Project
PRJEB1986: zebrafish developmental stages transcriptome
Navigation
Downloads
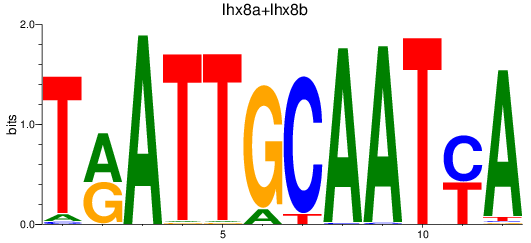
Results for lhx8a+lhx8b
Z-value: 0.85
Transcription factors associated with lhx8a+lhx8b
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
lhx8a
|
ENSDARG00000002330 | LIM homeobox 8a |
|
lhx8b
|
ENSDARG00000042145 | LIM homeobox 8b |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| lhx8b | dr11_v1_chr8_-_17926814_17926814 | -0.74 | 2.9e-04 | Click! |
| lhx8a | dr11_v1_chr2_+_11205795_11205795 | 0.62 | 4.8e-03 | Click! |
Activity profile of lhx8a+lhx8b motif
Sorted Z-values of lhx8a+lhx8b motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr9_-_22318511 | 3.63 |
ENSDART00000129295
|
crygm2d2
|
crystallin, gamma M2d2 |
| chr12_-_4756478 | 3.09 |
ENSDART00000152181
|
mapta
|
microtubule-associated protein tau a |
| chr7_+_26326462 | 3.08 |
ENSDART00000173515
|
zanl
|
zonadhesin, like |
| chr18_+_924949 | 3.02 |
ENSDART00000170888
ENSDART00000193163 |
pkma
|
pyruvate kinase M1/2a |
| chr9_-_22310919 | 2.68 |
ENSDART00000108719
|
crygm2d10
|
crystallin, gamma M2d10 |
| chr18_-_8877077 | 2.65 |
ENSDART00000137266
|
si:dkey-95h12.2
|
si:dkey-95h12.2 |
| chr2_-_293036 | 2.54 |
ENSDART00000171629
|
si:ch73-40a17.3
|
si:ch73-40a17.3 |
| chr17_-_6730247 | 2.38 |
ENSDART00000031091
|
vsnl1b
|
visinin-like 1b |
| chr6_+_40661703 | 2.30 |
ENSDART00000142492
|
eno1b
|
enolase 1b, (alpha) |
| chr4_+_5531583 | 2.27 |
ENSDART00000137500
ENSDART00000042080 |
si:dkey-14d8.6
|
si:dkey-14d8.6 |
| chr18_-_898870 | 2.25 |
ENSDART00000151777
ENSDART00000062654 |
parp6a
|
poly (ADP-ribose) polymerase family, member 6a |
| chr19_+_10339538 | 2.24 |
ENSDART00000151808
ENSDART00000151235 |
rcvrn3
|
recoverin 3 |
| chr1_+_56274613 | 2.19 |
ENSDART00000052683
|
c3a.3
|
complement component c3a, duplicate 3 |
| chr13_+_23988442 | 2.12 |
ENSDART00000010918
|
agt
|
angiotensinogen |
| chr16_+_45739193 | 1.93 |
ENSDART00000184852
ENSDART00000156851 ENSDART00000154704 |
paqr6
|
progestin and adipoQ receptor family member VI |
| chr7_+_53152108 | 1.83 |
ENSDART00000171350
|
cdh29
|
cadherin 29 |
| chr7_+_2467057 | 1.81 |
ENSDART00000154517
|
si:dkey-125e8.3
|
si:dkey-125e8.3 |
| chr19_-_40191358 | 1.81 |
ENSDART00000183919
|
grn1
|
granulin 1 |
| chr9_-_41277347 | 1.81 |
ENSDART00000181213
|
glsb
|
glutaminase b |
| chr5_-_54197084 | 1.67 |
ENSDART00000163640
|
grk1b
|
G protein-coupled receptor kinase 1 b |
| chr24_+_792429 | 1.64 |
ENSDART00000082523
|
impa2
|
inositol(myo)-1(or 4)-monophosphatase 2 |
| chr3_-_28428198 | 1.62 |
ENSDART00000151546
|
rbfox1
|
RNA binding fox-1 homolog 1 |
| chr5_+_32247310 | 1.62 |
ENSDART00000182649
|
myha
|
myosin, heavy chain a |
| chr19_-_31042570 | 1.60 |
ENSDART00000144337
ENSDART00000136213 ENSDART00000133101 ENSDART00000190949 |
bzw2
|
basic leucine zipper and W2 domains 2 |
| chr24_+_39137001 | 1.60 |
ENSDART00000181086
ENSDART00000183724 ENSDART00000193466 |
tbc1d24
|
TBC1 domain family, member 24 |
| chr6_+_2195625 | 1.59 |
ENSDART00000155659
|
acvr1bb
|
activin A receptor type 1Bb |
| chr3_-_15264698 | 1.58 |
ENSDART00000111948
ENSDART00000142594 |
sez6l2
|
seizure related 6 homolog (mouse)-like 2 |
| chr2_-_34555945 | 1.58 |
ENSDART00000056671
|
brinp2
|
bone morphogenetic protein/retinoic acid inducible neural-specific 2 |
| chr8_+_3820134 | 1.56 |
ENSDART00000122454
|
citb
|
citron rho-interacting serine/threonine kinase b |
| chr8_-_53198154 | 1.55 |
ENSDART00000083416
|
gabrd
|
gamma-aminobutyric acid (GABA) A receptor, delta |
| chr11_+_30981777 | 1.50 |
ENSDART00000148949
|
cacna1ab
|
calcium channel, voltage-dependent, P/Q type, alpha 1A subunit, b |
| chr5_+_27137473 | 1.48 |
ENSDART00000181833
|
unc5db
|
unc-5 netrin receptor Db |
| chr14_+_17197132 | 1.48 |
ENSDART00000054598
|
rtn4rl2b
|
reticulon 4 receptor-like 2b |
| chr4_-_17785120 | 1.48 |
ENSDART00000024775
|
mybpc1
|
myosin binding protein C, slow type |
| chr1_+_7546259 | 1.46 |
ENSDART00000015732
|
mylz3
|
myosin, light polypeptide 3, skeletal muscle |
| chr18_+_7286788 | 1.39 |
ENSDART00000022998
|
ANO2 (1 of many)
|
si:ch73-86n2.1 |
| chr3_-_32169754 | 1.37 |
ENSDART00000179010
|
tnnt1
|
troponin T type 1 (skeletal, slow) |
| chr14_-_33105434 | 1.36 |
ENSDART00000163795
|
dlg3
|
discs, large homolog 3 (Drosophila) |
| chr9_+_49712868 | 1.30 |
ENSDART00000192969
ENSDART00000183310 |
CSRNP3
|
cysteine and serine rich nuclear protein 3 |
| chr5_-_22882210 | 1.21 |
ENSDART00000182232
|
si:ch211-26b3.4
|
si:ch211-26b3.4 |
| chr3_+_25154078 | 1.19 |
ENSDART00000156973
|
si:ch211-256m1.8
|
si:ch211-256m1.8 |
| chr17_-_2386569 | 1.19 |
ENSDART00000121614
|
PLCB2
|
phospholipase C beta 2 |
| chr6_+_11250033 | 1.11 |
ENSDART00000065411
ENSDART00000132677 |
atg9a
|
ATG9 autophagy related 9 homolog A (S. cerevisiae) |
| chr2_+_20332044 | 1.06 |
ENSDART00000112131
|
plppr4a
|
phospholipid phosphatase related 4a |
| chr18_+_9635178 | 1.06 |
ENSDART00000183486
|
pclob
|
piccolo presynaptic cytomatrix protein b |
| chr10_-_7974155 | 1.06 |
ENSDART00000147368
