Project
PRJEB1986: zebrafish developmental stages transcriptome
Navigation
Downloads
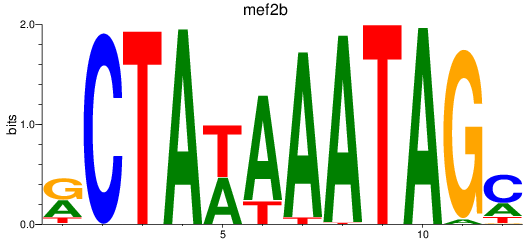
Results for mef2b
Z-value: 1.18
Transcription factors associated with mef2b
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
mef2b
|
ENSDARG00000093170 | myocyte enhancer factor 2b |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| mef2b | dr11_v1_chr22_+_18187857_18187857 | 0.36 | 1.3e-01 | Click! |
Activity profile of mef2b motif
Sorted Z-values of mef2b motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr3_-_61205711 | 10.52 |
ENSDART00000055062
|
pvalb1
|
parvalbumin 1 |
| chr5_+_51597677 | 7.06 |
ENSDART00000048210
ENSDART00000184797 |
ckmt2b
|
creatine kinase, mitochondrial 2b (sarcomeric) |
| chr25_+_29160102 | 7.01 |
ENSDART00000162854
|
pkmb
|
pyruvate kinase M1/2b |
| chr8_+_22930627 | 3.30 |
ENSDART00000187860
|
sypa
|
synaptophysin a |
| chr9_+_31795343 | 3.29 |
ENSDART00000139584
|
itgbl1
|
integrin, beta-like 1 |
| chr10_-_22845485 | 3.25 |
ENSDART00000079454
|
vamp2
|
vesicle-associated membrane protein 2 |
| chr23_+_44614056 | 3.11 |
ENSDART00000188379
|
eno3
|
enolase 3, (beta, muscle) |
| chr6_-_14139503 | 3.08 |
ENSDART00000089577
|
cacnb4b
|
calcium channel, voltage-dependent, beta 4b subunit |
| chr7_+_14291323 | 3.04 |
ENSDART00000053521
|
rhcga
|
Rh family, C glycoprotein a |
| chr6_-_40722480 | 2.92 |
ENSDART00000188187
|
kbtbd12
|
kelch repeat and BTB (POZ) domain containing 12 |
| chr14_+_35901249 | 2.89 |
ENSDART00000105604
|
zgc:77938
|
zgc:77938 |
| chr20_-_26042070 | 2.88 |
ENSDART00000140255
|
si:dkey-12h9.6
|
si:dkey-12h9.6 |
| chr5_-_72125551 | 2.87 |
ENSDART00000149412
|
smyd1a
|
SET and MYND domain containing 1a |
| chr20_-_26001288 | 2.84 |
ENSDART00000136518
ENSDART00000063177 |
capn3b
|
calpain 3b |
| chr7_-_7420301 | 2.83 |
ENSDART00000102620
|
six7
|
SIX homeobox 7 |
| chr10_+_37145007 | 2.82 |
ENSDART00000131777
|
cuedc1a
|
CUE domain containing 1a |
| chr1_+_8601935 | 2.79 |
ENSDART00000152367
|
si:ch211-160d14.6
|
si:ch211-160d14.6 |
| chr17_-_12336987 | 2.78 |
ENSDART00000172001
|
snap25b
|
synaptosomal-associated protein, 25b |
| chr6_-_40722200 | 2.70 |
ENSDART00000035101
|
kbtbd12
|
kelch repeat and BTB (POZ) domain containing 12 |
| chr11_+_11201096 | 2.66 |
ENSDART00000171916
ENSDART00000171521 ENSDART00000087105 ENSDART00000159603 |
myom2a
|
myomesin 2a |
| chr6_-_35472923 | 2.59 |
ENSDART00000185907
|
rgs8
|
regulator of G protein signaling 8 |
| chr3_-_32818607 | 2.58 |
ENSDART00000075465
|
mylpfa
|
myosin light chain, phosphorylatable, fast skeletal muscle a |
| chr25_-_29363934 | 2.44 |
ENSDART00000166889
|
nptna
|
neuroplastin a |
| chr13_+_9432501 | 2.43 |
ENSDART00000058064
|
zgc:123321
|
zgc:123321 |
| chr10_-_24371312 | 2.35 |
ENSDART00000149362
|
pitpnab
|
phosphatidylinositol transfer protein, alpha b |
| chr23_-_7826849 | 2.32 |
ENSDART00000157612
|
myt1b
|
myelin transcription factor 1b |
| chr19_+_41169996 | 2.25 |
ENSDART00000048438
|
asb4
|
ankyrin repeat and SOCS box containing 4 |
| chr23_-_32157865 | 2.22 |
ENSDART00000000876
|
nr4a1
|
nuclear receptor subfamily 4, group A, member 1 |
| chr10_+_29698467 | 2.22 |
ENSDART00000163402
|
dlg2
|
discs, large homolog 2 (Drosophila) |
| chr1_+_45080897 | 2.13 |
ENSDART00000129819
|
si:ch211-151p13.8
|
si:ch211-151p13.8 |
| chr21_-_12272543 | 2.11 |
ENSDART00000081510
ENSDART00000151297 |
celf4
|
CUGBP, Elav-like family member 4 |
| chr19_-_31035155 | 2.03 |
ENSDART00000161882
|
bzw2
|
basic leucine zipper and W2 domains 2 |
| chr2_+_1202347 | 1.95 |
ENSDART00000075837
|
CABZ01084566.1
|
|
| chr21_+_11684830 | 1.93 |
ENSDART00000147473
|
pcsk1
|
proprotein convertase subtilisin/kexin type 1 |
| chr19_-_31035325 | 1.93 |
ENSDART00000147504
|
bzw2
|
basic leucine zipper and W2 domains 2 |
| chr3_+_26135502 | 1.92 |
ENSDART00000146979
|
atp2a1
|
ATPase sarcoplasmic/endoplasmic reticulum Ca2+ transporting 1 |
