Project
PRJEB1986: zebrafish developmental stages transcriptome
Navigation
Downloads
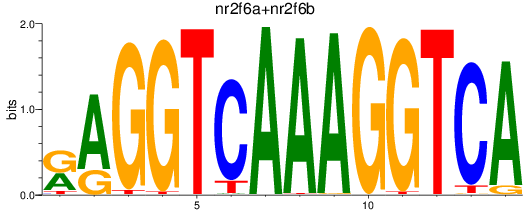
Results for nr2f6a+nr2f6b
Z-value: 0.60
Transcription factors associated with nr2f6a+nr2f6b
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
nr2f6b
|
ENSDARG00000003165 | nuclear receptor subfamily 2, group F, member 6b |
|
nr2f6a
|
ENSDARG00000003607 | nuclear receptor subfamily 2, group F, member 6a |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| nr2f6a | dr11_v1_chr2_+_24374305_24374305 | 0.95 | 2.2e-10 | Click! |
| nr2f6b | dr11_v1_chr11_+_6116503_6116503 | 0.94 | 1.7e-09 | Click! |
Activity profile of nr2f6a+nr2f6b motif
Sorted Z-values of nr2f6a+nr2f6b motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr4_-_16451375 | 4.13 |
ENSDART00000192700
ENSDART00000128835 |
wu:fc23c09
|
wu:fc23c09 |
| chr5_+_43870389 | 2.00 |
ENSDART00000141002
|
zgc:112966
|
zgc:112966 |
| chr4_+_12031958 | 1.95 |
ENSDART00000044154
|
tnnt2c
|
troponin T2c, cardiac |
| chr21_-_44512893 | 1.89 |
ENSDART00000166853
|
zgc:136410
|
zgc:136410 |
| chr23_+_2914577 | 1.85 |
ENSDART00000184897
|
DHX35
|
zgc:158828 |
| chr5_+_22067570 | 1.58 |
ENSDART00000045574
|
shisa2a
|
shisa family member 2a |
| chr13_-_37127970 | 1.37 |
ENSDART00000135510
|
syne2b
|
spectrin repeat containing, nuclear envelope 2b |
| chr22_-_7050 | 1.27 |
ENSDART00000127829
|
atad3
|
ATPase family, AAA domain containing 3 |
| chr23_+_30730121 | 1.25 |
ENSDART00000134141
|
asxl1
|
additional sex combs like transcriptional regulator 1 |
| chr23_+_44634187 | 1.23 |
ENSDART00000143688
|
si:ch73-265d7.2
|
si:ch73-265d7.2 |
| chr7_+_25000060 | 1.19 |
ENSDART00000039265
ENSDART00000141814 |
naa40
|
N(alpha)-acetyltransferase 40, NatD catalytic subunit, homolog (S. cerevisiae) |
| chr25_+_418932 | 1.17 |
ENSDART00000059193
|
prtgb
|
protogenin homolog b (Gallus gallus) |
| chr13_-_39160018 | 1.14 |
ENSDART00000168795
|
col9a1b
|
collagen, type IX, alpha 1b |
| chr12_+_49125510 | 1.12 |
ENSDART00000185804
|
FO704607.1
|
|
| chr13_-_39159810 | 1.12 |
ENSDART00000131508
|
col9a1b
|
collagen, type IX, alpha 1b |
| chr19_-_3056235 | 1.12 |
ENSDART00000137020
|
bop1
|
block of proliferation 1 |
| chr2_+_6067371 | 1.11 |
ENSDART00000053868
ENSDART00000145244 |
aldh9a1b
|
aldehyde dehydrogenase 9 family, member A1b |
| chr9_-_25255490 | 1.08 |
ENSDART00000141502
|
htr2aa
|
5-hydroxytryptamine (serotonin) receptor 2A, genome duplicate a |
| chr12_+_26670778 | 1.07 |
ENSDART00000144355
|
arhgap12b
|
Rho GTPase activating protein 12b |
| chr9_+_32859967 | 1.06 |
ENSDART00000168992
|
si:dkey-145p14.5
|
si:dkey-145p14.5 |
| chr25_+_18587338 | 1.03 |
ENSDART00000180287
|
met
|
MET proto-oncogene, receptor tyrosine kinase |
| chr7_+_38267136 | 1.00 |
ENSDART00000173613
|
gpatch1
|
G patch domain containing 1 |
| chr6_+_40775800 | 0.95 |
ENSDART00000085090
|
si:ch211-157b11.8
|
si:ch211-157b11.8 |
| chr6_+_1787160 | 0.94 |
ENSDART00000113505
|
myl9b
|
myosin, light chain 9b, regulatory |
| chr3_-_34561624 | 0.88 |
ENSDART00000129313
|
sept9a
|
septin 9a |
| chr16_+_54875530 | 0.88 |
ENSDART00000149795
|
nr0b2a
|