ENSDART00000075524 |
osbp2
|
oxysterol binding protein 2 |
| chr19_+_4139065 | 1.03 |
ENSDART00000172524
|
si:dkey-218f9.10
|
si:dkey-218f9.10 |
| chr8_+_7737062 | 1.01 |
ENSDART00000166712
|
fgd1
|
FYVE, RhoGEF and PH domain containing 1 |
| chr20_-_51727860 | 1.01 |
ENSDART00000147044
|
brox
|
BRO1 domain and CAAX motif containing |
| chr21_-_36972127 | 0.98 |
ENSDART00000100310
|
dbn1
|
drebrin 1 |
| chr6_+_11249706 | 0.98 |
ENSDART00000186547
ENSDART00000193287 |
atg9a
|
ATG9 autophagy related 9 homolog A (S. cerevisiae) |
| chr22_+_1779401 | 0.96 |
ENSDART00000170126
|
znf1154
|
zinc finger protein 1154 |
| chr9_-_18568927 | 0.96 |
ENSDART00000084668
|
enox1
|
ecto-NOX disulfide-thiol exchanger 1 |
| chr21_-_36571804 | 0.94 |
ENSDART00000138129
|
wwc1
|
WW and C2 domain containing 1 |
| chr20_-_39273505 | 0.92 |
ENSDART00000153114
|
clu
|
clusterin |
| chr1_-_58064738 | 0.92 |
ENSDART00000073778
|
caspb
|
caspase b |
| chr23_-_21797517 | 0.91 |
ENSDART00000110041
|
lrrc38a
|
leucine rich repeat containing 38a |
| chr8_+_25351863 | 0.91 |
ENSDART00000034092
|
dnase1l1l
|
deoxyribonuclease I-like 1-like |
| chr11_-_43200994 | 0.89 |
ENSDART00000164700
|
sptbn1
|
spectrin, beta, non-erythrocytic 1 |
| chr21_-_13662237 | 0.89 |
ENSDART00000091647
ENSDART00000151547 |
pnpla7a
|
patatin-like phospholipase domain containing 7a |
| chr6_-_57938043 | 0.88 |
ENSDART00000171073
|
tox2
|
TOX high mobility group box family member 2 |
| chr11_+_21050326 | 0.88 |
ENSDART00000065984
|
zgc:113307
|
zgc:113307 |
| chr16_-_27749172 | 0.88 |
ENSDART00000145198
|
steap4
|
STEAP family member 4 |
| chr6_+_11250316 | 0.87 |
ENSDART00000137122
|
atg9a
|
ATG9 autophagy related 9 homolog A (S. cerevisiae) |
| chr2_+_20331445 | 0.86 |
ENSDART00000186880
|
plppr4a
|
phospholipid phosphatase related 4a |
| chr11_+_25276748 | 0.86 |
ENSDART00000126211
|
cyldb
|
cylindromatosis (turban tumor syndrome), b |
| chr2_-_14387335 | 0.86 |
ENSDART00000189332
ENSDART00000164786 ENSDART00000188572 |
sgip1b
|
SH3-domain GRB2-like (endophilin) interacting protein 1b |
| chr23_-_41965557 | 0.84 |
ENSDART00000144183
|
slc1a7b
|
solute carrier family 1 (glutamate transporter), member 7b |
| chr21_+_26612777 | 0.84 |
ENSDART00000142667
|
esrra
|
estrogen-related receptor alpha |
| chr6_-_957830 | 0.84 |
ENSDART00000090019
ENSDART00000184286 |
zeb2b
|
zinc finger E-box binding homeobox 2b |
| chr16_-_12319822 | 0.84 |
ENSDART00000127453
ENSDART00000184526 |
trpv6
|
transient receptor potential cation channel, subfamily V, member 6 |
| chr20_-_38610360 | 0.83 |
ENSDART00000168783
ENSDART00000164638 |
slc30a2
|
solute carrier family 30 (zinc transporter), member 2 |
| chr8_+_6967108 | 0.83 |
ENSDART00000004588
|
asic1a
|
acid-sensing (proton-gated) ion channel 1a |
| chr3_+_15773991 | 0.81 |
ENSDART00000089923
|
znf652
|
zinc finger protein 652 |
| chr21_-_10488379 | 0.81 |
ENSDART00000163878
|
hcn1
|
hyperpolarization activated cyclic nucleotide-gated potassium channel 1 |
| chr7_+_26224211 | 0.80 |
ENSDART00000173999
|
vgf
|
VGF nerve growth factor inducible |
| chr2_-_20470125 | 0.80 |
ENSDART00000114899
ENSDART00000147806 |
frrs1a
|
ferric-chelate reductase 1a |
| chr19_-_6303099 | 0.78 |
ENSDART00000133371
|
pou2f2a
|
POU class 2 homeobox 2a |
| chr4_-_287425 | 0.78 |
ENSDART00000159128
|
echdc3
|
enoyl CoA hydratase domain containing 3 |
| chr7_-_39378903 | 0.78 |
ENSDART00000173659
|
slc8b1
|
solute carrier family 8 (sodium/lithium/calcium exchanger), member B1 |
| chr20_+_18551657 | 0.77 |
ENSDART00000147001
|
si:dkeyp-72h1.1
|
si:dkeyp-72h1.1 |
| chr10_+_29199172 | 0.77 |
ENSDART00000148828
|
picalma
|
phosphatidylinositol binding clathrin assembly protein a |
| chr16_-_35329803 | 0.75 |
ENSDART00000161729
ENSDART00000157700 ENSDART00000184584 ENSDART00000174713 ENSDART00000162518 |
ptprub
|
protein tyrosine phosphatase, receptor type, U, b |
| chr10_+_21804772 | 0.75 |
ENSDART00000162194
|
pcdh1g31
|
protocadherin 1 gamma 31 |
| chr23_+_34321237 | 0.75 |
ENSDART00000173272
|
plxna1a
|
plexin A1a |
| chr22_-_15562933 | 0.74 |
ENSDART00000141528
|
ankmy1
|
ankyrin repeat and MYND domain containing 1 |
| chr7_+_72469724 | 0.73 |
ENSDART00000187629
|
HECTD4
|
HECT domain E3 ubiquitin protein ligase 4 |
| chr25_-_35147698 | 0.73 |
ENSDART00000121963
|
zgc:165555
|
zgc:165555 |
| chr9_+_6079528 | 0.72 |
ENSDART00000142167
|
st6gal2a
|
ST6 beta-galactosamide alpha-2,6-sialyltranferase 2a |
| chr20_+_42563618 | 0.71 |
ENSDART00000153441
|
igf2r
|
insulin-like growth factor 2 receptor |
| chr15_+_25683069 | 0.71 |
ENSDART00000148190
|
hic1
|
hypermethylated in cancer 1 |
| chr7_+_2228276 | 0.70 |
ENSDART00000064294
|
si:dkey-187j14.4
|
si:dkey-187j14.4 |
| chr3_+_16724614 | 0.70 |
ENSDART00000182135
|
gys1
|
glycogen synthase 1 (muscle) |
| chr17_-_8173380 | 0.70 |
ENSDART00000148520
|
syne1b
|
spectrin repeat containing, nuclear envelope 1b |
| chr14_+_19258702 | 0.69 |
ENSDART00000187087
ENSDART00000005738 |
slitrk2
|
SLIT and NTRK-like family, member 2 |
| chr9_-_34986827 | 0.69 |
ENSDART00000137862
