Project
PRJEB1986: zebrafish developmental stages transcriptome
Navigation
Downloads
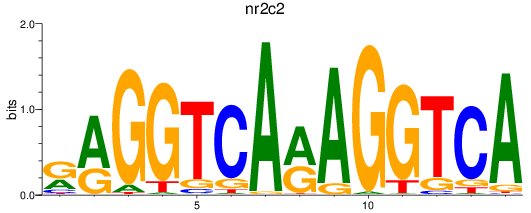
Results for nr2c2
Z-value: 0.95
Transcription factors associated with nr2c2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
nr2c2
|
ENSDARG00000042477 | nuclear receptor subfamily 2, group C, member 2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| nr2c2 | dr11_v1_chr8_+_53120278_53120278 | -0.60 | 6.3e-03 | Click! |
Activity profile of nr2c2 motif
Sorted Z-values of nr2c2 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr23_-_45705525 | 5.99 |
ENSDART00000148959
|
ednrab
|
endothelin receptor type Ab |
| chr4_-_16451375 | 5.65 |
ENSDART00000192700
ENSDART00000128835 |
wu:fc23c09
|
wu:fc23c09 |
| chr21_-_44512893 | 4.27 |
ENSDART00000166853
|
zgc:136410
|
zgc:136410 |
| chr16_-_5115993 | 3.58 |
ENSDART00000138654
|
ttk
|
ttk protein kinase |
| chr24_-_1021318 | 3.26 |
ENSDART00000181403
|
ralaa
|
v-ral simian leukemia viral oncogene homolog Aa (ras related) |
| chr22_-_7050 | 3.11 |
ENSDART00000127829
|
atad3
|
ATPase family, AAA domain containing 3 |
| chr4_+_12031958 | 2.99 |
ENSDART00000044154
|
tnnt2c
|
troponin T2c, cardiac |
| chr14_+_989733 | 2.99 |
ENSDART00000161487
ENSDART00000127317 |
si:ch73-308l14.2
|
si:ch73-308l14.2 |
| chr19_-_8534985 | 2.94 |
ENSDART00000191064
|
dpm3
|
dolichyl-phosphate mannosyltransferase polypeptide 3 |
| chr8_+_15251448 | 2.66 |
ENSDART00000063717
|
zgc:171480
|
zgc:171480 |
| chr19_-_3056235 | 2.66 |
ENSDART00000137020
|
bop1
|
block of proliferation 1 |
| chr3_-_31804481 | 2.43 |
ENSDART00000028270
|
gfap
|
glial fibrillary acidic protein |
| chr13_-_37127970 | 2.42 |
ENSDART00000135510
|
syne2b
|
spectrin repeat containing, nuclear envelope 2b |
| chr22_-_17595310 | 2.37 |
ENSDART00000099056
|
gpx4a
|
glutathione peroxidase 4a |
| chr23_-_10696626 | 2.30 |
ENSDART00000177571
|
foxp1a
|
forkhead box P1a |
| chr1_-_44933094 | 2.27 |
ENSDART00000147527
|
si:dkey-9i23.14
|
si:dkey-9i23.14 |
| chr5_+_43870389 | 2.10 |
ENSDART00000141002
|
zgc:112966
|
zgc:112966 |
| chr22_-_15587360 | 2.02 |
ENSDART00000142717
ENSDART00000138978 |
tpm4a
|
tropomyosin 4a |
| chr1_+_15062 | 2.01 |
ENSDART00000169005
|
cep97
|
centrosomal protein 97 |
| chr3_-_25119839 | 1.99 |
ENSDART00000154724
|
chadla
|
chondroadherin-like a |
| chr25_+_27410352 | 1.98 |
ENSDART00000154362
|
pot1
|
protection of telomeres 1 homolog |
| chr18_+_1615 | 1.95 |
ENSDART00000082450
|
homer2
|
homer scaffolding protein 2 |
| chr23_+_44634187 | 1.93 |
ENSDART00000143688
|
si:ch73-265d7.2
|
si:ch73-265d7.2 |
| chr21_-_308852 | 1.89 |
ENSDART00000151613
|
lhfpl2a
|
LHFPL tetraspan subfamily member 2a |
| chr6_-_11523987 | 1.85 |
ENSDART00000189363
|
gulp1b
|
GULP, engulfment adaptor PTB domain containing 1b |
| chr14_+_97017 | 1.79 |
ENSDART00000159300
ENSDART00000169523 |
mcm7
|
minichromosome maintenance complex component 7 |
| chr23_+_2914577 | 1.78 |
ENSDART00000184897
|
DHX35
|
zgc:158828 |
| chr2_-_42960353 | 1.78 |
ENSDART00000098303
|
oc90
|
otoconin 90 |
| chr9_-_33121535 | 1.77 |
ENSDART00000166371
ENSDART00000138052 |
zgc:172014
|
zgc:172014 |
| chr9_-_32300783 | 1.76 |
ENSDART00000078596
|
hspd1
|
heat shock 60 protein 1 |
| chr20_+_36806398 | 1.74 |
ENSDART00000153317
|
abracl
|
ABRA C-terminal like |
| chr19_-_27564458 | 1.73 |
ENSDART00000123155
|
si:dkeyp-46h3.6
|
si:dkeyp-46h3.6 |
| chr7_+_25000060 | 1.72 |
ENSDART00000039265
ENSDART00000141814 |
naa40
|
N(alpha)-acetyltransferase 40, NatD catalytic subunit, homolog (S. cerevisiae) |
| chr9_-_32300611 | 1.69 |
ENSDART00000127938
|
hspd1
|
heat shock 60 protein 1 |
| chr16_+_54875530 | 1.68 |
ENSDART00000149795
|
nr0b2a
|
nuclear receptor subfamily 0, group B, member 2a |
| chr21_-_22317920 | 1.67 |
ENSDART00000191083
ENSDART00000108701 |
gdpd4b
|
glycerophosphodiester phosphodiesterase domain containing 4b |
| chr3_+_39566999 | 1.64 |
ENSDART00000146867
|
aldoaa
|
aldolase a, fructose-bisphosphate, a |
| chr18_+_30441740 | 1.64 |
ENSDART00000189074
|
gse1
|
Gse1 coiled-coil protein |
| chr2_+_6067371 | 1.63 |
ENSDART00000053868
ENSDART00000145244 |
aldh9a1b
|
aldehyde dehydrogenase 9 family, member A1b |
| chr3_+_301479 | 1.62 |
ENSDART00000165169
|
CABZ01075611.1
|
|
| chr9_+_22782027 | 1.58 |
ENSDART00000090816
|
rif1
|
replication timing regulatory factor 1 |
| chr25_+_418932 | 1.57 |
ENSDART00000059193
|
prtgb
|
protogenin homolog b (Gallus gallus) |
| chr23_-_16737161 | 1.55 |
ENSDART00000132573
|
si:ch211-224l10.4
|
si:ch211-224l10.4 |
| chr9_+_33216945 | 1.53 |
ENSDART00000134029
|
si:ch211-125e6.12
|
si:ch211-125e6.12 |
| chr25_+_18587338 | 1.50 |
ENSDART00000180287
|
met
|
MET proto-oncogene, receptor tyrosine kinase |
| chr3_-_34561624 | 1.46 |
ENSDART00000129313
|
sept9a
|
septin 9a |
| chr3_+_34121156 | 1.44 |
ENSDART00000174929
|
aldh3b1
|
aldehyde dehydrogenase 3 family, member B1 |
| chr20_+_26943072 | 1.43 |
ENSDART00000153215
|
cdca4
|
cell division cycle associated 4 |
| chr8_+_39802506 | 1.43 |
ENSDART00000018862
|
hnf1a
|
HNF1 homeobox a |
| chr23_+_30730121 | 1.40 |
ENSDART00000134141
|
asxl1
|
additional sex combs like transcriptional regulator 1 |
| chr14_+_22498757 | 1.39 |
ENSDART00000021657
|
smyd5
|
SMYD family member 5 |
| chr3_-_36419641 | 1.37 |
ENSDART00000173545
|
cog1
|
component of oligomeric golgi complex 1 |
| chr1_-_59232267 | 1.37 |
ENSDART00000169658
ENSDART00000163257 |
akap8l
|
A kinase (PRKA) anchor protein 8-like |
| chr8_+_36570508 | 1.35 |
ENSDART00000145868
|
pold2
|
polymerase (DNA directed), delta 2, regulatory subunit |
| chr18_-_43866526 | 1.35 |
ENSDART00000111309
|
treh
|
trehalase (brush-border membrane glycoprotein) |
| chr7_+_22718251 | 1.32 |
ENSDART00000027718
ENSDART00000143341 |
fxr2
|
fragile X mental retardation, autosomal homolog 2 |
| chr20_-_25877793 | 1.32 |
ENSDART00000177333
ENSDART00000184619 |
exoc1
|
exocyst complex component 1 |
| chr20_-_45807982 | 1.32 |
ENSDART00000074546
|
fermt1
|
fermitin family member 1 |
| chr1_-_51720633 | 1.31 |
ENSDART00000045894
|
rnaseh2a
|
ribonuclease H2, subunit A |
| chr7_+_19374683 | 1.29 |
ENSDART00000162700
|
snrpf
|
small nuclear ribonucleoprotein polypeptide F |
| chr4_+_47656992 | 1.29 |
ENSDART00000161148
|
znf1040
|
zinc finger protein 1040 |
| chr6_+_40775800 | 1.27 |
ENSDART00000085090
|
si:ch211-157b11.8
|
si:ch211-157b11.8 |
| chr11_+_6116503 | 1.27 |
ENSDART00000176170
|
nr2f6b
|
nuclear receptor subfamily 2, group F, member 6b |
| chr19_+_20274944 | 1.27 |