| chr23_+_6077503 | 1.85 |
ENSDART00000081714
ENSDART00000139834 |
mybpha
|
myosin binding protein Ha |
| chr7_-_27686021 | 1.85 |
ENSDART00000079112
ENSDART00000100989 |
calca
|
calcitonin/calcitonin-related polypeptide, alpha |
| chr20_+_41549200 | 1.84 |
ENSDART00000135715
|
fam184a
|
family with sequence similarity 184, member A |
| chr5_-_10946232 | 1.83 |
ENSDART00000163139
ENSDART00000031265 |
rtn4r
|
reticulon 4 receptor |
| chr3_+_23092762 | 1.80 |
ENSDART00000142884
ENSDART00000024136 |
gngt2a
|
guanine nucleotide binding protein (G protein), gamma transducing activity polypeptide 2a |
| chr13_-_27767330 | 1.78 |
ENSDART00000131631
ENSDART00000112553 ENSDART00000189911 |
rims1a
|
regulating synaptic membrane exocytosis 1a |
| chr2_-_4787566 | 1.76 |
ENSDART00000160663
ENSDART00000157808 |
tnk2b
|
tyrosine kinase, non-receptor, 2b |
| chr21_+_11685009 | 1.76 |
ENSDART00000014668
|
pcsk1
|
proprotein convertase subtilisin/kexin type 1 |
| chr25_-_13381854 | 1.74 |
ENSDART00000164621
ENSDART00000169129 |
ndrg4
|
NDRG family member 4 |
| chr18_+_7264961 | 1.74 |
ENSDART00000188461
|
CABZ01015105.1
|
|
| chr20_+_34455645 | 1.70 |
ENSDART00000135789
|
mettl11b
|
methyltransferase like 11B |
| chr21_-_27185915 | 1.67 |
ENSDART00000135052
|
slc8a4a
|
solute carrier family 8 (sodium/calcium exchanger), member 4a |
| chr21_+_12010505 | 1.63 |
ENSDART00000123522
|
aqp7
|
aquaporin 7 |
| chr23_-_38497705 | 1.57 |
ENSDART00000109493
|
tshz2
|
teashirt zinc finger homeobox 2 |
| chr23_+_20687340 | 1.56 |
ENSDART00000143503
|
usp21
|
ubiquitin specific peptidase 21 |
| chr25_+_3677650 | 1.51 |
ENSDART00000154348
|
prnprs3
|
prion protein, related sequence 3 |
| chr23_+_22658700 | 1.50 |
ENSDART00000192248
|
eno1a
|
enolase 1a, (alpha) |
| chr4_-_23908802 | 1.47 |
ENSDART00000138873
|
celf2
|
cugbp, Elav-like family member 2 |
| chr3_-_49504023 | 1.43 |
ENSDART00000168108
|
prkacaa
|
protein kinase, cAMP-dependent, catalytic, alpha, genome duplicate a |
| chr12_-_34035364 | 1.41 |
ENSDART00000087065
|
timp2a
|
TIMP metallopeptidase inhibitor 2a |
| chr6_-_10780698 | 1.41 |
ENSDART00000151714
|
gpr155b
|
G protein-coupled receptor 155b |
| chr5_-_51819027 | 1.35 |
ENSDART00000164267
|
homer1b
|
homer scaffolding protein 1b |
| chr10_-_2522588 | 1.34 |
ENSDART00000081926
|
CU856539.1
|
|
| chr21_-_25669820 | 1.34 |
ENSDART00000148236
|
tmem179b
|
transmembrane protein 179B |
| chr9_-_52814204 | 1.30 |
ENSDART00000140771
ENSDART00000007401 |
MAP3K13
|
si:ch211-45c16.2 |
| chr16_+_29663809 | 1.25 |
ENSDART00000191336
|
tmod4
|
tropomodulin 4 (muscle) |
| chr1_-_21483832 | 1.23 |
ENSDART00000102790
|
glrba
|
glycine receptor, beta a |
| chr13_+_47710434 | 1.20 |
ENSDART00000188724
|
TMEM87B
|
transmembrane protein 87B |
| chr24_-_20444844 | 1.18 |
ENSDART00000048940
|
vill
|
villin-like |
| chr17_-_50234004 | 1.12 |
ENSDART00000058706
|
fosaa
|
v-fos FBJ murine osteosarcoma viral oncogene homolog Aa |
| chr19_-_8096984 | 1.12 |
ENSDART00000146987
|
si:dkey-266f7.9
|
si:dkey-266f7.9 |
| chr14_+_24215046 | 1.09 |
ENSDART00000079215
|
stc2a
|
stanniocalcin 2a |
| chr4_+_77060861 | 1.07 |
ENSDART00000174271
ENSDART00000174393 ENSDART00000150450 |
si:dkey-240n22.8
|
si:dkey-240n22.8 |
| chr1_+_19708508 | 1.05 |
ENSDART00000054581
ENSDART00000131206 |
march1
|
membrane-associated ring finger (C3HC4) 1 |
| chr14_-_7245971 | 1.04 |
ENSDART00000108796
|
stox2b
|
storkhead box 2b |
| chr6_-_46403475 | 0.99 |
ENSDART00000154148
|
camk1a
|
calcium/calmodulin-dependent protein kinase Ia |
| chr7_+_33457148 | 0.98 |
ENSDART00000133562
ENSDART00000074587 |
paqr5b
|
progestin and adipoQ receptor family member Vb |
| chr15_+_20543770 | 0.98 |
ENSDART00000092357
|
sgsm2
|
small G protein signaling modulator 2 |
| chr16_-_17188294 | 0.95 |
ENSDART00000165883
|
opn9
|
opsin 9 |
| chr13_-_37647209 | 0.91 |
ENSDART00000189102
|
si:dkey-188i13.10
|