nuclear receptor subfamily 0, group B, member 2a |
| chr9_+_32860345 | 0.85 |
ENSDART00000121751
|
si:dkey-145p14.5
|
si:dkey-145p14.5 |
| chr24_-_1021318 | 0.83 |
ENSDART00000181403
|
ralaa
|
v-ral simian leukemia viral oncogene homolog Aa (ras related) |
| chr1_-_58868306 | 0.83 |
ENSDART00000166615
|
dnm2b
|
dynamin 2b |
| chr24_+_81527 | 0.75 |
ENSDART00000192139
|
reck
|
reversion-inducing-cysteine-rich protein with kazal motifs |
| chr3_+_25999477 | 0.72 |
ENSDART00000024316
|
mcm5
|
minichromosome maintenance complex component 5 |
| chr5_+_49744713 | 0.71 |
ENSDART00000133384
|
nr2f1a
|
nuclear receptor subfamily 2, group F, member 1a |
| chr12_+_48681601 | 0.69 |
ENSDART00000187831
|
uros
|
uroporphyrinogen III synthase |
| chr8_+_39802506 | 0.68 |
ENSDART00000018862
|
hnf1a
|
HNF1 homeobox a |
| chr5_-_26330313 | 0.67 |
ENSDART00000148656
|
arvcfb
|
ARVCF, delta catenin family member b |
| chr22_-_3595439 | 0.67 |
ENSDART00000083308
|
ptprsa
|
protein tyrosine phosphatase, receptor type, s, a |
| chr1_-_51720633 | 0.65 |
ENSDART00000045894
|
rnaseh2a
|
ribonuclease H2, subunit A |
| chr11_-_6974022 | 0.65 |
ENSDART00000172851
|
COMP
|
si:ch211-43f4.1 |
| chr5_-_3839285 | 0.64 |
ENSDART00000122292
|
mlxipl
|
MLX interacting protein like |
| chr22_-_15587360 | 0.64 |
ENSDART00000142717
ENSDART00000138978 |
tpm4a
|
tropomyosin 4a |
| chr9_-_28990649 | 0.62 |
ENSDART00000078823
|
ptpn4a
|
protein tyrosine phosphatase, non-receptor type 4a |
| chr1_+_12195700 | 0.57 |
ENSDART00000040307
|
tdrd7a
|
tudor domain containing 7 a |
| chr24_-_21172122 | 0.53 |
ENSDART00000154259
|
atp6v1ab
|
ATPase H+ transporting V1 subunit Ab |
| chr19_+_2631565 | 0.50 |
ENSDART00000171487
|
fam126a
|
family with sequence similarity 126, member A |
| chr4_-_75172216 | 0.49 |
ENSDART00000127522
|
naa40
|
N(alpha)-acetyltransferase 40, NatD catalytic subunit, homolog (S. cerevisiae) |
| chr10_+_20128267 | 0.46 |
ENSDART00000064615
|
dmtn
|
dematin actin binding protein |
| chr14_+_30413758 | 0.46 |
ENSDART00000092953
|
cnot7
|
CCR4-NOT transcription complex, subunit 7 |
| chr9_+_54644626 | 0.45 |
ENSDART00000190609
|
egfl6
|
EGF-like-domain, multiple 6 |
| chr1_+_23783349 | 0.43 |
ENSDART00000007531
|
slit2
|
slit homolog 2 (Drosophila) |
| chr18_-_18850302 | 0.41 |
ENSDART00000131965
ENSDART00000167624 |
tgm2l
|
transglutaminase 2, like |
| chr19_+_2877079 | 0.41 |
ENSDART00000160847
|
hgh1
|
HGH1 homolog (S. cerevisiae) |
| chr20_+_46286082 | 0.41 |
ENSDART00000073519
|
taar14f
|
trace amine associated receptor 14f |
| chr8_+_16676894 | 0.41 |
ENSDART00000076586
|
si:ch211-198n5.11
|
si:ch211-198n5.11 |
| chr2_-_56649883 | 0.38 |
ENSDART00000191786
|
gpx4b
|
glutathione peroxidase 4b |
| chr18_-_43866526 | 0.38 |
ENSDART00000111309
|
treh
|
trehalase (brush-border membrane glycoprotein) |
| chr24_+_39034090 | 0.36 |
ENSDART00000185763
|
capn15
|
calpain 15 |
| chr25_+_15647993 | 0.36 |
ENSDART00000186578
ENSDART00000031828 |
spon1b
|
spondin 1b |
| chr13_+_45980163 | 0.35 |
ENSDART00000074547
ENSDART00000005195 |
bsdc1
|
BSD domain containing 1 |
| chr16_-_24605969 | 0.35 |
ENSDART00000163305
ENSDART00000167121 |
