Project
PRJEB1986: zebrafish developmental stages transcriptome
Navigation
Downloads
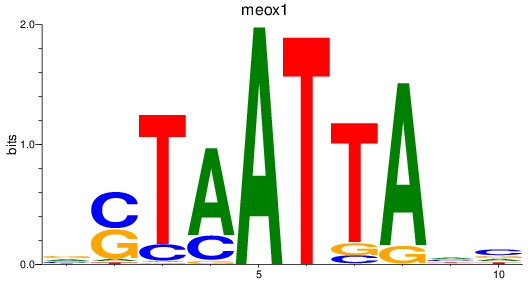
Results for meox1
Z-value: 0.63
Transcription factors associated with meox1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
meox1
|
ENSDARG00000007891 | mesenchyme homeobox 1 |
|
meox1
|
ENSDARG00000115382 | mesenchyme homeobox 1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| meox1 | dr11_v1_chr12_+_27462225_27462225 | -0.70 | 7.5e-04 | Click! |
Activity profile of meox1 motif
Sorted Z-values of meox1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr1_+_17676745 | 2.81 |
ENSDART00000030665
|
slc25a4
|
solute carrier family 25 (mitochondrial carrier; adenine nucleotide translocator), member 4 |
| chr11_+_30244356 | 2.58 |
ENSDART00000036050
ENSDART00000150080 |
rs1a
|
retinoschisin 1a |
| chr3_-_50443607 | 2.44 |
ENSDART00000074036
|
rcvrna
|
recoverin a |
| chr25_+_29160102 | 2.38 |
ENSDART00000162854
|
pkmb
|
pyruvate kinase M1/2b |
| chr16_+_5774977 | 2.31 |
ENSDART00000134202
|
ccka
|
cholecystokinin a |
| chr19_+_40856534 | 2.28 |
ENSDART00000051950
|
gngt1
|
guanine nucleotide binding protein (G protein), gamma transducing activity polypeptide 1 |
| chr19_+_40856807 | 2.09 |
ENSDART00000139083
|
gngt1
|
guanine nucleotide binding protein (G protein), gamma transducing activity polypeptide 1 |
| chr1_-_23308225 | 1.77 |
ENSDART00000137567
ENSDART00000008201 |
smim14
|
small integral membrane protein 14 |
| chr17_-_37395460 | 1.68 |
ENSDART00000148160
ENSDART00000075975 |
crip1
|
cysteine-rich protein 1 |
| chr1_-_50859053 | 1.63 |
ENSDART00000132779
ENSDART00000137648 |
si:dkeyp-123h10.2
|
si:dkeyp-123h10.2 |
| chr20_-_40755614 | 1.60 |
ENSDART00000061247
|
cx32.3
|
connexin 32.3 |
| chr2_+_50608099 | 1.55 |
ENSDART00000185805
ENSDART00000111135 |
neurod6b
|
neuronal differentiation 6b |
| chr3_-_31079186 | 1.54 |
ENSDART00000145636
ENSDART00000140569 |
ELOB (1 of many)
elob
|
elongin B elongin B |
| chr15_+_47903864 | 1.31 |
ENSDART00000063835
|
otx5
|
orthodenticle homolog 5 |
| chr20_-_9462433 | 1.29 |
ENSDART00000152674
ENSDART00000040557 |
zgc:101840
|
zgc:101840 |
| chr5_-_38155005 | 1.22 |
ENSDART00000097770
|
gucy2d
|
guanylate cyclase 2D, retinal |
| chr16_+_31804590 | 1.21 |
ENSDART00000167321
|
wnt4b
|
wingless-type MMTV integration site family, member 4b |
| chr16_+_46111849 | 1.19 |
ENSDART00000172232
|
sv2a
|
synaptic vesicle glycoprotein 2A |
| chr8_-_23780334 | 1.17 |
ENSDART00000145179
ENSDART00000145894 |
zgc:195245
|
zgc:195245 |
| chr17_-_28198099 | 1.11 |
ENSDART00000156143
|
htr1d
|
5-hydroxytryptamine (serotonin) receptor 1D, G protein-coupled |
| chr6_-_35472923 | 1.10 |
ENSDART00000185907
|
rgs8
|
regulator of G protein signaling 8 |
| chr23_+_40460333 | 1.05 |
ENSDART00000184658
|
soga3b
|
SOGA family member 3b |
| chr7_-_27685365 | 1.03 |
ENSDART00000188342
|
calca
|
calcitonin/calcitonin-related polypeptide, alpha |
| chr20_+_4060839 | 1.00 |
ENSDART00000178565
|
TRIM67
|
tripartite motif containing 67 |
| chr20_-_9436521 | 1.00 |
ENSDART00000133000
|
zgc:101840
|
zgc:101840 |
| chr18_-_1185772 | 0.98 |
ENSDART00000143245
|
nptnb
|
neuroplastin b |
| chr19_-_5103141 | 0.96 |
ENSDART00000150952
|
tpi1a
|
triosephosphate isomerase 1a |
| chr25_+_3326885 | 0.95 |
ENSDART00000104866
|
ldhbb
|
lactate dehydrogenase Bb |
| chr24_-_41320037 | 0.95 |
ENSDART00000129058
|
rheb
|
Ras homolog, mTORC1 binding |
| chr22_+_17205608 | 0.94 |
ENSDART00000181267
|
rab3b
|
RAB3B, member RAS oncogene family |
| chr2_-_33645411 | 0.94 |
ENSDART00000114663
|
b4galt2
|
UDP-Gal:betaGlcNAc beta 1,4- galactosyltransferase, polypeptide 2 |
| chr16_+_17389116 | 0.92 |
ENSDART00000103750
ENSDART00000173448 |
fam131bb
|
family with sequence similarity 131, member Bb |
| chr4_+_21129752 | 0.92 |
ENSDART00000169764
|
syt1a
|
synaptotagmin Ia |
| chr14_-_30587814 | 0.90 |
ENSDART00000144912
ENSDART00000149714 |
tmem265
|
transmembrane protein 265 |
| chr7_+_30787903 | 0.90 |
ENSDART00000174000
|
apba2b
|
amyloid beta (A4) precursor protein-binding, family A, member 2b |
| chr22_+_20427170 | 0.89 |
ENSDART00000136744
|
foxq2
|
forkhead box Q2 |
| chr18_+_1154189 | 0.85 |
ENSDART00000135090
|
si:ch1073-75f15.2
|
si:ch1073-75f15.2 |
| chr4_+_12612723 | 0.84 |
ENSDART00000133767
|
lmo3
|
LIM domain only 3 |
| chr13_+_33117528 | 0.83 |
ENSDART00000085719
|
si:ch211-10a23.2
|
si:ch211-10a23.2 |
| chr3_+_33341640 | 0.83 |
ENSDART00000186352
|
pyya
|
peptide YYa |
| chr2_-_15324837 | 0.83 |
ENSDART00000015655
|
tecrl2b
|
trans-2,3-enoyl-CoA reductase-like 2b |
| chr9_-_44295071 | 0.83 |
ENSDART00000011837
|
neurod1
|
neuronal differentiation 1 |
| chr24_+_16547035 | 0.81 |
ENSDART00000164319
|
sema5a
|
sema domain, seven thrombospondin repeats (type 1 and type 1-like), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 5A |
| chr10_+_29698467 | 0.80 |
ENSDART00000163402
|
dlg2
|
discs, large homolog 2 (Drosophila) |
| chr3_-_61162750 | 0.80 |
ENSDART00000055064
|
pvalb8
|
parvalbumin 8 |
| chr15_+_21276735 | 0.80 |
ENSDART00000111213
|
ubash3bb
|
ubiquitin associated and SH3 domain containing Bb |
| chr21_-_22827548 | 0.80 |
ENSDART00000079161
|
angptl5
|
angiopoietin-like 5 |
| chr8_-_30338872 | 0.79 |
ENSDART00000137583
|
dock8
|
dedicator of cytokinesis 8 |
| chr18_+_8833251 | 0.78 |
ENSDART00000143519
|
impdh1a
|
IMP (inosine 5'-monophosphate) dehydrogenase 1a |
| chr4_+_11384891 | 0.77 |
ENSDART00000092381
ENSDART00000186577 ENSDART00000191054 ENSDART00000191584 |
pcloa
|
piccolo presynaptic cytomatrix protein a |
| chr13_+_25449681 | 0.75 |
ENSDART00000101328
|
atoh7
|
atonal bHLH transcription factor 7 |
| chr11_+_38280454 | 0.74 |
ENSDART00000171496
|
CDK18
|
si:dkey-166c18.1 |
| chr9_+_11532025 | 0.74 |
ENSDART00000109037
|
cdk5r2b
|
cyclin-dependent kinase 5, regulatory subunit 2b (p39) |
| chr9_-_43538328 | 0.73 |
ENSDART00000140526
|
znf385b
|
zinc finger protein 385B |
| chr5_+_38263240 | 0.73 |
ENSDART00000051231
|
gnb2
|
guanine nucleotide binding protein (G protein), beta polypeptide 2 |
| chr14_-_858985 | 0.73 |
ENSDART00000148687
ENSDART00000149375 |
slc34a1a
|
solute carrier family 34 (type II sodium/phosphate cotransporter), member 1a |
| chr15_-_36533322 | 0.72 |
ENSDART00000156466
ENSDART00000121755 |
si:dkey-262k9.4
|
si:dkey-262k9.4 |
| chr7_-_28148310 | 0.71 |
ENSDART00000044208
|
lmo1
|
LIM domain only 1 |
| chr2_-_28671139 | 0.70 |
ENSDART00000165272
ENSDART00000164657 |
dhcr7
|
7-dehydrocholesterol reductase |
| chr13_-_29420885 | 0.70 |
ENSDART00000024225
|
chata
|
choline O-acetyltransferase a |
| chr11_+_28476298 | 0.69 |
ENSDART00000122319
|
lrrc38b
|
leucine rich repeat containing 38b |
| chr25_+_3327071 | 0.68 |
ENSDART00000136131
ENSDART00000133243 |
ldhbb
|
lactate dehydrogenase Bb |
| chr23_+_28731379 | 0.67 |
ENSDART00000047378
|
cort
|
cortistatin |
| chr21_+_11244068 | 0.67 |
ENSDART00000163432
|
arid6
|
AT-rich interaction domain 6 |
| chr24_-_28648949 | 0.66 |
ENSDART00000180227
|
tecrl2a
|
trans-2,3-enoyl-CoA reductase-like 2a |
| chr1_+_40023640 | 0.66 |
ENSDART00000101623
|
lgi2b
|
leucine-rich repeat LGI family, member 2b |
| chr19_-_32641725 | 0.64 |
ENSDART00000165006
ENSDART00000188185 |
hpca
|
hippocalcin |
| chr21_-_37790727 | 0.64 |
ENSDART00000162907
|
gabrb4
|
gamma-aminobutyric acid (GABA) A receptor, beta 4 |
| chr20_-_14925281 | 0.63 |
ENSDART00000152641
|
dnm3a
|
dynamin 3a |
| chr6_-_55585423 | 0.61 |
ENSDART00000157129
|
slc12a5a
|
solute carrier family 12 (potassium/chloride transporter), member 5a |
| chr7_+_21841037 | 0.61 |
ENSDART00000077503
|
tm4sf5
|
transmembrane 4 L six family member 5 |
| chr19_-_5103313 | 0.61 |
ENSDART00000037007
|
tpi1a
|
triosephosphate isomerase 1a |
| chr5_+_64732270 | 0.60 |
ENSDART00000134241
|
olfm1a
|
olfactomedin 1a |
| chr15_-_23376541 | 0.59 |
ENSDART00000078570
|
c1qtnf5
|
C1q and TNF related 5 |
| chr1_-_10647484 | 0.59 |
ENSDART00000164541
ENSDART00000188958 ENSDART00000190904 |
si:dkey-31e10.1
|
si:dkey-31e10.1 |
| chr22_-_11137268 | 0.58 |
ENSDART00000178882
