Project
PRJEB1986: zebrafish developmental stages transcriptome
Navigation
Downloads
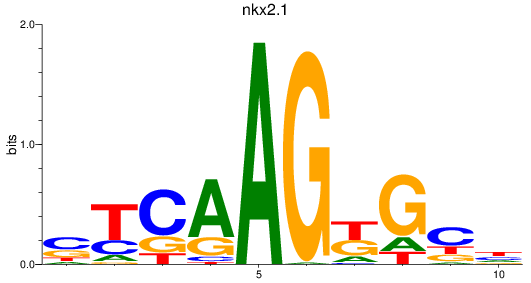
Results for nkx2.1
Z-value: 0.40
Transcription factors associated with nkx2.1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
nkx2.1
|
ENSDARG00000019835 | NK2 homeobox 1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| nkx2.1 | dr11_v1_chr17_+_38262408_38262408 | 0.69 | 1.0e-03 | Click! |
Activity profile of nkx2.1 motif
Sorted Z-values of nkx2.1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_-_59571758 | 1.03 |
ENSDART00000193546
ENSDART00000167087 |
wfikkn1
|
WAP, follistatin/kazal, immunoglobulin, kunitz and netrin domain containing 1 |
| chr20_+_48782068 | 0.97 |
ENSDART00000159275
|
nkx2.2b
|
NK2 homeobox 2b |
| chr13_+_10023256 | 0.94 |
ENSDART00000110035
|
srbd1
|
S1 RNA binding domain 1 |
| chr5_-_54712159 | 0.85 |
ENSDART00000149207
|
ccnb1
|
cyclin B1 |
| chr2_+_19195841 | 0.78 |
ENSDART00000163137
ENSDART00000161095 |
elovl1a
|
ELOVL fatty acid elongase 1a |
| chr11_-_308838 | 0.65 |
ENSDART00000112538
|
poc1a
|
POC1 centriolar protein A |
| chr15_-_23508214 | 0.65 |
ENSDART00000115051
|
abcg4b
|
ATP-binding cassette, sub-family G (WHITE), member 4b |
| chr4_-_2052687 | 0.61 |
ENSDART00000138291
ENSDART00000150844 |
cpsf6
|
cleavage and polyadenylation specific factor 6 |
| chr17_+_48314724 | 0.59 |
ENSDART00000125617
|
smoc1
|
SPARC related modular calcium binding 1 |
| chr6_+_38880166 | 0.57 |
ENSDART00000019939
ENSDART00000144286 |
bin2b
|
bridging integrator 2b |
| chr22_-_24248420 | 0.53 |
ENSDART00000165433
|
rgs2
|
regulator of G protein signaling 2 |
| chr3_+_24207243 | 0.51 |
ENSDART00000023454
ENSDART00000136400 |
adsl
|
adenylosuccinate lyase |
| chr6_+_38879961 | 0.48 |
ENSDART00000184798
|
bin2b
|
bridging integrator 2b |
| chr7_-_24875421 | 0.47 |
ENSDART00000173920
|
adad2
|
adenosine deaminase domain containing 2 |
| chr19_+_7292654 | 0.47 |
ENSDART00000140459
|
ccdc127b
|
coiled-coil domain containing 127b |
| chr11_-_3987885 | 0.47 |
ENSDART00000058735
|
glt8d1
|
glycosyltransferase 8 domain containing 1 |
| chr4_-_20314749 | 0.46 |
ENSDART00000066894
ENSDART00000188123 |
dcp1b
|
decapping mRNA 1B |
| chr2_-_59345920 | 0.46 |
ENSDART00000134662
|
ftr37
|
finTRIM family, member 37 |
| chr8_-_44298964 | 0.45 |
ENSDART00000098520
|
fzd10
|
frizzled class receptor 10 |
| chr1_+_57040472 | 0.44 |
ENSDART00000181365
|
si:ch211-1f22.16
|
si:ch211-1f22.16 |
| chr7_+_24390939 | 0.42 |
ENSDART00000087494
ENSDART00000125463 |
haus3
|
HAUS augmin-like complex, subunit 3 |
| chr3_-_23461954 | 0.41 |
ENSDART00000040065
|
casc3
|
cancer susceptibility candidate 3 |
| chr1_+_27977297 | 0.40 |
ENSDART00000180692
ENSDART00000166819 |
sugt1
|
SGT1 homolog, MIS12 kinetochore complex assembly cochaperone |
| chr3_-_33422738 | 0.40 |
ENSDART00000075493
|
ccdc103
|
coiled-coil domain containing 103 |
| chr7_+_16991711 | 0.39 |
ENSDART00000173660
|
nav2a
|
neuron navigator 2a |
| chr3_+_22335030 | 0.39 |
ENSDART00000055676
|
zgc:103564
|
zgc:103564 |
| chr25_-_28674739 | 0.37 |
ENSDART00000067073
|
lrrc10
|
leucine rich repeat containing 10 |
| chr21_+_34088377 | 0.37 |
ENSDART00000170070
|
mtmr1b
|
myotubularin related protein 1b |
| chr8_-_20243389 | 0.36 |
ENSDART00000184904
|
acer1
|
alkaline ceramidase 1 |
| chr3_+_17933132 | 0.36 |
ENSDART00000104299
ENSDART00000162144 ENSDART00000162242 ENSDART00000166289 ENSDART00000171101 ENSDART00000164853 |
cnp
|
2',3'-cyclic nucleotide 3' phosphodiesterase |
| chr17_+_23975762 | 0.35 |
ENSDART00000155941
|
xpo1b
|
exportin 1 (CRM1 homolog, yeast) b |
| chr6_+_52931841 | 0.35 |
ENSDART00000174358
|
si:dkeyp-3f10.12
|
si:dkeyp-3f10.12 |
| chr3_+_39600562 | 0.34 |
ENSDART00000134309
ENSDART00000007170 |
prpsap2
|
phosphoribosyl pyrophosphate synthetase-associated protein 2 |
| chr2_-_37401600 | 0.33 |
ENSDART00000015723
|
prkci
|
protein kinase C, iota |
| chr15_-_23482088 | 0.33 |
ENSDART00000185823
ENSDART00000185523 |
nlrx1
|
NLR family member X1 |
| chr11_-_24063196 | 0.33 |
ENSDART00000036513
|
trib3
|
tribbles pseudokinase 3 |
| chr7_-_30639385 | 0.32 |
ENSDART00000173618
|
myo1ea
|
myosin IE, a |
| chr2_+_26237322 | 0.30 |
ENSDART00000030520
|
palm1b
|
paralemmin 1b |
| chr23_-_33558161 | 0.29 |
ENSDART00000018301
|
itga5
|
integrin, alpha 5 (fibronectin receptor, alpha polypeptide) |
| chr4_-_9054947 | 0.29 |
ENSDART00000109764
|
si:dkey-48p11.3
|
si:dkey-48p11.3 |
| chr18_+_44631789 | 0.28 |
ENSDART00000144271
|
bicra
|
BRD4 interacting chromatin remodeling complex associated protein |
| chr20_-_38827623 | 0.26 |
ENSDART00000153310
|
cad
|
carbamoyl-phosphate synthetase 2, aspartate transcarbamylase, and dihydroorotase |
| chr3_+_32118670 | 0.26 |
ENSDART00000055287
ENSDART00000111688 |
zgc:109934
|
zgc:109934 |
| chr3_-_7464250 | 0.24 |
ENSDART00000159873
|
znf1001
|
zinc finger protein 1001 |
| chr20_+_46255057 | 0.24 |
ENSDART00000100536
|
taar14i
|
trace amine associated receptor 14i |
| chr11_-_8271374 | 0.24 |
ENSDART00000168253
|
pimr202
|
Pim proto-oncogene, serine/threonine kinase, related 202 |
| chr4_+_76748500 | 0.24 |
ENSDART00000075607
|
ms4a17a.10
|
membrane-spanning 4-domains, subfamily A, member 17A.10 |
| chr13_-_36579086 | 0.24 |
ENSDART00000146671
|
lgals3a
|
lectin, galactoside binding soluble 3a |
| chr1_+_16621345 | 0.24 |
ENSDART00000149026
|
pcm1
|
pericentriolar material 1 |
| chr11_+_25596038 | 0.23 |
ENSDART00000140856
|
ccdc120
|
coiled-coil domain containing 120 |
| chr17_-_12249990 | 0.23 |
ENSDART00000177889
ENSDART00000155545 |
ahctf1
|
AT hook containing transcription factor 1 |
| chr4_+_37406676 | 0.22 |
ENSDART00000130981
|
si:ch73-134f24.1
|
si:ch73-134f24.1 |
| chr2_+_314249 | 0.21 |
ENSDART00000082086
|
zgc:113452
|
zgc:113452 |
| chr6_-_18531760 | 0.21 |
ENSDART00000167167
|
utp6
|
UTP6, small subunit (SSU) processome component, homolog (yeast) |
| chr17_+_24597001 | 0.21 |
ENSDART00000191834
|
rlf
|
rearranged L-myc fusion |
| chr7_-_3894831 | 0.20 |
ENSDART00000172921
|
si:dkey-88n24.11
|
si:dkey-88n24.11 |
| chr22_+_38762693 | 0.19 |
ENSDART00000015016
ENSDART00000150187 |
alpi.1
|
alkaline phosphatase, intestinal, tandem duplicate 1 |
| chr6_+_18531932 | 0.19 |
ENSDART00000165271
|
suz12b
|
SUZ12 polycomb repressive complex 2b subunit |
| chr15_-_31357634 | 0.19 |
ENSDART00000127485
|
or111-2
|
odorant receptor, family D, subfamily 111, member 2 |
| chr14_-_11507211 | 0.19 |
ENSDART00000186873
ENSDART00000109181 ENSDART00000186166 ENSDART00000186986 |
zgc:174917
|
zgc:174917 |
| chr14_+_10656975 | 0.19 |
ENSDART00000127594
ENSDART00000125865 |
atrx
|
alpha thalassemia/mental retardation syndrome X-linked homolog (human) |
| chr19_+_7292445 | 0.18 |
ENSDART00000026634
|
ccdc127b
|
coiled-coil domain containing 127b |
| chr14_-_33478963 | 0.17 |
ENSDART00000132813
|
lamp2
|
lysosomal-associated membrane protein 2 |
| chr1_+_46026457 | 0.17 |
ENSDART00000132705
|
si:ch211-138g9.2
|
si:ch211-138g9.2 |
| chr18_+_808911 | 0.17 |
ENSDART00000172518
|
cox5ab
|
cytochrome c oxidase subunit Vab |
| chr14_-_16476863 | 0.16 |
ENSDART00000089021
|
canx
|
calnexin |
| chr15_-_29012493 | 0.16 |
ENSDART00000060018
|
dharma
|
dharma |
| chr11_+_12879635 | 0.16 |
ENSDART00000182515
ENSDART00000081296 |
si:dkey-11m19.5
|
si:dkey-11m19.5 |
| chr1_-_23294753 | 0.16 |
ENSDART00000013263
|
ugdh
|
UDP-glucose 6-dehydrogenase |
| chr4_-_3805992 | 0.16 |
ENSDART00000190125
|
si:dkey-61f9.1
|
si:dkey-61f9.1 |
| chr22_+_39096911 | 0.15 |
ENSDART00000157127
ENSDART00000153841 |
lmcd1
|
LIM and cysteine-rich domains 1 |
| chr12_-_19862912 | 0.15 |
ENSDART00000145788
|
shisa9a
|
shisa family member 9a |
| chr25_-_17590971 | 0.14 |
ENSDART00000189942
|
mmp15a
|
matrix metallopeptidase 15a |
| chr19_-_24224142 | 0.14 |
ENSDART00000136409
ENSDART00000114390 |
prf1.8
|
perforin 1.8 |
| chr13_+_46927350 | 0.13 |
ENSDART00000165041
ENSDART00000167931 |
mtrf1l
|
mitochondrial translational release factor 1-like |
| chr7_-_48733662 | 0.12 |
ENSDART00000191675
|
traf6
|
TNF receptor-associated factor 6 |
| chr25_-_20691075 | 0.11 |
ENSDART00000067373
|
edc3
|
enhancer of mRNA decapping 3 homolog (S. cerevisiae) |
| chr25_+_25124684 | 0.11 |
ENSDART00000167542
|
ldha
|
lactate dehydrogenase A4 |
| chr1_+_54835131 | 0.10 |
ENSDART00000145070
|
si:ch211-196h16.4
|
si:ch211-196h16.4 |
| chr15_-_4053149 | 0.10 |
ENSDART00000189076
|
CR626884.4
|
|
| chr4_-_50926767 | 0.10 |
ENSDART00000183430
|
si:ch211-208f21.3
|
si:ch211-208f21.3 |
| chr18_-_22701800 | 0.10 |
ENSDART00000135098
|
si:ch73-113g13.3
|
si:ch73-113g13.3 |
| chr13_+_27040887 | 0.09 |
ENSDART00000132714
|
soul2
|
heme-binding protein soul2 |
| chr11_-_36040549 | 0.09 |
ENSDART00000112684
|
setmar
|
SET domain and mariner transposase fusion gene |
| chr11_-_8226088 | 0.09 |
ENSDART00000173364
|
si:cabz01021067.1
|
si:cabz01021067.1 |
| chr12_-_30583668 | 0.09 |
ENSDART00000153406
|
casp7
|
caspase 7, apoptosis-related cysteine peptidase |
| chr1_-_58000438 | 0.08 |
ENSDART00000163761
|
si:ch211-114l13.9
|
si:ch211-114l13.9 |
| chr21_+_45502621 | 0.08 |
ENSDART00000166719
|
si:dkey-223p19.2
|
si:dkey-223p19.2 |
| chr13_-_10727550 | 0.08 |
ENSDART00000190925
|
ppm1ba
|
protein phosphatase, Mg2+/Mn2+ dependent, 1Ba |
| chr25_+_19485198 | 0.07 |
ENSDART00000156730
|
glsl
|
glutaminase like |
| chr1_+_55662491 | 0.07 |
ENSDART00000152386
|
adgre8
|
adhesion G protein-coupled receptor E8 |
| chr1_+_55583116 | 0.07 |
ENSDART00000152163
|
adgre19
|
adhesion G protein-coupled receptor E19 |
| chr12_+_20506197 | 0.06 |
ENSDART00000153010
|
si:zfos-754c12.2
|
si:zfos-754c12.2 |
| chr5_-_35456269 | 0.06 |
ENSDART00000051312
|
ttc33
|
tetratricopeptide repeat domain 33 |
| chr8_-_36469117 | 0.05 |
ENSDART00000111240
|
mhc2dab
|
major histocompatibility complex class II DAB gene |
| chr4_+_76403698 | 0.05 |
ENSDART00000184821
ENSDART00000169373 |
FP074874.2
FP074874.1
|
|
| chr11_+_583725 | 0.04 |
ENSDART00000189415
|
mkrn2os.2
|
MKRN2 opposite strand, tandem duplicate 2 |
| chr12_+_30788912 | 0.04 |
ENSDART00000160422
|
aldh18a1
|
aldehyde dehydrogenase 18 family, member A1 |
| chr7_-_59123066 | 0.04 |
ENSDART00000175438
|
dennd4c
|
DENN/MADD domain containing 4C |
| chr7_+_48555626 | 0.04 |
ENSDART00000125483
ENSDART00000083514 |
kcnq1
|
potassium voltage-gated channel, KQT-like subfamily, member 1 |
| chr4_-_17838179 | 0.04 |
ENSDART00000146931
|
avpr2l
|
arginine vasopressin receptor 2, like |
| chr9_-_7089303 | 0.04 |
ENSDART00000146609
|
coa5
|
cytochrome C oxidase assembly factor 5 |
| chr2_-_11119303 | 0.03 |
ENSDART00000135450
ENSDART00000131836 |
cryz
|
crystallin, zeta (quinone reductase) |
| chr11_-_5939861 | 0.03 |
ENSDART00000110033
|
abhd8b
|
abhydrolase domain containing 8b |
| chr21_-_2348838 | 0.02 |
ENSDART00000160337
|
si:ch73-299h12.8
|
si:ch73-299h12.8 |
| chr11_+_37909654 | 0.01 |
ENSDART00000172211
|
si:ch211-112f3.4
|
si:ch211-112f3.4 |
Network of associatons between targets according to the STRING database.
First level regulatory network of nkx2.1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.6 | GO:0046831 | regulation of nucleobase-containing compound transport(GO:0032239) positive regulation of nucleobase-containing compound transport(GO:0032241) positive regulation of nucleocytoplasmic transport(GO:0046824) regulation of RNA export from nucleus(GO:0046831) positive regulation of RNA export from nucleus(GO:0046833) messenger ribonucleoprotein complex assembly(GO:1990120) |
| 0.2 | 1.1 | GO:0071800 | podosome assembly(GO:0071800) |
| 0.1 | 0.4 | GO:0036159 | inner dynein arm assembly(GO:0036159) |
| 0.1 | 0.5 | GO:0090299 | regulation of neural crest formation(GO:0090299) |
| 0.1 | 0.6 | GO:0031087 | nuclear-transcribed mRNA catabolic process, deadenylation-independent decay(GO:0031086) deadenylation-independent decapping of nuclear-transcribed mRNA(GO:0031087) |
| 0.1 | 0.8 | GO:0007080 | mitotic metaphase plate congression(GO:0007080) |
| 0.1 | 0.3 | GO:0021563 | glossopharyngeal nerve development(GO:0021563) |
| 0.1 | 0.3 | GO:0046391 | 5-phosphoribose 1-diphosphate biosynthetic process(GO:0006015) 5-phosphoribose 1-diphosphate metabolic process(GO:0046391) |
| 0.1 | 0.3 | GO:0045217 | establishment or maintenance of polarity of embryonic epithelium(GO:0016332) cell-cell junction maintenance(GO:0045217) |
| 0.1 | 0.2 | GO:0048320 | axial mesoderm formation(GO:0048320) |
| 0.1 | 0.5 | GO:0006167 | AMP biosynthetic process(GO:0006167) |
| 0.0 | 1.0 | GO:0021508 | floor plate formation(GO:0021508) floor plate morphogenesis(GO:0033505) |
| 0.0 | 0.2 | GO:0048245 | eosinophil chemotaxis(GO:0048245) eosinophil migration(GO:0072677) |
| 0.0 | 0.2 | GO:0034454 | microtubule anchoring at centrosome(GO:0034454) |
| 0.0 | 0.3 | GO:0090245 | axis elongation involved in somitogenesis(GO:0090245) |
| 0.0 | 0.2 | GO:0051292 | nuclear pore complex assembly(GO:0051292) |
| 0.0 | 0.1 | GO:0070126 | mitochondrial translational termination(GO:0070126) |
| 0.0 | 0.5 | GO:0006382 | adenosine to inosine editing(GO:0006382) |
| 0.0 | 0.3 | GO:0000056 | ribosomal small subunit export from nucleus(GO:0000056) |
| 0.0 | 0.8 | GO:0034625 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.0 | 0.4 | GO:0034312 | diol biosynthetic process(GO:0034312) sphingosine biosynthetic process(GO:0046512) |
| 0.0 | 0.4 | GO:0061026 | cardiac muscle tissue regeneration(GO:0061026) |
| 0.0 | 0.6 | GO:0033344 | cholesterol efflux(GO:0033344) |
| 0.0 | 0.2 | GO:0001561 | fatty acid alpha-oxidation(GO:0001561) |
| 0.0 | 0.2 | GO:0003190 | heart valve formation(GO:0003188) atrioventricular valve formation(GO:0003190) |
| 0.0 | 0.4 | GO:0043124 | negative regulation of I-kappaB kinase/NF-kappaB signaling(GO:0043124) |
| 0.0 | 0.2 | GO:0048172 | regulation of short-term neuronal synaptic plasticity(GO:0048172) |
| 0.0 | 0.4 | GO:0000184 | nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:0000184) |
| 0.0 | 0.2 | GO:0006622 | protein targeting to lysosome(GO:0006622) |
| 0.0 | 0.1 | GO:0070498 | interleukin-1-mediated signaling pathway(GO:0070498) |
| 0.0 | 0.6 | GO:0002072 | optic cup morphogenesis involved in camera-type eye development(GO:0002072) |
| 0.0 | 0.2 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
| 0.0 | 0.2 | GO:0007198 | adenylate cyclase-inhibiting serotonin receptor signaling pathway(GO:0007198) |
| 0.0 | 0.4 | GO:0050821 | protein stabilization(GO:0050821) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 0.8 | GO:0097125 | cyclin B1-CDK1 complex(GO:0097125) |
| 0.2 | 0.6 | GO:0042382 | paraspeckles(GO:0042382) |
| 0.2 | 1.1 | GO:0001891 | phagocytic cup(GO:0001891) |
| 0.1 | 0.3 | GO:0002189 | ribose phosphate diphosphokinase complex(GO:0002189) |
| 0.1 | 0.4 | GO:0071012 | U2-type catalytic step 1 spliceosome(GO:0071006) catalytic step 1 spliceosome(GO:0071012) |
| 0.1 | 0.2 | GO:0031166 | integral component of vacuolar membrane(GO:0031166) intrinsic component of vacuolar membrane(GO:0031310) |
| 0.1 | 0.3 | GO:0005915 | cell-cell adherens junction(GO:0005913) zonula adherens(GO:0005915) |
| 0.0 | 0.4 | GO:0070652 | HAUS complex(GO:0070652) |
| 0.0 | 0.4 | GO:0036157 | outer dynein arm(GO:0036157) |
| 0.0 | 0.2 | GO:0034388 | Pwp2p-containing subcomplex of 90S preribosome(GO:0034388) |
| 0.0 | 0.2 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.0 | 0.2 | GO:0036156 | inner dynein arm(GO:0036156) |
| 0.0 | 0.2 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.0 | 0.2 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.0 | 0.2 | GO:0005751 | mitochondrial respiratory chain complex IV(GO:0005751) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.5 | GO:0016842 | amidine-lyase activity(GO:0016842) |
| 0.1 | 0.3 | GO:0031434 | mitogen-activated protein kinase kinase binding(GO:0031434) |
| 0.1 | 0.3 | GO:0016743 | carboxyl- or carbamoyltransferase activity(GO:0016743) |
| 0.1 | 0.3 | GO:0004113 | 2',3'-cyclic-nucleotide 3'-phosphodiesterase activity(GO:0004113) |
| 0.1 | 0.3 | GO:0004749 | ribose phosphate diphosphokinase activity(GO:0004749) |
| 0.1 | 1.0 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.1 | 0.4 | GO:0017040 | ceramidase activity(GO:0017040) |
| 0.0 | 0.2 | GO:0019865 | IgE binding(GO:0019863) immunoglobulin binding(GO:0019865) |
| 0.0 | 0.5 | GO:0003726 | double-stranded RNA adenosine deaminase activity(GO:0003726) |
| 0.0 | 0.2 | GO:0003979 | UDP-glucose 6-dehydrogenase activity(GO:0003979) |
| 0.0 | 0.8 | GO:0009922 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.0 | 0.1 | GO:1990174 | phosphodiesterase decapping endonuclease activity(GO:1990174) |
| 0.0 | 0.4 | GO:0052629 | phosphatidylinositol-3,5-bisphosphate 3-phosphatase activity(GO:0052629) |
| 0.0 | 0.3 | GO:0005049 | nuclear export signal receptor activity(GO:0005049) |
| 0.0 | 0.4 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.0 | 0.2 | GO:0004035 | alkaline phosphatase activity(GO:0004035) |
| 0.0 | 0.3 | GO:0001968 | fibronectin binding(GO:0001968) |
| 0.0 | 0.6 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.0 | 0.1 | GO:0004459 | L-lactate dehydrogenase activity(GO:0004459) |
| 0.0 | 0.8 | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity(GO:0016538) |
| 0.0 | 0.1 | GO:0003747 | translation release factor activity(GO:0003747) translation termination factor activity(GO:0008079) |
| 0.0 | 0.3 | GO:0004698 | protein kinase C activity(GO:0004697) calcium-dependent protein kinase C activity(GO:0004698) |
| 0.0 | 0.2 | GO:0030515 | snoRNA binding(GO:0030515) |
| 0.0 | 0.0 | GO:0003960 | NADPH:quinone reductase activity(GO:0003960) |
| 0.0 | 0.2 | GO:0051378 | amine binding(GO:0043176) serotonin binding(GO:0051378) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
| 0.0 | 0.8 | PID FOXM1 PATHWAY | FOXM1 transcription factor network |
| 0.0 | 0.3 | PID SYNDECAN 2 PATHWAY | Syndecan-2-mediated signaling events |
| 0.0 | 0.3 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.8 | REACTOME RECRUITMENT OF NUMA TO MITOTIC CENTROSOMES | Genes involved in Recruitment of NuMA to mitotic centrosomes |
| 0.1 | 0.3 | REACTOME NEGATIVE REGULATION OF THE PI3K AKT NETWORK | Genes involved in Negative regulation of the PI3K/AKT network |
| 0.0 | 0.5 | REACTOME P75NTR RECRUITS SIGNALLING COMPLEXES | Genes involved in p75NTR recruits signalling complexes |
| 0.0 | 0.5 | REACTOME PURINE RIBONUCLEOSIDE MONOPHOSPHATE BIOSYNTHESIS | Genes involved in Purine ribonucleoside monophosphate biosynthesis |
| 0.0 | 0.6 | REACTOME MRNA DECAY BY 5 TO 3 EXORIBONUCLEASE | Genes involved in mRNA Decay by 5' to 3' Exoribonuclease |
| 0.0 | 0.6 | REACTOME SIGNALING BY FGFR1 FUSION MUTANTS | Genes involved in Signaling by FGFR1 fusion mutants |
| 0.0 | 0.4 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.0 | 0.3 | REACTOME PYRIMIDINE METABOLISM | Genes involved in Pyrimidine metabolism |
| 0.0 | 0.3 | REACTOME NEGATIVE REGULATORS OF RIG I MDA5 SIGNALING | Genes involved in Negative regulators of RIG-I/MDA5 signaling |