si:dkey-188i13.10 |
| chr23_+_20110086 | 0.91 |
ENSDART00000054664
|
tnnc1b
|
troponin C type 1b (slow) |
| chr12_+_41697664 | 0.90 |
ENSDART00000162302
|
bnip3
|
BCL2 interacting protein 3 |
| chr14_-_32405387 | 0.87 |
ENSDART00000184647
|
fgf13a
|
fibroblast growth factor 13a |
| chr16_+_25245857 | 0.87 |
ENSDART00000155220
|
klhl38b
|
kelch-like family member 38b |
| chr23_+_3721042 | 0.87 |
ENSDART00000143323
|
smim29
|
small integral membrane protein 29 |
| chr4_+_5180650 | 0.86 |
ENSDART00000067390
|
fgf6b
|
fibroblast growth factor 6b |
| chr9_-_23891102 | 0.82 |
ENSDART00000186799
|
asb18
|
ankyrin repeat and SOCS box containing 18 |
| chr17_-_10025234 | 0.80 |
ENSDART00000008355
|
cfl2
|
cofilin 2 (muscle) |
| chr17_+_45648836 | 0.78 |
ENSDART00000155037
|
zgc:162184
|
zgc:162184 |
| chr7_-_31938938 | 0.78 |
ENSDART00000132353
|
bdnf
|
brain-derived neurotrophic factor |
| chr8_+_17987215 | 0.77 |
ENSDART00000113605
|
lrriq3
|
leucine-rich repeats and IQ motif containing 3 |
| chr21_+_43559123 | 0.76 |
ENSDART00000151212
|
gpr185a
|
G protein-coupled receptor 185 a |
| chr13_-_12667220 | 0.75 |
ENSDART00000079594
|
fam241a
|
family with sequence similarity 241 member A |
| chr11_-_26701611 | 0.74 |
ENSDART00000083010
|
acad9
|
acyl-CoA dehydrogenase family, member 9 |
| chr19_-_7321221 | 0.72 |
ENSDART00000092375
|
oxr1b
|
oxidation resistance 1b |
| chr5_+_16580739 | 0.69 |
ENSDART00000135140
|
htr7c
|
5-hydroxytryptamine (serotonin) receptor 7c |
| chr25_-_29988352 | 0.68 |
ENSDART00000067059
|
fam19a5b
|
family with sequence similarity 19 (chemokine (C-C motif)-like), member A5b |
| chr10_-_41450367 | 0.68 |
ENSDART00000122682
ENSDART00000189549 |
cabp1b
|
calcium binding protein 1b |
| chr9_-_23894392 | 0.67 |
ENSDART00000133417
|
asb18
|
ankyrin repeat and SOCS box containing 18 |
| chr8_+_4337312 | 0.66 |
ENSDART00000182228
|
myl2b
|
myosin, light chain 2b, regulatory, cardiac, slow |
| chr4_-_5863906 | 0.66 |
ENSDART00000169424
|
slc5a8
|
solute carrier family 5 (sodium/monocarboxylate cotransporter), member 8 |
| chr12_+_8373525 | 0.66 |
ENSDART00000152180
|
arid5b
|
AT-rich interaction domain 5B |
| chr5_-_30151815 | 0.61 |
ENSDART00000156048
|
zbtb44
|
zinc finger and BTB domain containing 44 |
| chr9_+_32178050 | 0.60 |
ENSDART00000169526
|
coq10b
|
coenzyme Q10B |
| chr24_-_7777389 | 0.59 |
ENSDART00000138541
|
rpgrip1
|
RPGR interacting protein 1 |
| chr8_-_53166975 | 0.59 |
ENSDART00000114683
|
rbsn
|
rabenosyn, RAB effector |
| chr23_-_25050329 | 0.56 |
ENSDART00000140216
|
avpr2aa
|
arginine vasopressin receptor 2a, duplicate a |
| chr8_+_44420108 | 0.56 |
ENSDART00000075381
|
CU571323.1
|
|
| chr10_-_17501528 | 0.55 |
ENSDART00000144847
|
slc2a11l
|
solute carrier family 2 (facilitated glucose transporter), member 11-like |
| chr25_-_12809361 | 0.52 |
ENSDART00000162750
|
ca5a
|
carbonic anhydrase Va |
| chr10_-_8033468 | 0.51 |
ENSDART00000140476
|
atp6v0a2a
|
ATPase H+ transporting V0 subunit a2a |
| chr15_-_36347858 | 0.50 |
ENSDART00000155274
ENSDART00000157936 |
si:dkey-23k10.2
|
si:dkey-23k10.2 |
| chr9_-_28399071 | 0.50 |
ENSDART00000104317
ENSDART00000064343 |
klf7b
|
Kruppel-like factor 7b |
| chr7_+_50464500 | 0.47 |
ENSDART00000191356
|
serinc4
|
serine incorporator 4 |
| chr5_+_64842730 | 0.47 |
ENSDART00000144732
|
lrrc8ab
|
leucine rich repeat containing 8 VRAC subunit Ab |
| chr10_-_2527342 | 0.46 |
ENSDART00000184168
|
CU856539.1
|
|
| chr1_-_58592866 | 0.46 |
ENSDART00000161039
|
adgre5b.2
|
adhesion G protein-coupled receptor E5b, duplicate 2 |
| chr11_-_1948784 | 0.46 |
ENSDART00000082475
|
nr1d4b
|
nuclear receptor subfamily 1, group D, member 4b |
| chr15_-_14552101 | 0.46 |
ENSDART00000171169
|
numbl
|
numb homolog (Drosophila)-like |
| chr7_-_48263516 | 0.46 |
ENSDART00000006619
ENSDART00000142370 ENSDART00000148273 ENSDART00000147968 |
rbpms2b
|
RNA binding protein with multiple splicing 2b |
| chr3_-_18189283 | 0.46 |
ENSDART00000049240
|
tob1a
|
transducer of ERBB2, 1a |
| chr3_-_13946446 | 0.45 |
ENSDART00000171249
|
gcdhb
|
glutaryl-CoA dehydrogenase b |
| chr3_+_32425202 | 0.44 |
ENSDART00000156464
|
prr12b
|
proline rich 12b |
| chr21_+_40589770 | 0.42 |
ENSDART00000164650
ENSDART00000161584 ENSDART00000161108 |
pdk3b
|
pyruvate dehydrogenase kinase, isozyme 3b |
| chr10_+_29259882 | 0.41 |
ENSDART00000180606
|
sytl2a
|
synaptotagmin-like 2a |
| chr22_+_38310957 | 0.40 |
ENSDART00000040550
|
traf5
|
Tnf receptor-associated factor 5 |
| chr16_-_17200120 | 0.38 |
ENSDART00000147739
|
gapdh
|
glyceraldehyde-3-phosphate dehydrogenase |
| chr9_-_33107237 | 0.37 |
ENSDART00000013918
|
casq2
|
calsequestrin 2 |
| chr25_+_35212919 | 0.37 |
ENSDART00000180127
|
ano5a
|
anoctamin 5a |
| chr8_+_30456161 | 0.36 |
ENSDART00000085894
|
pgm5
|
phosphoglucomutase 5 |
| chr7_+_39006837 | 0.35 |
ENSDART00000173735
|
dgkza
|
diacylglycerol kinase, zeta a |
| chr19_+_24394560 | 0.33 |
ENSDART00000142506
|
si:dkey-81h8.1
|
si:dkey-81h8.1 |
| chr11_-_34478225 | 0.32 |
ENSDART00000189604
|
xxylt1
|
xyloside xylosyltransferase 1 |
| chr17_-_50233493 | 0.32 |
ENSDART00000172266
|
fosaa
|
v-fos FBJ murine osteosarcoma viral oncogene homolog Aa |
| chr2_+_37975026 | 0.31 |
ENSDART00000034802
|
si:rp71-1g18.13
|
si:rp71-1g18.13 |
| chr22_-_20259309 | 0.30 |
ENSDART00000139160
|
si:dkey-110c1.10
|
si:dkey-110c1.10 |
| chr11_-_18601955 | 0.29 |
ENSDART00000180565
|
zmynd8
|
zinc finger, MYND-type containing 8 |
| chr9_-_48937240 | 0.29 |
ENSDART00000075627
|
cers6
|
ceramide synthase 6 |
| chr1_-_45662774 | 0.28 |
ENSDART00000042158
|
serhl
|
serine hydrolase-like |
| chr10_+_29260096 | 0.23 |
ENSDART00000088973
|
sytl2a
|
synaptotagmin-like 2a |
| chr1_-_6494384 | 0.23 |
ENSDART00000109356
|
klf7a
|
Kruppel-like factor 7a |
| chr17_-_13026634 | 0.23 |
ENSDART00000113713
|
fam177a1
|
family with sequence similarity 177, member A1 |
| chr5_-_31926906 | 0.22 |
ENSDART00000187340
|
ssh1b
|
slingshot protein phosphatase 1b |
| chr24_+_8904135 | 0.22 |
ENSDART00000066782
|
gnal
|
guanine nucleotide binding protein (G protein), alpha activating activity polypeptide, olfactory type |
| chr6_+_43353271 | 0.21 |
ENSDART00000191775
|
BX927362.2
|
|
| chr24_+_26088143 | 0.19 |
ENSDART00000171129
|
CR352209.1
|
|
| chr23_-_3721444 | 0.19 |
ENSDART00000141682
|
nudt3a
|
nudix (nucleoside diphosphate linked moiety X)-type motif 3a |
| chr5_-_18911114 | 0.19 |
ENSDART00000014434
|
bri3bp
|
bri3 binding protein |
| chr16_-_16225260 | 0.18 |
ENSDART00000165790
|
gra
|
granulito |
| chr7_-_52334840 | 0.18 |
ENSDART00000174173
|
CR938716.1
|
|
| chr4_+_14343706 | 0.17 |
ENSDART00000142845
|
prl2
|
prolactin 2 |
| chr17_+_1992495 | 0.16 |
ENSDART00000192937
|
CABZ01062996.1
|
|
| chr2_-_37353098 | 0.13 |
ENSDART00000056522
|
skila
|
SKI-like proto-oncogene a |
| chr6_+_36942966 | 0.13 |
ENSDART00000028895
|
negr1
|
neuronal growth regulator 1 |
| chr6_-_42003780 | 0.13 |
ENSDART00000032527
|
cav3
|
caveolin 3 |
| chr10_-_8032885 | 0.12 |
ENSDART00000188619
|
atp6v0a2a
|
ATPase H+ transporting V0 subunit a2a |
| chr5_-_25236340 | 0.11 |
ENSDART00000162774
|
abca2
|
ATP-binding cassette, sub-family A (ABC1), member 2 |
| chr2_-_6182098 | 0.10 |
ENSDART00000156167
|
si:ch73-182a11.2
|
si:ch73-182a11.2 |
| chr19_-_14155781 | 0.09 |
ENSDART00000169232
|
nr0b2b
|
nuclear receptor subfamily 0, group B, member 2b |
| chr24_-_28333029 | 0.08 |
ENSDART00000149015
ENSDART00000129174 |
prkag2a
|
protein kinase, AMP-activated, gamma 2 non-catalytic subunit a |
| chr6_-_6248893 | 0.07 |
ENSDART00000124662
|
rtn4a
|
reticulon 4a |
| chr19_-_41069573 | 0.06 |
ENSDART00000111982
ENSDART00000193142 |
sgce
|
sarcoglycan, epsilon |
| chr4_-_13509946 | 0.04 |
ENSDART00000134720
|
ifng1-2
|
interferon, gamma 1-2 |
| chr20_-_7000225 | 0.04 |
ENSDART00000100098
|
adcy1a
|
adenylate cyclase 1a |
| chr20_-_24443680 | 0.03 |
ENSDART00000191337
|
CR385053.3
|
|
| chr11_+_29965822 | 0.03 |
ENSDART00000127049
|
il1fma
|
interleukin-1 family member A |
| chr6_-_46768040 | 0.02 |
ENSDART00000154071
|
igfn1.2
|
immunoglobulin-like and fibronectin type III domain containing 1, tandem duplicate 2 |
| chr5_+_61657702 | 0.02 |
ENSDART00000134387
ENSDART00000171248 |
ap2b1
|
adaptor-related protein complex 2, beta 1 subunit |
| chr6_+_51713076 | 0.01 |
ENSDART00000146281
|
ripor3
|
RIPOR family member 3 |
| chr13_+_42011287 | 0.01 |
ENSDART00000131147
|
cyp1b1
|
cytochrome P450, family 1, subfamily B, polypeptide 1 |
| chr10_+_8688678 | 0.01 |
ENSDART00000147568
|
si:dkey-27b3.4
|
si:dkey-27b3.4 |
| chr3_+_17456428 | 0.01 |
ENSDART00000090676
ENSDART00000182082 |
si:ch211-210g13.5
|
si:ch211-210g13.5 |
Network of associatons between targets according to the STRING database.
First level regulatory network of mef2b
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 2.8 | GO:0097264 | self proteolysis(GO:0097264) |
| 0.6 | 2.6 | GO:0060159 | regulation of dopamine receptor signaling pathway(GO:0060159) |
| 0.6 | 2.3 | GO:0003322 | pancreatic A cell development(GO:0003322) |
| 0.6 | 2.9 | GO:0023041 | neuronal signal transduction(GO:0023041) |
| 0.6 | 4.6 | GO:0032889 | regulation of vacuole fusion, non-autophagic(GO:0032889) |
| 0.6 | 1.7 | GO:0006480 | N-terminal protein amino acid methylation(GO:0006480) |
| 0.5 | 7.1 | GO:0042396 | phosphagen metabolic process(GO:0006599) phosphocreatine metabolic process(GO:0006603) phosphagen biosynthetic process(GO:0042396) phosphocreatine biosynthetic process(GO:0046314) |
| 0.5 | 1.9 | GO:0031446 | regulation of twitch skeletal muscle contraction(GO:0014724) regulation of fast-twitch skeletal muscle fiber contraction(GO:0031446) positive regulation of fast-twitch skeletal muscle fiber contraction(GO:0031448) negative regulation of striated muscle contraction(GO:0045988) relaxation of skeletal muscle(GO:0090076) |
| 0.4 | 2.9 | GO:0046292 | formaldehyde metabolic process(GO:0046292) formaldehyde catabolic process(GO:0046294) |
| 0.4 | 3.0 | GO:0072488 | ammonium transmembrane transport(GO:0072488) |
| 0.3 | 4.5 | GO:0045162 | clustering of voltage-gated sodium channels(GO:0045162) |
| 0.3 | 1.5 | GO:0021731 | trigeminal motor nucleus development(GO:0021731) |
| 0.3 | 0.8 | GO:0030043 | actin filament fragmentation(GO:0030043) |
| 0.3 | 1.0 | GO:0002495 | antigen processing and presentation of peptide antigen via MHC class II(GO:0002495) |
| 0.3 | 2.9 | GO:0030240 | skeletal muscle thin filament assembly(GO:0030240) |
| 0.2 | 2.2 | GO:0043435 | response to corticotropin-releasing hormone(GO:0043435) cellular response to corticotropin-releasing hormone stimulus(GO:0071376) |
| 0.2 | 3.7 | GO:0016486 | peptide hormone processing(GO:0016486) |
| 0.2 | 1.4 | GO:0051045 | negative regulation of membrane protein ectodomain proteolysis(GO:0051045) |
| 0.2 | 3.2 | GO:0035493 | SNARE complex assembly(GO:0035493) |
| 0.2 | 1.3 | GO:2001256 | regulation of store-operated calcium entry(GO:2001256) |
| 0.2 | 2.8 | GO:1901642 | nucleoside transport(GO:0015858) nucleoside transmembrane transport(GO:1901642) |
| 0.2 | 2.8 | GO:0046549 | retinal cone cell development(GO:0046549) |
| 0.2 | 1.4 | GO:0071326 | cellular response to carbohydrate stimulus(GO:0071322) cellular response to monosaccharide stimulus(GO:0071326) cellular response to hexose stimulus(GO:0071331) cellular response to glucose stimulus(GO:0071333) |
| 0.2 | 1.3 | GO:0051694 | pointed-end actin filament capping(GO:0051694) |
| 0.1 | 1.7 | GO:0042044 | fluid transport(GO:0042044) |
| 0.1 | 7.2 | GO:0006096 | glycolytic process(GO:0006096) |
| 0.1 | 2.1 | GO:0048026 | positive regulation of mRNA splicing, via spliceosome(GO:0048026) |
| 0.1 | 3.3 | GO:0033627 | cell adhesion mediated by integrin(GO:0033627) |
| 0.1 | 2.2 | GO:0010923 | negative regulation of phosphatase activity(GO:0010923) |
| 0.1 | 0.9 | GO:0097345 | mitochondrial outer membrane permeabilization(GO:0097345) |
| 0.1 | 0.4 | GO:0010882 | regulation of cardiac muscle contraction by calcium ion signaling(GO:0010882) |
| 0.1 | 0.5 | GO:0051148 | smooth muscle cell differentiation(GO:0051145) negative regulation of muscle cell differentiation(GO:0051148) |
| 0.1 | 2.4 | GO:0070593 | dendrite self-avoidance(GO:0070593) |
| 0.1 | 1.9 | GO:0048936 | peripheral nervous system neuron axonogenesis(GO:0048936) |
| 0.1 | 1.2 | GO:0060012 | synaptic transmission, glycinergic(GO:0060012) |
| 0.1 | 1.8 | GO:0048791 | calcium ion-regulated exocytosis of neurotransmitter(GO:0048791) |
| 0.1 | 3.1 | GO:1901385 | regulation of voltage-gated calcium channel activity(GO:1901385) |
| 0.1 | 1.6 | GO:0021551 | central nervous system morphogenesis(GO:0021551) |
| 0.1 | 0.7 | GO:0060612 | adipose tissue development(GO:0060612) |
| 0.1 | 0.8 | GO:0048672 | positive regulation of collateral sprouting(GO:0048672) |
| 0.1 | 0.5 | GO:0021634 | optic nerve formation(GO:0021634) |
| 0.1 | 0.9 | GO:1905150 | regulation of voltage-gated sodium channel activity(GO:1905150) |
| 0.1 | 1.9 | GO:0055064 | chloride ion homeostasis(GO:0055064) |
| 0.1 | 0.6 | GO:0046548 | retinal rod cell development(GO:0046548) |
| 0.1 | 0.5 | GO:0033539 | fatty acid beta-oxidation using acyl-CoA dehydrogenase(GO:0033539) |
| 0.1 | 0.6 | GO:0001992 | regulation of systemic arterial blood pressure by vasopressin(GO:0001992) |
| 0.0 | 0.6 | GO:0007035 | vacuolar acidification(GO:0007035) pH reduction(GO:0045851) intracellular pH reduction(GO:0051452) |
| 0.0 | 1.5 | GO:0006376 | mRNA splice site selection(GO:0006376) |
| 0.0 | 0.2 | GO:0007191 | adenylate cyclase-activating dopamine receptor signaling pathway(GO:0007191) |
| 0.0 | 1.2 | GO:0051014 | actin filament severing(GO:0051014) |
| 0.0 | 0.2 | GO:1901909 | diphosphoinositol polyphosphate metabolic process(GO:0071543) diadenosine pentaphosphate metabolic process(GO:1901906) diadenosine pentaphosphate catabolic process(GO:1901907) diadenosine hexaphosphate metabolic process(GO:1901908) diadenosine hexaphosphate catabolic process(GO:1901909) adenosine 5'-(hexahydrogen pentaphosphate) metabolic process(GO:1901910) adenosine 5'-(hexahydrogen pentaphosphate) catabolic process(GO:1901911) |
| 0.0 | 2.4 | GO:0015914 | phospholipid transport(GO:0015914) |
| 0.0 | 0.9 | GO:0003009 | skeletal muscle contraction(GO:0003009) |
| 0.0 | 0.4 | GO:0033209 | tumor necrosis factor-mediated signaling pathway(GO:0033209) |
| 0.0 | 0.1 | GO:0042693 | muscle cell fate commitment(GO:0042693) |
| 0.0 | 0.2 | GO:0045947 | negative regulation of translational initiation(GO:0045947) |
| 0.0 | 0.9 | GO:0007602 | phototransduction(GO:0007602) |
| 0.0 | 0.7 | GO:0015908 | fatty acid transport(GO:0015908) |
| 0.0 | 2.1 | GO:0006936 | muscle contraction(GO:0006936) |
| 0.0 | 0.4 | GO:0007205 | protein kinase C-activating G-protein coupled receptor signaling pathway(GO:0007205) |
| 0.0 | 0.5 | GO:0045746 | negative regulation of Notch signaling pathway(GO:0045746) |
| 0.0 | 1.0 | GO:0048477 | oogenesis(GO:0048477) |
| 0.0 | 0.8 | GO:0006006 | glucose metabolic process(GO:0006006) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 4.6 | GO:0000015 | phosphopyruvate hydratase complex(GO:0000015) |
| 0.5 | 1.9 | GO:0031673 | H zone(GO:0031673) |
| 0.5 | 2.8 | GO:0070032 | synaptobrevin 2-SNAP-25-syntaxin-1a-complexin I complex(GO:0070032) |
| 0.3 | 5.3 | GO:0031430 | M band(GO:0031430) |
| 0.1 | 2.6 | GO:0044298 | neuronal cell body membrane(GO:0032809) cell body membrane(GO:0044298) |
| 0.1 | 1.4 | GO:0001669 | acrosomal vesicle(GO:0001669) |
| 0.1 | 2.9 | GO:0030867 | rough endoplasmic reticulum membrane(GO:0030867) |
| 0.1 | 1.8 | GO:0031680 | G-protein beta/gamma-subunit complex(GO:0031680) |
| 0.1 | 3.2 | GO:0097610 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.1 | 2.2 | GO:0031594 | neuromuscular junction(GO:0031594) |
| 0.1 | 3.3 | GO:0008305 | integrin complex(GO:0008305) |
| 0.1 | 0.6 | GO:0000220 | vacuolar proton-transporting V-type ATPase, V0 domain(GO:0000220) |
| 0.1 | 5.1 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.1 | 0.4 | GO:0033018 | sarcoplasmic reticulum lumen(GO:0033018) |
| 0.0 | 3.1 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.0 | 0.6 | GO:0032391 | photoreceptor connecting cilium(GO:0032391) |
| 0.0 | 2.2 | GO:0005865 | striated muscle thin filament(GO:0005865) myofilament(GO:0036379) |
| 0.0 | 2.2 | GO:0042383 | sarcolemma(GO:0042383) |
| 0.0 | 2.0 | GO:0031234 | extrinsic component of cytoplasmic side of plasma membrane(GO:0031234) |
| 0.0 | 0.9 | GO:0030426 | growth cone(GO:0030426) |
| 0.0 | 1.0 | GO:0031902 | late endosome membrane(GO:0031902) |
| 0.0 | 0.4 | GO:0009898 | cytoplasmic side of plasma membrane(GO:0009898) |
| 0.0 | 1.3 | GO:0014069 | postsynaptic density(GO:0014069) |
| 0.0 | 3.8 | GO:0030133 | transport vesicle(GO:0030133) |
| 0.0 | 2.3 | GO:0030424 | axon(GO:0030424) |
| 0.0 | 3.6 | GO:0005667 | transcription factor complex(GO:0005667) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 7.0 | GO:0004743 | pyruvate kinase activity(GO:0004743) |
| 0.6 | 1.9 | GO:0031716 | calcitonin receptor binding(GO:0031716) |
| 0.6 | 2.9 | GO:0004024 | alcohol dehydrogenase activity, zinc-dependent(GO:0004024) S-(hydroxymethyl)glutathione dehydrogenase activity(GO:0051903) |
| 0.6 | 4.6 | GO:0004634 | phosphopyruvate hydratase activity(GO:0004634) |
| 0.6 | 1.7 | GO:0071885 | N-terminal protein N-methyltransferase activity(GO:0071885) |
| 0.5 | 7.1 | GO:0004111 | creatine kinase activity(GO:0004111) phosphotransferase activity, nitrogenous group as acceptor(GO:0016775) |
| 0.3 | 2.8 | GO:0008504 | monoamine transmembrane transporter activity(GO:0008504) |
| 0.3 | 2.4 | GO:0008526 | phosphatidylinositol transporter activity(GO:0008526) |
| 0.3 | 1.4 | GO:0004679 | AMP-activated protein kinase activity(GO:0004679) |
| 0.3 | 1.3 | GO:0035256 | G-protein coupled glutamate receptor binding(GO:0035256) |
| 0.2 | 0.7 | GO:0004466 | long-chain-acyl-CoA dehydrogenase activity(GO:0004466) |
| 0.2 | 9.3 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.2 | 1.6 | GO:0015168 | glycerol transmembrane transporter activity(GO:0015168) |
| 0.2 | 2.2 | GO:0035259 | glucocorticoid receptor binding(GO:0035259) |
| 0.2 | 3.0 | GO:0008519 | ammonium transmembrane transporter activity(GO:0008519) |
| 0.1 | 1.2 | GO:0016934 | extracellular-glycine-gated ion channel activity(GO:0016933) extracellular-glycine-gated chloride channel activity(GO:0016934) |
| 0.1 | 1.7 | GO:0005432 | calcium:sodium antiporter activity(GO:0005432) |
| 0.1 | 3.6 | GO:0004198 | calcium-dependent cysteine-type endopeptidase activity(GO:0004198) |
| 0.1 | 1.8 | GO:0048495 | Roundabout binding(GO:0048495) |
| 0.1 | 0.5 | GO:0004361 | glutaryl-CoA dehydrogenase activity(GO:0004361) |
| 0.1 | 1.9 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.1 | 1.4 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.1 | 3.1 | GO:0008331 | high voltage-gated calcium channel activity(GO:0008331) |
| 0.1 | 0.3 | GO:0004377 | GDP-Man:Man3GlcNAc2-PP-Dol alpha-1,2-mannosyltransferase activity(GO:0004377) |
| 0.1 | 0.4 | GO:0004365 | glyceraldehyde-3-phosphate dehydrogenase (NAD+) (phosphorylating) activity(GO:0004365) glyceraldehyde-3-phosphate dehydrogenase (NAD(P)+) (phosphorylating) activity(GO:0043891) |
| 0.1 | 1.8 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.1 | 2.4 | GO:0098632 | protein binding involved in cell-cell adhesion(GO:0098632) |
| 0.1 | 0.8 | GO:0005165 | nerve growth factor receptor binding(GO:0005163) neurotrophin receptor binding(GO:0005165) |
| 0.1 | 1.3 | GO:0005523 | tropomyosin binding(GO:0005523) |
| 0.1 | 3.3 | GO:0005178 | integrin binding(GO:0005178) |