|
atp6ap2
|
ATPase H+ transporting accessory protein 2 |
| chr21_-_7035599 | 0.58 |
ENSDART00000139777
|
si:ch211-93g21.1
|
si:ch211-93g21.1 |
| chr3_-_49110710 | 0.58 |
ENSDART00000160404
|
trim35-12
|
tripartite motif containing 35-12 |
| chr15_+_44366556 | 0.57 |
ENSDART00000133449
|
GUCY1A2
|
guanylate cyclase 1 soluble subunit alpha 2 |
| chr21_-_27185915 | 0.57 |
ENSDART00000135052
|
slc8a4a
|
solute carrier family 8 (sodium/calcium exchanger), member 4a |
| chr23_+_20110086 | 0.56 |
ENSDART00000054664
|
tnnc1b
|
troponin C type 1b (slow) |
| chr1_+_16397063 | 0.56 |
ENSDART00000159794
|
micu3a
|
mitochondrial calcium uptake family, member 3a |
| chr1_-_50590626 | 0.56 |
ENSDART00000146892
ENSDART00000022733 |
abcg2d
|
ATP-binding cassette, sub-family G (WHITE), member 2d |
| chr3_+_34919810 | 0.56 |
ENSDART00000055264
|
ca10b
|
carbonic anhydrase Xb |
| chr12_-_20795867 | 0.56 |
ENSDART00000152835
ENSDART00000153424 |
nme2a
|
NME/NM23 nucleoside diphosphate kinase 2a |
| chr20_+_52442870 | 0.55 |
ENSDART00000163100
|
arhgap39
|
Rho GTPase activating protein 39 |
| chr18_+_9615147 | 0.55 |
ENSDART00000160284
|
pclob
|
piccolo presynaptic cytomatrix protein b |
| chr18_+_9637744 | 0.55 |
ENSDART00000190171
|
pclob
|
piccolo presynaptic cytomatrix protein b |
| chr20_-_46362606 | 0.53 |
ENSDART00000153087
|
bmf2
|
BCL2 modifying factor 2 |
| chr10_-_34772211 | 0.51 |
ENSDART00000145450
ENSDART00000134307 |
dclk1a
|
doublecortin-like kinase 1a |
| chr14_+_25817628 | 0.51 |
ENSDART00000047680
|
glra1
|
glycine receptor, alpha 1 |
| chr3_-_60827402 | 0.50 |
ENSDART00000053494
|
anks4b
|
ankyrin repeat and sterile alpha motif domain containing 4B |
| chr15_-_34785594 | 0.50 |
ENSDART00000154256
|
gabbr1a
|
gamma-aminobutyric acid (GABA) B receptor, 1a |
| chr20_-_35040041 | 0.49 |
ENSDART00000131919
|
kif26bb
|
kinesin family member 26Bb |
| chr21_+_1381276 | 0.49 |
ENSDART00000192907
|
TCF4
|
transcription factor 4 |
| chr11_+_45323455 | 0.47 |
ENSDART00000189249
|
mafgb
|
v-maf avian musculoaponeurotic fibrosarcoma oncogene homolog Gb |
| chr19_-_9662958 | 0.47 |
ENSDART00000041094
|
clcn1a
|
chloride channel, voltage-sensitive 1a |
| chr10_+_3049636 | 0.47 |
ENSDART00000081794
ENSDART00000183167 ENSDART00000191634 ENSDART00000183514 |
rasgrf2a
|
Ras protein-specific guanine nucleotide-releasing factor 2a |
| chr22_-_10121880 | 0.47 |
ENSDART00000002348
|
rdh5
|
retinol dehydrogenase 5 (11-cis/9-cis) |
| chr1_-_44701313 | 0.46 |
ENSDART00000193926
|
si:dkey-28b4.8
|
si:dkey-28b4.8 |
| chr19_+_9295244 | 0.46 |
ENSDART00000132255
ENSDART00000144299 |
si:ch73-15n24.1
|
si:ch73-15n24.1 |
| chr10_+_2582254 | 0.45 |
ENSDART00000016103
|
nxnl2
|
nucleoredoxin like 2 |
| chr2_-_11258547 | 0.45 |
ENSDART00000165803
ENSDART00000193817 |
slc44a5a
|
solute carrier family 44, member 5a |
| chr22_+_17828267 | 0.44 |
ENSDART00000136016
|
hapln4
|
hyaluronan and proteoglycan link protein 4 |
| chr19_-_20113696 | 0.44 |
ENSDART00000188813
|
npy
|
neuropeptide Y |
| chr19_-_27542277 | 0.44 |
ENSDART00000114301
|
si:ch211-152p11.4
|
si:ch211-152p11.4 |
| chr23_+_27740788 | 0.43 |
ENSDART00000053871
|
dhh
|
desert hedgehog |
| chr20_-_28800999 | 0.42 |
ENSDART00000049462
|
rab15
|
RAB15, member RAS oncogene family |
| chr21_+_28445052 | 0.42 |
ENSDART00000077871
|
pygma
|
phosphorylase, glycogen, muscle A |
| chr22_-_21897203 | 0.41 |
ENSDART00000158501
ENSDART00000105566 ENSDART00000136795 |
gna11a
|
guanine nucleotide binding protein (G protein), alpha 11a (Gq class) |
| chr20_+_23185530 | 0.40 |
ENSDART00000142346
|
lrrc66
|
leucine rich repeat containing 66 |
| chr4_-_15420452 | 0.40 |
ENSDART00000016230
|
plxna4
|
plexin A4 |
| chr17_+_11675362 | 0.39 |
ENSDART00000157911
|
kif26ba
|
kinesin family member 26Ba |
| chr5_-_51619742 | 0.39 |
ENSDART00000188537
|
otpb
|
orthopedia homeobox b |
| chr13_-_44630111 | 0.39 |
ENSDART00000110092
|
mdga1
|
MAM domain containing glycosylphosphatidylinositol anchor 1 |
| chr12_+_20352400 | 0.39 |
ENSDART00000066383
|
hbae5
|
hemoglobin, alpha embryonic 5 |
| chr20_-_14924858 | 0.39 |
ENSDART00000047039
|
dnm3a
|
dynamin 3a |
| chr15_-_38154616 | 0.38 |
ENSDART00000099392
|
irgq2
|
immunity-related GTPase family, q2 |
| chr7_+_54642005 | 0.38 |
ENSDART00000171864
|
fgf19
|
fibroblast growth factor 19 |
| chr16_-_26296477 | 0.38 |
ENSDART00000157553
|
erfl1
|
Ets2 repressor factor like 1 |
| chr15_+_7057050 | 0.38 |
ENSDART00000061828
|
foxl2a
|
forkhead box L2a |
| chr23_-_44219902 | 0.38 |
ENSDART00000185874
|
zgc:158659
|
zgc:158659 |
| chr3_+_17537352 | 0.37 |
ENSDART00000104549
|
hcrt
|
hypocretin (orexin) neuropeptide precursor |
| chr6_-_6487876 | 0.37 |
ENSDART00000137642
|
cep170ab
|
centrosomal protein 170Ab |
| chr5_+_56023186 | 0.37 |
ENSDART00000156230
|
fzd9a
|
frizzled class receptor 9a |
| chr6_-_24392426 | 0.36 |
ENSDART00000163965
|
brdt
|
bromodomain, testis-specific |
| chr15_+_9053059 | 0.35 |
ENSDART00000012039
|
ppm1na
|
protein phosphatase, Mg2+/Mn2+ dependent, 1Na (putative) |
| chr14_+_40874608 | 0.35 |
ENSDART00000168448
|
si:ch211-106m9.1
|
si:ch211-106m9.1 |
| chr12_-_33357655 | 0.35 |
ENSDART00000066233
ENSDART00000148165 |
slc16a3
|
solute carrier family 16 (monocarboxylate transporter), member 3 |
| chr11_-_2478374 | 0.35 |
ENSDART00000173205
|
si:ch73-267c23.10
|
si:ch73-267c23.10 |
| chr22_-_10459880 | 0.34 |
ENSDART00000064801
|
ogn
|
osteoglycin |
| chr18_+_11506561 | 0.34 |
ENSDART00000121647
|
PRMT8
|
protein arginine methyltransferase 8 |
| chr5_+_20147830 | 0.34 |
ENSDART00000098727
|
svopa
|
SV2 related protein a |
| chr20_+_37294112 | 0.34 |
ENSDART00000076293
ENSDART00000140450 |
cx23
|
connexin 23 |
| chr20_-_45060241 | 0.34 |
ENSDART00000185227
|
klhl29
|
kelch-like family member 29 |
| chr12_-_46112892 | 0.33 |
ENSDART00000187128
ENSDART00000114268 |
zgc:153932
|
zgc:153932 |
| chr4_-_28158335 | 0.33 |
ENSDART00000134605
|
gramd4a
|
GRAM domain containing 4a |
| chr5_+_30635309 | 0.33 |
ENSDART00000183769
|
abcg4a
|
ATP-binding cassette, sub-family G (WHITE), member 4a |
| chr19_-_657439 | 0.32 |
ENSDART00000167100
|
slc6a18
|
solute carrier family 6 (neutral amino acid transporter), member 18 |
| chr20_-_1378514 | 0.32 |
ENSDART00000181830
|
scara5
|
scavenger receptor class A, member 5 (putative) |
| chr4_+_74929427 | 0.32 |
ENSDART00000174082
|
nup50
|
nucleoporin 50 |
| chr14_-_17622080 | 0.32 |
ENSDART00000112149
|
si:ch211-159i8.4
|
si:ch211-159i8.4 |
| chr9_-_38021889 | 0.31 |
ENSDART00000183482
ENSDART00000124333 |
adcy5
|
adenylate cyclase 5 |
| chr19_-_205104 | 0.31 |
ENSDART00000011890
|
zbtb22a
|
zinc finger and BTB domain containing 22a |
| chr19_-_27542433 | 0.31 |
ENSDART00000136414
|
si:ch211-152p11.4
|
si:ch211-152p11.4 |
| chr2_-_16380283 | 0.31 |
ENSDART00000149992
|
si:dkey-231j24.3
|
si:dkey-231j24.3 |
| chr6_+_28208973 | 0.30 |
ENSDART00000171216
ENSDART00000171377 ENSDART00000167389 ENSDART00000166988 |
LSM2 (1 of many)
|
si:ch73-14h10.2 |
| chr19_+_5543072 | 0.30 |
ENSDART00000082080
|
jupb
|
junction plakoglobin b |
| chr13_+_255067 | 0.30 |
ENSDART00000102505
|
foxg1d
|
forkhead box G1d |
| chr23_+_33963619 | 0.30 |
ENSDART00000140666
ENSDART00000084792 |
plpbp
|
pyridoxal phosphate binding protein |
| chr9_+_50001746 | 0.30 |
ENSDART00000058892
|
slc38a11
|
solute carrier family 38, member 11 |
| chr25_+_5035343 | 0.30 |
ENSDART00000011751
|
parvb
|
parvin, beta |
| chr1_-_44161417 | 0.29 |
ENSDART00000083213
|
slc43a3a
|
solute carrier family 43, member 3a |
| chr20_-_23226453 | 0.28 |
ENSDART00000142721
|
dcun1d4
|
DCN1, defective in cullin neddylation 1, domain containing 4 (S. cerevisiae) |
| chr5_-_30074332 | 0.28 |
ENSDART00000147963
|
bco2a
|
beta-carotene oxygenase 2a |
| chr10_-_26512742 | 0.28 |
ENSDART00000135951
|
si:dkey-5g14.1
|
si:dkey-5g14.1 |
| chr7_+_23292133 | 0.28 |
ENSDART00000134489
|
htr2cl1
|
5-hydroxytryptamine (serotonin) receptor 2C, G protein-coupled-like 1 |
| chr18_-_16590056 | 0.28 |
ENSDART00000143744
|
mgat4c
|
mgat4 family, member C |
| chr2_+_21229909 | 0.27 |
ENSDART00000046127
|
vipr1a
|
vasoactive intestinal peptide receptor 1a |
| chr11_+_25129013 | 0.26 |
ENSDART00000132879
|
ndrg3a
|
ndrg family member 3a |
| chr11_-_2131280 | 0.26 |
ENSDART00000008409
|
calcoco1b
|
calcium binding and coiled-coil domain 1b |
| chr16_+_43347966 | 0.26 |
ENSDART00000171308
|
zmp:0000000930
|
zmp:0000000930 |
| chr10_+_35952532 | 0.25 |
ENSDART00000184730
|
rtn4rl1a
|
reticulon 4 receptor-like 1a |
| chr24_+_40905100 | 0.25 |
ENSDART00000167854
|
scn12ab
|
sodium channel, voltage gated, type XII, alpha b |
| chr16_-_26255877 | 0.25 |
ENSDART00000146214
|
erfl1
|
Ets2 repressor factor like 1 |
| chr8_-_21142550 | 0.25 |
ENSDART00000143192
ENSDART00000186820 ENSDART00000135938 |
cpt2
|
carnitine palmitoyltransferase 2 |
| chr2_-_17235891 | 0.25 |
ENSDART00000144251
|
artnb
|
artemin b |
| chr12_+_22580579 | 0.24 |
ENSDART00000171725
ENSDART00000192290 |
capgb
|
capping protein (actin filament), gelsolin-like b |
| chr14_+_23717165 | 0.24 |
ENSDART00000006373
|
ndfip1
|
Nedd4 family interacting protein 1 |
| chr11_-_42554290 | 0.24 |
ENSDART00000130573
|
atp6ap1la
|
ATPase H+ transporting accessory protein 1 like a |
| chr16_-_40043322 | 0.24 |
ENSDART00000145278
|
si:dkey-29b11.3
|
si:dkey-29b11.3 |
| chr2_-_10896745 | 0.24 |
ENSDART00000114609
|
cdcp2
|
CUB domain containing protein 2 |
| chr1_-_10647307 | 0.24 |
ENSDART00000103548
|
si:dkey-31e10.1
|
si:dkey-31e10.1 |
| chr15_+_15856178 | 0.23 |
ENSDART00000080338
|
dusp14
|
dual specificity phosphatase 14 |
| chr23_-_35790235 | 0.23 |
ENSDART00000142369
ENSDART00000141141 ENSDART00000011004 |
mfsd5
|
major facilitator superfamily domain containing 5 |
| chr24_-_25004553 | 0.23 |
ENSDART00000080997
ENSDART00000136860 |
zdhhc20b
|
zinc finger, DHHC-type containing 20b |
| chr20_-_29864390 | 0.23 |
ENSDART00000161834
ENSDART00000132278 |
rnf144ab
|
ring finger protein 144ab |
| chr3_+_1637358 | 0.22 |
ENSDART00000180266
|
CR394546.5
|
|
| chr25_-_3469576 | 0.21 |
ENSDART00000186738
|
hbp1
|
HMG-box transcription factor 1 |
| chr24_+_40860320 | 0.21 |
ENSDART00000161351
|
gorasp1b
|
golgi reassembly stacking protein 1b |
| chr15_-_9272328 | 0.21 |
ENSDART00000172114
|
calm2a
|
calmodulin 2a (phosphorylase kinase, delta) |
| chr4_-_17629444 | 0.21 |
ENSDART00000108814
|
nrip2
|
nuclear receptor interacting protein 2 |
| chr10_-_26512993 | 0.20 |
ENSDART00000188549
ENSDART00000193316 |
si:dkey-5g14.1
|
si:dkey-5g14.1 |
| chr11_+_14284866 | 0.20 |
ENSDART00000163729
|
si:ch211-262i1.3
|
si:ch211-262i1.3 |
| chr23_+_27740592 | 0.20 |
ENSDART00000137875
|
dhh
|
desert hedgehog |
| chr11_-_29768054 | 0.20 |
ENSDART00000079117
|
plekha3
|
pleckstrin homology domain containing, family A (phosphoinositide binding specific) member 3 |
| chr11_-_472547 | 0.19 |
ENSDART00000005923
|
zgc:77375
|
zgc:77375 |
| chr8_+_20776654 | 0.19 |
ENSDART00000135850
|
nfic
|
nuclear factor I/C |
| chr1_-_17715493 | 0.18 |
ENSDART00000133027
|
si:dkey-256e7.8
|
si:dkey-256e7.8 |
| chr2_+_47249179 | 0.18 |
ENSDART00000144438
|
si:ch211-284d12.3
|
si:ch211-284d12.3 |
| chr15_-_22074315 | 0.18 |
ENSDART00000149830
|
drd2a
|
dopamine receptor D2a |
| chr16_+_45424907 | 0.18 |
ENSDART00000134471
|
phf1
|
PHD finger protein 1 |
| chr5_-_68093169 | 0.18 |
ENSDART00000051849
|
slc25a11
|
solute carrier family 25 (mitochondrial carrier; oxoglutarate carrier), member 11 |
| chr25_+_11456696 | 0.18 |
ENSDART00000171408
|
AGBL1
|
si:ch73-141f14.1 |
| chr16_-_21038015 | 0.17 |
ENSDART00000059239
|
snx10b
|
sorting nexin 10b |
| chr25_-_4713461 | 0.17 |
ENSDART00000155302
|
drd4a
|
dopamine receptor D4a |
| chr3_-_30388525 | 0.17 |
ENSDART00000183790
|
lrrc4ba
|
leucine rich repeat containing 4Ba |
| chr9_-_19161982 | 0.16 |
ENSDART00000081878
|
pou1f1
|
POU class 1 homeobox 1 |
| chr17_+_30545895 | 0.15 |
ENSDART00000076739
|
nhsl1a
|
NHS-like 1a |
| chr11_+_36231248 | 0.14 |
ENSDART00000131104
|
GPR62 (1 of many)
|
si:ch211-213o11.11 |
Network of associatons between targets according to the STRING database.
First level regulatory network of meox1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 2.8 | GO:0015860 | intracellular nucleoside transport(GO:0015859) purine nucleoside transmembrane transport(GO:0015860) purine nucleotide transport(GO:0015865) ATP transport(GO:0015867) purine ribonucleotide transport(GO:0015868) adenine nucleotide transport(GO:0051503) mitochondrial ATP transmembrane transport(GO:1990544) |
| 0.6 | 1.7 | GO:0097237 | cellular response to antibiotic(GO:0071236) cellular response to toxic substance(GO:0097237) |
| 0.4 | 1.6 | GO:0046166 | methylglyoxal biosynthetic process(GO:0019242) glyceraldehyde-3-phosphate biosynthetic process(GO:0046166) |
| 0.3 | 1.1 | GO:0060159 | regulation of dopamine receptor signaling pathway(GO:0060159) |
| 0.2 | 0.7 | GO:1900620 | acetylcholine biosynthetic process(GO:0008292) acetate ester biosynthetic process(GO:1900620) |
| 0.2 | 2.4 | GO:0036368 | cone photoresponse recovery(GO:0036368) |
| 0.2 | 1.9 | GO:1904071 | presynaptic active zone assembly(GO:1904071) |
| 0.2 | 0.5 | GO:0071361 | cellular response to ethanol(GO:0071361) |
| 0.1 | 0.4 | GO:0052576 | carbohydrate localization(GO:0052575) carbohydrate storage(GO:0052576) |
| 0.1 | 0.8 | GO:0042745 | circadian sleep/wake cycle process(GO:0022410) circadian sleep/wake cycle(GO:0042745) |
| 0.1 | 0.4 | GO:0007414 | axonal defasciculation(GO:0007414) |
| 0.1 | 0.4 | GO:0048496 | maintenance of organ identity(GO:0048496) |
| 0.1 | 0.7 | GO:0044341 | sodium-dependent phosphate transport(GO:0044341) |
| 0.1 | 0.4 | GO:0051039 | histone displacement(GO:0001207) regulation of transcription involved in meiotic cell cycle(GO:0051037) positive regulation of transcription involved in meiotic cell cycle(GO:0051039) |
| 0.1 | 0.3 | GO:2000434 | regulation of protein neddylation(GO:2000434) |
| 0.1 | 0.6 | GO:0016539 | intein-mediated protein splicing(GO:0016539) protein splicing(GO:0030908) |
| 0.1 | 0.3 | GO:0034163 | regulation of toll-like receptor 9 signaling pathway(GO:0034163) negative regulation of toll-like receptor 9 signaling pathway(GO:0034164) |
| 0.1 | 0.8 | GO:0046037 | GMP biosynthetic process(GO:0006177) GMP metabolic process(GO:0046037) |
| 0.1 | 0.5 | GO:0042362 | fat-soluble vitamin biosynthetic process(GO:0042362) |
| 0.1 | 1.1 | GO:0007631 | feeding behavior(GO:0007631) |
| 0.1 | 0.5 | GO:0045604 | regulation of epidermal cell differentiation(GO:0045604) |
| 0.1 | 0.6 | GO:0061075 | positive regulation of neural retina development(GO:0061075) positive regulation of retina development in camera-type eye(GO:1902868) |
| 0.1 | 0.4 | GO:0021767 | mammillary body development(GO:0021767) |
| 0.1 | 0.3 | GO:0035994 | response to muscle stretch(GO:0035994) |
| 0.1 | 0.3 | GO:0002159 | desmosome assembly(GO:0002159) |
| 0.1 | 1.1 | GO:0050795 | regulation of behavior(GO:0050795) |
| 0.1 | 1.0 | GO:0098884 | postsynaptic neurotransmitter receptor internalization(GO:0098884) |
| 0.1 | 1.6 | GO:0055064 | chloride ion homeostasis(GO:0055064) |
| 0.0 | 0.2 | GO:0014707 | branchiomeric skeletal muscle development(GO:0014707) |
| 0.0 | 0.8 | GO:0048923 | posterior lateral line neuromast hair cell differentiation(GO:0048923) |
| 0.0 | 0.6 | GO:0042044 | fluid transport(GO:0042044) |
| 0.0 | 0.3 | GO:0070207 | protein homotrimerization(GO:0070207) |
| 0.0 | 1.5 | GO:0042761 | very long-chain fatty acid biosynthetic process(GO:0042761) |
| 0.0 | 1.1 | GO:0007168 | receptor guanylyl cyclase signaling pathway(GO:0007168) |
| 0.0 | 0.9 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
| 0.0 | 0.6 | GO:0006183 | GTP biosynthetic process(GO:0006183) guanosine-containing compound biosynthetic process(GO:1901070) |
| 0.0 | 1.2 | GO:0045839 | negative regulation of mitotic nuclear division(GO:0045839) |
| 0.0 | 0.8 | GO:0010923 | negative regulation of phosphatase activity(GO:0010923) |
| 0.0 | 2.8 | GO:0032869 | cellular response to insulin stimulus(GO:0032869) |
| 0.0 | 0.2 | GO:0015743 | thiosulfate transport(GO:0015709) oxaloacetate transport(GO:0015729) malate transport(GO:0015743) succinate transport(GO:0015744) succinate transmembrane transport(GO:0071422) malate transmembrane transport(GO:0071423) |
| 0.0 | 1.0 | GO:0070593 | dendrite self-avoidance(GO:0070593) |
| 0.0 | 0.8 | GO:0060158 | phospholipase C-activating dopamine receptor signaling pathway(GO:0060158) |
| 0.0 | 0.3 | GO:0016121 | carotene metabolic process(GO:0016119) carotene catabolic process(GO:0016121) terpene metabolic process(GO:0042214) terpene catabolic process(GO:0046247) |
| 0.0 | 0.6 | GO:0036444 | calcium ion transmembrane import into mitochondrion(GO:0036444) |
| 0.0 | 1.5 | GO:0006368 | transcription elongation from RNA polymerase II promoter(GO:0006368) |
| 0.0 | 0.7 | GO:0042446 | hormone biosynthetic process(GO:0042446) |
| 0.0 | 0.2 | GO:0035627 | ceramide transport(GO:0035627) |
| 0.0 | 1.2 | GO:0007586 | digestion(GO:0007586) |
| 0.0 | 0.9 | GO:0006904 | vesicle docking involved in exocytosis(GO:0006904) |
| 0.0 | 1.2 | GO:0036269 | swimming behavior(GO:0036269) |
| 0.0 | 0.3 | GO:0045471 | response to ethanol(GO:0045471) |
| 0.0 | 0.4 | GO:0006658 | phosphatidylserine metabolic process(GO:0006658) |
| 0.0 | 1.0 | GO:0032922 | circadian regulation of gene expression(GO:0032922) |
| 0.0 | 0.5 | GO:0010507 | negative regulation of autophagy(GO:0010507) |
| 0.0 | 0.7 | GO:0072332 | intrinsic apoptotic signaling pathway by p53 class mediator(GO:0072332) |
| 0.0 | 0.4 | GO:0015671 | oxygen transport(GO:0015671) |
| 0.0 | 0.5 | GO:0007214 | gamma-aminobutyric acid signaling pathway(GO:0007214) |
| 0.0 | 0.6 | GO:1902476 | chloride transmembrane transport(GO:1902476) |
| 0.0 | 0.3 | GO:0031060 | regulation of histone methylation(GO:0031060) |
| 0.0 | 0.5 | GO:0045494 | photoreceptor cell maintenance(GO:0045494) |
| 0.0 | 0.4 | GO:0035622 | intrahepatic bile duct development(GO:0035622) |
| 0.0 | 0.4 | GO:0043124 | negative regulation of I-kappaB kinase/NF-kappaB signaling(GO:0043124) |
| 0.0 | 0.4 | GO:0032008 | positive regulation of TOR signaling(GO:0032008) |
| 0.0 | 0.8 | GO:0048843 | negative regulation of axon extension involved in axon guidance(GO:0048843) negative regulation of axon guidance(GO:1902668) |
| 0.0 | 0.3 | GO:0033344 | cholesterol efflux(GO:0033344) |
| 0.0 | 0.3 | GO:0030032 | lamellipodium assembly(GO:0030032) |
| 0.0 | 1.1 | GO:0010506 | regulation of autophagy(GO:0010506) |
| 0.0 | 0.6 | GO:0003009 | skeletal muscle contraction(GO:0003009) |
| 0.0 | 0.3 | GO:0048665 | neuron fate specification(GO:0048665) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 4.4 | GO:0031680 | G-protein beta/gamma-subunit complex(GO:0031680) |
| 0.3 | 1.5 | GO:0030891 | VCB complex(GO:0030891) |
| 0.2 | 1.9 | GO:0098982 | GABA-ergic synapse(GO:0098982) |
| 0.2 | 0.9 | GO:0042584 | chromaffin granule membrane(GO:0042584) |
| 0.1 | 0.7 | GO:0016533 | cyclin-dependent protein kinase 5 holoenzyme complex(GO:0016533) |
| 0.1 | 0.5 | GO:0038037 | G-protein coupled receptor dimeric complex(GO:0038037) G-protein coupled receptor heterodimeric complex(GO:0038039) G-protein coupled receptor complex(GO:0097648) |
| 0.1 | 1.1 | GO:0032809 | neuronal cell body membrane(GO:0032809) cell body membrane(GO:0044298) |
| 0.1 | 1.0 | GO:0098844 | postsynaptic endocytic zone membrane(GO:0098844) |
| 0.0 | 0.6 | GO:1990246 | uniplex complex(GO:1990246) |
| 0.0 | 0.4 | GO:0032593 | insulin-responsive compartment(GO:0032593) |
| 0.0 | 0.2 | GO:0033181 | plasma membrane proton-transporting V-type ATPase complex(GO:0033181) |
| 0.0 | 0.7 | GO:0005903 | brush border(GO:0005903) |
| 0.0 | 0.4 | GO:0005943 | phosphatidylinositol 3-kinase complex, class IA(GO:0005943) |
| 0.0 | 0.6 | GO:0032591 | dendritic spine membrane(GO:0032591) |
| 0.0 | 0.4 | GO:0031838 | haptoglobin-hemoglobin complex(GO:0031838) |
| 0.0 | 1.2 | GO:0043204 | perikaryon(GO:0043204) |
| 0.0 | 1.1 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.0 | 0.4 | GO:0002116 | semaphorin receptor complex(GO:0002116) |
| 0.0 | 0.6 | GO:1902710 | GABA receptor complex(GO:1902710) GABA-A receptor complex(GO:1902711) |
| 0.0 | 3.9 | GO:0030424 | axon(GO:0030424) |
| 0.0 | 1.5 | GO:0008021 | synaptic vesicle(GO:0008021) |
| 0.0 | 1.2 | GO:0099501 | synaptic vesicle membrane(GO:0030672) exocytic vesicle membrane(GO:0099501) |
| 0.0 | 0.7 | GO:0034705 | voltage-gated potassium channel complex(GO:0008076) potassium channel complex(GO:0034705) |
| 0.0 | 0.6 | GO:0005861 | troponin complex(GO:0005861) |
| 0.0 | 0.9 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.0 | 0.1 | GO:0030670 | phagocytic vesicle membrane(GO:0030670) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 2.8 | GO:0005471 | ATP:ADP antiporter activity(GO:0005471) |
| 0.4 | 2.4 | GO:0004743 | pyruvate kinase activity(GO:0004743) |
| 0.4 | 1.6 | GO:0004807 | triose-phosphate isomerase activity(GO:0004807) methylglyoxal synthase activity(GO:0008929) |
| 0.3 | 1.0 | GO:0031716 | calcitonin receptor binding(GO:0031716) |
| 0.3 | 1.3 | GO:0031841 | neuropeptide Y receptor binding(GO:0031841) type 2 neuropeptide Y receptor binding(GO:0031843) |
| 0.2 | 1.6 | GO:0004459 | L-lactate dehydrogenase activity(GO:0004459) |
| 0.2 | 4.4 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.1 | 0.6 | GO:0008559 | xenobiotic-transporting ATPase activity(GO:0008559) |
| 0.1 | 0.8 | GO:0003938 | IMP dehydrogenase activity(GO:0003938) |
| 0.1 | 0.7 | GO:0005436 | sodium:phosphate symporter activity(GO:0005436) |
| 0.1 | 1.9 | GO:0098882 | structural constituent of presynaptic active zone(GO:0098882) |
| 0.1 | 0.4 | GO:0004645 | phosphorylase activity(GO:0004645) glycogen phosphorylase activity(GO:0008184) |
| 0.1 | 2.7 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.1 | 0.6 | GO:0005113 | patched binding(GO:0005113) |
| 0.1 | 0.5 | GO:0004965 | G-protein coupled GABA receptor activity(GO:0004965) |
| 0.1 | 1.1 | GO:0004383 | guanylate cyclase activity(GO:0004383) |
| 0.1 | 0.5 | GO:0016934 | extracellular-glycine-gated ion channel activity(GO:0016933) extracellular-glycine-gated chloride channel activity(GO:0016934) |
| 0.1 | 0.7 | GO:0061575 | cyclin-dependent protein serine/threonine kinase activator activity(GO:0061575) |
| 0.1 | 0.7 | GO:0016413 | O-acetyltransferase activity(GO:0016413) |
| 0.0 | 0.2 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.0 | 0.7 | GO:0031682 | G-protein gamma-subunit binding(GO:0031682) |
| 0.0 | 1.1 | GO:0043176 | amine binding(GO:0043176) serotonin binding(GO:0051378) |
| 0.0 | 0.4 | GO:0031826 | type 2A serotonin receptor binding(GO:0031826) |
| 0.0 | 0.3 | GO:0004999 | vasoactive intestinal polypeptide receptor activity(GO:0004999) |
| 0.0 | 0.6 | GO:0005432 | calcium:sodium antiporter activity(GO:0005432) |
| 0.0 | 0.1 | GO:0005252 | open rectifier potassium channel activity(GO:0005252) |
| 0.0 | 0.2 | GO:1902388 | ceramide transporter activity(GO:0035620) ceramide 1-phosphate binding(GO:1902387) ceramide 1-phosphate transporter activity(GO:1902388) |
