr3_-_28120092 | 1.18 |
ENSDART00000151143
|
rbfox1
|
RNA binding fox-1 homolog 1 |
| chr12_+_26512569 | 1.14 |
ENSDART00000066264
|
chad
|
chondroadherin |
| chr1_+_37391716 | 1.07 |
ENSDART00000191986
|
sparcl1
|
SPARC-like 1 |
| chr11_+_6819050 | 1.05 |
ENSDART00000104289
|
rab3ab
|
RAB3A, member RAS oncogene family, b |
| chr17_-_20897250 | 0.99 |
ENSDART00000088106
|
ank3b
|
ankyrin 3b |
| chr16_-_12173554 | 0.98 |
ENSDART00000110567
ENSDART00000155935 |
clstn3
|
calsyntenin 3 |
| chr11_-_10798021 | 0.97 |
ENSDART00000167112
ENSDART00000179725 ENSDART00000091923 ENSDART00000185825 |
slc4a10a
|
solute carrier family 4, sodium bicarbonate transporter, member 10a |
| chr2_-_32505091 | 0.95 |
ENSDART00000141884
ENSDART00000056639 |
faim2a
|
Fas apoptotic inhibitory molecule 2a |
| chr8_-_23081511 | 0.94 |
ENSDART00000142015
ENSDART00000135764 ENSDART00000147021 |
si:dkey-70p6.1
|
si:dkey-70p6.1 |
| chr21_+_4116437 | 0.90 |
ENSDART00000167791
ENSDART00000123759 |
rapgef1b
|
Rap guanine nucleotide exchange factor (GEF) 1b |
| chr2_+_47582488 | 0.90 |
ENSDART00000149967
|
scg2b
|
secretogranin II (chromogranin C), b |
| chr12_+_10116912 | 0.90 |
ENSDART00000189630
|
si:dkeyp-118b1.2
|
si:dkeyp-118b1.2 |
| chr5_-_71460556 | 0.89 |
ENSDART00000108804
|
brinp1
|
bone morphogenetic protein/retinoic acid inducible neural-specific 1 |
| chr8_-_14052349 | 0.89 |
ENSDART00000135811
|
atp2b3a
|
ATPase plasma membrane Ca2+ transporting 3a |
| chr5_+_19309877 | 0.87 |
ENSDART00000190338
|
rusc2
|
RUN and SH3 domain containing 2 |
| chr6_+_3828560 | 0.85 |
ENSDART00000185273
ENSDART00000179091 |
gad1b
|
glutamate decarboxylase 1b |
| chr10_-_33156789 | 0.82 |
ENSDART00000192268
ENSDART00000182065 ENSDART00000081170 |
cux1a
|
cut-like homeobox 1a |
| chr19_-_17526735 | 0.81 |
ENSDART00000189391
|
thrb
|
thyroid hormone receptor beta |
| chr1_-_23157583 | 0.81 |
ENSDART00000144208
|
adgrl3.1
|
adhesion G protein-coupled receptor L3.1 |
| chr8_+_40477264 | 0.79 |
ENSDART00000085559
|
gck
|
glucokinase (hexokinase 4) |
| chr22_+_17359346 | 0.79 |
ENSDART00000145434
|
gpr52
|
G protein-coupled receptor 52 |
| chr14_+_49376011 | 0.78 |
ENSDART00000020961
|
tnip1
|
TNFAIP3 interacting protein 1 |
| chr24_-_23320223 | 0.78 |
ENSDART00000135846
|
zfhx4
|
zinc finger homeobox 4 |
| chr4_+_7827261 | 0.78 |
ENSDART00000129568
|
phyh
|
phytanoyl-CoA 2-hydroxylase |
| chr20_+_22130284 | 0.77 |
ENSDART00000025575
|
clocka
|
clock circadian regulator a |
| chr2_-_56387041 | 0.77 |
ENSDART00000036240
|
cers4b
|
ceramide synthase 4b |
| chr25_-_9805269 | 0.76 |
ENSDART00000192048
|
lrrc4c
|
leucine rich repeat containing 4C |
| chr16_-_12173399 | 0.76 |
ENSDART00000142574
|
clstn3
|
calsyntenin 3 |
| chr18_+_3579829 | 0.75 |
ENSDART00000158763
ENSDART00000182850 ENSDART00000162754 ENSDART00000178789 ENSDART00000172656 |
lrch3
|
leucine-rich repeats and calponin homology (CH) domain containing 3 |
| chr4_-_9214288 | 0.74 |
ENSDART00000067309
|
avpr1ab
|
arginine vasopressin receptor 1Ab |
| chr7_-_68419830 | 0.74 |
ENSDART00000191606
|
CU468723.2
|
|
| chr21_-_23475361 | 0.73 |
ENSDART00000156658
ENSDART00000157454 |
ncam1a
|
neural cell adhesion molecule 1a |
| chr2_+_5887626 | 0.73 |
ENSDART00000147831
|
si:ch211-168b3.1
|
si:ch211-168b3.1 |
| chr10_-_31782616 | 0.73 |
ENSDART00000128839
|
fez1
|
fasciculation and elongation protein zeta 1 (zygin I) |
| chr23_-_15216654 | 0.70 |
ENSDART00000131649
|
sulf2b
|
sulfatase 2b |
| chr2_+_25860344 | 0.70 |
ENSDART00000178841
|
slc7a14a
|
solute carrier family 7, member 14a |
| chr15_-_7598294 | 0.70 |