fxyd6l
|
FXYD domain containing ion transport regulator 6 like |
| chr17_-_25303486 | 0.35 |
ENSDART00000162235
|
ppie
|
peptidylprolyl isomerase E (cyclophilin E) |
| chr10_+_29852677 | 0.33 |
ENSDART00000141549
|
hspa8
|
heat shock protein 8 |
| chr14_+_30413312 | 0.31 |
ENSDART00000186864
|
cnot7
|
CCR4-NOT transcription complex, subunit 7 |
| chr17_+_23926796 | 0.27 |
ENSDART00000021177
|
pex13
|
peroxisomal biogenesis factor 13 |
| chr21_-_1640547 | 0.25 |
ENSDART00000151041
|
zgc:152948
|
zgc:152948 |
| chr16_-_50229193 | 0.23 |
ENSDART00000161782
ENSDART00000010081 |
etfb
|
electron-transfer-flavoprotein, beta polypeptide |
| chr3_+_39566999 | 0.23 |
ENSDART00000146867
|
aldoaa
|
aldolase a, fructose-bisphosphate, a |
| chr25_+_22017182 | 0.22 |
ENSDART00000156517
|
si:dkey-217l24.1
|
si:dkey-217l24.1 |
| chr21_+_1119046 | 0.22 |
ENSDART00000184678
|
CABZ01088049.1
|
|
| chr11_-_44999858 | 0.20 |
ENSDART00000167759
ENSDART00000126845 |
ldb1b
|
LIM-domain binding 1b |
| chr3_+_1179601 | 0.19 |
ENSDART00000173378
|
triobpb
|
TRIO and F-actin binding protein b |
| chr25_+_20119466 | 0.19 |
ENSDART00000104304
|
bpgm
|
2,3-bisphosphoglycerate mutase |
| chr17_+_33418475 | 0.17 |
ENSDART00000169145
|
snap23.1
|
synaptosomal-associated protein 23.1 |
| chr23_-_45319592 | 0.17 |
ENSDART00000189861
|
ccdc171
|
coiled-coil domain containing 171 |
| chr7_+_48297842 | 0.17 |
ENSDART00000052123
|
slc25a44b
|
solute carrier family 25, member 44 b |
| chr2_-_22659450 | 0.17 |
ENSDART00000115025
|
thap4
|
THAP domain containing 4 |
| chr19_+_2275019 | 0.15 |
ENSDART00000136138
|
itgb8
|
integrin, beta 8 |
| chr20_+_2281933 | 0.14 |
ENSDART00000137579
|
si:ch73-18b11.2
|
si:ch73-18b11.2 |
| chr12_-_34580988 | 0.14 |
ENSDART00000152926
|
bahcc1b
|
BAH domain and coiled-coil containing 1b |
| chr10_+_11265387 | 0.13 |
ENSDART00000038888
|
hsdl2
|
hydroxysteroid dehydrogenase like 2 |
| chr8_+_999421 | 0.11 |
ENSDART00000149528
|
fabp1b.1
|
fatty acid binding protein 1b, tandem duplicate 1 |
| chr9_-_37613792 | 0.09 |
ENSDART00000138345
|
sema5ba
|
sema domain, seven thrombospondin repeats (type 1 and type 1-like), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 5Ba |
| chr24_-_7587401 | 0.09 |
ENSDART00000093163
|
galnt11
|
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 11 (GalNAc-T11) |
| chr2_-_41571454 | 0.09 |
ENSDART00000022643
|
znf622
|
zinc finger protein 622 |
| chr23_+_44633858 | 0.07 |
ENSDART00000180728
|
si:ch73-265d7.2
|
si:ch73-265d7.2 |
| chr22_-_11833317 | 0.07 |
ENSDART00000125423
ENSDART00000000192 |
ptpn4b
|
protein tyrosine phosphatase, non-receptor type 4b |
| chr8_-_8698607 | 0.05 |
ENSDART00000046712
|
zgc:86609
|
zgc:86609 |
| chr3_-_41292275 | 0.04 |
ENSDART00000144088
|
sdk1a
|
sidekick cell adhesion molecule 1a |
| chr1_+_58353661 | 0.02 |
ENSDART00000140074
|
si:dkey-222h21.2
|
si:dkey-222h21.2 |
| chr1_+_58119117 | 0.01 |
ENSDART00000146788
|
si:ch211-15j1.5
|
si:ch211-15j1.5 |
| chr8_+_8699085 | 0.01 |
ENSDART00000021209
|
uxt
|
ubiquitously-expressed, prefoldin-like chaperone |
Network of associatons between targets according to the STRING database.
First level regulatory network of nr2f6a+nr2f6b
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.8 | GO:1905048 | regulation of metallopeptidase activity(GO:1905048) |
| 0.1 | 0.4 | GO:0007414 | axonal defasciculation(GO:0007414) |
| 0.1 | 0.4 | GO:0005991 | trehalose metabolic process(GO:0005991) |
| 0.1 | 1.7 | GO:0006474 | N-terminal protein amino acid acetylation(GO:0006474) |
| 0.1 | 0.7 | GO:0043137 | DNA replication, removal of RNA primer(GO:0043137) |
| 0.1 | 1.0 | GO:0021707 | cerebellar granular layer development(GO:0021681) cerebellar granular layer morphogenesis(GO:0021683) cerebellar granular layer formation(GO:0021684) cerebellar granule cell differentiation(GO:0021707) |
| 0.1 | 0.7 | GO:0055011 | atrial cardiac muscle cell differentiation(GO:0055011) |
| 0.1 | 0.5 | GO:0045329 | carnitine biosynthetic process(GO:0045329) |
| 0.1 | 0.8 | GO:0043928 | exonucleolytic nuclear-transcribed mRNA catabolic process involved in deadenylation-dependent decay(GO:0043928) |
| 0.1 | 0.7 | GO:1902299 | pre-replicative complex assembly involved in nuclear cell cycle DNA replication(GO:0006267) pre-replicative complex assembly(GO:0036388) pre-replicative complex assembly involved in cell cycle DNA replication(GO:1902299) |
| 0.1 | 1.1 | GO:0000463 | maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000463) |
| 0.1 | 0.4 | GO:0006552 | leucine catabolic process(GO:0006552) |
| 0.1 | 0.3 | GO:0048260 | positive regulation of receptor-mediated endocytosis(GO:0048260) protein localization to early endosome(GO:1902946) |
| 0.1 | 1.3 | GO:0007168 | receptor guanylyl cyclase signaling pathway(GO:0007168) |
| 0.1 | 0.6 | GO:0030719 | P granule organization(GO:0030719) |
| 0.0 | 0.7 | GO:0099560 | synaptic membrane adhesion(GO:0099560) |
| 0.0 | 0.8 | GO:0098884 | postsynaptic neurotransmitter receptor internalization(GO:0098884) |
| 0.0 | 0.7 | GO:0006782 | protoporphyrinogen IX biosynthetic process(GO:0006782) protoporphyrinogen IX metabolic process(GO:0046501) |
| 0.0 | 0.9 | GO:0043433 | negative regulation of sequence-specific DNA binding transcription factor activity(GO:0043433) |
| 0.0 | 2.6 | GO:0060048 | cardiac muscle contraction(GO:0060048) |
| 0.0 | 1.1 | GO:0051209 | release of sequestered calcium ion into cytosol(GO:0051209) |
| 0.0 | 0.6 | GO:0010811 | positive regulation of cell-substrate adhesion(GO:0010811) |
| 0.0 | 0.3 | GO:0022615 | protein import into peroxisome matrix, docking(GO:0016560) protein to membrane docking(GO:0022615) |
| 0.0 | 0.7 | GO:0003323 | type B pancreatic cell development(GO:0003323) |
| 0.0 | 0.5 | GO:0030032 | lamellipodium assembly(GO:0030032) |
| 0.0 | 0.7 | GO:0016339 | calcium-dependent cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0016339) |
| 0.0 | 0.4 | GO:0018149 | peptide cross-linking(GO:0018149) |
| 0.0 | 0.3 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.0 | 0.2 | GO:0001966 | thigmotaxis(GO:0001966) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.1 | GO:0070545 | PeBoW complex(GO:0070545) |
| 0.2 | 0.8 | GO:1990909 | Wnt signalosome(GO:1990909) |
| 0.2 | 1.2 | GO:0035517 | PR-DUB complex(GO:0035517) |
| 0.2 | 0.7 | GO:0032299 | ribonuclease H2 complex(GO:0032299) |
| 0.1 | 0.3 | GO:1990429 | Pex17p-Pex14p docking complex(GO:1990415) peroxisomal importomer complex(GO:1990429) |
| 0.1 | 0.8 | GO:0030015 | CCR4-NOT core complex(GO:0030015) |
| 0.0 | 1.0 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.0 | 2.0 | GO:0005861 | troponin complex(GO:0005861) |
| 0.0 | 0.8 | GO:0098844 | postsynaptic endocytic zone membrane(GO:0098844) |
| 0.0 | 0.7 | GO:0042555 | MCM complex(GO:0042555) |
| 0.0 | 0.5 | GO:0033180 | proton-transporting V-type ATPase, V1 domain(GO:0033180) |
| 0.0 | 0.9 | GO:0032156 | septin ring(GO:0005940) septin complex(GO:0031105) septin cytoskeleton(GO:0032156) |
| 0.0 | 0.6 | GO:0043186 | P granule(GO:0043186) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.7 | GO:0043998 | H2A histone acetyltransferase activity(GO:0043998) |
| 0.4 | 1.1 | GO:0043878 | glyceraldehyde-3-phosphate dehydrogenase (NAD+) (non-phosphorylating) activity(GO:0043878) |
| 0.2 | 1.2 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.2 | 2.0 | GO:0030172 | troponin C binding(GO:0030172) troponin I binding(GO:0031013) |
| 0.1 | 0.4 | GO:0004485 | methylcrotonoyl-CoA carboxylase activity(GO:0004485) |
| 0.1 | 0.4 | GO:0015927 | alpha,alpha-trehalase activity(GO:0004555) trehalase activity(GO:0015927) |
| 0.1 | 0.7 | GO:0004523 | RNA-DNA hybrid ribonuclease activity(GO:0004523) |
| 0.1 | 0.4 | GO:0047066 | phospholipid-hydroperoxide glutathione peroxidase activity(GO:0047066) |
| 0.1 | 0.7 | GO:0043142 | single-stranded DNA-dependent ATP-dependent DNA helicase activity(GO:0017116) single-stranded DNA-dependent ATPase activity(GO:0043142) |
| 0.1 | 0.8 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.1 | 0.8 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.0 | 0.5 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 0.0 | 0.2 | GO:0004619 | bisphosphoglycerate mutase activity(GO:0004082) phosphoglycerate mutase activity(GO:0004619) |
| 0.0 | 1.0 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.0 | 0.2 | GO:0004332 | fructose-bisphosphate aldolase activity(GO:0004332) |
| 0.0 | 0.2 | GO:0030274 | LIM domain binding(GO:0030274) |
| 0.0 | 0.4 | GO:0048495 | Roundabout binding(GO:0048495) |
| 0.0 | 0.8 | GO:0019003 | GDP binding(GO:0019003) |
| 0.0 | 0.3 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.0 | 0.3 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 0.0 | 0.4 | GO:0003810 | protein-glutamine gamma-glutamyltransferase activity(GO:0003810) |
| 0.0 | 1.1 | GO:0004993 | G-protein coupled serotonin receptor activity(GO:0004993) |
| 0.0 | 1.1 | GO:0043021 | ribonucleoprotein complex binding(GO:0043021) |
| 0.0 | 0.3 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.0 | 0.4 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.0 | 0.6 | GO:0005178 | integrin binding(GO:0005178) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 1.0 | PID ARF6 PATHWAY | Arf6 signaling events |
| 0.0 | 0.4 | PID GLYPICAN 1PATHWAY | Glypican 1 network |
| 0.0 | 0.7 | PID PS1 PATHWAY | Presenilin action in Notch and Wnt signaling |
| 0.0 | 0.2 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.4 | REACTOME DIGESTION OF DIETARY CARBOHYDRATE | Genes involved in Digestion of dietary carbohydrate |
| 0.1 | 0.7 | REACTOME UNWINDING OF DNA | Genes involved in Unwinding of DNA |
| 0.0 | 0.7 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.0 | 0.8 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.0 | 1.0 | REACTOME SEMA4D IN SEMAPHORIN SIGNALING | Genes involved in Sema4D in semaphorin signaling |
| 0.0 | 0.7 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.0 | 0.4 | REACTOME ACTIVATION OF RAC | Genes involved in Activation of Rac |
| 0.0 | 0.3 | REACTOME GABA SYNTHESIS RELEASE REUPTAKE AND DEGRADATION | Genes involved in GABA synthesis, release, reuptake and degradation |
| 0.0 | 0.6 | REACTOME INTEGRATION OF ENERGY METABOLISM | Genes involved in Integration of energy metabolism |