ENSDART00000151237
|
oxnad1
|
oxidoreductase NAD-binding domain containing 1 |
| chr4_-_25858620 | 1.27 |
ENSDART00000128968
|
nr2c1
|
nuclear receptor subfamily 2, group C, member 1 |
| chr5_+_35786141 | 1.27 |
ENSDART00000022043
ENSDART00000127383 |
stard8
|
StAR-related lipid transfer (START) domain containing 8 |
| chr13_-_39159810 | 1.27 |
ENSDART00000131508
|
col9a1b
|
collagen, type IX, alpha 1b |
| chr15_-_35246742 | 1.26 |
ENSDART00000131479
|
mff
|
mitochondrial fission factor |
| chr18_-_127558 | 1.26 |
ENSDART00000149556
|
trpm7
|
transient receptor potential cation channel, subfamily M, member 7 |
| chr12_+_19138452 | 1.23 |
ENSDART00000141346
ENSDART00000066397 |
phf5a
|
PHD finger protein 5A |
| chr9_-_32684008 | 1.23 |
ENSDART00000041751
|
ercc1
|
excision repair cross-complementation group 1 |
| chr12_+_48681601 | 1.22 |
ENSDART00000187831
|
uros
|
uroporphyrinogen III synthase |
| chr18_-_48508585 | 1.20 |
ENSDART00000133364
|
kcnj1a.4
|
potassium inwardly-rectifying channel, subfamily J, member 1a, tandem duplicate 4 |
| chr12_+_27024676 | 1.19 |
ENSDART00000153104
|
msl1b
|
male-specific lethal 1 homolog b (Drosophila) |
| chr17_+_53424415 | 1.19 |
ENSDART00000157022
|
slc9a1b
|
solute carrier family 9 member A1b |
| chr8_+_23802384 | 1.18 |
ENSDART00000137820
|
si:ch211-163l21.8
|
si:ch211-163l21.8 |
| chr10_+_31244619 | 1.18 |
ENSDART00000145562
ENSDART00000184412 |
robo4
|
roundabout, axon guidance receptor, homolog 4 (Drosophila) |
| chr6_+_1787160 | 1.17 |
ENSDART00000113505
|
myl9b
|
myosin, light chain 9b, regulatory |
| chr9_+_32860345 | 1.16 |
ENSDART00000121751
|
si:dkey-145p14.5
|
si:dkey-145p14.5 |
| chr6_+_52931841 | 1.14 |
ENSDART00000174358
|
si:dkeyp-3f10.12
|
si:dkeyp-3f10.12 |
| chr2_-_56649883 | 1.13 |
ENSDART00000191786
|
gpx4b
|
glutathione peroxidase 4b |
| chr18_-_19819812 | 1.13 |
ENSDART00000060344
|
aagab
|
alpha and gamma adaptin binding protein |
| chr9_+_32978302 | 1.13 |
ENSDART00000007630
|
nhlh2
|
nescient helix loop helix 2 |
| chr3_+_25999477 | 1.12 |
ENSDART00000024316
|
mcm5
|
minichromosome maintenance complex component 5 |
| chr10_+_43994471 | 1.11 |
ENSDART00000138242
ENSDART00000186359 |
cldn5b
|
claudin 5b |
| chr5_-_3839285 | 1.10 |
ENSDART00000122292
|
mlxipl
|
MLX interacting protein like |
| chr21_+_8198652 | 1.09 |
ENSDART00000011096
|
nr6a1b
|
nuclear receptor subfamily 6, group A, member 1b |
| chr17_-_25303486 | 1.08 |
ENSDART00000162235
|
ppie
|
peptidylprolyl isomerase E (cyclophilin E) |
| chr16_-_39131666 | 1.08 |
ENSDART00000075517
|
gdf6a
|
growth differentiation factor 6a |
| chr23_+_30736895 | 1.08 |
ENSDART00000042944
|
asxl1
|
additional sex combs like transcriptional regulator 1 |
| chr2_-_25143373 | 1.07 |
ENSDART00000160108
|
nceh1a
|
neutral cholesterol ester hydrolase 1a |
| chr13_-_39160018 | 1.06 |
ENSDART00000168795
|
col9a1b
|
collagen, type IX, alpha 1b |
| chr2_-_7666021 | 1.06 |
ENSDART00000180007
|
CABZ01021592.1
|
|
| chr20_-_34164278 | 1.04 |
ENSDART00000153128
|
hmcn1
|
hemicentin 1 |
| chr5_-_32890807 | 1.04 |
ENSDART00000007512
|
pole3
|
polymerase (DNA directed), epsilon 3 (p17 subunit) |
| chr17_-_22137199 | 1.04 |
ENSDART00000089689
|
mrps5
|
mitochondrial ribosomal protein S5 |
| chr15_+_8394819 | 1.03 |
ENSDART00000147086
|
tmem160
|
transmembrane protein 160 |
| chr21_-_1640547 | 1.01 |
ENSDART00000151041
|
zgc:152948
|
zgc:152948 |
| chr18_-_34549721 | 1.01 |
ENSDART00000137101
ENSDART00000021880 |
ssr3
|
signal sequence receptor, gamma |
| chr18_-_48517040 | 1.00 |
ENSDART00000143645
|
kcnj1a.3
|
potassium inwardly-rectifying channel, subfamily J, member 1a, tandem duplicate 3 |
| chr18_-_18850302 | 1.00 |
ENSDART00000131965
ENSDART00000167624 |
tgm2l
|
transglutaminase 2, like |
| chr3_+_40289418 | 0.98 |
ENSDART00000017304
|
cpsf4
|
cleavage and polyadenylation specific factor 4 |
| chr24_+_81527 | 0.98 |
ENSDART00000192139
|
reck
|
reversion-inducing-cysteine-rich protein with kazal motifs |
| chr3_-_18792492 | 0.98 |
ENSDART00000134208
ENSDART00000034373 |
hagh
|
hydroxyacylglutathione hydrolase |
| chr6_-_14004772 | 0.98 |
ENSDART00000185629
|
zgc:92027
|
zgc:92027 |
| chr5_-_26330313 | 0.97 |
ENSDART00000148656
|
arvcfb
|
ARVCF, delta catenin family member b |
| chr22_-_14262115 | 0.97 |
ENSDART00000168264
|
si:ch211-246m6.5
|
si:ch211-246m6.5 |
| chr13_-_36566260 | 0.96 |
ENSDART00000030133
|
synj2bp
|
synaptojanin 2 binding protein |
| chr4_-_56157199 | 0.96 |
ENSDART00000169806
|
si:ch211-207e19.15
|
si:ch211-207e19.15 |
| chr4_+_38981587 | 0.96 |
ENSDART00000142713
|
si:dkey-66k12.3
|
si:dkey-66k12.3 |
| chr8_+_36570791 | 0.95 |
ENSDART00000145566
ENSDART00000180527 |
pold2
|
polymerase (DNA directed), delta 2, regulatory subunit |
| chr1_+_12195700 | 0.95 |
ENSDART00000040307
|
tdrd7a
|
tudor domain containing 7 a |
| chr13_-_25199260 | 0.94 |
ENSDART00000057605
|
adka
|
adenosine kinase a |
| chr3_+_31680592 | 0.94 |
ENSDART00000172456
|
mylk5
|
myosin, light chain kinase 5 |
| chr8_+_32742650 | 0.94 |
ENSDART00000138117
|
hmcn2
|
hemicentin 2 |
| chr24_-_21172122 | 0.93 |
ENSDART00000154259
|
atp6v1ab
|
ATPase H+ transporting V1 subunit Ab |
| chr13_+_713941 | 0.93 |
ENSDART00000109737
|
cox15
|
cytochrome c oxidase assembly homolog 15 (yeast) |
| chr9_-_25255490 | 0.93 |
ENSDART00000141502
|
htr2aa
|
5-hydroxytryptamine (serotonin) receptor 2A, genome duplicate a |
| chr1_+_24387659 | 0.92 |
ENSDART00000130356
|
qdprb2
|
quinoid dihydropteridine reductase b2 |
| chr25_-_25575717 | 0.92 |
ENSDART00000067138
|
hic1l
|
hypermethylated in cancer 1 like |
| chr7_+_35193832 | 0.92 |
ENSDART00000189002
|
zdhhc1
|
zinc finger, DHHC-type containing 1 |
| chr9_+_32859967 | 0.90 |
ENSDART00000168992
|
si:dkey-145p14.5
|
si:dkey-145p14.5 |
| chr9_+_32301456 | 0.90 |
ENSDART00000078608
ENSDART00000185153 ENSDART00000144947 |
hspe1
|
heat shock 10 protein 1 |
| chr14_-_48103207 | 0.89 |
ENSDART00000056712
|
etfdh
|
electron-transferring-flavoprotein dehydrogenase |
| chr17_+_32374876 | 0.89 |
ENSDART00000183851
|
ywhaqb
|
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, theta polypeptide b |
| chr4_-_16833518 | 0.89 |
ENSDART00000179867
|
ldhba
|
lactate dehydrogenase Ba |
| chr8_+_23439340 | 0.89 |
ENSDART00000109932
ENSDART00000185469 |
foxp3b
|
forkhead box P3b |
| chr23_-_24234371 | 0.88 |
ENSDART00000124539
|
ddost
|
dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit (non-catalytic) |
| chr5_+_37035978 | 0.88 |
ENSDART00000167418
|
nfkbib
|