ENSDART00000165898
|
gbe1b
|
glucan (1,4-alpha-), branching enzyme 1b |
| chr13_-_31166544 | 0.70 |
ENSDART00000146250
ENSDART00000132129 ENSDART00000139591 |
mapk8a
|
mitogen-activated protein kinase 8a |
| chr15_-_7598542 | 0.68 |
ENSDART00000173092
|
gbe1b
|
glucan (1,4-alpha-), branching enzyme 1b |
| chr15_+_22394074 | 0.66 |
ENSDART00000109931
|
oafa
|
OAF homolog a (Drosophila) |
| chr17_-_26507289 | 0.65 |
ENSDART00000155616
|
ccser2a
|
coiled-coil serine-rich protein 2a |
| chr16_+_21738194 | 0.64 |
ENSDART00000163688
|
FP085428.1
|
Danio rerio si:ch211-154o6.4 (si:ch211-154o6.4), mRNA. |
| chr16_-_44127307 | 0.61 |
ENSDART00000104583
ENSDART00000151936 ENSDART00000058685 ENSDART00000190830 |
zfpm2a
|
zinc finger protein, FOG family member 2a |
| chr15_-_34785594 | 0.60 |
ENSDART00000154256
|
gabbr1a
|
gamma-aminobutyric acid (GABA) B receptor, 1a |
| chr21_+_6212844 | 0.60 |
ENSDART00000150301
|
fnbp1b
|
formin binding protein 1b |
| chr7_-_41693004 | 0.59 |
ENSDART00000121509
|
malrd1
|
MAM and LDL receptor class A domain containing 1 |
| chr18_-_46380471 | 0.59 |
ENSDART00000144444
ENSDART00000086772 |
slc19a3b
|
solute carrier family 19 (thiamine transporter), member 3b |
| chr2_+_47581997 | 0.58 |
ENSDART00000112579
|
scg2b
|
secretogranin II (chromogranin C), b |
| chr14_+_21783229 | 0.57 |
ENSDART00000170784
|
ankrd13d
|
ankyrin repeat domain 13 family, member D |
| chr22_-_3092136 | 0.57 |
ENSDART00000157620
|
ing5a
|
inhibitor of growth family, member 5a |
| chr7_-_36096582 | 0.56 |
ENSDART00000188507
|
CR848715.1
|
|
| chr14_-_31893996 | 0.55 |
ENSDART00000173222
|
gpr101
|
G protein-coupled receptor 101 |
| chr3_+_39853788 | 0.55 |
ENSDART00000154869
|
cacna1ha
|
calcium channel, voltage-dependent, T type, alpha 1H subunit a |
| chr9_+_32358514 | 0.54 |
ENSDART00000144608
|
plcl1
|
phospholipase C like 1 |
| chr15_+_17446796 | 0.54 |
ENSDART00000157189
|
snx19b
|
sorting nexin 19b |
| chr2_-_6262441 | 0.53 |
ENSDART00000092190
|
arl14
|
ADP-ribosylation factor-like 14 |
| chr6_+_40714811 | 0.52 |
ENSDART00000153868
|
ccdc36
|
coiled-coil domain containing 36 |
| chr6_+_40992409 | 0.52 |
ENSDART00000151419
|
tgfa
|
transforming growth factor, alpha |
| chr3_-_30434016 | 0.52 |
ENSDART00000150958
|
lrrc4ba
|
leucine rich repeat containing 4Ba |
| chr6_-_33707278 | 0.50 |
ENSDART00000188103
|
mast2
|
microtubule associated serine/threonine kinase 2 |
| chr24_-_36680261 | 0.50 |
ENSDART00000059507
|
ccr10
|
chemokine (C-C motif) receptor 10 |
| chr21_-_17482465 | 0.49 |
ENSDART00000004548
|
barhl1b
|
BarH-like homeobox 1b |
| chr16_-_26255877 | 0.49 |
ENSDART00000146214
|
erfl1
|
Ets2 repressor factor like 1 |
| chr18_+_15876385 | 0.49 |
ENSDART00000142527
|
eea1
|
early endosome antigen 1 |
| chr10_-_7785930 | 0.47 |
ENSDART00000043961
ENSDART00000111058 |
mpx
|
myeloid-specific peroxidase |
| chr1_-_36151377 | 0.45 |
ENSDART00000037516
|
znf827
|
zinc finger protein 827 |
| chr20_-_40367493 | 0.44 |
ENSDART00000075096
|
smpdl3a
|
sphingomyelin phosphodiesterase, acid-like 3A |
| chr1_+_40140498 | 0.43 |
ENSDART00000101640
|
zgc:152658
|
zgc:152658 |
| chr19_+_19512515 | 0.43 |
ENSDART00000180065
|
jazf1a
|
JAZF zinc finger 1a |
| chr12_+_32729470 | 0.43 |
ENSDART00000175712
|
rbfox3a
|
RNA binding fox-1 homolog 3a |
| chr18_-_26894732 | 0.42 |
ENSDART00000147735
ENSDART00000188938 |
BX470164.1
si:dkey-24l11.2
|
si:dkey-24l11.2 |
| chr21_-_39058490 | 0.42 |
ENSDART00000114885
|
aldh3a2b
|
aldehyde dehydrogenase 3 family, member A2b |
| chr17_-_8169774 | 0.42 |
ENSDART00000091828
|
syne1b
|