|
si:ch211-160b11.4
|
si:ch211-160b11.4 |
| chr23_-_29668286 | 0.69 |
ENSDART00000129248
|
clstn1
|
calsyntenin 1 |
| chr2_+_45548890 | 0.69 |
ENSDART00000113994
|
fndc7a
|
fibronectin type III domain containing 7a |
| chr8_-_18316482 | 0.67 |
ENSDART00000190156
|
rnf220b
|
ring finger protein 220b |
| chr15_+_32268790 | 0.66 |
ENSDART00000154457
|
fhdc4
|
FH2 domain containing 4 |
| chr3_+_49021079 | 0.66 |
ENSDART00000162012
|
zgc:163083
|
zgc:163083 |
| chr2_-_34483597 | 0.66 |
ENSDART00000133224
|
brinp2
|
bone morphogenetic protein/retinoic acid inducible neural-specific 2 |
| chr21_+_9997418 | 0.65 |
ENSDART00000181454
ENSDART00000171579 |
herc7
|
hect domain and RLD 7 |
| chr20_-_39273987 | 0.64 |
ENSDART00000127173
|
clu
|
clusterin |
| chr21_+_13861589 | 0.64 |
ENSDART00000015629
ENSDART00000171306 |
stxbp1a
|
syntaxin binding protein 1a |
| chr1_-_23308225 | 0.64 |
ENSDART00000137567
ENSDART00000008201 |
smim14
|
small integral membrane protein 14 |
| chr17_+_12138715 | 0.64 |
ENSDART00000112888
|
SCCPDH (1 of many)
|
zgc:174379 |
| chr18_+_31410652 | 0.63 |
ENSDART00000098504
|
def8
|
differentially expressed in FDCP 8 homolog (mouse) |
| chr14_-_8625845 | 0.63 |
ENSDART00000054760
|
zgc:162144
|
zgc:162144 |
| chr18_-_7539166 | 0.63 |
ENSDART00000133541
|
si:dkey-30c15.2
|
si:dkey-30c15.2 |
| chr4_-_12766418 | 0.62 |
ENSDART00000024312
|
dera
|
deoxyribose-phosphate aldolase (putative) |
| chr3_-_55197881 | 0.61 |
ENSDART00000179903
|
kank2
|
KN motif and ankyrin repeat domains 2 |
| chr3_-_30384353 | 0.61 |
ENSDART00000186832
|
lrrc4ba
|
leucine rich repeat containing 4Ba |
| chr9_-_52847881 | 0.61 |
ENSDART00000166407
|
trpm2
|
transient receptor potential cation channel, subfamily M, member 2 |
| chr18_+_8340886 | 0.60 |
ENSDART00000081132
|
cpt1b
|
carnitine palmitoyltransferase 1B (muscle) |
| chr2_-_32551178 | 0.60 |
ENSDART00000145603
|
smarcd3a
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily d, member 3a |
| chr10_-_7666810 | 0.60 |
ENSDART00000191479
|
pcyox1
|
prenylcysteine oxidase 1 |
| chr1_-_2457546 | 0.58 |
ENSDART00000103795
|
ggact.1
|
gamma-glutamylamine cyclotransferase, tandem duplicate 1 |
| chr19_+_20801486 | 0.58 |
ENSDART00000038202
|
cndp2
|
carnosine dipeptidase 2 |
| chr2_+_37227011 | 0.58 |
ENSDART00000126587
ENSDART00000084958 |
samd7
|
sterile alpha motif domain containing 7 |
| chr20_+_23173710 | 0.57 |
ENSDART00000074172
|
sgcb
|
sarcoglycan, beta (dystrophin-associated glycoprotein) |
| chr11_-_891674 | 0.57 |
ENSDART00000172904
|
atg7
|
ATG7 autophagy related 7 homolog (S. cerevisiae) |
| chr22_+_38164486 | 0.57 |
ENSDART00000137521
|
tm4sf18
|
transmembrane 4 L six family member 18 |
| chr20_-_25436389 | 0.57 |
ENSDART00000153266
|
itsn2a
|
intersectin 2a |
| chr22_-_7129631 | 0.55 |
ENSDART00000171359
|
asic1b
|
acid-sensing (proton-gated) ion channel 1b |
| chr1_+_52929185 | 0.55 |
ENSDART00000147683
|
inpp4b
|
inositol polyphosphate-4-phosphatase type II B |
| chr23_+_4362463 | 0.55 |
ENSDART00000039829
|
zgc:112175
|
zgc:112175 |
| chr25_-_28768489 | 0.55 |
ENSDART00000088315
|
washc4
|
WASH complex subunit 4 |
| chr15_-_45356635 | 0.53 |
ENSDART00000192000
|
CABZ01068246.1
|
|
| chr4_+_612363 | 0.52 |
ENSDART00000049154
|
pthlha
|
parathyroid hormone-like hormone a |
| chr6_-_12456077 | 0.52 |
ENSDART00000190107
|
gpd2
|
glycerol-3-phosphate dehydrogenase 2 (mitochondrial) |
| chr21_-_26918901 | 0.52 |
ENSDART00000100685
|
lrfn4a
|
leucine rich repeat and fibronectin type III domain containing 4a |
| chr5_+_51909740 | 0.52 |
ENSDART00000162541
|
thbs4a
|
thrombospondin 4a |
| chr24_+_36339242 | 0.51 |
ENSDART00000105686
ENSDART00000142264 |
grnb
|
granulin b |
| chr5_-_30080332 | 0.51 |
ENSDART00000140049
|
bco2a
|
beta-carotene oxygenase 2a |
| chr20_+_28434196 | 0.50 |
ENSDART00000034245
|
dpf3
|
D4, zinc and double PHD fingers, family 3 |
| chr18_-_46957840 | 0.50 |
ENSDART00000182754
|
gramd1bb
|
GRAM domain containing 1Bb |
| chr16_-_45209684 | 0.50 |
ENSDART00000184595
|
fxyd1
|
FXYD domain containing ion transport regulator 1 (phospholemman) |
| chr5_-_23749348 | 0.49 |
ENSDART00000140766
|
gbgt1l2
|
globoside alpha-1,3-N-acetylgalactosaminyltransferase 1, like 2 |
| chr8_-_2289264 | 0.49 |
ENSDART00000189397
|
plat
|
plasminogen activator, tissue |
| chr15_-_4094888 | 0.48 |
ENSDART00000166307
|
TM4SF19
|
si:dkey-83h2.3 |
| chr7_+_5959203 | 0.47 |
ENSDART00000110813
|
zgc:165555
|
zgc:165555 |
| chr9_-_16877456 | 0.47 |
ENSDART00000161105
ENSDART00000160869 |
fbxl3a
|
F-box and leucine-rich repeat protein 3a |
| chr12_+_42574148 | 0.47 |
ENSDART00000157855
|
ebf3a
|
early B cell factor 3a |
| chr10_-_42131408 | 0.47 |
ENSDART00000076693
|
stambpa
|
STAM binding protein a |
| chr15_-_25613114 | 0.46 |
ENSDART00000182431
ENSDART00000187405 |
taok1a
|
TAO kinase 1a |
| chr11_-_10828539 | 0.46 |
ENSDART00000040180
|
tbr1a
|
T-box, brain, 1a |
| chr4_+_11479705 | 0.46 |
ENSDART00000019458
ENSDART00000150587 ENSDART00000139370 ENSDART00000135826 |
asb13a.1
|
ankyrin repeat and SOCS box containing 13a, tandem duplicate 1 |