| 0.1 | 0.4 | GO:0004740 | pyruvate dehydrogenase (acetyl-transferring) kinase activity(GO:0004740) |
| 0.1 | 0.6 | GO:0005000 | vasopressin receptor activity(GO:0005000) |
| 0.0 | 0.3 | GO:0035252 | UDP-xylosyltransferase activity(GO:0035252) |
| 0.0 | 0.6 | GO:0051117 | ATPase binding(GO:0051117) |
| 0.0 | 0.9 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.0 | 2.9 | GO:0018024 | histone-lysine N-methyltransferase activity(GO:0018024) |
| 0.0 | 0.2 | GO:0034432 | endopolyphosphatase activity(GO:0000298) diphosphoinositol-polyphosphate diphosphatase activity(GO:0008486) bis(5'-adenosyl)-hexaphosphatase activity(GO:0034431) bis(5'-adenosyl)-pentaphosphatase activity(GO:0034432) |
| 0.0 | 1.0 | GO:0042287 | MHC protein binding(GO:0042287) |
| 0.0 | 1.9 | GO:0044325 | ion channel binding(GO:0044325) |
| 0.0 | 0.9 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.0 | 0.4 | GO:0016868 | intramolecular transferase activity, phosphotransferases(GO:0016868) |
| 0.0 | 0.2 | GO:0008190 | eukaryotic initiation factor 4E binding(GO:0008190) |
| 0.0 | 0.6 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 0.0 | 1.2 | GO:0005546 | phosphatidylinositol-4,5-bisphosphate binding(GO:0005546) |
| 0.0 | 1.8 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.0 | 1.0 | GO:0004683 | calmodulin-dependent protein kinase activity(GO:0004683) |
| 0.0 | 0.9 | GO:0008020 | G-protein coupled photoreceptor activity(GO:0008020) |
| 0.0 | 0.3 | GO:0050291 | sphingosine N-acyltransferase activity(GO:0050291) |
| 0.0 | 0.7 | GO:0005343 | organic acid:sodium symporter activity(GO:0005343) |
| 0.0 | 3.7 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.0 | 0.4 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.0 | 13.4 | GO:0005509 | calcium ion binding(GO:0005509) |
| 0.0 | 1.5 | GO:0003730 | mRNA 3'-UTR binding(GO:0003730) |
| 0.0 | 1.6 | GO:0004843 | thiol-dependent ubiquitin-specific protease activity(GO:0004843) thiol-dependent ubiquitinyl hydrolase activity(GO:0036459) ubiquitinyl hydrolase activity(GO:0101005) |
| 0.0 | 0.4 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.0 | 3.6 | GO:0051015 | actin filament binding(GO:0051015) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 3.2 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
| 0.1 | 2.2 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.0 | 2.6 | PID P75 NTR PATHWAY | p75(NTR)-mediated signaling |
| 0.0 | 0.8 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.0 | 0.9 | PID MTOR 4PATHWAY | mTOR signaling pathway |
| 0.0 | 0.2 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 0.0 | 0.4 | SIG CD40PATHWAYMAP | Genes related to CD40 signaling |
| 0.0 | 0.4 | ST JNK MAPK PATHWAY | JNK MAPK Pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.1 | 3.2 | REACTOME ACETYLCHOLINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Acetylcholine Neurotransmitter Release Cycle |
| 0.5 | 1.6 | REACTOME PASSIVE TRANSPORT BY AQUAPORINS | Genes involved in Passive Transport by Aquaporins |
| 0.4 | 3.7 | REACTOME PEPTIDE HORMONE BIOSYNTHESIS | Genes involved in Peptide hormone biosynthesis |
| 0.2 | 3.3 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.1 | 1.9 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.1 | 1.9 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.1 | 2.2 | REACTOME PIP3 ACTIVATES AKT SIGNALING | Genes involved in PIP3 activates AKT signaling |
| 0.1 | 0.2 | REACTOME OLFACTORY SIGNALING PATHWAY | Genes involved in Olfactory Signaling Pathway |
| 0.0 | 2.6 | REACTOME G ALPHA I SIGNALLING EVENTS | Genes involved in G alpha (i) signalling events |
| 0.0 | 0.7 | REACTOME AMINO ACID AND OLIGOPEPTIDE SLC TRANSPORTERS | Genes involved in Amino acid and oligopeptide SLC transporters |
| 0.0 | 2.3 | REACTOME ANTIGEN PROCESSING UBIQUITINATION PROTEASOME DEGRADATION | Genes involved in Antigen processing: Ubiquitination & Proteasome degradation |