spectrin repeat containing, nuclear envelope 1b |
| chr12_-_30338779 | 0.41 |
ENSDART00000192511
|
vwa2
|
von Willebrand factor A domain containing 2 |
| chr10_-_41352502 | 0.41 |
ENSDART00000052971
ENSDART00000128156 |
rab11fip1b
|
RAB11 family interacting protein 1 (class I) b |
| chr13_+_36144341 | 0.41 |
ENSDART00000182930
ENSDART00000187327 |
TTC9
|
si:ch211-259k16.3 |
| chr21_+_3901775 | 0.41 |
ENSDART00000053609
|
dolpp1
|
dolichyldiphosphatase 1 |
| chr5_-_67661102 | 0.40 |
ENSDART00000013605
|
zbtb20
|
zinc finger and BTB domain containing 20 |
| chr3_-_44753325 | 0.40 |
ENSDART00000074543
|
hs3st4
|
heparan sulfate (glucosamine) 3-O-sulfotransferase 4 |
| chr3_-_32071468 | 0.39 |
ENSDART00000121979
|
BX511080.1
|
|
| chr13_+_49727333 | 0.39 |
ENSDART00000168799
ENSDART00000037559 |
ggps1
|
geranylgeranyl diphosphate synthase 1 |
| chr25_+_22017182 | 0.36 |
ENSDART00000156517
|
si:dkey-217l24.1
|
si:dkey-217l24.1 |
| chr5_+_42136359 | 0.36 |
ENSDART00000083731
|
trpv1
|
transient receptor potential cation channel, subfamily V, member 1 |
| chr21_+_17768174 | 0.34 |
ENSDART00000141380
|
rxraa
|
retinoid X receptor, alpha a |
| chr22_-_7669624 | 0.34 |
ENSDART00000189617
|
AL929276.2
|
|
| chr3_-_8399774 | 0.34 |
ENSDART00000190238
ENSDART00000113140 ENSDART00000132949 |
rbfox3b
|
RNA binding fox-1 homolog 3b |
| chr23_+_43870886 | 0.33 |
ENSDART00000102658
ENSDART00000149088 |
nfxl1
|
nuclear transcription factor, X-box binding-like 1 |
| chr21_+_18877130 | 0.33 |
ENSDART00000136893
|
si:dkey-65l23.2
|
si:dkey-65l23.2 |
| chr14_+_11946395 | 0.33 |
ENSDART00000193290
|
frmpd3
|
FERM and PDZ domain containing 3 |
| chr2_+_21309272 | 0.32 |
ENSDART00000141322
|
zbtb47a
|
zinc finger and BTB domain containing 47a |
| chr7_+_40658003 | 0.32 |
ENSDART00000172435
|
lmbr1
|
limb development membrane protein 1 |
| chr15_+_17441734 | 0.31 |
ENSDART00000153729
|
snx19b
|
sorting nexin 19b |
| chr19_-_21766461 | 0.31 |
ENSDART00000104279
|
znf516
|
zinc finger protein 516 |
| chr10_-_28477023 | 0.31 |
ENSDART00000137008
|
bbx
|
bobby sox homolog (Drosophila) |
| chr15_-_33325019 | 0.30 |
ENSDART00000181092
ENSDART00000168920 |
nbeab
|
neurobeachin b |
| chr13_-_37383625 | 0.30 |
ENSDART00000192888
ENSDART00000100322 ENSDART00000138934 |
kcnh5b
|
potassium voltage-gated channel, subfamily H (eag-related), member 5b |
| chr2_-_17044959 | 0.30 |
ENSDART00000090260
|
clcn2a
|
chloride channel, voltage-sensitive 2a |
| chr7_+_30933867 | 0.29 |
ENSDART00000156325
|
tjp1a
|
tight junction protein 1a |
| chr10_-_15405564 | 0.29 |
ENSDART00000020665
|
sgtb
|
small glutamine-rich tetratricopeptide repeat (TPR)-containing, beta |
| chr25_-_35046114 | 0.28 |
ENSDART00000185267
|
zgc:165555
|
zgc:165555 |
| chr20_+_42978499 | 0.27 |
ENSDART00000138793
|
si:ch211-203k16.3
|
si:ch211-203k16.3 |
| chr5_-_1999417 | 0.27 |
ENSDART00000155437
ENSDART00000145781 |
si:ch211-160e1.5
|
si:ch211-160e1.5 |
| chr14_-_4044545 | 0.26 |
ENSDART00000169527
|
snx25
|
sorting nexin 25 |
| chr2_+_30281444 | 0.26 |
ENSDART00000172756
|
si:dkey-82k12.13
|
si:dkey-82k12.13 |
| chr23_+_25832689 | 0.26 |
ENSDART00000138907
|
hnf4a
|
hepatocyte nuclear factor 4, alpha |
| chr16_+_22587661 | 0.26 |
ENSDART00000129612
ENSDART00000142241 |
she
|
Src homology 2 domain containing E |
| chr23_-_20002459 | 0.25 |
ENSDART00000163396
|
b4galt3
|
UDP-Gal:betaGlcNAc beta 1,4- galactosyltransferase, polypeptide 3 |
| chr16_-_26435431 | 0.25 |
ENSDART00000187526
|
megf8
|
multiple EGF-like-domains 8 |
| chr14_+_36628131 | 0.25 |
ENSDART00000188625
ENSDART00000125345 |
TENM3
|