| chr6_+_12865137 | 0.45 |
ENSDART00000090065
|
fam117ba
|
family with sequence similarity 117, member Ba |
| chr4_+_31405646 | 0.44 |
ENSDART00000128732
|
si:rp71-5o12.3
|
si:rp71-5o12.3 |
| chr22_-_910926 | 0.44 |
ENSDART00000180075
|
FP016205.1
|
|
| chr21_-_33995710 | 0.44 |
ENSDART00000100508
ENSDART00000179622 |
ebf1b
|
early B cell factor 1b |
| chr13_-_33667922 | 0.43 |
ENSDART00000141631
|
si:dkey-76k16.5
|
si:dkey-76k16.5 |
| chr22_-_23000815 | 0.43 |
ENSDART00000137111
|
ptprc
|
protein tyrosine phosphatase, receptor type, C |
| chr2_-_3403020 | 0.43 |
ENSDART00000092741
|
snap47
|
synaptosomal-associated protein, 47 |
| chr18_+_19120984 | 0.43 |
ENSDART00000141501
|
si:dkey-242h9.3
|
si:dkey-242h9.3 |
| chr25_+_19238175 | 0.43 |
ENSDART00000110730
ENSDART00000193619 ENSDART00000154420 |
ppip5k1b
|
diphosphoinositol pentakisphosphate kinase 1b |
| chr1_+_10019653 | 0.43 |
ENSDART00000190227
|
trim2b
|
tripartite motif containing 2b |
| chr9_-_4606463 | 0.43 |
ENSDART00000179110
|
galnt13
|
UDP-N-acetyl-alpha-D-galactosamine:polypeptide N-acetylgalactosaminyltransferase 13 |
| chr25_-_27621268 | 0.42 |
ENSDART00000146205
ENSDART00000073511 |
hyal6
|
hyaluronoglucosaminidase 6 |
| chr16_+_4838808 | 0.42 |
ENSDART00000179363
|
ibtk
|
inhibitor of Bruton agammaglobulinemia tyrosine kinase |
| chr22_-_29586608 | 0.41 |
ENSDART00000059869
|
adra2a
|
adrenoceptor alpha 2A |
| chr20_-_49681850 | 0.41 |
ENSDART00000025926
|
col12a1b
|
collagen, type XII, alpha 1b |
| chr17_+_15559046 | 0.40 |
ENSDART00000187126
|
bach2a
|
BTB and CNC homology 1, basic leucine zipper transcription factor 2a |
| chr8_+_26254903 | 0.39 |
ENSDART00000113763
|
slc26a6
|
solute carrier family 26, member 6 |
| chr3_-_24456451 | 0.38 |
ENSDART00000024480
ENSDART00000156814 |
baiap2l2a
|
BAI1-associated protein 2-like 2a |
| chr10_+_21656654 | 0.38 |
ENSDART00000160464
|
pcdh1g2
|
protocadherin 1 gamma 2 |
| chr4_+_45458619 | 0.38 |
ENSDART00000124892
|
si:dkey-256i11.3
|
si:dkey-256i11.3 |
| chr5_-_35888499 | 0.37 |
ENSDART00000193932
|
rxfp2l
|
relaxin/insulin-like family peptide receptor 2, like |
| chr2_+_49644803 | 0.37 |
ENSDART00000160342
|
si:ch211-209f23.7
|
si:ch211-209f23.7 |
| chr7_-_51727760 | 0.37 |
ENSDART00000174180
|
hdac8
|
histone deacetylase 8 |
| chr6_+_9289802 | 0.36 |
ENSDART00000188650
|
kalrnb
|
kalirin RhoGEF kinase b |
| chr16_-_35427060 | 0.36 |
ENSDART00000172294
|
ctps1b
|
CTP synthase 1b |
| chr3_+_464127 | 0.36 |
ENSDART00000169510
ENSDART00000144886 |
dicp1.1
|
diverse immunoglobulin domain-containing protein 1.1 |
| chr4_+_42878556 | 0.36 |
ENSDART00000171047
|
si:zfos-451d2.2
|
si:zfos-451d2.2 |
| chr23_+_21278948 | 0.36 |
ENSDART00000156701
ENSDART00000033970 |
ubr4
|
ubiquitin protein ligase E3 component n-recognin 4 |
| chr7_-_1101694 | 0.36 |
ENSDART00000183017
ENSDART00000182681 |
dctn1a
|
dynactin 1a |
| chr13_+_17702522 | 0.35 |
ENSDART00000057913
|
si:dkey-27m7.4
|
si:dkey-27m7.4 |
| chr13_-_1085961 | 0.35 |
ENSDART00000171872
ENSDART00000166598 |
plek
|
pleckstrin |
| chr13_-_8892514 | 0.35 |
ENSDART00000144553
|
plekhh2
|
pleckstrin homology domain containing, family H (with MyTH4 domain) member 2 |
| chr16_+_25535993 | 0.35 |
ENSDART00000077436
|
mylipb
|
myosin regulatory light chain interacting protein b |
| chr4_+_15605844 | 0.34 |
ENSDART00000101619
ENSDART00000021384 |
exoc4
|
exocyst complex component 4 |
| chr22_-_23706771 | 0.34 |
ENSDART00000159771
|
cfhl1
|
complement factor H like 1 |
| chr7_+_27253063 | 0.34 |
ENSDART00000191138
|
sox6
|
SRY (sex determining region Y)-box 6 |
| chr1_+_7414318 | 0.34 |
ENSDART00000127426
|
si:ch73-383l1.1
|
si:ch73-383l1.1 |
| chr18_-_7539469 | 0.34 |
ENSDART00000101296
|
si:dkey-30c15.2
|
si:dkey-30c15.2 |
| chr4_-_4751981 | 0.33 |
ENSDART00000147436
ENSDART00000092984 ENSDART00000158466 |
creb3l2
|
cAMP responsive element binding protein 3-like 2 |
| chr16_+_49088698 | 0.33 |
ENSDART00000182037
|
plcl2
|
phospholipase C like 2 |
| chr25_+_28893615 | 0.32 |
ENSDART00000156994
ENSDART00000075151 |
amn1
|
antagonist of mitotic exit network 1 homolog (S. cerevisiae) |
| chr22_+_9069081 | 0.32 |
ENSDART00000187842
|
BX248395.1
|
|
| chr22_-_8720794 | 0.31 |
ENSDART00000191236
ENSDART00000185738 |
si:ch73-27e22.1
si:ch73-27e22.8
|
si:ch73-27e22.1 si:ch73-27e22.8 |
| chr6_-_16456093 | 0.31 |
ENSDART00000083305
ENSDART00000181640 |
slc19a2
|
solute carrier family 19 (thiamine transporter), member 2 |
| chr22_+_6254194 | 0.31 |
ENSDART00000112388
ENSDART00000135176 |
rnasel4
|
ribonuclease like 4 |
| chr4_+_65614183 | 0.31 |
ENSDART00000180387
|
si:dkey-205i10.1
|
si:dkey-205i10.1 |
| chr24_+_12945803 | 0.30 |
ENSDART00000005105
|
psme1
|
proteasome activator subunit 1 |
| chr22_+_31295791 | 0.29 |
ENSDART00000092447
|
grip2b
|
glutamate receptor interacting protein 2b |
| chr1_-_59077650 | 0.29 |
ENSDART00000043516
|
MFAP4 (1 of many)
|
si:zfos-2330d3.1 |
| chr22_-_9694433 | 0.29 |
ENSDART00000180101
|
si:dkey-286j17.4
|
si:dkey-286j17.4 |
| chr15_-_23645810 | 0.28 |