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, beta |
| chr17_-_49407091 | 0.87 |
ENSDART00000021950
|
mthfd1b
|
methylenetetrahydrofolate dehydrogenase (NADP+ dependent) 1b |
| chr7_+_38267136 | 0.86 |
ENSDART00000173613
|
gpatch1
|
G patch domain containing 1 |
| chr20_+_25445826 | 0.86 |
ENSDART00000012581
|
pfas
|
phosphoribosylformylglycinamidine synthase |
| chr9_-_28990649 | 0.86 |
ENSDART00000078823
|
ptpn4a
|
protein tyrosine phosphatase, non-receptor type 4a |
| chr16_+_23972126 | 0.86 |
ENSDART00000132742
ENSDART00000145330 |
apoc1
|
apolipoprotein C-I |
| chr19_+_2877079 | 0.85 |
ENSDART00000160847
|
hgh1
|
HGH1 homolog (S. cerevisiae) |
| chr1_+_58353661 | 0.85 |
ENSDART00000140074
|
si:dkey-222h21.2
|
si:dkey-222h21.2 |
| chr1_-_26664840 | 0.85 |
ENSDART00000102500
|
anp32b
|
acidic (leucine-rich) nuclear phosphoprotein 32 family, member B |
| chr16_-_29146624 | 0.85 |
ENSDART00000159814
ENSDART00000009826 |
mef2d
|
myocyte enhancer factor 2d |
| chr14_-_6286966 | 0.85 |
ENSDART00000168174
|
elp1
|
elongator complex protein 1 |
| chr13_-_35808904 | 0.84 |
ENSDART00000171667
|
map3k4
|
mitogen-activated protein kinase kinase kinase 4 |
| chr17_-_45733401 | 0.84 |
ENSDART00000185727
|
arf6b
|
ADP-ribosylation factor 6b |
| chr2_-_22659450 | 0.83 |
ENSDART00000115025
|
thap4
|
THAP domain containing 4 |
| chr7_+_23940933 | 0.83 |
ENSDART00000173628
|
si:dkey-183c6.7
|
si:dkey-183c6.7 |
| chr20_-_21994901 | 0.82 |
ENSDART00000004984
|
daam1b
|
dishevelled associated activator of morphogenesis 1b |
| chr23_-_29394505 | 0.81 |
ENSDART00000017728
|
pgd
|
phosphogluconate dehydrogenase |
| chr25_+_25766033 | 0.81 |
ENSDART00000103638
ENSDART00000039952 |
idh3a
|
isocitrate dehydrogenase 3 (NAD+) alpha |
| chr25_-_16076257 | 0.81 |
ENSDART00000140780
|
ovch2
|
ovochymase 2 |
| chr18_-_7448047 | 0.80 |
ENSDART00000193213
ENSDART00000131940 ENSDART00000186944 ENSDART00000052803 |
si:dkey-30c15.10
|
si:dkey-30c15.10 |
| chr21_-_21515466 | 0.80 |
ENSDART00000147593
|
nectin3b
|
nectin cell adhesion molecule 3b |
| chr13_-_24260609 | 0.80 |
ENSDART00000138747
|
urb2
|
URB2 ribosome biogenesis 2 homolog (S. cerevisiae) |
| chr17_+_24036791 | 0.80 |
ENSDART00000140767
|
b3gnt2b
|
UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 2b |
| chr20_+_46286082 | 0.79 |
ENSDART00000073519
|
taar14f
|
trace amine associated receptor 14f |
| chr1_-_44581937 | 0.79 |
ENSDART00000009858
|
tmx2b
|
thioredoxin-related transmembrane protein 2b |
| chr14_+_30413758 | 0.78 |
ENSDART00000092953
|
cnot7
|
CCR4-NOT transcription complex, subunit 7 |
| chr18_-_29972709 | 0.78 |
ENSDART00000131207
|
si:ch211-220f16.2
|
si:ch211-220f16.2 |
| chr9_-_1939232 | 0.78 |
ENSDART00000146131
|
hoxd3a
|
homeobox D3a |
| chr10_+_20128267 | 0.78 |
ENSDART00000064615
|
dmtn
|
dematin actin binding protein |
| chr10_-_21362071 | 0.76 |
ENSDART00000125167
|
avd
|
avidin |
| chr11_-_6974022 | 0.76 |
ENSDART00000172851
|
COMP
|
si:ch211-43f4.1 |
| chr4_+_37503994 | 0.76 |
ENSDART00000158217
|
znf1102
|
zinc finger protein 1102 |
| chr23_-_28808480 | 0.76 |
ENSDART00000078167
|
dffa
|
DNA fragmentation factor, alpha polypeptide |
| chr13_+_7241170 | 0.75 |
ENSDART00000109434
|
aifm2
|
apoptosis-inducing factor, mitochondrion-associated, 2 |
| chr25_+_15647993 | 0.75 |
ENSDART00000186578
ENSDART00000031828 |
spon1b
|
spondin 1b |
| chr5_-_41860556 | 0.74 |
ENSDART00000140154
|
si:dkey-65b12.12
|
si:dkey-65b12.12 |
| chr7_+_17543487 | 0.73 |
ENSDART00000126652
|
CU672228.2
|
Danio rerio novel immune-type receptor 1l (nitr1l), mRNA. |
| chr3_-_7134113 | 0.73 |
ENSDART00000180849
|
BX005085.6
|
|
| chr7_+_5936601 | 0.72 |
ENSDART00000181693
|
si:dkey-23a13.11
|
si:dkey-23a13.11 |
| chr2_+_58377395 | 0.72 |
ENSDART00000193511
|
vapal
|
VAMP (vesicle-associated membrane protein)-associated protein A, like |
| chr19_+_32158010 | 0.71 |
ENSDART00000005255
|
mrpl53
|
mitochondrial ribosomal protein L53 |
| chr18_+_14529005 | 0.71 |
ENSDART00000186379
|
kcng4a
|
potassium voltage-gated channel, subfamily G, member 4a |
| chr3_+_50201240 | 0.71 |
ENSDART00000156347
|
epn3a
|
epsin 3a |
| chr2_+_35880600 | 0.71 |
ENSDART00000004277
|
lamc1
|
laminin, gamma 1 |
| chr12_-_33568174 | 0.71 |
ENSDART00000142329
|
tdrkh
|
tudor and KH domain containing |
| chr12_+_26670778 | 0.71 |
ENSDART00000144355
|
arhgap12b
|
Rho GTPase activating protein 12b |
| chr20_-_49117438 | 0.71 |
ENSDART00000057700
|
naa20
|
N(alpha)-acetyltransferase 20, NatB catalytic subunit |
| chr11_+_11751006 | 0.71 |
ENSDART00000180372
ENSDART00000081018 |
jupa
|
junction plakoglobin a |
| chr24_+_36389536 | 0.70 |
ENSDART00000179898
|
map3k14b
|
mitogen-activated protein kinase kinase kinase 14b |
| chr3_-_32590164 | 0.70 |
ENSDART00000151151
|
tspan4b
|
tetraspanin 4b |
| chr17_-_40110782 | 0.69 |
ENSDART00000126929
|
si:dkey-187k19.2
|
si:dkey-187k19.2 |
| chr3_+_27770110 | 0.69 |
ENSDART00000017962
|
eci1
|
enoyl-CoA delta isomerase 1 |
| chr10_+_24692076 | 0.69 |
ENSDART00000181600
|
tpte
|
transmembrane phosphatase with tensin homology |
| chr8_+_2530065 | 0.69 |
ENSDART00000063943
|
mrpl40
|
mitochondrial ribosomal protein L40 |
| chr19_-_874888 | 0.69 |
ENSDART00000007206
|
eomesa
|
eomesodermin homolog a |
| chr16_-_24605969 | 0.68 |
ENSDART00000163305
ENSDART00000167121 |
fxyd6l
|
FXYD domain containing ion transport regulator 6 like |
| chr10_+_17235370 | 0.68 |
ENSDART00000038780
|
sppl3
|
signal peptide peptidase 3 |
| chr16_-_32128345 | 0.68 |
ENSDART00000102100
ENSDART00000137699 |
cnot3a
|
CCR4-NOT transcription complex, subunit 3a |
| chr1_+_54737353 | 0.67 |
ENSDART00000130675
ENSDART00000162075 |
pi4k2a
|
phosphatidylinositol 4-kinase type 2 alpha |
| chr14_+_34501245 | 0.67 |
ENSDART00000131424
|
lcp2b
|
lymphocyte cytosolic protein 2b |
| chr16_+_42829735 | 0.67 |
ENSDART00000014956
|
polr3glb
|
polymerase (RNA) III (DNA directed) polypeptide G like b |
| chr19_-_24484758 | 0.67 |
ENSDART00000052422
|
mtx1b
|
metaxin 1b |
| chr10_+_589501 | 0.65 |
ENSDART00000188415
|
LO018557.1
|
|
| chr9_+_24936496 | 0.65 |
ENSDART00000157474
|
slc39a10
|
solute carrier family 39 (zinc transporter), member 10 |
| chr2_-_22660232 | 0.65 |
ENSDART00000174742
|
thap4
|
THAP domain containing 4 |
| chr23_+_8797143 | 0.65 |
ENSDART00000132992
|
sox18
|
SRY (sex determining region Y)-box 18 |
| chr9_-_43082945 | 0.64 |
ENSDART00000142257
|
ccdc141
|
coiled-coil domain containing 141 |
| chr6_-_21873266 | 0.64 |
ENSDART00000151658
ENSDART00000151152 ENSDART00000151179 |