| 0.0 | 0.9 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.0 | 0.2 | GO:0004095 | carnitine O-palmitoyltransferase activity(GO:0004095) O-palmitoyltransferase activity(GO:0016416) |
| 0.0 | 0.2 | GO:0015131 | thiosulfate transmembrane transporter activity(GO:0015117) oxaloacetate transmembrane transporter activity(GO:0015131) malate transmembrane transporter activity(GO:0015140) |
| 0.0 | 0.9 | GO:0008378 | galactosyltransferase activity(GO:0008378) |
| 0.0 | 0.4 | GO:0052812 | phosphatidylinositol-4,5-bisphosphate 3-kinase activity(GO:0046934) phosphatidylinositol-3,4-bisphosphate 5-kinase activity(GO:0052812) |
| 0.0 | 0.3 | GO:0045294 | alpha-catenin binding(GO:0045294) |
| 0.0 | 0.4 | GO:0031720 | haptoglobin binding(GO:0031720) |
| 0.0 | 0.3 | GO:0010436 | beta-carotene 15,15'-monooxygenase activity(GO:0003834) carotenoid dioxygenase activity(GO:0010436) |
| 0.0 | 1.0 | GO:0098632 | protein binding involved in cell-cell adhesion(GO:0098632) |
| 0.0 | 2.2 | GO:0016627 | oxidoreductase activity, acting on the CH-CH group of donors(GO:0016627) |
| 0.0 | 0.3 | GO:0015129 | lactate transmembrane transporter activity(GO:0015129) |
| 0.0 | 0.4 | GO:0070577 | lysine-acetylated histone binding(GO:0070577) |
| 0.0 | 0.5 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.0 | 1.0 | GO:0019003 | GDP binding(GO:0019003) |
| 0.0 | 1.0 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.0 | 0.3 | GO:0001591 | dopamine neurotransmitter receptor activity, coupled via Gi/Go(GO:0001591) |
| 0.0 | 0.6 | GO:0015379 | cation:chloride symporter activity(GO:0015377) potassium:chloride symporter activity(GO:0015379) |
| 0.0 | 0.4 | GO:0042813 | Wnt-activated receptor activity(GO:0042813) |
| 0.0 | 1.0 | GO:0001786 | phosphatidylserine binding(GO:0001786) |
| 0.0 | 0.6 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.0 | 0.6 | GO:0004550 | nucleoside diphosphate kinase activity(GO:0004550) |
| 0.0 | 0.6 | GO:0016917 | GABA-A receptor activity(GO:0004890) GABA receptor activity(GO:0016917) |
| 0.0 | 0.3 | GO:0097602 | cullin family protein binding(GO:0097602) |
| 0.0 | 0.8 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.0 | 0.1 | GO:0050431 | transforming growth factor beta binding(GO:0050431) |
| 0.0 | 0.4 | GO:0005104 | fibroblast growth factor receptor binding(GO:0005104) |
| 0.0 | 0.4 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.0 | 0.7 | GO:0044325 | ion channel binding(GO:0044325) |
| 0.0 | 0.1 | GO:0004142 | diacylglycerol cholinephosphotransferase activity(GO:0004142) ethanolaminephosphotransferase activity(GO:0004307) |
| 0.0 | 0.3 | GO:0004016 | adenylate cyclase activity(GO:0004016) |
| 0.0 | 0.4 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.0 | 0.2 | GO:0033549 | MAP kinase tyrosine/serine/threonine phosphatase activity(GO:0017017) MAP kinase phosphatase activity(GO:0033549) |
| 0.0 | 0.2 | GO:0070001 | aspartic-type endopeptidase activity(GO:0004190) aspartic-type peptidase activity(GO:0070001) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 6.0 | PID RHODOPSIN PATHWAY | Visual signal transduction: Rods |
| 0.0 | 0.6 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.0 | 0.6 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
| 0.0 | 0.8 | NABA PROTEOGLYCANS | Genes encoding proteoglycans |
| 0.0 | 1.8 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 0.3 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 0.0 | 1.0 | PID MTOR 4PATHWAY | mTOR signaling pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 4.4 | REACTOME OLFACTORY SIGNALING PATHWAY | Genes involved in Olfactory Signaling Pathway |
| 0.2 | 1.1 | REACTOME SEROTONIN RECEPTORS | Genes involved in Serotonin receptors |
| 0.2 | 1.0 | REACTOME REGULATION OF RHEB GTPASE ACTIVITY BY AMPK | Genes involved in Regulation of Rheb GTPase activity by AMPK |
| 0.1 | 1.0 | REACTOME INHIBITION OF INSULIN SECRETION BY ADRENALINE NORADRENALINE | Genes involved in Inhibition of Insulin Secretion by Adrenaline/Noradrenaline |
| 0.1 | 2.5 | REACTOME INTERACTIONS OF VPR WITH HOST CELLULAR PROTEINS | Genes involved in Interactions of Vpr with host cellular proteins |
| 0.0 | 1.3 | REACTOME KERATAN SULFATE BIOSYNTHESIS | Genes involved in Keratan sulfate biosynthesis |
| 0.0 | 0.8 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.0 | 0.8 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.0 | 1.0 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.0 | 0.4 | REACTOME FGFR4 LIGAND BINDING AND ACTIVATION | Genes involved in FGFR4 ligand binding and activation |
| 0.0 | 0.7 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.0 | 0.6 | REACTOME NITRIC OXIDE STIMULATES GUANYLATE CYCLASE | Genes involved in Nitric oxide stimulates guanylate cyclase |
| 0.0 | 0.3 | REACTOME BILE SALT AND ORGANIC ANION SLC TRANSPORTERS | Genes involved in Bile salt and organic anion SLC transporters |
| 0.0 | 0.3 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.0 | 0.3 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.0 | 0.2 | REACTOME ACTIVATED AMPK STIMULATES FATTY ACID OXIDATION IN MUSCLE | Genes involved in Activated AMPK stimulates fatty-acid oxidation in muscle |
| 0.0 | 0.5 | REACTOME LIGAND GATED ION CHANNEL TRANSPORT | Genes involved in Ligand-gated ion channel transport |
| 0.0 | 0.3 | REACTOME N GLYCAN ANTENNAE ELONGATION | Genes involved in N-Glycan antennae elongation |
| 0.0 | 0.6 | REACTOME CLASS B 2 SECRETIN FAMILY RECEPTORS | Genes involved in Class B/2 (Secretin family receptors) |
| 0.0 | 1.5 | REACTOME G ALPHA I SIGNALLING EVENTS | Genes involved in G alpha (i) signalling events |