si:dkey-237h12.3 |
| chr1_+_51391082 | 0.25 |
ENSDART00000063936
|
atg4da
|
autophagy related 4D, cysteine peptidase a |
| chr17_-_45247332 | 0.25 |
ENSDART00000016815
|
ttbk2a
|
tau tubulin kinase 2a |
| chr25_+_31267268 | 0.24 |
ENSDART00000181239
|
tnni2a.3
|
troponin I type 2a (skeletal, fast), tandem duplicate 3 |
| chr16_-_17541890 | 0.23 |
ENSDART00000131328
|
clcn1b
|
chloride channel, voltage-sensitive 1b |
| chr12_-_47793857 | 0.23 |
ENSDART00000161294
|
dydc2
|
DPY30 domain containing 2 |
| chr8_+_9866351 | 0.22 |
ENSDART00000133985
|
kcnd1
|
potassium voltage-gated channel, Shal-related subfamily, member 1 |
| chr17_+_23937262 | 0.22 |
ENSDART00000113276
|
si:ch211-189k9.2
|
si:ch211-189k9.2 |
| chr22_-_23067859 | 0.22 |
ENSDART00000137873
|
ptprc
|
protein tyrosine phosphatase, receptor type, C |
| chr9_+_51147380 | 0.22 |
ENSDART00000003913
|
ifih1
|
interferon induced with helicase C domain 1 |
| chr22_-_38360205 | 0.21 |
ENSDART00000162055
|
mark1
|
MAP/microtubule affinity-regulating kinase 1 |
| chr3_+_32443395 | 0.21 |
ENSDART00000188447
|
prr12b
|
proline rich 12b |
| chr16_+_19732543 | 0.21 |
ENSDART00000149901
ENSDART00000052927 |
twist1b
|
twist family bHLH transcription factor 1b |
| chr25_-_11088839 | 0.20 |
ENSDART00000154748
|
sv2bb
|
synaptic vesicle glycoprotein 2Bb |
| chr12_+_47794089 | 0.19 |
ENSDART00000160726
|
polr3a
|
polymerase (RNA) III (DNA directed) polypeptide A |
| chr18_-_20560007 | 0.19 |
ENSDART00000141367
ENSDART00000090186 |
si:ch211-238n5.4
|
si:ch211-238n5.4 |
| chr23_-_35195908 | 0.19 |
ENSDART00000122429
|
klf15
|
Kruppel-like factor 15 |
| chr18_-_44611252 | 0.18 |
ENSDART00000173095
|
spred3
|
sprouty-related, EVH1 domain containing 3 |
| chr18_+_8937718 | 0.18 |
ENSDART00000138115
|
si:dkey-5i3.1
|
si:dkey-5i3.1 |
| chr7_+_63325819 | 0.18 |
ENSDART00000085612
ENSDART00000161436 |
pcdh7b
|
protocadherin 7b |
| chr4_+_5842433 | 0.18 |
ENSDART00000124085
ENSDART00000179848 |
usp18
|
ubiquitin specific peptidase 18 |
| chr9_+_32930622 | 0.17 |
ENSDART00000100928
|
cnga4
|
cyclic nucleotide gated channel alpha 4 |
| chr5_+_56268436 | 0.17 |
ENSDART00000021159
|
lhx1b
|
LIM homeobox 1b |
| chr20_-_13660600 | 0.17 |
ENSDART00000063826
|
TAGAP
|
si:ch211-122h15.4 |
| chr6_-_17999776 | 0.17 |
ENSDART00000183048
ENSDART00000181577 ENSDART00000170597 |
rptor
|
regulatory associated protein of MTOR, complex 1 |
| chr14_-_41243180 | 0.16 |
ENSDART00000184548
|
drp2
|
dystrophin related protein 2 |
| chr2_+_42871831 | 0.16 |
ENSDART00000171393
|
efr3a
|
EFR3 homolog A (S. cerevisiae) |
| chr12_-_18883079 | 0.16 |
ENSDART00000178619
|
shisa8b
|
shisa family member 8b |
| chr20_-_3403033 | 0.16 |
ENSDART00000092264
|
rev3l
|
REV3-like, polymerase (DNA directed), zeta, catalytic subunit |
| chr25_+_10929790 | 0.16 |
ENSDART00000045182
|
CR339041.2
|
|
| chr3_-_58165254 | 0.15 |
ENSDART00000093031
|
snu13a
|
SNU13 homolog, small nuclear ribonucleoprotein a (U4/U6.U5) |
| chr24_-_15131831 | 0.15 |
ENSDART00000028410
|
cd226
|
CD226 molecule |
| chr6_-_16456093 | 0.15 |
ENSDART00000083305
ENSDART00000181640 |
slc19a2
|
solute carrier family 19 (thiamine transporter), member 2 |
| chr1_-_46663997 | 0.15 |
ENSDART00000134450
|
ebpl
|
emopamil binding protein-like |
| chr20_-_13660767 | 0.13 |
ENSDART00000127654
|
TAGAP
|
si:ch211-122h15.4 |
| chr11_-_22605981 | 0.13 |
ENSDART00000186923
|
myog
|
myogenin |
| chr18_+_13248956 | 0.13 |
ENSDART00000080709
|
plcg2
|
phospholipase C, gamma 2 |
| chr22_-_6144428 | 0.13 |
ENSDART00000106118
|
si:dkey-19a16.4
|
si:dkey-19a16.4 |
| chr16_-_31525290 | 0.12 |
ENSDART00000109996
|
wisp1b
|
WNT1 inducible signaling pathway protein 1b |
| chr3_-_29506960 | 0.12 |
ENSDART00000141720
|
cyth4a
|
cytohesin 4a |
| chr12_+_48204891 | 0.12 |
ENSDART00000190534
ENSDART00000164427 |
ndr2
|
nodal-related 2 |
| chr21_-_43482426 | 0.12 |
ENSDART00000192901
|
ankrd46a
|
ankyrin repeat domain 46a |
| chr24_-_39826865 | 0.11 |
ENSDART00000089232
|
slc12a7b
|
solute carrier family 12 (potassium/chloride transporter), member 7b |
| chr1_+_47391404 | 0.11 |
ENSDART00000113559
|
bcl9
|
B cell CLL/lymphoma 9 |
| chr23_+_16908012 | 0.10 |
ENSDART00000133946
|
si:dkey-147f3.8
|
si:dkey-147f3.8 |
| chr8_-_14484599 | 0.10 |
ENSDART00000057644
|
lhx4
|
LIM homeobox 4 |
| chr16_-_2650341 | 0.10 |
ENSDART00000128169
ENSDART00000155432 ENSDART00000103722 |
lyplal1
|
lysophospholipase-like 1 |
| chr5_-_3118346 | 0.10 |
ENSDART00000167554
|
hsf5
|
heat shock transcription factor family member 5 |
| chr7_+_2228276 | 0.10 |
ENSDART00000064294
|
si:dkey-187j14.4
|
si:dkey-187j14.4 |
| chr2_+_22042745 | 0.09 |
ENSDART00000132039
|
tox
|
thymocyte selection-associated high mobility group box |
| chr15_-_39948635 | 0.09 |
ENSDART00000114836
|
msh5
|
mutS homolog 5 |
| chr5_+_56026031 | 0.09 |
ENSDART00000050970
|
fzd9a
|
frizzled class receptor 9a |
| chr7_+_73649686 | 0.08 |
ENSDART00000185589
|
PCP4L1
|
si:dkey-46i9.1 |
| chr10_-_11840353 | 0.08 |
ENSDART00000127581
|
trim23
|
tripartite motif containing 23 |
| chr12_-_35582683 | 0.08 |
ENSDART00000167933
|
sec24c
|
SEC24 homolog C, COPII coat complex component |
| chr20_+_18992679 | 0.08 |
ENSDART00000147105
ENSDART00000152811 |
tdh
|
L-threonine dehydrogenase |
| chr2_-_45510223 | 0.08 |
ENSDART00000113058
|
gpsm2
|
G protein signaling modulator 2 |
| chr9_+_19421841 | 0.06 |
ENSDART00000159090
ENSDART00000084771 |
pde9a
|
phosphodiesterase 9A |
| chr7_+_33372680 | 0.05 |
ENSDART00000193436
ENSDART00000099988 |
glceb
|
glucuronic acid epimerase b |
| chr21_-_40799404 | 0.05 |
ENSDART00000147405
ENSDART00000157512 |
zgc:162239
|
zgc:162239 |
| chr16_+_11683548 | 0.05 |
ENSDART00000138000
|
si:dkey-250k15.7
|
si:dkey-250k15.7 |
| chr7_-_4461104 | 0.05 |
ENSDART00000023090
ENSDART00000140770 |
slc12a10.1
|
solute carrier family 12 (sodium/potassium/chloride transporters), member 10, tandem duplicate 1 |
| chr5_-_14211487 | 0.04 |
ENSDART00000135731
|
npffr1l2
|
neuropeptide FF receptor 1 like 2 |
| chr10_-_39052264 | 0.04 |
ENSDART00000144036
|
igsf5a
|
immunoglobulin superfamily, member 5a |
| chr8_-_621142 | 0.04 |
ENSDART00000187576
ENSDART00000190316 |
si:ch211-87m7.2
|
si:ch211-87m7.2 |
| chr2_+_44781185 | 0.03 |
ENSDART00000187103
ENSDART00000147292 ENSDART00000098123 |
ece2a
|
endothelin converting enzyme 2a |
| chr19_+_43256986 | 0.02 |
ENSDART00000182336
|
DGKD
|
diacylglycerol kinase delta |
| chr17_-_45378473 | 0.02 |
ENSDART00000132969
|
znf106a
|
zinc finger protein 106a |
| chr6_-_40352215 | 0.02 |
ENSDART00000103992
|
ttll3
|
tubulin tyrosine ligase-like family, member 3 |
| chr19_-_38539670 | 0.01 |
ENSDART00000136775
|
col16a1
|
collagen, type XVI, alpha 1 |
| chr20_+_52546186 | 0.00 |
ENSDART00000110777
ENSDART00000153377 ENSDART00000153013 ENSDART00000042704 |
eef1db
|
eukaryotic translation elongation factor 1 delta b (guanine nucleotide exchange protein) |
| chr20_-_3004126 | 0.00 |
ENSDART00000064390
|
ppil4
|
peptidylprolyl isomerase (cyclophilin)-like 4 |
| chr23_+_25993686 | 0.00 |
ENSDART00000053993
|
wisp2
|
WNT1 inducible signaling pathway protein 2 |
Network of associatons between targets according to the STRING database.
First level regulatory network of onecut3a+onecut3b
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.1 | GO:0031630 | regulation of synaptic vesicle fusion to presynaptic membrane(GO:0031630) |
| 0.3 | 0.9 | GO:0009449 | gamma-aminobutyric acid biosynthetic process(GO:0009449) |
| 0.3 | 1.3 | GO:0046485 | ether lipid metabolic process(GO:0046485) |
| 0.2 | 0.7 | GO:0021961 | posterior commissure morphogenesis(GO:0021961) |
| 0.2 | 1.2 | GO:0097066 | response to thyroid hormone(GO:0097066) |
| 0.2 | 0.8 | GO:0046551 | retinal cone cell fate determination(GO:0042671) eye photoreceptor cell fate commitment(GO:0042706) photoreceptor cell fate determination(GO:0043703) retinal cone cell fate commitment(GO:0046551) photoreceptor cell fate commitment(GO:0046552) camera-type eye photoreceptor cell fate commitment(GO:0060220) |
| 0.2 | 1.7 | GO:0051965 | positive regulation of synapse assembly(GO:0051965) |
| 0.1 | 0.8 | GO:0001561 | fatty acid alpha-oxidation(GO:0001561) |
| 0.1 | 0.5 | GO:0032655 | interleukin-12 production(GO:0032615) regulation of interleukin-12 production(GO:0032655) |
| 0.1 | 0.3 | GO:1903332 | regulation of protein folding(GO:1903332) positive regulation of protein folding(GO:1903334) regulation of chaperone-mediated protein folding(GO:1903644) positive regulation of chaperone-mediated protein folding(GO:1903646) |
| 0.1 | 0.9 | GO:0071300 | cellular response to retinoic acid(GO:0071300) |
| 0.1 | 1.3 | GO:0048899 | anterior lateral line development(GO:0048899) |
| 0.1 | 0.8 | GO:0032024 | positive regulation of insulin secretion(GO:0032024) |
| 0.1 | 0.7 | GO:0001992 | regulation of systemic arterial blood pressure by vasopressin(GO:0001992) |
| 0.1 | 0.5 | GO:0042138 | meiotic DNA double-strand break formation(GO:0042138) |
| 0.1 | 0.5 | GO:0032228 | regulation of synaptic transmission, GABAergic(GO:0032228) |
| 0.1 | 0.5 | GO:2000273 | positive regulation of epidermal growth factor-activated receptor activity(GO:0045741) positive regulation of receptor activity(GO:2000273) |
| 0.1 | 0.7 | GO:0035860 | esophagus smooth muscle contraction(GO:0014846) glial cell-derived neurotrophic factor receptor signaling pathway(GO:0035860) |
| 0.1 | 0.2 | GO:0002833 | response to tumor cell(GO:0002347) natural killer cell cytokine production(GO:0002370) immune response to tumor cell(GO:0002418) natural killer cell mediated cytotoxicity directed against tumor cell target(GO:0002420) natural killer cell mediated immune response to tumor cell(GO:0002423) regulation of natural killer cell cytokine production(GO:0002727) positive regulation of natural killer cell cytokine production(GO:0002729) positive regulation of response to biotic stimulus(GO:0002833) regulation of response to tumor cell(GO:0002834) positive regulation of response to tumor cell(GO:0002836) regulation of immune response to tumor cell(GO:0002837) positive regulation of immune response to tumor cell(GO:0002839) regulation of natural killer cell mediated immune response to tumor cell(GO:0002855) positive regulation of natural killer cell mediated immune response to tumor cell(GO:0002857) regulation of natural killer cell mediated cytotoxicity directed against tumor cell target(GO:0002858) positive regulation of natural killer cell mediated cytotoxicity directed against tumor cell target(GO:0002860) positive regulation of Fc receptor mediated stimulatory signaling pathway(GO:0060369) |
| 0.0 | 0.4 | GO:0006685 | sphingomyelin catabolic process(GO:0006685) |
| 0.0 | 1.0 | GO:0015701 | bicarbonate transport(GO:0015701) |
| 0.0 | 0.3 | GO:0019375 | galactosylceramide biosynthetic process(GO:0006682) galactolipid biosynthetic process(GO:0019375) |
| 0.0 | 1.5 | GO:1903670 | regulation of sprouting angiogenesis(GO:1903670) |
| 0.0 | 0.2 | GO:0032608 | interferon-beta production(GO:0032608) regulation of interferon-beta production(GO:0032648) positive regulation of interferon-beta production(GO:0032728) |
| 0.0 | 0.5 | GO:2000377 | regulation of reactive oxygen species metabolic process(GO:2000377) |
| 0.0 | 0.8 | GO:0043124 | negative regulation of I-kappaB kinase/NF-kappaB signaling(GO:0043124) |
| 0.0 | 0.5 | GO:0086010 | membrane depolarization during action potential(GO:0086010) |
| 0.0 | 0.2 | GO:0010801 | regulation of peptidyl-threonine phosphorylation(GO:0010799) negative regulation of peptidyl-threonine phosphorylation(GO:0010801) |
| 0.0 | 1.4 | GO:0005978 | glycogen biosynthetic process(GO:0005978) glucan biosynthetic process(GO:0009250) |
| 0.0 | 1.3 | GO:0021854 | hypothalamus development(GO:0021854) |
| 0.0 | 0.6 | GO:2001235 | positive regulation of apoptotic signaling pathway(GO:2001235) |
| 0.0 | 0.3 | GO:0036058 | filtration diaphragm assembly(GO:0036058) slit diaphragm assembly(GO:0036060) |
| 0.0 | 0.1 | GO:0048743 | positive regulation of skeletal muscle fiber development(GO:0048743) |
| 0.0 | 0.2 | GO:0006348 | chromatin silencing at telomere(GO:0006348) |
| 0.0 | 0.9 | GO:1903038 | negative regulation of T cell activation(GO:0050868) negative regulation of leukocyte cell-cell adhesion(GO:1903038) |
| 0.0 | 0.8 | GO:0009648 | photoperiodism(GO:0009648) |
| 0.0 | 0.3 | GO:0070365 | hepatocyte differentiation(GO:0070365) |
| 0.0 | 0.2 | GO:0021694 | cerebellar Purkinje cell layer formation(GO:0021694) cerebellar Purkinje cell differentiation(GO:0021702) |
| 0.0 | 0.6 | GO:0007214 | gamma-aminobutyric acid signaling pathway(GO:0007214) |
| 0.0 | 1.3 | GO:0031146 | SCF-dependent proteasomal ubiquitin-dependent protein catabolic process(GO:0031146) |
| 0.0 | 0.1 | GO:0046069 | cGMP catabolic process(GO:0046069) |
| 0.0 | 0.4 | GO:0098703 | calcium ion import across plasma membrane(GO:0098703) calcium ion import into cell(GO:1990035) |
| 0.0 | 0.2 | GO:0048172 | regulation of short-term neuronal synaptic plasticity(GO:0048172) |
| 0.0 | 0.2 | GO:0042276 | error-prone translesion synthesis(GO:0042276) |
| 0.0 | 0.4 | GO:0007097 | nuclear migration(GO:0007097) establishment of nucleus localization(GO:0040023) |
| 0.0 | 0.4 | GO:0010669 | epithelial structure maintenance(GO:0010669) |
| 0.0 | 0.8 | GO:0071391 | cellular response to estrogen stimulus(GO:0071391) |
| 0.0 | 0.3 | GO:0032526 | response to retinoic acid(GO:0032526) |
| 0.0 | 1.0 | GO:0050919 | negative chemotaxis(GO:0050919) |
| 0.0 | 0.1 | GO:0045117 | azole transport(GO:0045117) |
| 0.0 | 0.8 | GO:0046513 | ceramide biosynthetic process(GO:0046513) |
| 0.0 | 0.6 | GO:0071880 | adenylate cyclase-activating adrenergic receptor signaling pathway(GO:0071880) |
| 0.0 | 0.1 | GO:1905066 | regulation of canonical Wnt signaling pathway involved in heart development(GO:1905066) |
| 0.0 | 0.1 | GO:0060573 | ventral spinal cord interneuron specification(GO:0021521) cell fate specification involved in pattern specification(GO:0060573) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.8 | GO:1990513 | CLOCK-BMAL transcription complex(GO:1990513) |
| 0.1 | 1.2 | GO:0045009 | melanosome membrane(GO:0033162) chitosome(GO:0045009) |
| 0.1 | 0.3 | GO:0072380 | TRC complex(GO:0072380) |
| 0.1 | 4.0 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.1 | 0.6 | GO:0097648 | G-protein coupled receptor dimeric complex(GO:0038037) G-protein coupled receptor heterodimeric complex(GO:0038039) G-protein coupled receptor complex(GO:0097648) |
| 0.1 | 0.4 | GO:0032591 | dendritic spine membrane(GO:0032591) |
| 0.0 | 0.6 | GO:0070775 | H3 histone acetyltransferase complex(GO:0070775) MOZ/MORF histone acetyltransferase complex(GO:0070776) |
| 0.0 | 0.2 | GO:0031428 | box C/D snoRNP complex(GO:0031428) |
| 0.0 | 0.4 | GO:0005640 | nuclear outer membrane(GO:0005640) |
| 0.0 | 0.5 | GO:0001518 | voltage-gated sodium channel complex(GO:0001518) sodium channel complex(GO:0034706) |
| 0.0 | 1.5 | GO:0030141 | secretory granule(GO:0030141) |
| 0.0 | 1.3 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.0 | 0.2 | GO:0048188 | Set1C/COMPASS complex(GO:0048188) |
| 0.0 | 0.2 | GO:0042575 | DNA polymerase complex(GO:0042575) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.4 | GO:0003844 | 1,4-alpha-glucan branching enzyme activity(GO:0003844) |
| 0.3 | 0.9 | GO:0004351 | glutamate decarboxylase activity(GO:0004351) |
| 0.3 | 3.9 | GO:0004971 | AMPA glutamate receptor activity(GO:0004971) |
| 0.2 | 0.8 | GO:0070324 | thyroid hormone binding(GO:0070324) |
| 0.1 | 1.3 | GO:0016714 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced pteridine as one donor, and incorporation of one atom of oxygen(GO:0016714) |
| 0.1 | 0.8 | GO:0004396 | glucokinase activity(GO:0004340) hexokinase activity(GO:0004396) glucose binding(GO:0005536) fructokinase activity(GO:0008865) mannokinase activity(GO:0019158) |
| 0.1 | 0.8 | GO:0070888 | E-box binding(GO:0070888) |
| 0.1 | 0.5 | GO:0008332 | low voltage-gated calcium channel activity(GO:0008332) |
| 0.1 | 0.6 | GO:0004965 | G-protein coupled GABA receptor activity(GO:0004965) |
| 0.1 | 0.7 | GO:0016909 | JUN kinase activity(GO:0004705) SAP kinase activity(GO:0016909) |
| 0.1 | 1.0 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 0.1 | 0.7 | GO:0005000 | vasopressin receptor activity(GO:0005000) |
| 0.1 | 1.0 | GO:0048495 | Roundabout binding(GO:0048495) |
| 0.1 | 0.8 | GO:0050291 | sphingosine N-acyltransferase activity(GO:0050291) |
| 0.0 | 0.4 | GO:0008467 | [heparan sulfate]-glucosamine 3-sulfotransferase 1 activity(GO:0008467) |
| 0.0 | 0.3 | GO:0001784 | phosphotyrosine binding(GO:0001784) |
| 0.0 | 0.4 | GO:0004028 | 3-chloroallyl aldehyde dehydrogenase activity(GO:0004028) |
| 0.0 | 0.7 | GO:0008449 | N-acetylglucosamine-6-sulfatase activity(GO:0008449) |
| 0.0 | 0.3 | GO:0044323 | retinoic acid-responsive element binding(GO:0044323) |
| 0.0 | 0.2 | GO:0005250 | A-type (transient outward) potassium channel activity(GO:0005250) |
| 0.0 | 0.4 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.0 | 0.9 | GO:0005388 | calcium-transporting ATPase activity(GO:0005388) |
| 0.0 | 0.5 | GO:0005247 | voltage-gated chloride channel activity(GO:0005247) |
| 0.0 | 1.1 | GO:0005518 | collagen binding(GO:0005518) |
| 0.0 | 0.5 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.0 | 1.3 | GO:0015485 | cholesterol binding(GO:0015485) |
| 0.0 | 0.5 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.0 | 0.1 | GO:0016863 | intramolecular oxidoreductase activity, transposing C=C bonds(GO:0016863) |
| 0.0 | 0.1 | GO:0045118 | azole transporter activity(GO:0045118) azole transmembrane transporter activity(GO:1901474) |
| 0.0 | 0.1 | GO:0070698 | type I activin receptor binding(GO:0070698) |
| 0.0 | 0.1 | GO:0008743 | L-threonine 3-dehydrogenase activity(GO:0008743) |
| 0.0 | 0.4 | GO:0004659 | prenyltransferase activity(GO:0004659) |
| 0.0 | 0.6 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.0 | 0.1 | GO:0047464 | heparosan-N-sulfate-glucuronate 5-epimerase activity(GO:0047464) |
| 0.0 | 0.2 | GO:0043855 | intracellular cyclic nucleotide activated cation channel activity(GO:0005221) intracellular cAMP activated cation channel activity(GO:0005222) intracellular cGMP activated cation channel activity(GO:0005223) cyclic nucleotide-gated ion channel activity(GO:0043855) |
| 0.0 | 0.5 | GO:0019957 | chemokine binding(GO:0019956) C-C chemokine binding(GO:0019957) |
| 0.0 | 0.2 | GO:0031726 | CCR1 chemokine receptor binding(GO:0031726) CCR4 chemokine receptor binding(GO:0031729) |
| 0.0 | 0.8 | GO:0016706 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, 2-oxoglutarate as one donor, and incorporation of one atom each of oxygen into both donors(GO:0016706) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.5 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.0 | 1.7 | PID IL6 7 PATHWAY | IL6-mediated signaling events |
| 0.0 | 0.8 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.0 | 1.1 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 1.1 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.0 | 0.4 | PID PI3K PLC TRK PATHWAY | Trk receptor signaling mediated by PI3K and PLC-gamma |
| 0.0 | 0.3 | SIG BCR SIGNALING PATHWAY | Members of the BCR signaling pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.3 | REACTOME CREATION OF C4 AND C2 ACTIVATORS | Genes involved in Creation of C4 and C2 activators |
| 0.1 | 0.5 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.1 | 1.1 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.0 | 0.8 | REACTOME PEROXISOMAL LIPID METABOLISM | Genes involved in Peroxisomal lipid metabolism |
| 0.0 | 0.4 | REACTOME HS GAG BIOSYNTHESIS | Genes involved in HS-GAG biosynthesis |
| 0.0 | 0.4 | REACTOME SYNTHESIS OF SUBSTRATES IN N GLYCAN BIOSYTHESIS | Genes involved in Synthesis of substrates in N-glycan biosythesis |
| 0.0 | 0.8 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.0 | 0.2 | REACTOME REGULATION OF IFNA SIGNALING | Genes involved in Regulation of IFNA signaling |
| 0.0 | 0.1 | REACTOME PHOSPHORYLATION OF CD3 AND TCR ZETA CHAINS | Genes involved in Phosphorylation of CD3 and TCR zeta chains |
| 0.0 | 0.2 | REACTOME MTORC1 MEDIATED SIGNALLING | Genes involved in mTORC1-mediated signalling |
| 0.0 | 0.4 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.0 | 0.2 | REACTOME TRAF3 DEPENDENT IRF ACTIVATION PATHWAY | Genes involved in TRAF3-dependent IRF activation pathway |
| 0.0 | 0.2 | REACTOME RNA POL III CHAIN ELONGATION | Genes involved in RNA Polymerase III Chain Elongation |