si:dkey-19e4.5
|
si:dkey-19e4.5 |
| chr19_+_2631565 | 0.64 |
ENSDART00000171487
|
fam126a
|
family with sequence similarity 126, member A |
| chr10_+_35417099 | 0.64 |
ENSDART00000063398
|
hhla2a.1
|
HERV-H LTR-associating 2a, tandem duplicate 1 |
| chr2_-_55853943 | 0.64 |
ENSDART00000122576
|
rx2
|
retinal homeobox gene 2 |
| chr4_-_42074960 | 0.64 |
ENSDART00000164406
|
zgc:174696
|
zgc:174696 |
Network of associatons between targets according to the STRING database.
First level regulatory network of nr2c2
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 3.6 | GO:1905133 | meiotic cell cycle phase transition(GO:0044771) metaphase/anaphase transition of meiotic cell cycle(GO:0044785) regulation of meiotic cell cycle phase transition(GO:1901993) negative regulation of meiotic cell cycle phase transition(GO:1901994) regulation of metaphase/anaphase transition of meiotic cell cycle(GO:1902102) negative regulation of metaphase/anaphase transition of meiotic cell cycle(GO:1902103) regulation of meiotic chromosome separation(GO:1905132) negative regulation of meiotic chromosome separation(GO:1905133) |
| 0.9 | 3.4 | GO:0045041 | positive regulation of interleukin-6 production(GO:0032755) protein import into mitochondrial intermembrane space(GO:0045041) |
| 0.9 | 6.0 | GO:0003319 | cardioblast migration to the midline involved in heart rudiment formation(GO:0003319) |
| 0.6 | 1.8 | GO:0010961 | cellular magnesium ion homeostasis(GO:0010961) |
| 0.6 | 1.2 | GO:0001954 | positive regulation of cell-matrix adhesion(GO:0001954) |
| 0.5 | 1.4 | GO:0005991 | trehalose metabolic process(GO:0005991) |
| 0.4 | 1.2 | GO:0070914 | UV-damage excision repair(GO:0070914) |
| 0.4 | 2.0 | GO:0051972 | regulation of telomerase activity(GO:0051972) |
| 0.3 | 2.1 | GO:0001955 | blood vessel maturation(GO:0001955) |
| 0.3 | 1.0 | GO:0006089 | lactate metabolic process(GO:0006089) |
| 0.3 | 6.5 | GO:0006271 | DNA strand elongation involved in DNA replication(GO:0006271) |
| 0.3 | 0.9 | GO:0050995 | negative regulation of lipid transport(GO:0032369) negative regulation of lipid catabolic process(GO:0050995) |
| 0.3 | 0.8 | GO:0002926 | tRNA wobble base 5-methoxycarbonylmethyl-2-thiouridine biosynthesis.(GO:0002926) |
| 0.3 | 0.8 | GO:0071918 | urea transmembrane transport(GO:0071918) |
| 0.2 | 1.9 | GO:0021627 | olfactory nerve development(GO:0021553) olfactory nerve morphogenesis(GO:0021627) olfactory nerve formation(GO:0021628) |
| 0.2 | 1.9 | GO:0046546 | development of primary male sexual characteristics(GO:0046546) |
| 0.2 | 0.7 | GO:0055109 | invagination involved in gastrulation with mouth forming second(GO:0055109) morphogenesis of an epithelial fold(GO:0060571) |
| 0.2 | 1.1 | GO:1902299 | pre-replicative complex assembly involved in nuclear cell cycle DNA replication(GO:0006267) pre-replicative complex assembly(GO:0036388) pre-replicative complex assembly involved in cell cycle DNA replication(GO:1902299) |
| 0.2 | 0.7 | GO:0050861 | positive regulation of B cell receptor signaling pathway(GO:0050861) |
| 0.2 | 2.9 | GO:0006474 | N-terminal protein amino acid acetylation(GO:0006474) |
| 0.2 | 0.6 | GO:0031441 | negative regulation of mRNA 3'-end processing(GO:0031441) negative regulation of mRNA polyadenylation(GO:1900364) |
| 0.2 | 1.2 | GO:0071908 | determination of intestine left/right asymmetry(GO:0071908) |
| 0.2 | 0.5 | GO:0033335 | anal fin development(GO:0033335) |
| 0.2 | 0.5 | GO:0019408 | dolichol biosynthetic process(GO:0019408) |
| 0.2 | 0.9 | GO:0044209 | AMP salvage(GO:0044209) |
| 0.1 | 1.6 | GO:0021707 | cerebellar granular layer development(GO:0021681) cerebellar granular layer morphogenesis(GO:0021683) cerebellar granular layer formation(GO:0021684) cerebellar granule cell differentiation(GO:0021707) |
| 0.1 | 2.7 | GO:0000463 | maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000463) |
| 0.1 | 0.4 | GO:0034375 | high-density lipoprotein particle remodeling(GO:0034375) |
| 0.1 | 0.6 | GO:0051645 | Golgi localization(GO:0051645) |
| 0.1 | 0.7 | GO:0002159 | desmosome assembly(GO:0002159) |
| 0.1 | 0.4 | GO:0007414 | axonal defasciculation(GO:0007414) |
| 0.1 | 0.9 | GO:0090497 | skin morphogenesis(GO:0043589) mesenchymal cell migration(GO:0090497) |
| 0.1 | 0.5 | GO:1904667 | negative regulation of ligase activity(GO:0051352) negative regulation of ubiquitin-protein transferase activity(GO:0051444) negative regulation of ubiquitin protein ligase activity(GO:1904667) |
| 0.1 | 1.3 | GO:0043984 | histone H4-K16 acetylation(GO:0043984) |
| 0.1 | 1.2 | GO:0044090 | positive regulation of vacuole organization(GO:0044090) |
| 0.1 | 2.0 | GO:0048096 | chromatin-mediated maintenance of transcription(GO:0048096) |
| 0.1 | 1.8 | GO:1903963 | icosanoid secretion(GO:0032309) arachidonic acid secretion(GO:0050482) arachidonate transport(GO:1903963) |
| 0.1 | 1.4 | GO:0045638 | negative regulation of myeloid cell differentiation(GO:0045638) |
| 0.1 | 0.6 | GO:0090527 | actin filament reorganization(GO:0090527) |
| 0.1 | 0.4 | GO:0021785 | branchiomotor neuron axon guidance(GO:0021785) |
| 0.1 | 0.5 | GO:0006526 | arginine biosynthetic process(GO:0006526) |
| 0.1 | 0.6 | GO:2000001 | regulation of DNA damage checkpoint(GO:2000001) |
| 0.1 | 3.5 | GO:0006506 | GPI anchor biosynthetic process(GO:0006506) |
| 0.1 | 0.8 | GO:0031106 | septin ring organization(GO:0031106) septin cytoskeleton organization(GO:0032185) |
| 0.1 | 1.0 | GO:0098787 | mRNA cleavage involved in mRNA processing(GO:0098787) pre-mRNA cleavage required for polyadenylation(GO:0098789) |
| 0.1 | 0.8 | GO:0030262 | apoptotic DNA fragmentation(GO:0006309) cellular component disassembly involved in execution phase of apoptosis(GO:0006921) apoptotic nuclear changes(GO:0030262) |
| 0.1 | 1.2 | GO:0097369 | sodium ion import(GO:0097369) sodium ion import across plasma membrane(GO:0098719) sodium ion import into cell(GO:1990118) |
| 0.1 | 1.1 | GO:0043928 | exonucleolytic nuclear-transcribed mRNA catabolic process involved in deadenylation-dependent decay(GO:0043928) |
| 0.1 | 0.8 | GO:0006102 | isocitrate metabolic process(GO:0006102) |
| 0.1 | 0.6 | GO:0048714 | positive regulation of glial cell differentiation(GO:0045687) positive regulation of oligodendrocyte differentiation(GO:0048714) |
| 0.1 | 2.1 | GO:0018149 | peptide cross-linking(GO:0018149) |
| 0.1 | 0.6 | GO:0035092 | spermatid nucleus differentiation(GO:0007289) sperm chromatin condensation(GO:0035092) spermatogenesis, exchange of chromosomal proteins(GO:0035093) |