ENSDART00000168845
|
ckmb
|
creatine kinase, muscle b |
| chr1_-_40086978 | 0.28 |
ENSDART00000136990
|
si:dkey-117m1.4
|
si:dkey-117m1.4 |
Network of associatons between targets according to the STRING database.
First level regulatory network of lhx8a+lhx8b
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 2.7 | GO:0048245 | eosinophil chemotaxis(GO:0048245) eosinophil migration(GO:0072677) |
| 0.4 | 3.9 | GO:0036368 | cone photoresponse recovery(GO:0036368) |
| 0.4 | 3.5 | GO:0044805 | late nucleophagy(GO:0044805) |
| 0.3 | 2.3 | GO:0032889 | regulation of vacuole fusion, non-autophagic(GO:0032889) |
| 0.2 | 1.8 | GO:0006537 | glutamate biosynthetic process(GO:0006537) glutamine catabolic process(GO:0006543) |
| 0.2 | 2.2 | GO:0071300 | cellular response to retinoic acid(GO:0071300) |
| 0.2 | 0.9 | GO:0015677 | copper ion import(GO:0015677) |
| 0.2 | 0.8 | GO:0070317 | G0 to G1 transition(GO:0045023) regulation of G0 to G1 transition(GO:0070316) negative regulation of G0 to G1 transition(GO:0070317) |
| 0.2 | 1.4 | GO:0070207 | protein homotrimerization(GO:0070207) |
| 0.2 | 1.0 | GO:0098974 | postsynaptic actin cytoskeleton organization(GO:0098974) |
| 0.2 | 1.4 | GO:0031444 | slow-twitch skeletal muscle fiber contraction(GO:0031444) |
| 0.2 | 1.0 | GO:0007624 | ultradian rhythm(GO:0007624) |
| 0.2 | 2.1 | GO:0006020 | inositol metabolic process(GO:0006020) |
| 0.2 | 0.5 | GO:0002076 | osteoblast development(GO:0002076) |
| 0.2 | 0.7 | GO:0030951 | establishment or maintenance of microtubule cytoskeleton polarity(GO:0030951) |
| 0.2 | 0.8 | GO:0032119 | sequestering of zinc ion(GO:0032119) regulation of sequestering of zinc ion(GO:0061088) |
| 0.1 | 1.5 | GO:0038007 | netrin-activated signaling pathway(GO:0038007) |
| 0.1 | 0.9 | GO:0032475 | otolith formation(GO:0032475) |
| 0.1 | 0.6 | GO:0033147 | negative regulation of intracellular estrogen receptor signaling pathway(GO:0033147) |
| 0.1 | 0.5 | GO:0014909 | smooth muscle cell migration(GO:0014909) |
| 0.1 | 0.6 | GO:0030329 | prenylated protein catabolic process(GO:0030327) prenylcysteine catabolic process(GO:0030328) prenylcysteine metabolic process(GO:0030329) |
| 0.1 | 2.1 | GO:0001990 | regulation of systemic arterial blood pressure by hormone(GO:0001990) |
| 0.1 | 0.3 | GO:0070278 | extracellular matrix constituent secretion(GO:0070278) |
| 0.1 | 1.1 | GO:1904071 | presynaptic active zone assembly(GO:1904071) |
| 0.1 | 0.4 | GO:0044210 | 'de novo' CTP biosynthetic process(GO:0044210) |
| 0.1 | 0.9 | GO:0070266 | necroptotic process(GO:0070266) |
| 0.1 | 1.2 | GO:0016540 | protein autoprocessing(GO:0016540) |
| 0.1 | 0.4 | GO:0071881 | adenylate cyclase-inhibiting adrenergic receptor signaling pathway(GO:0071881) |
| 0.1 | 0.5 | GO:0021877 | forebrain neuron fate commitment(GO:0021877) |
| 0.1 | 0.8 | GO:0035677 | posterior lateral line neuromast hair cell development(GO:0035677) |
| 0.1 | 1.5 | GO:0007274 | neuromuscular synaptic transmission(GO:0007274) |
| 0.1 | 0.6 | GO:0032986 | nucleosome disassembly(GO:0006337) chromatin disassembly(GO:0031498) protein-DNA complex disassembly(GO:0032986) |
| 0.1 | 0.4 | GO:0051454 | pH elevation(GO:0045852) intracellular pH elevation(GO:0051454) |
| 0.1 | 2.3 | GO:0006471 | protein ADP-ribosylation(GO:0006471) |
| 0.1 | 1.0 | GO:0000737 | DNA catabolic process, endonucleolytic(GO:0000737) |
| 0.1 | 2.3 | GO:0030199 | collagen fibril organization(GO:0030199) |
| 0.1 | 0.5 | GO:0016121 | carotene metabolic process(GO:0016119) carotene catabolic process(GO:0016121) terpene metabolic process(GO:0042214) terpene catabolic process(GO:0046247) |
| 0.1 | 3.0 | GO:0006096 | glycolytic process(GO:0006096) |
| 0.1 | 0.6 | GO:0009264 | deoxyribonucleotide catabolic process(GO:0009264) deoxyribose phosphate catabolic process(GO:0046386) |
| 0.0 | 0.1 | GO:0001779 | natural killer cell differentiation(GO:0001779) |
| 0.0 | 0.4 | GO:0070933 | histone H4 deacetylation(GO:0070933) |
| 0.0 | 1.4 | GO:0048268 | clathrin coat assembly(GO:0048268) |
| 0.0 | 5.7 | GO:0002088 | lens development in camera-type eye(GO:0002088) |
| 0.0 | 0.5 | GO:0006072 | glycerol-3-phosphate metabolic process(GO:0006072) |
| 0.0 | 0.4 | GO:2000251 | positive regulation of actin cytoskeleton reorganization(GO:2000251) |
| 0.0 | 1.1 | GO:0040023 | nuclear migration(GO:0007097) establishment of nucleus localization(GO:0040023) |
| 0.0 | 0.2 | GO:0019483 | beta-alanine metabolic process(GO:0019482) beta-alanine biosynthetic process(GO:0019483) |
| 0.0 | 0.3 | GO:0032228 | regulation of synaptic transmission, GABAergic(GO:0032228) |
| 0.0 | 1.2 | GO:0061035 | regulation of cartilage development(GO:0061035) |
| 0.0 | 0.5 | GO:0042396 | phosphagen metabolic process(GO:0006599) phosphocreatine metabolic process(GO:0006603) phosphagen biosynthetic process(GO:0042396) phosphocreatine biosynthetic process(GO:0046314) |
| 0.0 | 2.2 | GO:0006956 | complement activation(GO:0006956) |
| 0.0 | 1.6 | GO:0021854 | hypothalamus development(GO:0021854) |
| 0.0 | 0.6 | GO:0009437 | carnitine metabolic process(GO:0009437) |
| 0.0 | 0.4 | GO:0030214 | hyaluronan catabolic process(GO:0030214) |
| 0.0 | 0.7 | GO:1902285 | semaphorin-plexin signaling pathway involved in neuron projection guidance(GO:1902285) semaphorin-plexin signaling pathway involved in axon guidance(GO:1902287) |
| 0.0 | 1.4 | GO:0097120 | receptor localization to synapse(GO:0097120) |
| 0.0 | 0.5 | GO:0030259 | lipid glycosylation(GO:0030259) |
| 0.0 | 0.8 | GO:0001757 | somite specification(GO:0001757) |
| 0.0 | 0.4 | GO:0007190 | activation of adenylate cyclase activity(GO:0007190) positive regulation of cAMP metabolic process(GO:0030816) positive regulation of cAMP biosynthetic process(GO:0030819) positive regulation of adenylate cyclase activity(GO:0045762) |
| 0.0 | 0.6 | GO:0007032 | endosome organization(GO:0007032) |
| 0.0 | 0.3 | GO:0099638 | neurotransmitter receptor transport, endosome to postsynaptic membrane(GO:0098887) endosome to plasma membrane protein transport(GO:0099638) neurotransmitter receptor transport, endosome to plasma membrane(GO:0099639) |
| 0.0 | 1.1 | GO:0032924 | activin receptor signaling pathway(GO:0032924) |
| 0.0 | 1.0 | GO:0046847 | filopodium assembly(GO:0046847) |
| 0.0 | 0.6 | GO:0051289 | protein homotetramerization(GO:0051289) |
| 0.0 | 1.6 | GO:0018107 | peptidyl-threonine phosphorylation(GO:0018107) |
| 0.0 | 1.0 | GO:0006904 | vesicle docking involved in exocytosis(GO:0006904) |
| 0.0 | 1.9 | GO:0046839 | phospholipid dephosphorylation(GO:0046839) |
| 0.0 | 1.6 | GO:1901214 | regulation of neuron death(GO:1901214) |
| 0.0 | 0.5 | GO:0043153 | entrainment of circadian clock by photoperiod(GO:0043153) |
| 0.0 | 0.4 | GO:0070672 | interleukin-15-mediated signaling pathway(GO:0035723) response to interleukin-15(GO:0070672) cellular response to interleukin-15(GO:0071350) |
| 0.0 | 0.5 | GO:0070536 | protein K63-linked deubiquitination(GO:0070536) |
| 0.0 | 0.4 | GO:0070570 | regulation of axon regeneration(GO:0048679) regulation of neuron projection regeneration(GO:0070570) |
| 0.0 | 0.6 | GO:0042219 | cellular modified amino acid catabolic process(GO:0042219) |
| 0.0 | 0.6 | GO:0007041 | lysosomal transport(GO:0007041) |
| 0.0 | 0.5 | GO:0097194 | execution phase of apoptosis(GO:0097194) |
| 0.0 | 0.5 | GO:0006919 | activation of cysteine-type endopeptidase activity involved in apoptotic process(GO:0006919) |
| 0.0 | 0.3 | GO:0086010 | membrane depolarization during action potential(GO:0086010) |
| 0.0 | 0.4 | GO:0016082 | synaptic vesicle priming(GO:0016082) |
| 0.0 | 0.7 | GO:0008630 | intrinsic apoptotic signaling pathway in response to DNA damage(GO:0008630) |
| 0.0 | 0.7 | GO:0005978 | glycogen biosynthetic process(GO:0005978) glucan biosynthetic process(GO:0009250) |
| 0.0 | 0.5 | GO:0055008 | cardiac muscle tissue morphogenesis(GO:0055008) |
| 0.0 | 0.3 | GO:0010950 | positive regulation of endopeptidase activity(GO:0010950) positive regulation of peptidase activity(GO:0010952) |
| 0.0 | 1.1 | GO:1902476 | chloride transmembrane transport(GO:1902476) |
| 0.0 | 0.3 | GO:0050829 | defense response to Gram-negative bacterium(GO:0050829) |
| 0.0 | 0.2 | GO:0017121 | phospholipid scrambling(GO:0017121) |
| 0.0 | 1.2 | GO:0006334 | nucleosome assembly(GO:0006334) |
| 0.0 | 3.0 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
| 0.0 | 0.3 | GO:0006942 | regulation of striated muscle contraction(GO:0006942) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 2.7 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.3 | 2.3 | GO:0000015 | phosphopyruvate hydratase complex(GO:0000015) |
| 0.2 | 0.9 | GO:0061702 | inflammasome complex(GO:0061702) |
| 0.2 | 0.9 | GO:0008091 | spectrin(GO:0008091) |
| 0.2 | 1.6 | GO:0042583 | chromaffin granule(GO:0042583) |
| 0.1 | 3.5 | GO:0000407 | pre-autophagosomal structure(GO:0000407) |
| 0.1 | 1.7 | GO:0048179 | activin receptor complex(GO:0048179) |
| 0.1 | 0.6 | GO:0071203 | WASH complex(GO:0071203) |
| 0.1 | 1.1 | GO:0098982 | GABA-ergic synapse(GO:0098982) |
| 0.1 | 0.4 | GO:0097268 | cytoophidium(GO:0097268) |
| 0.1 | 0.5 | GO:0009331 | glycerol-3-phosphate dehydrogenase complex(GO:0009331) |
| 0.1 | 0.3 | GO:0032584 | growth cone membrane(GO:0032584) |
| 0.1 | 0.3 | GO:0008537 | proteasome activator complex(GO:0008537) |
| 0.1 | 1.1 | GO:0097038 | perinuclear endoplasmic reticulum(GO:0097038) |
| 0.1 | 0.8 | GO:0098833 | presynaptic endocytic zone(GO:0098833) presynaptic endocytic zone membrane(GO:0098835) extrinsic component of presynaptic membrane(GO:0098888) extrinsic component of presynaptic endocytic zone(GO:0098894) |
| 0.0 | 1.5 | GO:1902710 | GABA receptor complex(GO:1902710) GABA-A receptor complex(GO:1902711) |
| 0.0 | 1.1 | GO:0071565 | nBAF complex(GO:0071565) |
| 0.0 | 0.7 | GO:0005640 | nuclear outer membrane(GO:0005640) |
| 0.0 | 1.6 | GO:0005861 | troponin complex(GO:0005861) |
| 0.0 | 0.7 | GO:0002116 | semaphorin receptor complex(GO:0002116) |
| 0.0 | 1.8 | GO:0016342 | catenin complex(GO:0016342) |
| 0.0 | 0.2 | GO:0030121 | AP-1 adaptor complex(GO:0030121) |
| 0.0 | 1.3 | GO:0005905 | clathrin-coated pit(GO:0005905) |
| 0.0 | 0.6 | GO:0090665 | dystrophin-associated glycoprotein complex(GO:0016010) glycoprotein complex(GO:0090665) |
| 0.0 | 1.0 | GO:0030864 | cortical actin cytoskeleton(GO:0030864) |
| 0.0 | 1.5 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.0 | 0.3 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.0 | 0.9 | GO:0034705 | voltage-gated potassium channel complex(GO:0008076) potassium channel complex(GO:0034705) |
| 0.0 | 2.1 | GO:0043025 | neuronal cell body(GO:0043025) |
| 0.0 | 1.4 | GO:0098878 | ionotropic glutamate receptor complex(GO:0008328) neurotransmitter receptor complex(GO:0098878) |
| 0.0 | 0.3 | GO:0034706 | voltage-gated sodium channel complex(GO:0001518) sodium channel complex(GO:0034706) |
| 0.0 | 1.9 | GO:0016459 | myosin complex(GO:0016459) |
| 0.0 | 0.2 | GO:0097651 | phosphatidylinositol 3-kinase complex, class IB(GO:0005944) phosphatidylinositol 3-kinase complex, class I(GO:0097651) |
| 0.0 | 0.4 | GO:0035371 | microtubule plus-end(GO:0035371) |
| 0.0 | 1.7 | GO:0005802 | trans-Golgi network(GO:0005802) |
| 0.0 | 0.7 | GO:0032580 | Golgi cisterna membrane(GO:0032580) |
| 0.0 | 0.5 | GO:0097610 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.0 | 0.3 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.0 | 1.2 | GO:0000786 | nucleosome(GO:0000786) |
| 0.0 | 0.2 | GO:0031083 | BLOC-1 complex(GO:0031083) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.6 | GO:0052834 | inositol monophosphate 4-phosphatase activity(GO:0052833) inositol monophosphate phosphatase activity(GO:0052834) |
| 0.5 | 2.7 | GO:0019865 | IgE binding(GO:0019863) immunoglobulin binding(GO:0019865) |
| 0.5 | 3.0 | GO:0004743 | pyruvate kinase activity(GO:0004743) |
| 0.3 | 1.7 | GO:0050254 | rhodopsin kinase activity(GO:0050254) |
| 0.3 | 2.3 | GO:0004634 | phosphopyruvate hydratase activity(GO:0004634) |
| 0.3 | 1.7 | GO:0016361 | activin receptor activity, type I(GO:0016361) |
| 0.3 | 1.4 | GO:0044736 | acid-sensing ion channel activity(GO:0044736) |
| 0.2 | 1.2 | GO:0008798 | beta-aspartyl-peptidase activity(GO:0008798) |
| 0.2 | 0.9 | GO:0052851 | cupric reductase activity(GO:0008823) ferric-chelate reductase (NADPH) activity(GO:0052851) |
| 0.2 | 1.8 | GO:0004359 | glutaminase activity(GO:0004359) |
| 0.2 | 0.7 | GO:0004373 | glycogen (starch) synthase activity(GO:0004373) |
| 0.2 | 0.6 | GO:0072571 | ADP-D-ribose binding(GO:0072570) mono-ADP-D-ribose binding(GO:0072571) |
| 0.1 | 1.5 | GO:0005042 | netrin receptor activity(GO:0005042) |
| 0.1 | 0.4 | GO:0047961 | glycine N-acyltransferase activity(GO:0047961) |
| 0.1 | 0.6 | GO:0001735 | prenylcysteine oxidase activity(GO:0001735) |
| 0.1 | 0.6 | GO:0070573 | metallodipeptidase activity(GO:0070573) |
| 0.1 | 0.6 | GO:0019779 | Atg12 activating enzyme activity(GO:0019778) Atg8 activating enzyme activity(GO:0019779) |
| 0.1 | 0.4 | GO:0033857 | inositol heptakisphosphate kinase activity(GO:0000829) diphosphoinositol-pentakisphosphate kinase activity(GO:0033857) |
| 0.1 | 1.0 | GO:0004530 | deoxyribonuclease I activity(GO:0004530) |
| 0.1 | 0.4 | GO:0003883 | CTP synthase activity(GO:0003883) |
| 0.1 | 0.7 | GO:0003835 | beta-galactoside alpha-2,6-sialyltransferase activity(GO:0003835) |
| 0.1 | 0.6 | GO:0016416 | carnitine O-palmitoyltransferase activity(GO:0004095) O-palmitoyltransferase activity(GO:0016416) |
| 0.1 | 1.3 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.1 | 0.6 | GO:0016316 | phosphatidylinositol-3,4-bisphosphate 4-phosphatase activity(GO:0016316) |
| 0.1 | 0.9 | GO:0001530 | lipopolysaccharide binding(GO:0001530) |
| 0.1 | 6.3 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.1 | 1.9 | GO:0042577 | lipid phosphatase activity(GO:0042577) |
| 0.1 | 1.6 | GO:0005523 | tropomyosin binding(GO:0005523) |
| 0.1 | 0.4 | GO:0004938 | alpha2-adrenergic receptor activity(GO:0004938) |
| 0.1 | 0.8 | GO:0005432 | calcium:sodium antiporter activity(GO:0005432) |
| 0.1 | 0.8 | GO:0005545 | 1-phosphatidylinositol binding(GO:0005545) |
| 0.1 | 1.1 | GO:0098882 | structural constituent of presynaptic active zone(GO:0098882) |
| 0.1 | 1.6 | GO:0051787 | misfolded protein binding(GO:0051787) |
| 0.1 | 0.5 | GO:0003834 | beta-carotene 15,15'-monooxygenase activity(GO:0003834) carotenoid dioxygenase activity(GO:0010436) |
| 0.1 | 0.7 | GO:0005537 | mannose binding(GO:0005537) |
| 0.1 | 1.5 | GO:0008331 | high voltage-gated calcium channel activity(GO:0008331) |
| 0.0 | 0.3 | GO:0061133 | endopeptidase activator activity(GO:0061133) |
| 0.0 | 1.5 | GO:0004890 | GABA-A receptor activity(GO:0004890) |
| 0.0 | 0.8 | GO:0030552 | cAMP binding(GO:0030552) |
| 0.0 | 2.3 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.0 | 0.3 | GO:1901474 | azole transporter activity(GO:0045118) azole transmembrane transporter activity(GO:1901474) |
| 0.0 | 1.1 | GO:0015248 | sterol transporter activity(GO:0015248) |
| 0.0 | 0.5 | GO:0004111 | creatine kinase activity(GO:0004111) phosphotransferase activity, nitrogenous group as acceptor(GO:0016775) |
| 0.0 | 0.6 | GO:0016832 | aldehyde-lyase activity(GO:0016832) |
| 0.0 | 0.4 | GO:0004415 | hyalurononglucosaminidase activity(GO:0004415) |
| 0.0 | 0.2 | GO:0015280 | ligand-gated sodium channel activity(GO:0015280) |
| 0.0 | 0.8 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.0 | 1.1 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.0 | 0.4 | GO:0032041 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.0 | 0.8 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.0 | 0.5 | GO:0051428 | peptide hormone receptor binding(GO:0051428) |
| 0.0 | 0.7 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.0 | 0.2 | GO:0004994 | somatostatin receptor activity(GO:0004994) |
| 0.0 | 0.4 | GO:0042289 | MHC class II protein binding(GO:0042289) |
| 0.0 | 0.3 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.0 | 1.8 | GO:0045296 | cadherin binding(GO:0045296) |
| 0.0 | 2.9 | GO:0061135 | endopeptidase inhibitor activity(GO:0004866) endopeptidase regulator activity(GO:0061135) |
| 0.0 | 0.9 | GO:0044325 | ion channel binding(GO:0044325) |
| 0.0 | 0.3 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.0 | 0.9 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.0 | 0.2 | GO:0015272 | ATP-activated inward rectifier potassium channel activity(GO:0015272) |
| 0.0 | 0.3 | GO:0005248 | voltage-gated sodium channel activity(GO:0005248) voltage-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1905030) |
| 0.0 | 0.2 | GO:0017128 | phospholipid scramblase activity(GO:0017128) |
| 0.0 | 0.3 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.0 | 0.2 | GO:0004656 | procollagen-proline 4-dioxygenase activity(GO:0004656) |
| 0.0 | 0.3 | GO:0030159 | receptor signaling complex scaffold activity(GO:0030159) |
| 0.0 | 1.7 | GO:0004222 | metalloendopeptidase activity(GO:0004222) |
| 0.0 | 0.4 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.2 | PID THROMBIN PAR4 PATHWAY | PAR4-mediated thrombin signaling events |
| 0.1 | 2.1 | PID PRL SIGNALING EVENTS PATHWAY | Signaling events mediated by PRL |
| 0.0 | 1.6 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
| 0.0 | 0.1 | ST JAK STAT PATHWAY | Jak-STAT Pathway |
| 0.0 | 0.5 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.0 | 0.2 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.0 | 1.0 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.0 | 0.6 | SIG PIP3 SIGNALING IN B LYMPHOCYTES | Genes related to PIP3 signaling in B lymphocytes |
| 0.0 | 0.8 | PID PTP1B PATHWAY | Signaling events mediated by PTP1B |
| 0.0 | 0.9 | PID TGFBR PATHWAY | TGF-beta receptor signaling |
| 0.0 | 0.3 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
| 0.0 | 0.3 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.2 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.1 | 2.8 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.1 | 1.4 | REACTOME IONOTROPIC ACTIVITY OF KAINATE RECEPTORS | Genes involved in Ionotropic activity of Kainate Receptors |
| 0.1 | 0.4 | REACTOME PHOSPHORYLATION OF CD3 AND TCR ZETA CHAINS | Genes involved in Phosphorylation of CD3 and TCR zeta chains |
| 0.1 | 0.8 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.1 | 0.9 | REACTOME INTERACTION BETWEEN L1 AND ANKYRINS | Genes involved in Interaction between L1 and Ankyrins |
| 0.0 | 0.9 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.0 | 0.6 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.0 | 0.6 | REACTOME ACTIVATED AMPK STIMULATES FATTY ACID OXIDATION IN MUSCLE | Genes involved in Activated AMPK stimulates fatty-acid oxidation in muscle |
| 0.0 | 0.6 | REACTOME SYNTHESIS OF PIPS AT THE EARLY ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the early endosome membrane |
| 0.0 | 0.2 | REACTOME DSCAM INTERACTIONS | Genes involved in DSCAM interactions |
| 0.0 | 0.4 | REACTOME AMINE LIGAND BINDING RECEPTORS | Genes involved in Amine ligand-binding receptors |
| 0.0 | 1.9 | REACTOME RESPONSE TO ELEVATED PLATELET CYTOSOLIC CA2 | Genes involved in Response to elevated platelet cytosolic Ca2+ |
| 0.0 | 2.1 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.0 | 0.2 | REACTOME PYRIMIDINE CATABOLISM | Genes involved in Pyrimidine catabolism |
| 0.0 | 0.3 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.0 | 0.1 | REACTOME AMINO ACID SYNTHESIS AND INTERCONVERSION TRANSAMINATION | Genes involved in Amino acid synthesis and interconversion (transamination) |
| 0.0 | 0.7 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.0 | 0.7 | REACTOME GLUCOSE METABOLISM | Genes involved in Glucose metabolism |
| 0.0 | 0.1 | REACTOME IL 7 SIGNALING | Genes involved in Interleukin-7 signaling |
| 0.0 | 0.7 | REACTOME POTASSIUM CHANNELS | Genes involved in Potassium Channels |
| 0.0 | 0.4 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |
| 0.0 | 0.5 | REACTOME NRAGE SIGNALS DEATH THROUGH JNK | Genes involved in NRAGE signals death through JNK |
| 0.0 | 0.2 | REACTOME G BETA GAMMA SIGNALLING THROUGH PI3KGAMMA | Genes involved in G beta:gamma signalling through PI3Kgamma |
| 0.0 | 0.1 | REACTOME ENOS ACTIVATION AND REGULATION | Genes involved in eNOS activation and regulation |