| 0.1 | 0.9 | GO:0046146 | tetrahydrobiopterin biosynthetic process(GO:0006729) tetrahydrobiopterin metabolic process(GO:0046146) |
| 0.1 | 0.9 | GO:0006189 | 'de novo' IMP biosynthetic process(GO:0006189) |
| 0.1 | 0.9 | GO:0030719 | P granule organization(GO:0030719) |
| 0.1 | 1.6 | GO:0043433 | negative regulation of sequence-specific DNA binding transcription factor activity(GO:0043433) |
| 0.1 | 0.5 | GO:0006574 | valine catabolic process(GO:0006574) |
| 0.1 | 0.6 | GO:0051673 | membrane disruption in other organism(GO:0051673) |
| 0.1 | 0.4 | GO:0045039 | protein import into mitochondrial inner membrane(GO:0045039) |
| 0.1 | 0.8 | GO:0030241 | skeletal muscle myosin thick filament assembly(GO:0030241) |
| 0.1 | 1.2 | GO:0046501 | protoporphyrinogen IX biosynthetic process(GO:0006782) protoporphyrinogen IX metabolic process(GO:0046501) |
| 0.1 | 0.7 | GO:0006465 | signal peptide processing(GO:0006465) |
| 0.1 | 0.4 | GO:0007252 | I-kappaB phosphorylation(GO:0007252) |
| 0.1 | 0.2 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 0.1 | 0.9 | GO:0035999 | tetrahydrofolate interconversion(GO:0035999) |
| 0.1 | 0.9 | GO:0006783 | heme biosynthetic process(GO:0006783) |
| 0.1 | 1.6 | GO:0001966 | thigmotaxis(GO:0001966) |
| 0.1 | 0.7 | GO:2000036 | regulation of stem cell population maintenance(GO:2000036) |
| 0.1 | 0.8 | GO:0006098 | pentose-phosphate shunt(GO:0006098) |
| 0.1 | 4.5 | GO:0060048 | cardiac muscle contraction(GO:0060048) |
| 0.1 | 0.3 | GO:0071926 | endocannabinoid signaling pathway(GO:0071926) retrograde trans-synaptic signaling by lipid(GO:0098920) retrograde trans-synaptic signaling by endocannabinoid(GO:0098921) |
| 0.1 | 0.5 | GO:0034633 | retinol transport(GO:0034633) isoprenoid transport(GO:0046864) terpenoid transport(GO:0046865) |
| 0.1 | 1.6 | GO:0051289 | protein homotetramerization(GO:0051289) |
| 0.1 | 0.2 | GO:0072020 | proximal straight tubule development(GO:0072020) |
| 0.1 | 1.0 | GO:0006614 | SRP-dependent cotranslational protein targeting to membrane(GO:0006614) |
| 0.1 | 0.7 | GO:0050908 | detection of light stimulus involved in visual perception(GO:0050908) detection of light stimulus involved in sensory perception(GO:0050962) |
| 0.1 | 0.6 | GO:0031294 | lymphocyte costimulation(GO:0031294) T cell costimulation(GO:0031295) |
| 0.1 | 0.3 | GO:0010873 | regulation of cholesterol esterification(GO:0010872) positive regulation of cholesterol esterification(GO:0010873) triglyceride-rich lipoprotein particle remodeling(GO:0034370) very-low-density lipoprotein particle remodeling(GO:0034372) reverse cholesterol transport(GO:0043691) |
| 0.1 | 0.8 | GO:0030309 | poly-N-acetyllactosamine metabolic process(GO:0030309) poly-N-acetyllactosamine biosynthetic process(GO:0030311) |
| 0.1 | 1.8 | GO:0006891 | intra-Golgi vesicle-mediated transport(GO:0006891) |
| 0.1 | 1.3 | GO:0007168 | receptor guanylyl cyclase signaling pathway(GO:0007168) |
| 0.0 | 0.9 | GO:0018279 | protein N-linked glycosylation via asparagine(GO:0018279) |
| 0.0 | 1.4 | GO:0003323 | type B pancreatic cell development(GO:0003323) |
| 0.0 | 0.5 | GO:0006607 | NLS-bearing protein import into nucleus(GO:0006607) |
| 0.0 | 0.1 | GO:0007060 | male meiosis chromosome segregation(GO:0007060) |
| 0.0 | 0.1 | GO:0050787 | response to mercury ion(GO:0046689) detoxification of mercury ion(GO:0050787) |
| 0.0 | 1.3 | GO:0000387 | spliceosomal snRNP assembly(GO:0000387) |
| 0.0 | 0.8 | GO:0030032 | lamellipodium assembly(GO:0030032) |
| 0.0 | 1.2 | GO:0006383 | transcription from RNA polymerase III promoter(GO:0006383) |
| 0.0 | 1.2 | GO:0051085 | 'de novo' protein folding(GO:0006458) 'de novo' posttranslational protein folding(GO:0051084) chaperone mediated protein folding requiring cofactor(GO:0051085) |
| 0.0 | 0.4 | GO:0050909 | sensory perception of taste(GO:0050909) |
| 0.0 | 0.7 | GO:0010811 | positive regulation of cell-substrate adhesion(GO:0010811) |
| 0.0 | 1.1 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.0 | 2.8 | GO:1990573 | potassium ion import across plasma membrane(GO:1990573) |
| 0.0 | 0.7 | GO:0010257 | NADH dehydrogenase complex assembly(GO:0010257) mitochondrial respiratory chain complex I assembly(GO:0032981) mitochondrial respiratory chain complex I biogenesis(GO:0097031) |
| 0.0 | 2.0 | GO:0002224 | toll-like receptor signaling pathway(GO:0002224) |
| 0.0 | 0.7 | GO:0007032 | endosome organization(GO:0007032) |
| 0.0 | 0.8 | GO:0035622 | intrahepatic bile duct development(GO:0035622) |
| 0.0 | 1.3 | GO:0045727 | positive regulation of translation(GO:0045727) |
| 0.0 | 0.9 | GO:0051209 | release of sequestered calcium ion into cytosol(GO:0051209) |
| 0.0 | 3.5 | GO:0006979 | response to oxidative stress(GO:0006979) |
| 0.0 | 1.0 | GO:0016339 | calcium-dependent cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0016339) |
| 0.0 | 1.0 | GO:0071711 | basement membrane organization(GO:0071711) |
| 0.0 | 1.0 | GO:0006635 | fatty acid beta-oxidation(GO:0006635) |
| 0.0 | 0.8 | GO:0008637 | apoptotic mitochondrial changes(GO:0008637) |
| 0.0 | 0.8 | GO:0007157 | heterophilic cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0007157) |
| 0.0 | 0.3 | GO:0070212 | protein poly-ADP-ribosylation(GO:0070212) |
| 0.0 | 0.5 | GO:0043252 | sodium-independent organic anion transport(GO:0043252) |
| 0.0 | 1.1 | GO:0070830 | apical junction assembly(GO:0043297) bicellular tight junction assembly(GO:0070830) |
| 0.0 | 1.6 | GO:0032200 | telomere maintenance(GO:0000723) telomere organization(GO:0032200) |
| 0.0 | 0.3 | GO:0071712 | ER-associated misfolded protein catabolic process(GO:0071712) |
| 0.0 | 1.0 | GO:0097120 | receptor localization to synapse(GO:0097120) |
| 0.0 | 0.9 | GO:0060219 | camera-type eye photoreceptor cell differentiation(GO:0060219) |
| 0.0 | 0.6 | GO:0019226 | transmission of nerve impulse(GO:0019226) |
| 0.0 | 0.2 | GO:0021680 | cerebellar Purkinje cell layer development(GO:0021680) |
| 0.0 | 0.1 | GO:0033292 | T-tubule organization(GO:0033292) |
| 0.0 | 0.6 | GO:0003094 | glomerular filtration(GO:0003094) |
| 0.0 | 0.8 | GO:0006081 | cellular aldehyde metabolic process(GO:0006081) |
| 0.0 | 0.2 | GO:0001952 | regulation of cell-matrix adhesion(GO:0001952) |
| 0.0 | 0.7 | GO:0043491 | protein kinase B signaling(GO:0043491) |
| 0.0 | 0.1 | GO:0070294 | renal sodium ion transport(GO:0003096) renal sodium ion absorption(GO:0070294) |
| 0.0 | 0.5 | GO:0045746 | negative regulation of Notch signaling pathway(GO:0045746) |
| 0.0 | 0.7 | GO:0060840 | artery development(GO:0060840) |
| 0.0 | 0.2 | GO:0072310 | glomerular visceral epithelial cell development(GO:0072015) glomerular epithelial cell development(GO:0072310) |
| 0.0 | 0.1 | GO:0007141 | male meiosis I(GO:0007141) |
| 0.0 | 0.9 | GO:1902600 | hydrogen ion transmembrane transport(GO:1902600) |
| 0.0 | 0.3 | GO:0032008 | positive regulation of TOR signaling(GO:0032008) |
| 0.0 | 0.9 | GO:0022904 | respiratory electron transport chain(GO:0022904) |
| 0.0 | 0.5 | GO:0071427 | mRNA export from nucleus(GO:0006406) mRNA-containing ribonucleoprotein complex export from nucleus(GO:0071427) |
| 0.0 | 0.9 | GO:0030917 | midbrain-hindbrain boundary development(GO:0030917) |
| 0.0 | 0.3 | GO:0045879 | negative regulation of smoothened signaling pathway(GO:0045879) |
| 0.0 | 0.3 | GO:0070972 | protein localization to endoplasmic reticulum(GO:0070972) |
| 0.0 | 0.3 | GO:0031629 | synaptic vesicle fusion to presynaptic active zone membrane(GO:0031629) vesicle fusion to plasma membrane(GO:0099500) |
| 0.0 | 3.4 | GO:0000122 | negative regulation of transcription from RNA polymerase II promoter(GO:0000122) |
| 0.0 | 0.4 | GO:1902036 | regulation of hematopoietic stem cell differentiation(GO:1902036) |
| 0.0 | 0.4 | GO:0018231 | peptidyl-L-cysteine S-palmitoylation(GO:0018230) peptidyl-S-diacylglycerol-L-cysteine biosynthetic process from peptidyl-cysteine(GO:0018231) |
| 0.0 | 1.4 | GO:0017157 | regulation of exocytosis(GO:0017157) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 2.9 | GO:0033185 | dolichol-phosphate-mannose synthase complex(GO:0033185) |
| 0.9 | 2.7 | GO:0070545 | PeBoW complex(GO:0070545) |
| 0.5 | 2.3 | GO:0043625 | delta DNA polymerase complex(GO:0043625) |
| 0.4 | 2.5 | GO:0035517 | PR-DUB complex(GO:0035517) |
| 0.3 | 1.3 | GO:0044326 | dendritic spine neck(GO:0044326) dendritic filopodium(GO:1902737) |
| 0.3 | 1.3 | GO:0032299 | ribonuclease H2 complex(GO:0032299) |
| 0.3 | 1.2 | GO:0000110 | nucleotide-excision repair factor 1 complex(GO:0000110) |
| 0.3 | 1.0 | GO:0008622 | epsilon DNA polymerase complex(GO:0008622) |
| 0.2 | 1.0 | GO:1990909 | Wnt signalosome(GO:1990909) |
| 0.2 | 0.6 | GO:1990072 | TRAPPIII protein complex(GO:1990072) |
| 0.2 | 0.7 | GO:0098553 | integral component of cytoplasmic side of endoplasmic reticulum membrane(GO:0071458) integral component of lumenal side of endoplasmic reticulum membrane(GO:0071556) lumenal side of endoplasmic reticulum membrane(GO:0098553) |
| 0.2 | 2.9 | GO:0042555 | MCM complex(GO:0042555) |
| 0.1 | 1.2 | GO:0072487 | MSL complex(GO:0072487) |
| 0.1 | 1.8 | GO:0030015 | CCR4-NOT core complex(GO:0030015) |
| 0.1 | 0.5 | GO:1904423 | dehydrodolichyl diphosphate synthase complex(GO:1904423) |
| 0.1 | 0.8 | GO:0005826 | actomyosin contractile ring(GO:0005826) |
| 0.1 | 0.3 | GO:0042721 | mitochondrial inner membrane protein insertion complex(GO:0042721) |
| 0.1 | 1.4 | GO:0017119 | Golgi transport complex(GO:0017119) |
| 0.1 | 4.1 | GO:0000315 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.1 | 3.6 | GO:0000781 | chromosome, telomeric region(GO:0000781) |
| 0.1 | 1.0 | GO:0030126 | COPI vesicle coat(GO:0030126) |
| 0.1 | 0.8 | GO:0033588 | Elongator holoenzyme complex(GO:0033588) |
| 0.1 | 0.7 | GO:0001401 | mitochondrial sorting and assembly machinery complex(GO:0001401) |
| 0.1 | 1.2 | GO:0034364 | high-density lipoprotein particle(GO:0034364) |
| 0.1 | 0.7 | GO:0031414 | N-terminal protein acetyltransferase complex(GO:0031414) |
| 0.1 | 0.5 | GO:0000347 | THO complex(GO:0000347) THO complex part of transcription export complex(GO:0000445) |
| 0.1 | 0.3 | GO:0034448 | EGO complex(GO:0034448) Gtr1-Gtr2 GTPase complex(GO:1990131) |
| 0.1 | 0.3 | GO:0034098 | VCP-NPL4-UFD1 AAA ATPase complex(GO:0034098) |
| 0.1 | 0.9 | GO:0008250 | oligosaccharyltransferase complex(GO:0008250) |
| 0.1 | 0.4 | GO:0008385 | IkappaB kinase complex(GO:0008385) |
| 0.1 | 1.3 | GO:0005732 | small nucleolar ribonucleoprotein complex(GO:0005732) |
| 0.1 | 1.2 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.1 | 2.4 | GO:0005861 | troponin complex(GO:0005861) |
| 0.1 | 0.9 | GO:0033180 | proton-transporting V-type ATPase, V1 domain(GO:0033180) |
| 0.1 | 7.0 | GO:0005759 | mitochondrial matrix(GO:0005759) |
| 0.1 | 0.5 | GO:0016272 | prefoldin complex(GO:0016272) |
| 0.1 | 1.2 | GO:0005686 | U2 snRNP(GO:0005686) U12-type spliceosomal complex(GO:0005689) |
| 0.1 | 1.2 | GO:0005666 | DNA-directed RNA polymerase III complex(GO:0005666) |
| 0.0 | 1.5 | GO:0043186 | P granule(GO:0043186) |
| 0.0 | 0.5 | GO:0072546 | ER membrane protein complex(GO:0072546) |
| 0.0 | 1.5 | GO:0032156 | septin ring(GO:0005940) septin complex(GO:0031105) septin cytoskeleton(GO:0032156) |
| 0.0 | 0.7 | GO:0030057 | desmosome(GO:0030057) |
| 0.0 | 3.4 | GO:0000776 | kinetochore(GO:0000776) |
| 0.0 | 0.7 | GO:0031083 | BLOC-1 complex(GO:0031083) |
| 0.0 | 0.1 | GO:0042719 | mitochondrial intermembrane space protein transporter complex(GO:0042719) |
| 0.0 | 0.7 | GO:0031594 | neuromuscular junction(GO:0031594) |
| 0.0 | 1.0 | GO:0031305 | integral component of mitochondrial inner membrane(GO:0031305) |
| 0.0 | 0.7 | GO:0030125 | clathrin vesicle coat(GO:0030125) |
| 0.0 | 0.1 | GO:0005854 | nascent polypeptide-associated complex(GO:0005854) |
| 0.0 | 0.9 | GO:0097517 | stress fiber(GO:0001725) contractile actin filament bundle(GO:0097517) |
| 0.0 | 0.3 | GO:0098843 | postsynaptic endocytic zone(GO:0098843) postsynaptic endocytic zone membrane(GO:0098844) |
| 0.0 | 0.5 | GO:0005901 | caveola(GO:0005901) plasma membrane raft(GO:0044853) |
| 0.0 | 0.6 | GO:0005811 | lipid particle(GO:0005811) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 6.0 | GO:0004962 | endothelin receptor activity(GO:0004962) |
| 0.7 | 3.5 | GO:0047066 | phospholipid-hydroperoxide glutathione peroxidase activity(GO:0047066) |
| 0.6 | 2.2 | GO:0043998 | H2A histone acetyltransferase activity(GO:0043998) |
| 0.5 | 1.6 | GO:0004174 | electron-transferring-flavoprotein dehydrogenase activity(GO:0004174) oxidoreductase activity, acting on the CH-NH group of donors, quinone or similar compound as acceptor(GO:0016649) |
| 0.5 | 1.6 | GO:0043878 | glyceraldehyde-3-phosphate dehydrogenase (NAD+) (non-phosphorylating) activity(GO:0043878) |
| 0.5 | 2.5 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.5 | 2.0 | GO:0010521 | telomerase inhibitor activity(GO:0010521) |
| 0.5 | 1.4 | GO:0004555 | alpha,alpha-trehalase activity(GO:0004555) trehalase activity(GO:0015927) |
| 0.4 | 1.8 | GO:0043140 | ATP-dependent 3'-5' DNA helicase activity(GO:0043140) single-stranded DNA-dependent ATP-dependent 3'-5' DNA helicase activity(GO:1990518) |
| 0.3 | 3.0 | GO:0031013 | troponin C binding(GO:0030172) troponin I binding(GO:0031013) |
| 0.2 | 0.9 | GO:0004155 | 6,7-dihydropteridine reductase activity(GO:0004155) NADH binding(GO:0070404) |
| 0.2 | 1.6 | GO:0004332 | fructose-bisphosphate aldolase activity(GO:0004332) |
| 0.2 | 2.8 | GO:0015272 | ATP-activated inward rectifier potassium channel activity(GO:0015272) |
| 0.2 | 0.9 | GO:0004001 | adenosine kinase activity(GO:0004001) |
| 0.2 | 1.3 | GO:0004523 | RNA-DNA hybrid ribonuclease activity(GO:0004523) |
| 0.2 | 1.1 | GO:0017116 | single-stranded DNA-dependent ATP-dependent DNA helicase activity(GO:0017116) single-stranded DNA-dependent ATPase activity(GO:0043142) |
| 0.2 | 0.7 | GO:0033149 | FFAT motif binding(GO:0033149) |
| 0.2 | 1.7 | GO:0004028 | 3-chloroallyl aldehyde dehydrogenase activity(GO:0004028) |
| 0.2 | 0.8 | GO:0004449 | isocitrate dehydrogenase (NAD+) activity(GO:0004449) |
| 0.2 | 0.8 | GO:0009374 | biotin binding(GO:0009374) |
| 0.1 | 0.5 | GO:0051377 | mannose-ethanolamine phosphotransferase activity(GO:0051377) |
| 0.1 | 1.8 | GO:0047631 | ADP-ribose diphosphatase activity(GO:0047631) |
| 0.1 | 0.5 | GO:0045547 | dehydrodolichyl diphosphate synthase activity(GO:0045547) |
| 0.1 | 0.9 | GO:0004459 | L-lactate dehydrogenase activity(GO:0004459) |
| 0.1 | 1.0 | GO:0004300 | enoyl-CoA hydratase activity(GO:0004300) |
| 0.1 | 0.9 | GO:0004329 | formate-tetrahydrofolate ligase activity(GO:0004329) |
| 0.1 | 0.5 | GO:0016842 | amidine-lyase activity(GO:0016842) |
| 0.1 | 0.5 | GO:0008442 | 3-hydroxyisobutyrate dehydrogenase activity(GO:0008442) |
| 0.1 | 0.6 | GO:0047192 | 1-alkylglycerophosphocholine O-acetyltransferase activity(GO:0047192) |
| 0.1 | 4.7 | GO:0051087 | chaperone binding(GO:0051087) |
| 0.1 | 0.7 | GO:0016314 | phosphatidylinositol-3,4,5-trisphosphate 3-phosphatase activity(GO:0016314) |
| 0.1 | 1.2 | GO:0005451 | monovalent cation:proton antiporter activity(GO:0005451) sodium:proton antiporter activity(GO:0015385) potassium:proton antiporter activity(GO:0015386) |
| 0.1 | 2.1 | GO:0003810 | protein-glutamine gamma-glutamyltransferase activity(GO:0003810) |
| 0.1 | 4.2 | GO:0004712 | protein serine/threonine/tyrosine kinase activity(GO:0004712) |
| 0.1 | 0.7 | GO:0004430 | 1-phosphatidylinositol 4-kinase activity(GO:0004430) |
| 0.1 | 3.3 | GO:0019003 | GDP binding(GO:0019003) |
| 0.1 | 0.4 | GO:1904121 | phosphatidylethanolamine transporter activity(GO:1904121) |
| 0.1 | 0.6 | GO:0004045 | aminoacyl-tRNA hydrolase activity(GO:0004045) |
| 0.1 | 0.7 | GO:0042500 | aspartic endopeptidase activity, intramembrane cleaving(GO:0042500) |
| 0.1 | 0.9 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 0.1 | 0.3 | GO:0005021 | vascular endothelial growth factor-activated receptor activity(GO:0005021) |
| 0.1 | 0.7 | GO:0004596 | peptide alpha-N-acetyltransferase activity(GO:0004596) |
| 0.1 | 0.7 | GO:0045294 | alpha-catenin binding(GO:0045294) |
| 0.1 | 0.9 | GO:0016653 | oxidoreductase activity, acting on NAD(P)H, heme protein as acceptor(GO:0016653) |
| 0.1 | 1.0 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.1 | 0.8 | GO:0015204 | urea transmembrane transporter activity(GO:0015204) |
| 0.1 | 1.1 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.1 | 0.9 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 0.1 | 0.4 | GO:0016415 | octanoyltransferase activity(GO:0016415) |
| 0.1 | 0.6 | GO:0034584 | piRNA binding(GO:0034584) |
| 0.1 | 1.1 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 0.1 | 0.4 | GO:0016863 | intramolecular oxidoreductase activity, transposing C=C bonds(GO:0016863) |
| 0.1 | 0.5 | GO:0034632 | retinol transporter activity(GO:0034632) |
| 0.1 | 0.9 | GO:0055102 | phospholipase inhibitor activity(GO:0004859) lipase inhibitor activity(GO:0055102) |
| 0.1 | 0.9 | GO:0016884 | carbon-nitrogen ligase activity, with glutamine as amido-N-donor(GO:0016884) |
| 0.1 | 0.5 | GO:0008467 | [heparan sulfate]-glucosamine 3-sulfotransferase 1 activity(GO:0008467) |
| 0.1 | 0.3 | GO:0004679 | AMP-activated protein kinase activity(GO:0004679) |
| 0.1 | 0.3 | GO:0060228 | phosphatidylcholine-sterol O-acyltransferase activator activity(GO:0060228) |
| 0.1 | 0.8 | GO:0008532 | N-acetyllactosaminide beta-1,3-N-acetylglucosaminyltransferase activity(GO:0008532) |
| 0.1 | 4.9 | GO:0001227 | transcriptional repressor activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001227) |
| 0.0 | 1.1 | GO:0004623 | phospholipase A2 activity(GO:0004623) |
| 0.0 | 1.3 | GO:0045182 | translation regulator activity(GO:0045182) |
| 0.0 | 0.1 | GO:0046964 | 3'-phosphoadenosine 5'-phosphosulfate transmembrane transporter activity(GO:0046964) |
| 0.0 | 2.7 | GO:0043021 | ribonucleoprotein complex binding(GO:0043021) |
| 0.0 | 0.5 | GO:0061608 | nuclear import signal receptor activity(GO:0061608) |
| 0.0 | 0.2 | GO:0042169 | SH2 domain binding(GO:0042169) |
| 0.0 | 3.6 | GO:0004879 | RNA polymerase II transcription factor activity, ligand-activated sequence-specific DNA binding(GO:0004879) |
| 0.0 | 1.2 | GO:0003684 | damaged DNA binding(GO:0003684) |
| 0.0 | 0.6 | GO:0008143 | poly(A) binding(GO:0008143) |
| 0.0 | 1.7 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 7.3 | GO:0030246 | carbohydrate binding(GO:0030246) |
| 0.0 | 0.3 | GO:1990404 | protein ADP-ribosylase activity(GO:1990404) |
| 0.0 | 0.5 | GO:0015347 | sodium-independent organic anion transmembrane transporter activity(GO:0015347) |
| 0.0 | 0.4 | GO:0048495 | Roundabout binding(GO:0048495) |
| 0.0 | 1.4 | GO:0016790 | thiolester hydrolase activity(GO:0016790) |
| 0.0 | 0.7 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.0 | 0.1 | GO:0004557 | alpha-galactosidase activity(GO:0004557) |
| 0.0 | 0.6 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.0 | 0.8 | GO:0015036 | disulfide oxidoreductase activity(GO:0015036) |
| 0.0 | 0.6 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.0 | 0.8 | GO:0050661 | NADP binding(GO:0050661) |
| 0.0 | 0.8 | GO:0000049 | tRNA binding(GO:0000049) |
| 0.0 | 1.2 | GO:0016836 | hydro-lyase activity(GO:0016836) |
| 0.0 | 0.2 | GO:0005332 | gamma-aminobutyric acid:sodium symporter activity(GO:0005332) |
| 0.0 | 2.3 | GO:0003714 | transcription corepressor activity(GO:0003714) |
| 0.0 | 0.5 | GO:0030374 | ligand-dependent nuclear receptor transcription coactivator activity(GO:0030374) |
| 0.0 | 0.7 | GO:0005251 | delayed rectifier potassium channel activity(GO:0005251) |
| 0.0 | 1.0 | GO:0031490 | chromatin DNA binding(GO:0031490) |
| 0.0 | 0.5 | GO:0098631 | protein binding involved in cell adhesion(GO:0098631) protein binding involved in cell-cell adhesion(GO:0098632) |
| 0.0 | 1.3 | GO:0001594 | trace-amine receptor activity(GO:0001594) |
| 0.0 | 0.4 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.0 | 2.8 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.0 | 0.2 | GO:0008422 | beta-glucosidase activity(GO:0008422) |
| 0.0 | 0.4 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.0 | 0.4 | GO:0043539 | protein serine/threonine kinase activator activity(GO:0043539) |
| 0.0 | 0.1 | GO:0005220 | inositol 1,4,5-trisphosphate-sensitive calcium-release channel activity(GO:0005220) |
| 0.0 | 0.1 | GO:0004366 | glycerol-3-phosphate O-acyltransferase activity(GO:0004366) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 3.8 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.1 | 0.7 | PID WNT CANONICAL PATHWAY | Canonical Wnt signaling pathway |
| 0.1 | 1.5 | PID ARF6 PATHWAY | Arf6 signaling events |
| 0.1 | 0.3 | PID VEGF VEGFR PATHWAY | VEGF and VEGFR signaling network |
| 0.0 | 0.8 | PID P38 MKK3 6PATHWAY | p38 MAPK signaling pathway |
| 0.0 | 1.2 | PID PS1 PATHWAY | Presenilin action in Notch and Wnt signaling |
| 0.0 | 0.8 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.0 | 0.9 | ST TUMOR NECROSIS FACTOR PATHWAY | Tumor Necrosis Factor Pathway. |
| 0.0 | 1.8 | PID ATR PATHWAY | ATR signaling pathway |
| 0.0 | 0.8 | SA CASPASE CASCADE | Apoptosis is mediated by caspases, cysteine proteases arranged in a proteolytic cascade. |
| 0.0 | 1.3 | PID AR TF PATHWAY | Regulation of Androgen receptor activity |
| 0.0 | 2.3 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.0 | 0.7 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
| 0.0 | 0.4 | PID GLYPICAN 1PATHWAY | Glypican 1 network |
| 0.0 | 0.6 | PID P53 REGULATION PATHWAY | p53 pathway |
| 0.0 | 0.8 | PID INSULIN PATHWAY | Insulin Pathway |
| 0.0 | 0.3 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.0 | 0.5 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.0 | 0.4 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
| 0.0 | 1.8 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 0.3 | PID VEGFR1 2 PATHWAY | Signaling events mediated by VEGFR1 and VEGFR2 |
| 0.0 | 0.5 | PID E2F PATHWAY | E2F transcription factor network |
| 0.0 | 0.4 | PID P73PATHWAY | p73 transcription factor network |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 1.4 | REACTOME DIGESTION OF DIETARY CARBOHYDRATE | Genes involved in Digestion of dietary carbohydrate |
| 0.2 | 2.9 | REACTOME UNWINDING OF DNA | Genes involved in Unwinding of DNA |
| 0.2 | 2.3 | REACTOME REMOVAL OF THE FLAP INTERMEDIATE FROM THE C STRAND | Genes involved in Removal of the Flap Intermediate from the C-strand |
| 0.2 | 2.9 | REACTOME SYNTHESIS OF SUBSTRATES IN N GLYCAN BIOSYTHESIS | Genes involved in Synthesis of substrates in N-glycan biosythesis |
| 0.2 | 0.8 | REACTOME APOPTOSIS INDUCED DNA FRAGMENTATION | Genes involved in Apoptosis induced DNA fragmentation |
| 0.1 | 2.0 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |
| 0.1 | 1.3 | REACTOME SLBP DEPENDENT PROCESSING OF REPLICATION DEPENDENT HISTONE PRE MRNAS | Genes involved in SLBP Dependent Processing of Replication-Dependent Histone Pre-mRNAs |
| 0.1 | 0.5 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 0.1 | 0.9 | REACTOME ADVANCED GLYCOSYLATION ENDPRODUCT RECEPTOR SIGNALING | Genes involved in Advanced glycosylation endproduct receptor signaling |
| 0.1 | 1.8 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.1 | 1.2 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.1 | 1.3 | REACTOME RIP MEDIATED NFKB ACTIVATION VIA DAI | Genes involved in RIP-mediated NFkB activation via DAI |
| 0.1 | 3.5 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
| 0.1 | 1.0 | REACTOME MITOCHONDRIAL FATTY ACID BETA OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.1 | 1.1 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.1 | 2.4 | REACTOME NUCLEAR SIGNALING BY ERBB4 | Genes involved in Nuclear signaling by ERBB4 |
| 0.1 | 1.5 | REACTOME SEMA4D IN SEMAPHORIN SIGNALING | Genes involved in Sema4D in semaphorin signaling |
| 0.1 | 1.3 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.1 | 0.9 | REACTOME PURINE RIBONUCLEOSIDE MONOPHOSPHATE BIOSYNTHESIS | Genes involved in Purine ribonucleoside monophosphate biosynthesis |
| 0.1 | 0.8 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.1 | 1.1 | REACTOME INTEGRATION OF ENERGY METABOLISM | Genes involved in Integration of energy metabolism |
| 0.1 | 0.7 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.1 | 1.2 | REACTOME FORMATION OF INCISION COMPLEX IN GG NER | Genes involved in Formation of incision complex in GG-NER |
| 0.0 | 0.3 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |
| 0.0 | 0.5 | REACTOME COPI MEDIATED TRANSPORT | Genes involved in COPI Mediated Transport |
| 0.0 | 0.3 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.0 | 0.7 | REACTOME SYNTHESIS OF PIPS AT THE EARLY ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the early endosome membrane |
| 0.0 | 0.6 | REACTOME ACYL CHAIN REMODELLING OF PC | Genes involved in Acyl chain remodelling of PC |
| 0.0 | 0.4 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.0 | 0.5 | REACTOME SYNTHESIS OF GLYCOSYLPHOSPHATIDYLINOSITOL GPI | Genes involved in Synthesis of glycosylphosphatidylinositol (GPI) |
| 0.0 | 0.8 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 1.1 | REACTOME TRANSPORT TO THE GOLGI AND SUBSEQUENT MODIFICATION | Genes involved in Transport to the Golgi and subsequent modification |
| 0.0 | 0.4 | REACTOME PEROXISOMAL LIPID METABOLISM | Genes involved in Peroxisomal lipid metabolism |
| 0.0 | 0.5 | REACTOME NETRIN1 SIGNALING | Genes involved in Netrin-1 signaling |
| 0.0 | 0.3 | REACTOME G1 S SPECIFIC TRANSCRIPTION | Genes involved in G1/S-Specific Transcription |
| 0.0 | 1.6 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 0.3 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.0 | 0.6 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.0 | 1.2 | REACTOME MRNA SPLICING | Genes involved in mRNA Splicing |
| 0.0 | 0.7 | REACTOME L1CAM INTERACTIONS | Genes involved in L1CAM interactions |
| 0.0 | 0.4 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |