Project
PRJEB1986: zebrafish developmental stages transcriptome
Navigation
Downloads
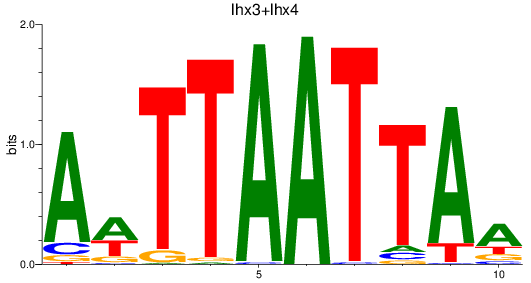
Results for lhx3+lhx4
Z-value: 0.89
Transcription factors associated with lhx3+lhx4
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
lhx3
|
ENSDARG00000003803 | LIM homeobox 3 |
|
lhx4
|
ENSDARG00000039458 | LIM homeobox 4 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| LHX3 | dr11_v1_chr5_+_71802014_71802014 | 0.78 | 9.1e-05 | Click! |
| lhx4 | dr11_v1_chr8_-_14484599_14484599 | 0.40 | 8.8e-02 | Click! |
Activity profile of lhx3+lhx4 motif
Sorted Z-values of lhx3+lhx4 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr9_+_34641237 | 5.90 |
ENSDART00000133996
|
shox
|
short stature homeobox |
| chr9_-_22099536 | 4.87 |
ENSDART00000101923
|
CR391987.1
|
|
| chr25_+_31276842 | 3.77 |
ENSDART00000187238
|
tnni2a.4
|
troponin I type 2a (skeletal, fast), tandem duplicate 4 |
| chr25_+_31227747 | 3.61 |
ENSDART00000033872
|
tnni2a.1
|
troponin I type 2a (skeletal, fast), tandem duplicate 1 |
| chr9_+_2574122 | 2.60 |
ENSDART00000166326
ENSDART00000191822 |
CIR1
|
si:ch73-167c12.2 |
| chr14_+_46313396 | 2.55 |
ENSDART00000047525
|
cryba1l1
|
crystallin, beta A1, like 1 |
| chr11_+_24002503 | 2.46 |
ENSDART00000164702
|
chia.2
|
chitinase, acidic.2 |
| chr11_+_24001993 | 2.32 |
ENSDART00000168215
|
chia.2
|
chitinase, acidic.2 |
| chr21_-_20939488 | 2.32 |
ENSDART00000039043
|
rgs7bpb
|
regulator of G protein signaling 7 binding protein b |
| chr20_+_4060839 | 2.29 |
ENSDART00000178565
|
TRIM67
|
tripartite motif containing 67 |
| chr15_-_37850969 | 2.17 |
ENSDART00000031418
|
hsc70
|
heat shock cognate 70 |
| chr5_-_41494831 | 2.15 |
ENSDART00000051081
|
eef2l2
|
eukaryotic translation elongation factor 2, like 2 |
| chr3_+_21225750 | 2.03 |
ENSDART00000139213
|
zgc:153968
|
zgc:153968 |
| chr5_+_2815021 | 1.99 |
ENSDART00000020472
|
hpda
|
4-hydroxyphenylpyruvate dioxygenase a |
| chr2_+_2223837 | 1.97 |
ENSDART00000101038
ENSDART00000129354 |
tmie
|
transmembrane inner ear |
| chr7_+_24573721 | 1.91 |
ENSDART00000173938
ENSDART00000173681 |
si:dkeyp-75h12.7
|
si:dkeyp-75h12.7 |
| chr15_-_46779934 | 1.87 |
ENSDART00000085136
|
clcn2c
|
chloride channel 2c |
| chr13_-_29421331 | 1.87 |
ENSDART00000150228
|
chata
|
choline O-acetyltransferase a |
| chr10_-_27049170 | 1.77 |
ENSDART00000143451
|
cnih2
|
cornichon family AMPA receptor auxiliary protein 2 |
| chr6_+_7444899 | 1.74 |
ENSDART00000053775
|
arf3b
|
ADP-ribosylation factor 3b |
| chr16_+_46111849 | 1.72 |
ENSDART00000172232
|
sv2a
|
synaptic vesicle glycoprotein 2A |
| chr15_-_44601331 | 1.69 |
ENSDART00000161514
|
zgc:165508
|
zgc:165508 |
| chr1_-_5455498 | 1.66 |
ENSDART00000040368
ENSDART00000114035 |
mnx2b
|
motor neuron and pancreas homeobox 2b |
| chr23_-_39666519 | 1.64 |
ENSDART00000110868
ENSDART00000190961 |
vwa1
|
von Willebrand factor A domain containing 1 |
| chr9_-_49493305 | 1.62 |
ENSDART00000148707
ENSDART00000148561 |
xirp2b
|
xin actin binding repeat containing 2b |
| chr3_+_29714775 | 1.60 |
ENSDART00000041388
|
cacng2a
|
calcium channel, voltage-dependent, gamma subunit 2a |
| chr7_-_66868543 | 1.55 |
ENSDART00000149680
|
ampd3a
|
adenosine monophosphate deaminase 3a |
| chr17_-_31483469 | 1.51 |
ENSDART00000062907
ENSDART00000061547 |
ltk
|
leukocyte receptor tyrosine kinase |
| chr18_-_38088099 | 1.48 |
ENSDART00000146120
|
luzp2
|
leucine zipper protein 2 |
| chr16_-_28856112 | 1.41 |
ENSDART00000078543
|
syt11b
|
synaptotagmin XIb |
| chr4_+_9669717 | 1.40 |
ENSDART00000004604
|
si:dkey-153k10.9
|
si:dkey-153k10.9 |
| chr5_+_71802014 | 1.39 |
ENSDART00000124939
ENSDART00000097164 |
LHX3
|
LIM homeobox 3 |
| chr17_+_13664442 | 1.39 |
ENSDART00000171689
|
lrfn5a
|
leucine rich repeat and fibronectin type III domain containing 5a |
| chr4_-_4387012 | 1.37 |
ENSDART00000191836
|
CU468826.3
|
Danio rerio U2 small nuclear ribonucleoprotein auxiliary factor 35 kDa subunit-related protein 1-like (LOC100331497), mRNA. |
| chr25_+_13406069 | 1.36 |
ENSDART00000010495
|
znrf1
|
zinc and ring finger 1 |
| chr9_+_54039006 | 1.35 |
ENSDART00000112441
|
tlr7
|
toll-like receptor 7 |
| chr8_-_34052019 | 1.34 |
ENSDART00000040126
ENSDART00000159208 ENSDART00000048994 ENSDART00000098822 |
pbx3b
|
pre-B-cell leukemia homeobox 3b |
| chr7_-_71829649 | 1.32 |
ENSDART00000160449
|
cacnb2a
|
calcium channel, voltage-dependent, beta 2a |
| chr20_+_41664350 | 1.30 |
ENSDART00000186247
|
fam184a
|
family with sequence similarity 184, member A |
| chr24_-_7697274 | 1.29 |
ENSDART00000186077
|
syt5b
|
synaptotagmin Vb |
| chr17_+_16564921 | 1.27 |
ENSDART00000151904
|
foxn3
|
forkhead box N3 |
| chr1_-_21901589 | 1.26 |
ENSDART00000140553
|
frmpd1a
|
FERM and PDZ domain containing 1a |
| chr15_-_46718759 | 1.24 |
ENSDART00000085926
ENSDART00000154577 |
zgc:162872
|
zgc:162872 |
| chr22_-_21897203 | 1.24 |
ENSDART00000158501
ENSDART00000105566 ENSDART00000136795 |
gna11a
|
guanine nucleotide binding protein (G protein), alpha 11a (Gq class) |
| chr3_-_48716422 | 1.23 |
ENSDART00000164979
|
si:ch211-114m9.1
|
si:ch211-114m9.1 |
| chr20_+_52389858 | 1.23 |
ENSDART00000185863
ENSDART00000166651 |
arhgap39
|
Rho GTPase activating protein 39 |
| chr18_+_1703984 | 1.23 |
ENSDART00000114010
|
slitrk3a
|
SLIT and NTRK-like family, member 3a |
| chr10_+_45089820 | 1.22 |
ENSDART00000175481
|
camk2b2
|
calcium/calmodulin-dependent protein kinase (CaM kinase) II beta 2 |
| chr10_+_22381802 | 1.21 |
ENSDART00000112484
|
nlgn2b
|
neuroligin 2b |
| chr16_-_27749172 | 1.20 |
ENSDART00000145198
|
steap4
|
STEAP family member 4 |
| chr9_+_38481780 | 1.19 |
ENSDART00000087241
|
lss
|
lanosterol synthase (2,3-oxidosqualene-lanosterol cyclase) |
| chr17_+_50524573 | 1.18 |
ENSDART00000187974
|
CR382385.1
|
|
| chr18_+_21408794 | 1.18 |
ENSDART00000140161
|
necab2
|
N-terminal EF-hand calcium binding protein 2 |
| chr23_-_30954738 | 1.15 |
ENSDART00000188996
|
osbpl2a
|
oxysterol binding protein-like 2a |
| chr1_+_33969015 | 1.15 |
ENSDART00000042984
ENSDART00000146530 |
epha6
|
eph receptor A6 |
| chr1_-_19215336 | 1.14 |
ENSDART00000162949
ENSDART00000170680 |
ptprdb
|
protein tyrosine phosphatase, receptor type, D, b |
| chr23_+_16638639 | 1.14 |
ENSDART00000143545
|
snphb
|
syntaphilin b |
| chr6_-_51386656 | 1.13 |
ENSDART00000154732
ENSDART00000177990 ENSDART00000184928 ENSDART00000180197 |
ptprt
|
protein tyrosine phosphatase, receptor type, t |
| chr18_+_924949 | 1.12 |
ENSDART00000170888
ENSDART00000193163 |
pkma
|
pyruvate kinase M1/2a |
| chr19_+_5480327 | 1.12 |
ENSDART00000148794
|
jupb
|
junction plakoglobin b |
| chr20_+_32523576 | 1.11 |
ENSDART00000147319
|
scml4
|
Scm polycomb group protein like 4 |
| chr14_-_4556896 | 1.11 |
ENSDART00000044678
ENSDART00000192863 |
GABRA2
|
gamma-aminobutyric acid type A receptor alpha2 subunit |
| chr11_+_30663300 | 1.10 |
ENSDART00000161662
|
ttbk1a
|
tau tubulin kinase 1a |
| chr20_+_32552912 | 1.08 |
ENSDART00000009691
|
scml4
|
Scm polycomb group protein like 4 |
| chr8_+_10561922 | 1.06 |
ENSDART00000133348
|
fam19a5l
|
family with sequence similarity 19 (chemokine (C-C motif)-like), member A5-like |
| chr25_-_4146947 | 1.05 |
ENSDART00000129268
|
fads2
|
fatty acid desaturase 2 |
| chr3_+_7808459 | 1.05 |
ENSDART00000162374
|
hook2
|
hook microtubule-tethering protein 2 |
| chr7_+_29954709 | 1.05 |
ENSDART00000173904
|
tpma
|
alpha-tropomyosin |
| chr10_+_26972755 | 1.05 |
ENSDART00000042162
|
tm7sf2
|
transmembrane 7 superfamily member 2 |
| chr7_+_25033924 | 1.05 |
ENSDART00000170873
|
sb:cb1058
|
sb:cb1058 |
| chr23_-_306796 | 1.05 |
ENSDART00000143125
|
anks1aa
|
ankyrin repeat and sterile alpha motif domain containing 1Aa |
| chr8_+_26141680 | 1.04 |
ENSDART00000078334
|
celsr3
|
cadherin, EGF LAG seven-pass G-type receptor 3 |
| chr20_-_5291012 | 1.02 |
ENSDART00000122892
|
cyp46a1.3
|
cytochrome P450, family 46, subfamily A, polypeptide 1, tandem duplicate 3 |
| chr21_+_45841731 | 1.02 |
ENSDART00000038657
|
faxdc2
|
fatty acid hydroxylase domain containing 2 |
| chr8_-_34051548 | 1.02 |
ENSDART00000105204
|
pbx3b
|
pre-B-cell leukemia homeobox 3b |
| chr19_+_43297546 | 1.01 |
ENSDART00000168002
|
laptm5
|
lysosomal protein transmembrane 5 |
| chr10_-_2524297 | 1.01 |
ENSDART00000192475
|
CU856539.1
|
|
| chr14_-_413273 | 1.00 |
ENSDART00000163976
ENSDART00000179907 |
FAT4
|
FAT atypical cadherin 4 |
| chr22_-_26865181 | 1.00 |
ENSDART00000138311
|
hmox2a
|
heme oxygenase 2a |
| chr9_+_54290896 | 1.00 |
ENSDART00000149175
|
pou4f3
|
POU class 4 homeobox 3 |
| chr6_+_39905021 | 0.99 |
ENSDART00000064904
|
endou
|
endonuclease, polyU-specific |
| chr14_-_2933185 | 0.99 |
ENSDART00000161677
ENSDART00000162446 ENSDART00000109378 |
si:dkey-201i24.6
|
si:dkey-201i24.6 |
| chr20_-_2949028 | 0.97 |
ENSDART00000104667
ENSDART00000193151 ENSDART00000131946 |
cdk19
|
cyclin-dependent kinase 19 |
| chr23_-_32100106 | 0.96 |
ENSDART00000044658
|
letmd1
|
LETM1 domain containing 1 |
| chr2_+_20430366 | 0.95 |
ENSDART00000155108
|
si:ch211-153l6.6
|
si:ch211-153l6.6 |
| chr8_+_52637507 | 0.93 |
ENSDART00000163830
|
si:dkey-90l8.3
|
si:dkey-90l8.3 |
| chr23_-_18024543 | 0.93 |
ENSDART00000139695
|
pm20d1.1
|
peptidase M20 domain containing 1, tandem duplicate 1 |
| chr17_+_15433518 | 0.92 |
ENSDART00000026180
|
fabp7a
|
fatty acid binding protein 7, brain, a |
| chr8_+_8459192 | 0.91 |
ENSDART00000140942
ENSDART00000014939 |
comta
|
catechol-O-methyltransferase a |
| chr3_-_59297532 | 0.91 |
ENSDART00000187991
|
CABZ01053748.1
|
|
| chr4_-_18309917 | 0.89 |
ENSDART00000189084
|
plxnc1
|
plexin C1 |
| chr17_+_38476300 | 0.89 |
ENSDART00000123298
|
stard9
|
StAR-related lipid transfer (START) domain containing 9 |
| chr2_-_1548330 | 0.89 |
ENSDART00000082155
ENSDART00000108481 ENSDART00000111272 |
dab1b
|
Dab, reelin signal transducer, homolog 1b (Drosophila) |
| chr6_+_51773873 | 0.87 |
ENSDART00000156516
|
tmem74b
|
transmembrane protein 74B |
| chr16_-_38118003 | 0.87 |
ENSDART00000058667
|
si:dkey-23o4.6
|
si:dkey-23o4.6 |
| chr22_-_31517300 | 0.86 |
ENSDART00000164799
|
slc6a6b
|
solute carrier family 6 (neurotransmitter transporter), member 6b |
| chr6_-_28980756 | 0.86 |
ENSDART00000014661
|
glmnb
|
glomulin, FKBP associated protein b |
| chr17_+_15433671 | 0.86 |
ENSDART00000149568
|
fabp7a
|
fatty acid binding protein 7, brain, a |
| chr24_+_13316737 | 0.85 |
ENSDART00000191658
|
SBSPON
|
somatomedin B and thrombospondin type 1 domain containing |
| chr7_-_28148310 | 0.85 |
ENSDART00000044208
|
lmo1
|
LIM domain only 1 |
| chr9_+_24159280 | 0.85 |
ENSDART00000184624
ENSDART00000178422 |
hecw2a
|
HECT, C2 and WW domain containing E3 ubiquitin protein ligase 2a |
| chr3_+_32553714 | 0.84 |
ENSDART00000165638
|
pax10
|
paired box 10 |
| chr12_-_48006835 | 0.84 |
ENSDART00000108989
|
adamts14
|
ADAM metallopeptidase with thrombospondin type 1 motif, 14 |
| chr6_-_11768198 | 0.83 |
ENSDART00000183463
|
march7
|
membrane-associated ring finger (C3HC4) 7 |
| chr9_+_33009284 | 0.83 |
ENSDART00000036926
|
vangl1
|
VANGL planar cell polarity protein 1 |
| chr16_-_45178430 | 0.83 |
ENSDART00000165186
|
si:dkey-33i11.9
|
si:dkey-33i11.9 |
| chr5_+_4436405 | 0.83 |
ENSDART00000167969
|
CABZ01079241.1
|
|
| chr19_-_5865766 | 0.83 |
ENSDART00000191007
|
LO018585.1
|
|
| chr7_+_10701938 | 0.82 |
ENSDART00000158162
|
arnt2
|
aryl-hydrocarbon receptor nuclear translocator 2 |
| chr21_-_14811058 | 0.81 |
ENSDART00000143100
|
phpt1
|
phosphohistidine phosphatase 1 |
| chr3_-_47876427 | 0.81 |
ENSDART00000180844
ENSDART00000124480 |
adgrl1a
|
adhesion G protein-coupled receptor L1a |
| chr18_+_39487486 | 0.80 |
ENSDART00000126978
|
acadl
|
acyl-CoA dehydrogenase long chain |
| chr1_+_6862917 | 0.80 |
ENSDART00000182953
|
erbb4a
|
erb-b2 receptor tyrosine kinase 4a |
| chr7_+_17229980 | 0.80 |
ENSDART00000184910
|
slc6a5
|
solute carrier family 6 (neurotransmitter transporter), member 5 |
| chr20_+_34933183 | 0.80 |
ENSDART00000062738
|
snap25a
|
synaptosomal-associated protein, 25a |
| chr13_+_1089942 | 0.80 |
ENSDART00000054322
|
cnrip1b
|
cannabinoid receptor interacting protein 1b |
| chr9_-_48407408 | 0.80 |
ENSDART00000058248
|
zgc:172182
|
zgc:172182 |
| chr2_+_37227011 | 0.79 |
ENSDART00000126587
ENSDART00000084958 |
samd7
|
sterile alpha motif domain containing 7 |
| chr1_+_38858399 | 0.79 |
ENSDART00000165454
|
CU915762.1
|
|
| chr12_-_26383242 | 0.79 |
ENSDART00000152941
|
usp54b
|
ubiquitin specific peptidase 54b |
| chr6_+_52350443 | 0.79 |
ENSDART00000151612
ENSDART00000151349 |
si:ch211-239j9.1
|
si:ch211-239j9.1 |
| chr18_+_15644559 | 0.79 |
ENSDART00000061794
|
nr1h4
|
nuclear receptor subfamily 1, group H, member 4 |
| chr13_-_29420885 | 0.78 |
ENSDART00000024225
|
chata
|
choline O-acetyltransferase a |
| chr14_-_858985 | 0.78 |
ENSDART00000148687
ENSDART00000149375 |
slc34a1a
|
solute carrier family 34 (type II sodium/phosphate cotransporter), member 1a |
| chr21_+_3093419 | 0.78 |
ENSDART00000162520
|
SHC3
|
SHC adaptor protein 3 |
| chr16_+_51326237 | 0.78 |
ENSDART00000168525
|
CU469384.1
|
|
| chr12_-_6880694 | 0.78 |
ENSDART00000171846
|
pcdh15b
|
protocadherin-related 15b |
| chr17_-_45040813 | 0.77 |
ENSDART00000075514
|
entpd5a
|
ectonucleoside triphosphate diphosphohydrolase 5a |
| chr21_-_2814709 | 0.77 |
ENSDART00000097664
|
SEMA4D
|
semaphorin 4D |
| chr4_-_2219705 | 0.76 |
ENSDART00000131046
|
si:ch73-278m9.1
|
si:ch73-278m9.1 |
| chr14_-_17306261 | 0.76 |
ENSDART00000191747
|
jakmip1
|
janus kinase and microtubule interacting protein 1 |
| chr12_+_1139690 | 0.75 |
ENSDART00000160442
|
CABZ01072885.1
|
|
| chr22_-_31060579 | 0.75 |
ENSDART00000182376
|
cand2
|
cullin-associated and neddylation-dissociated 2 (putative) |
| chr9_+_27339505 | 0.75 |
ENSDART00000011822
|
slc10a2
|
solute carrier family 10 (sodium/bile acid cotransporter), member 2 |
| chr9_+_29643036 | 0.75 |
ENSDART00000023210
ENSDART00000175160 |
trim13
|
tripartite motif containing 13 |
| chr8_+_43340995 | 0.75 |
ENSDART00000038566
|
rflna
|
refilin A |
| chr8_-_39952727 | 0.74 |
ENSDART00000181310
|
cabp1a
|
calcium binding protein 1a |
| chr15_+_9327252 | 0.73 |
ENSDART00000144381
|
sgcg
|
sarcoglycan, gamma |
| chr1_-_34750169 | 0.72 |
ENSDART00000149380
|
si:dkey-151m22.8
|
si:dkey-151m22.8 |
| chr2_+_29491314 | 0.72 |
ENSDART00000181774
|
dlgap1a
|
discs, large (Drosophila) homolog-associated protein 1a |
| chr5_-_21422390 | 0.71 |
ENSDART00000144198
|
tenm1
|
teneurin transmembrane protein 1 |
| chr10_-_43404027 | 0.71 |
ENSDART00000086227
|
edil3b
|
EGF-like repeats and discoidin I-like domains 3b |
| chr2_+_42072231 | 0.70 |
ENSDART00000084517
|
vcpip1
|
valosin containing protein (p97)/p47 complex interacting protein 1 |
| chr12_+_1000323 | 0.70 |
ENSDART00000054363
|
si:ch1073-272o11.3
|
si:ch1073-272o11.3 |
| chr23_-_20051369 | 0.70 |
ENSDART00000049836
|
bgnb
|
biglycan b |
| chr6_-_57539141 | 0.69 |
ENSDART00000156967
|
itcha
|
itchy E3 ubiquitin protein ligase a |
| chr6_-_43677125 | 0.69 |
ENSDART00000150128
|
foxp1b
|
forkhead box P1b |
| chr11_+_28476298 | 0.69 |
ENSDART00000122319
|
lrrc38b
|
leucine rich repeat containing 38b |
| chr9_+_45227028 | 0.69 |
ENSDART00000185579
|
adarb1b
|
adenosine deaminase, RNA-specific, B1b |
| chr7_+_31891110 | 0.69 |
ENSDART00000173883
|
mybpc3
|
myosin binding protein C, cardiac |
| chr21_-_275377 | 0.68 |
ENSDART00000157509
|
rln1
|
relaxin 1 |
| chr10_-_5844915 | 0.68 |
ENSDART00000185929
|
ankrd55
|
ankyrin repeat domain 55 |
| chr3_-_56456527 | 0.68 |
ENSDART00000156553
|
cyth1a
|
cytohesin 1a |
| chr21_+_21195487 | 0.67 |
ENSDART00000181746
ENSDART00000184832 |
rictorb
|
RPTOR independent companion of MTOR, complex 2b |
| chr10_-_5847655 | 0.66 |
ENSDART00000192773
|
ankrd55
|
ankyrin repeat domain 55 |
| chr18_+_19972853 | 0.66 |
ENSDART00000180071
|
skor1b
|
SKI family transcriptional corepressor 1b |
| chr1_-_22757145 | 0.66 |
ENSDART00000134719
|
prom1b
|
prominin 1 b |
| chr5_+_69747417 | 0.65 |
ENSDART00000153717
|
si:ch211-275j6.5
|
si:ch211-275j6.5 |
| chr7_+_66822229 | 0.65 |
ENSDART00000112109
|
lyve1a
|
lymphatic vessel endothelial hyaluronic receptor 1a |
| chr19_-_41371978 | 0.65 |
ENSDART00000166063
ENSDART00000170343 |
slc25a13
|
solute carrier family 25 (aspartate/glutamate carrier), member 13 |
| chr11_-_25733910 | 0.65 |
ENSDART00000171935
|
brpf3a
|
bromodomain and PHD finger containing, 3a |
| chr7_+_13382852 | 0.64 |
ENSDART00000166318
|
dagla
|
diacylglycerol lipase, alpha |
| chr4_-_5019113 | 0.64 |
ENSDART00000189321
ENSDART00000081990 |
strip2
|
striatin interacting protein 2 |
| chr6_-_56103718 | 0.64 |
ENSDART00000191002
|
gcnt7
|
glucosaminyl (N-acetyl) transferase family member 7 |
| chr11_+_40812590 | 0.63 |
ENSDART00000186690
|
errfi1a
|
ERBB receptor feedback inhibitor 1a |
| chr15_+_34592215 | 0.63 |
ENSDART00000099776
|
tspan13a
|
tetraspanin 13a |
| chr10_-_17083180 | 0.63 |
ENSDART00000170083
|
fam166b
|
family with sequence similarity 166, member B |
| chr7_-_11596450 | 0.62 |
ENSDART00000173863
|
stard5
|
StAR-related lipid transfer (START) domain containing 5 |
| chr21_+_27278120 | 0.62 |
ENSDART00000193882
|
si:dkey-175m17.7
|
si:dkey-175m17.7 |
| chr2_-_9059955 | 0.62 |
ENSDART00000022768
|
ak5
|
adenylate kinase 5 |
| chr17_+_43623598 | 0.62 |
ENSDART00000154138
|
znf365
|
zinc finger protein 365 |
| chr17_+_28533102 | 0.61 |
ENSDART00000156218
|
mdga2a
|
MAM domain containing glycosylphosphatidylinositol anchor 2a |
| chr25_+_7671640 | 0.61 |
ENSDART00000145367
|
kcnj11l
|
potassium inwardly-rectifying channel, subfamily J, member 11, like |
| chr3_-_32337653 | 0.61 |
ENSDART00000156918
ENSDART00000156551 |
si:dkey-16p21.8
|
si:dkey-16p21.8 |
| chr22_-_14739491 | 0.61 |
ENSDART00000133385
|
lrp1ba
|
low density lipoprotein receptor-related protein 1Ba |
| chr7_+_71547747 | 0.61 |
ENSDART00000180869
|
adcyap1a
|
adenylate cyclase activating polypeptide 1a |
| chr2_+_38554260 | 0.60 |
ENSDART00000171527
|
cdh24b
|
cadherin 24, type 2b |
| chr20_-_9462433 | 0.60 |
ENSDART00000152674
ENSDART00000040557 |
zgc:101840
|
zgc:101840 |
| chr4_+_12111154 | 0.60 |
ENSDART00000036779
|
tmem178b
|
transmembrane protein 178B |
| chr6_+_23887314 | 0.59 |
ENSDART00000163188
|
znf648
|
zinc finger protein 648 |
| chr14_-_45512807 | 0.59 |
ENSDART00000173172
|
si:ch211-114c17.1
|
si:ch211-114c17.1 |
| chr21_-_39327223 | 0.59 |
ENSDART00000115097
|
aifm5
|
apoptosis-inducing factor, mitochondrion-associated, 5 |
| chr9_-_38021889 | 0.58 |
ENSDART00000183482
ENSDART00000124333 |
adcy5
|
adenylate cyclase 5 |
| chr25_-_13839743 | 0.58 |
ENSDART00000158780
|
mapk8ip1a
|
mitogen-activated protein kinase 8 interacting protein 1a |
| chr17_-_2386569 | 0.58 |
ENSDART00000121614
|
PLCB2
|
phospholipase C beta 2 |
| chr3_+_39853788 | 0.57 |
ENSDART00000154869
|
cacna1ha
|
calcium channel, voltage-dependent, T type, alpha 1H subunit a |
| chr13_+_18371208 | 0.57 |
ENSDART00000138172
|
ccar1
|
cell division cycle and apoptosis regulator 1 |
| chr10_-_34870667 | 0.56 |
ENSDART00000161272
|
dclk1a
|
doublecortin-like kinase 1a |
| chr1_-_15797663 | 0.56 |
ENSDART00000177122
|
SGCZ
|
sarcoglycan zeta |
| chr24_+_24461341 | 0.56 |
ENSDART00000147658
|
bhlhe22
|
basic helix-loop-helix family, member e22 |
| chr10_-_34871737 | 0.56 |
ENSDART00000138755
|
dclk1a
|
doublecortin-like kinase 1a |
| chr22_+_34784075 | 0.56 |
ENSDART00000167538
|
lcor
|
ligand dependent nuclear receptor corepressor |
| chr23_-_37113215 | 0.55 |
ENSDART00000146835
|
zgc:193690
|
zgc:193690 |
| chr3_-_30061985 | 0.55 |
ENSDART00000189583
|
ppfia3
|
protein tyrosine phosphatase, receptor type, f polypeptide (PTPRF), interacting protein (liprin), alpha 3 |
| chr17_+_11675362 | 0.55 |
ENSDART00000157911
|
kif26ba
|
kinesin family member 26Ba |
| chr25_-_25058508 | 0.55 |
ENSDART00000087570
ENSDART00000178891 |
FQ311928.1
|
|
| chr5_+_19712011 | 0.54 |
ENSDART00000131924
|
fam222a
|
family with sequence similarity 222, member A |
| chr5_-_34185115 | 0.54 |
ENSDART00000192771
|
fibcd1
|
fibrinogen C domain containing 1 |
| chr21_-_5799122 | 0.54 |
ENSDART00000129351
ENSDART00000151202 |
ccni
|
cyclin I |
| chr6_-_59271270 | 0.53 |
ENSDART00000126870
|
ndufa4l2b
|
NADH dehydrogenase (ubiquinone) 1 alpha subcomplex, 4-like 2b |
Network of associatons between targets according to the STRING database.
First level regulatory network of lhx3+lhx4
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 2.7 | GO:1900620 | acetylcholine biosynthetic process(GO:0008292) acetate ester biosynthetic process(GO:1900620) |
| 0.5 | 1.4 | GO:0034154 | toll-like receptor 7 signaling pathway(GO:0034154) |
| 0.4 | 4.8 | GO:0006032 | chitin catabolic process(GO:0006032) |
| 0.3 | 1.0 | GO:1901187 | regulation of ephrin receptor signaling pathway(GO:1901187) |
| 0.3 | 2.0 | GO:0006572 | tyrosine catabolic process(GO:0006572) |
| 0.3 | 1.2 | GO:0099563 | modification of synaptic structure(GO:0099563) |
| 0.3 | 1.2 | GO:0015677 | copper ion import(GO:0015677) |
| 0.3 | 0.8 | GO:0035971 | peptidyl-histidine dephosphorylation(GO:0035971) |
| 0.3 | 0.8 | GO:0010265 | SCF complex assembly(GO:0010265) |
| 0.2 | 0.7 | GO:0061182 | negative regulation of chondrocyte differentiation(GO:0032331) negative regulation of cartilage development(GO:0061037) negative regulation of chondrocyte development(GO:0061182) regulation of bone mineralization involved in bone maturation(GO:1900157) negative regulation of bone mineralization involved in bone maturation(GO:1900158) negative regulation of bone development(GO:1903011) |
| 0.2 | 0.9 | GO:2000275 | regulation of oxidative phosphorylation uncoupler activity(GO:2000275) |
| 0.2 | 1.1 | GO:0051659 | maintenance of mitochondrion location(GO:0051659) |
| 0.2 | 1.1 | GO:0002159 | desmosome assembly(GO:0002159) |
| 0.2 | 1.5 | GO:0070285 | pigment cell development(GO:0070285) |
| 0.2 | 0.6 | GO:0021611 | facial nerve formation(GO:0021611) |
| 0.2 | 1.1 | GO:0006788 | heme oxidation(GO:0006788) |
| 0.2 | 0.7 | GO:0048313 | organelle inheritance(GO:0048308) Golgi inheritance(GO:0048313) |
| 0.2 | 1.2 | GO:0042983 | amyloid precursor protein biosynthetic process(GO:0042983) regulation of amyloid precursor protein biosynthetic process(GO:0042984) |
| 0.2 | 1.3 | GO:2000650 | negative regulation of sodium ion transmembrane transporter activity(GO:2000650) |
| 0.2 | 0.6 | GO:0071926 | endocannabinoid signaling pathway(GO:0071926) retrograde trans-synaptic signaling by lipid(GO:0098920) retrograde trans-synaptic signaling by endocannabinoid(GO:0098921) |
| 0.2 | 0.8 | GO:0019254 | carnitine metabolic process, CoA-linked(GO:0019254) |
| 0.2 | 1.5 | GO:0072506 | phosphate ion homeostasis(GO:0055062) trivalent inorganic anion homeostasis(GO:0072506) |
| 0.2 | 7.9 | GO:0003009 | skeletal muscle contraction(GO:0003009) |
| 0.1 | 2.2 | GO:0042026 | protein refolding(GO:0042026) |
| 0.1 | 0.4 | GO:0009750 | response to fructose(GO:0009750) |
| 0.1 | 0.7 | GO:0003210 | cardiac atrium formation(GO:0003210) |
| 0.1 | 0.5 | GO:0051876 | pigment granule dispersal(GO:0051876) |
| 0.1 | 0.6 | GO:0043508 | negative regulation of JUN kinase activity(GO:0043508) |
| 0.1 | 2.0 | GO:0060088 | auditory receptor cell stereocilium organization(GO:0060088) |
| 0.1 | 1.7 | GO:0021520 | spinal cord motor neuron cell fate specification(GO:0021520) |
| 0.1 | 5.8 | GO:0030282 | bone mineralization(GO:0030282) |
| 0.1 | 0.9 | GO:0097476 | spinal cord motor neuron migration(GO:0097476) lateral motor column neuron migration(GO:0097477) |
| 0.1 | 1.6 | GO:0032264 | IMP salvage(GO:0032264) |
| 0.1 | 1.2 | GO:0097104 | presynaptic membrane organization(GO:0097090) postsynaptic membrane assembly(GO:0097104) presynaptic membrane assembly(GO:0097105) |
| 0.1 | 0.4 | GO:0070983 | dendrite guidance(GO:0070983) regulation of homophilic cell adhesion(GO:1903385) |
| 0.1 | 0.9 | GO:0042135 | neurotransmitter catabolic process(GO:0042135) |
| 0.1 | 0.8 | GO:0043092 | amino acid import(GO:0043090) L-amino acid import(GO:0043092) |
| 0.1 | 0.6 | GO:0000395 | mRNA 5'-splice site recognition(GO:0000395) |
| 0.1 | 1.6 | GO:0098943 | neurotransmitter receptor transport, postsynaptic endosome to lysosome(GO:0098943) |
| 0.1 | 1.0 | GO:0016127 | cholesterol catabolic process(GO:0006707) sterol catabolic process(GO:0016127) |
| 0.1 | 0.5 | GO:0097272 | ammonia homeostasis(GO:0097272) |
| 0.1 | 0.6 | GO:0045604 | regulation of epidermal cell differentiation(GO:0045604) |
| 0.1 | 0.8 | GO:2000272 | negative regulation of receptor activity(GO:2000272) |
| 0.1 | 1.8 | GO:2000311 | regulation of alpha-amino-3-hydroxy-5-methyl-4-isoxazole propionate selective glutamate receptor activity(GO:2000311) |
| 0.1 | 2.7 | GO:0014059 | dopamine secretion(GO:0014046) regulation of dopamine secretion(GO:0014059) |
| 0.1 | 0.5 | GO:0021877 | forebrain neuron fate commitment(GO:0021877) |
| 0.1 | 0.4 | GO:0050938 | regulation of xanthophore differentiation(GO:0050938) |
| 0.1 | 1.5 | GO:0060158 | phospholipase C-activating dopamine receptor signaling pathway(GO:0060158) |
| 0.1 | 0.8 | GO:0048796 | swim bladder maturation(GO:0048796) swim bladder inflation(GO:0048798) |
| 0.1 | 0.5 | GO:1900029 | positive regulation of ruffle assembly(GO:1900029) |
| 0.1 | 0.2 | GO:0046379 | extracellular polysaccharide biosynthetic process(GO:0045226) extracellular polysaccharide metabolic process(GO:0046379) |
| 0.1 | 3.3 | GO:0016126 | sterol biosynthetic process(GO:0016126) |
| 0.1 | 2.0 | GO:0055064 | chloride ion homeostasis(GO:0055064) |
| 0.1 | 0.3 | GO:0060092 | regulation of synaptic transmission, glycinergic(GO:0060092) |
| 0.1 | 0.8 | GO:0034394 | protein localization to cell surface(GO:0034394) |
| 0.1 | 0.7 | GO:0015813 | acidic amino acid transport(GO:0015800) aspartate transport(GO:0015810) L-glutamate transport(GO:0015813) malate-aspartate shuttle(GO:0043490) |
| 0.1 | 0.8 | GO:0050962 | detection of light stimulus involved in visual perception(GO:0050908) detection of light stimulus involved in sensory perception(GO:0050962) |
| 0.1 | 1.3 | GO:1901385 | regulation of voltage-gated calcium channel activity(GO:1901385) |
| 0.1 | 0.7 | GO:0038203 | TORC2 signaling(GO:0038203) |
| 0.1 | 2.0 | GO:0070936 | protein K48-linked ubiquitination(GO:0070936) |
| 0.1 | 0.5 | GO:0071340 | skeletal muscle acetylcholine-gated channel clustering(GO:0071340) |
| 0.1 | 0.7 | GO:0006382 | adenosine to inosine editing(GO:0006382) |
| 0.1 | 0.4 | GO:0032475 | otolith formation(GO:0032475) |
| 0.1 | 0.9 | GO:0006825 | copper ion transport(GO:0006825) |
| 0.0 | 0.4 | GO:0098700 | equilibrioception(GO:0050957) neurotransmitter loading into synaptic vesicle(GO:0098700) |
| 0.0 | 0.8 | GO:0050728 | negative regulation of inflammatory response(GO:0050728) |
| 0.0 | 0.4 | GO:0034063 | stress granule assembly(GO:0034063) |
| 0.0 | 0.7 | GO:0044154 | histone H3-K14 acetylation(GO:0044154) |
| 0.0 | 1.3 | GO:0003323 | type B pancreatic cell development(GO:0003323) |
| 0.0 | 0.4 | GO:0019430 | response to superoxide(GO:0000303) response to oxygen radical(GO:0000305) removal of superoxide radicals(GO:0019430) cellular response to oxygen radical(GO:0071450) cellular response to superoxide(GO:0071451) cellular oxidant detoxification(GO:0098869) cellular detoxification(GO:1990748) |
| 0.0 | 1.3 | GO:0007157 | heterophilic cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0007157) |
| 0.0 | 0.7 | GO:0006415 | translational termination(GO:0006415) |
| 0.0 | 0.6 | GO:0055117 | regulation of cardiac muscle contraction(GO:0055117) |
| 0.0 | 0.7 | GO:0098962 | regulation of postsynaptic neurotransmitter receptor activity(GO:0098962) |
| 0.0 | 0.3 | GO:0070584 | mitochondrion morphogenesis(GO:0070584) |
| 0.0 | 0.6 | GO:0045471 | response to ethanol(GO:0045471) |
| 0.0 | 1.2 | GO:0035118 | embryonic pectoral fin morphogenesis(GO:0035118) |
| 0.0 | 0.1 | GO:0006526 | arginine biosynthetic process(GO:0006526) |
| 0.0 | 0.5 | GO:0005980 | polysaccharide catabolic process(GO:0000272) glycogen catabolic process(GO:0005980) glucan catabolic process(GO:0009251) cellular polysaccharide catabolic process(GO:0044247) |
| 0.0 | 0.2 | GO:0048011 | neurotrophin TRK receptor signaling pathway(GO:0048011) |
| 0.0 | 1.0 | GO:0007214 | gamma-aminobutyric acid signaling pathway(GO:0007214) |
| 0.0 | 0.4 | GO:0007026 | negative regulation of microtubule depolymerization(GO:0007026) negative regulation of microtubule polymerization or depolymerization(GO:0031111) |
| 0.0 | 0.5 | GO:0016973 | poly(A)+ mRNA export from nucleus(GO:0016973) |
| 0.0 | 0.2 | GO:0030326 | embryonic limb morphogenesis(GO:0030326) |
| 0.0 | 0.6 | GO:0086010 | membrane depolarization during action potential(GO:0086010) |
| 0.0 | 2.1 | GO:0006414 | translational elongation(GO:0006414) |
| 0.0 | 0.4 | GO:0060122 | inner ear receptor stereocilium organization(GO:0060122) |
| 0.0 | 0.7 | GO:1902285 | semaphorin-plexin signaling pathway involved in neuron projection guidance(GO:1902285) semaphorin-plexin signaling pathway involved in axon guidance(GO:1902287) |
| 0.0 | 0.2 | GO:0070207 | protein trimerization(GO:0070206) protein homotrimerization(GO:0070207) |
| 0.0 | 0.5 | GO:0030216 | keratinocyte differentiation(GO:0030216) |
| 0.0 | 0.3 | GO:0031294 | lymphocyte costimulation(GO:0031294) T cell costimulation(GO:0031295) |
| 0.0 | 0.6 | GO:0050829 | defense response to Gram-negative bacterium(GO:0050829) |
| 0.0 | 3.9 | GO:0006836 | neurotransmitter transport(GO:0006836) |
| 0.0 | 0.2 | GO:0071260 | cellular response to mechanical stimulus(GO:0071260) |
| 0.0 | 0.4 | GO:0003181 | atrioventricular valve morphogenesis(GO:0003181) |
| 0.0 | 0.2 | GO:0006685 | sphingomyelin catabolic process(GO:0006685) |
| 0.0 | 0.1 | GO:1990845 | adaptive thermogenesis(GO:1990845) |
| 0.0 | 1.1 | GO:0006636 | unsaturated fatty acid biosynthetic process(GO:0006636) |
| 0.0 | 0.3 | GO:0030214 | hyaluronan catabolic process(GO:0030214) |
| 0.0 | 0.7 | GO:0035845 | photoreceptor cell outer segment organization(GO:0035845) |
| 0.0 | 1.8 | GO:0006165 | nucleoside diphosphate phosphorylation(GO:0006165) |
| 0.0 | 1.1 | GO:0031122 | cytoplasmic microtubule organization(GO:0031122) |
| 0.0 | 0.1 | GO:0036462 | TRAIL-activated apoptotic signaling pathway(GO:0036462) |
| 0.0 | 0.5 | GO:0051289 | protein homotetramerization(GO:0051289) |
| 0.0 | 0.3 | GO:0042119 | granulocyte activation(GO:0036230) neutrophil activation(GO:0042119) |
| 0.0 | 0.3 | GO:0097324 | melanocyte migration(GO:0097324) |
| 0.0 | 0.5 | GO:0070536 | protein K63-linked deubiquitination(GO:0070536) |
| 0.0 | 0.3 | GO:0032483 | regulation of Rab protein signal transduction(GO:0032483) |
| 0.0 | 1.1 | GO:0018108 | peptidyl-tyrosine phosphorylation(GO:0018108) |
| 0.0 | 0.2 | GO:0001992 | regulation of systemic arterial blood pressure by vasopressin(GO:0001992) |
| 0.0 | 0.9 | GO:1990573 | potassium ion import across plasma membrane(GO:1990573) |
| 0.0 | 0.8 | GO:0000079 | regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0000079) regulation of cyclin-dependent protein kinase activity(GO:1904029) |
| 0.0 | 0.7 | GO:0032011 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.0 | 0.1 | GO:0010692 | regulation of alkaline phosphatase activity(GO:0010692) negative regulation of alkaline phosphatase activity(GO:0010693) |
| 0.0 | 0.1 | GO:0035138 | pectoral fin morphogenesis(GO:0035138) |
| 0.0 | 0.2 | GO:0046599 | regulation of centriole replication(GO:0046599) |
| 0.0 | 1.8 | GO:0002088 | lens development in camera-type eye(GO:0002088) |
| 0.0 | 0.6 | GO:0034332 | adherens junction organization(GO:0034332) |
| 0.0 | 0.3 | GO:0006828 | manganese ion transport(GO:0006828) |
| 0.0 | 0.4 | GO:0060319 | primitive erythrocyte differentiation(GO:0060319) |
| 0.0 | 0.1 | GO:0000492 | box C/D snoRNP assembly(GO:0000492) |
| 0.0 | 0.9 | GO:0060216 | definitive hemopoiesis(GO:0060216) |
| 0.0 | 0.1 | GO:0036498 | IRE1-mediated unfolded protein response(GO:0036498) |
| 0.0 | 0.1 | GO:0099565 | regulation of postsynaptic membrane potential(GO:0060078) chemical synaptic transmission, postsynaptic(GO:0099565) |
| 0.0 | 0.2 | GO:0043114 | regulation of vascular permeability(GO:0043114) |
| 0.0 | 1.0 | GO:0061515 | myeloid cell development(GO:0061515) |
| 0.0 | 0.1 | GO:0045743 | positive regulation of fibroblast growth factor receptor signaling pathway(GO:0045743) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.2 | GO:0060077 | inhibitory synapse(GO:0060077) |
| 0.2 | 7.4 | GO:0005861 | troponin complex(GO:0005861) |
| 0.1 | 0.8 | GO:0070032 | synaptobrevin 2-SNAP-25-syntaxin-1a-complexin I complex(GO:0070032) |
| 0.1 | 0.4 | GO:0031673 | H zone(GO:0031673) |
| 0.1 | 1.3 | GO:0031045 | dense core granule(GO:0031045) |
| 0.1 | 0.7 | GO:0016012 | sarcoglycan complex(GO:0016012) |
| 0.1 | 0.3 | GO:0097124 | cyclin A2-CDK2 complex(GO:0097124) |
| 0.1 | 1.1 | GO:0044295 | axonal growth cone(GO:0044295) |
| 0.1 | 0.7 | GO:0071914 | prominosome(GO:0071914) |
| 0.1 | 1.1 | GO:0030057 | desmosome(GO:0030057) |
| 0.0 | 3.5 | GO:0005891 | voltage-gated calcium channel complex(GO:0005891) |
| 0.0 | 0.4 | GO:0036449 | microtubule minus-end(GO:0036449) |
| 0.0 | 0.8 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.0 | 0.6 | GO:0000243 | commitment complex(GO:0000243) |
| 0.0 | 0.8 | GO:0005903 | brush border(GO:0005903) |
| 0.0 | 0.2 | GO:0032983 | kainate selective glutamate receptor complex(GO:0032983) |
| 0.0 | 0.5 | GO:0005751 | mitochondrial respiratory chain complex IV(GO:0005751) |
| 0.0 | 0.8 | GO:0032420 | stereocilium(GO:0032420) |
| 0.0 | 0.7 | GO:0070776 | H3 histone acetyltransferase complex(GO:0070775) MOZ/MORF histone acetyltransferase complex(GO:0070776) |
| 0.0 | 0.7 | GO:0002116 | semaphorin receptor complex(GO:0002116) |
| 0.0 | 1.0 | GO:0042734 | presynaptic membrane(GO:0042734) |
| 0.0 | 2.1 | GO:0099501 | synaptic vesicle membrane(GO:0030672) exocytic vesicle membrane(GO:0099501) |
| 0.0 | 0.5 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.0 | 1.8 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.0 | 1.0 | GO:0005637 | nuclear inner membrane(GO:0005637) |
| 0.0 | 0.8 | GO:0010494 | cytoplasmic stress granule(GO:0010494) |
| 0.0 | 0.3 | GO:0034751 | aryl hydrocarbon receptor complex(GO:0034751) |
| 0.0 | 0.7 | GO:0098978 | glutamatergic synapse(GO:0098978) |
| 0.0 | 0.1 | GO:0097255 | R2TP complex(GO:0097255) |
| 0.0 | 0.3 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.0 | 0.1 | GO:1990604 | IRE1-TRAF2-ASK1 complex(GO:1990604) |
| 0.0 | 1.0 | GO:0016592 | mediator complex(GO:0016592) |
| 0.0 | 0.4 | GO:0031932 | TORC2 complex(GO:0031932) |
| 0.0 | 0.6 | GO:0034705 | voltage-gated potassium channel complex(GO:0008076) potassium channel complex(GO:0034705) |
| 0.0 | 0.6 | GO:0043204 | perikaryon(GO:0043204) |
| 0.0 | 0.7 | GO:0005762 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.0 | 0.6 | GO:0031970 | mitochondrial intermembrane space(GO:0005758) organelle envelope lumen(GO:0031970) |
| 0.0 | 0.6 | GO:0016342 | catenin complex(GO:0016342) |
| 0.0 | 0.5 | GO:0032154 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.0 | 0.1 | GO:0017146 | NMDA selective glutamate receptor complex(GO:0017146) |
| 0.0 | 0.5 | GO:0005902 | microvillus(GO:0005902) |
| 0.0 | 0.5 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.0 | 0.7 | GO:0032432 | actin filament bundle(GO:0032432) |
| 0.0 | 1.1 | GO:0005884 | actin filament(GO:0005884) |
| 0.0 | 0.4 | GO:0042470 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.0 | 0.6 | GO:0031225 | anchored component of membrane(GO:0031225) |
| 0.0 | 8.4 | GO:0043005 | neuron projection(GO:0043005) |
| 0.0 | 0.8 | GO:0045211 | postsynaptic membrane(GO:0045211) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 2.0 | GO:0003868 | 4-hydroxyphenylpyruvate dioxygenase activity(GO:0003868) |
| 0.4 | 4.8 | GO:0004568 | chitinase activity(GO:0004568) |
| 0.3 | 1.2 | GO:0008823 | cupric reductase activity(GO:0008823) ferric-chelate reductase (NADPH) activity(GO:0052851) |
| 0.3 | 0.9 | GO:0055105 | ubiquitin-protein transferase inhibitor activity(GO:0055105) |
| 0.3 | 0.8 | GO:0004466 | long-chain-acyl-CoA dehydrogenase activity(GO:0004466) |
| 0.3 | 0.8 | GO:0033130 | acetylcholine receptor binding(GO:0033130) |
| 0.2 | 1.4 | GO:0032052 | bile acid binding(GO:0032052) |
| 0.2 | 0.9 | GO:0016206 | catechol O-methyltransferase activity(GO:0016206) |
| 0.2 | 0.6 | GO:0030882 | lipid antigen binding(GO:0030882) |
| 0.2 | 2.7 | GO:0016413 | O-acetyltransferase activity(GO:0016413) |
| 0.2 | 0.8 | GO:0101006 | protein histidine phosphatase activity(GO:0101006) |
| 0.2 | 1.1 | GO:0004743 | pyruvate kinase activity(GO:0004743) |
| 0.2 | 0.7 | GO:0031005 | filamin binding(GO:0031005) |
| 0.2 | 1.1 | GO:0004392 | heme oxygenase (decyclizing) activity(GO:0004392) |
| 0.2 | 1.0 | GO:0033781 | cholesterol 24-hydroxylase activity(GO:0033781) |
| 0.2 | 0.5 | GO:0004087 | carbamoyl-phosphate synthase (ammonia) activity(GO:0004087) |
| 0.2 | 0.6 | GO:0016521 | pituitary adenylate cyclase activating polypeptide activity(GO:0016521) |
| 0.1 | 1.0 | GO:0000254 | C-4 methylsterol oxidase activity(GO:0000254) |
| 0.1 | 2.2 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.1 | 1.6 | GO:0003876 | AMP deaminase activity(GO:0003876) adenosine-phosphate deaminase activity(GO:0047623) |
| 0.1 | 1.2 | GO:0031826 | type 2A serotonin receptor binding(GO:0031826) |
| 0.1 | 0.5 | GO:0072571 | ADP-D-ribose binding(GO:0072570) mono-ADP-D-ribose binding(GO:0072571) |
| 0.1 | 0.8 | GO:0005436 | sodium:phosphate symporter activity(GO:0005436) |
| 0.1 | 0.5 | GO:0004135 | glycogen debranching enzyme activity(GO:0004133) 4-alpha-glucanotransferase activity(GO:0004134) amylo-alpha-1,6-glucosidase activity(GO:0004135) |
| 0.1 | 0.4 | GO:0019777 | Atg12 transferase activity(GO:0019777) |
| 0.1 | 0.7 | GO:0008508 | bile acid:sodium symporter activity(GO:0008508) |
| 0.1 | 0.6 | GO:0016531 | copper chaperone activity(GO:0016531) |
| 0.1 | 1.8 | GO:0005504 | fatty acid binding(GO:0005504) |
| 0.1 | 1.9 | GO:0005247 | voltage-gated chloride channel activity(GO:0005247) |
| 0.1 | 0.5 | GO:0005412 | glucose:sodium symporter activity(GO:0005412) |
| 0.1 | 0.9 | GO:0004046 | aminoacylase activity(GO:0004046) |
| 0.1 | 0.6 | GO:0008332 | low voltage-gated calcium channel activity(GO:0008332) |
| 0.1 | 0.4 | GO:0019960 | C-X3-C chemokine binding(GO:0019960) protein binding involved in cell-matrix adhesion(GO:0098634) collagen binding involved in cell-matrix adhesion(GO:0098639) |
| 0.1 | 0.2 | GO:0050501 | hyaluronan synthase activity(GO:0050501) |
| 0.1 | 1.1 | GO:0015187 | glycine transmembrane transporter activity(GO:0015187) |
| 0.1 | 0.3 | GO:0005153 | interleukin-8 receptor binding(GO:0005153) |
| 0.1 | 0.9 | GO:0008353 | RNA polymerase II carboxy-terminal domain kinase activity(GO:0008353) |
| 0.1 | 1.1 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.1 | 0.7 | GO:0008079 | translation release factor activity(GO:0003747) translation termination factor activity(GO:0008079) |
| 0.1 | 0.4 | GO:0008379 | thioredoxin peroxidase activity(GO:0008379) |
| 0.1 | 0.9 | GO:0015272 | ATP-activated inward rectifier potassium channel activity(GO:0015272) |
| 0.1 | 0.7 | GO:0003726 | double-stranded RNA adenosine deaminase activity(GO:0003726) |
| 0.1 | 0.7 | GO:0032036 | myosin heavy chain binding(GO:0032036) |
| 0.1 | 3.1 | GO:0043022 | ribosome binding(GO:0043022) |
| 0.1 | 0.6 | GO:0004465 | lipoprotein lipase activity(GO:0004465) |
| 0.1 | 1.0 | GO:0005313 | L-glutamate transmembrane transporter activity(GO:0005313) acidic amino acid transmembrane transporter activity(GO:0015172) |
| 0.1 | 1.2 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 0.1 | 2.4 | GO:0001786 | phosphatidylserine binding(GO:0001786) |
| 0.1 | 0.2 | GO:0001635 | adrenomedullin receptor activity(GO:0001605) calcitonin gene-related peptide receptor activity(GO:0001635) |
| 0.1 | 0.4 | GO:0030274 | LIM domain binding(GO:0030274) |
| 0.0 | 1.0 | GO:0046875 | ephrin receptor binding(GO:0046875) |
| 0.0 | 0.1 | GO:0034618 | arginine binding(GO:0034618) |
| 0.0 | 1.9 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.0 | 0.4 | GO:0045294 | alpha-catenin binding(GO:0045294) |
| 0.0 | 0.7 | GO:0043994 | H3 histone acetyltransferase activity(GO:0010484) histone acetyltransferase activity (H3-K23 specific)(GO:0043994) |
| 0.0 | 0.2 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 0.0 | 0.8 | GO:0045134 | guanosine-diphosphatase activity(GO:0004382) uridine-diphosphatase activity(GO:0045134) |
| 0.0 | 0.6 | GO:0008330 | protein tyrosine/threonine phosphatase activity(GO:0008330) |
| 0.0 | 1.3 | GO:0008331 | high voltage-gated calcium channel activity(GO:0008331) |
| 0.0 | 0.2 | GO:0044736 | acid-sensing ion channel activity(GO:0044736) |
| 0.0 | 1.0 | GO:0016628 | oxidoreductase activity, acting on the CH-CH group of donors, NAD or NADP as acceptor(GO:0016628) |
| 0.0 | 0.2 | GO:0015277 | kainate selective glutamate receptor activity(GO:0015277) |
| 0.0 | 0.9 | GO:0005343 | organic acid:sodium symporter activity(GO:0005343) |
| 0.0 | 0.8 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.0 | 0.4 | GO:0070095 | fructose-6-phosphate binding(GO:0070095) |
| 0.0 | 0.3 | GO:0017110 | nucleoside-diphosphatase activity(GO:0017110) |
| 0.0 | 0.6 | GO:0070122 | isopeptidase activity(GO:0070122) |
| 0.0 | 0.4 | GO:0030507 | spectrin binding(GO:0030507) |
| 0.0 | 1.6 | GO:0005245 | voltage-gated calcium channel activity(GO:0005245) |
| 0.0 | 0.8 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.0 | 0.1 | GO:0001596 | angiotensin type I receptor activity(GO:0001596) |
| 0.0 | 0.5 | GO:0004724 | magnesium-dependent protein serine/threonine phosphatase activity(GO:0004724) |
| 0.0 | 2.4 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.0 | 0.8 | GO:0015248 | sterol transporter activity(GO:0015248) |
| 0.0 | 0.4 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.0 | 0.2 | GO:0042834 | peptidoglycan binding(GO:0042834) |
| 0.0 | 1.2 | GO:0004683 | calmodulin-dependent protein kinase activity(GO:0004683) |
| 0.0 | 0.5 | GO:0005242 | inward rectifier potassium channel activity(GO:0005242) |
| 0.0 | 0.5 | GO:0097153 | cysteine-type endopeptidase activity involved in apoptotic process(GO:0097153) |
| 0.0 | 0.6 | GO:0004016 | adenylate cyclase activity(GO:0004016) |
| 0.0 | 0.7 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.0 | 0.3 | GO:0004415 | hyalurononglucosaminidase activity(GO:0004415) |
| 0.0 | 0.6 | GO:0004550 | nucleoside diphosphate kinase activity(GO:0004550) |
| 0.0 | 0.7 | GO:0043539 | protein serine/threonine kinase activator activity(GO:0043539) |
| 0.0 | 0.7 | GO:0015485 | cholesterol binding(GO:0015485) |
| 0.0 | 0.3 | GO:0004322 | ferroxidase activity(GO:0004322) oxidoreductase activity, oxidizing metal ions, oxygen as acceptor(GO:0016724) |
| 0.0 | 0.5 | GO:0015002 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors(GO:0016675) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.0 | 0.6 | GO:0051537 | 2 iron, 2 sulfur cluster binding(GO:0051537) |
| 0.0 | 0.3 | GO:0001591 | dopamine neurotransmitter receptor activity, coupled via Gi/Go(GO:0001591) |
| 0.0 | 0.4 | GO:0022848 | acetylcholine-gated cation channel activity(GO:0022848) |
| 0.0 | 0.7 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.0 | 0.2 | GO:0005000 | vasopressin receptor activity(GO:0005000) |
| 0.0 | 0.2 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.0 | 0.6 | GO:0030374 | ligand-dependent nuclear receptor transcription coactivator activity(GO:0030374) |
| 0.0 | 0.8 | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity(GO:0016538) |
| 0.0 | 0.1 | GO:0016212 | kynurenine-oxoglutarate transaminase activity(GO:0016212) kynurenine aminotransferase activity(GO:0036137) |
| 0.0 | 0.4 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.0 | 0.3 | GO:0015279 | store-operated calcium channel activity(GO:0015279) |
| 0.0 | 0.4 | GO:0004629 | phosphatidylinositol phospholipase C activity(GO:0004435) phospholipase C activity(GO:0004629) |
| 0.0 | 2.0 | GO:0042803 | protein homodimerization activity(GO:0042803) |
| 0.0 | 0.2 | GO:0036312 | phosphatidylinositol 3-kinase regulatory subunit binding(GO:0036312) |
| 0.0 | 0.2 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.0 | 2.4 | GO:0042393 | histone binding(GO:0042393) |
| 0.0 | 1.3 | GO:0004843 | thiol-dependent ubiquitin-specific protease activity(GO:0004843) thiol-dependent ubiquitinyl hydrolase activity(GO:0036459) ubiquitinyl hydrolase activity(GO:0101005) |
| 0.0 | 0.2 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.0 | 0.6 | GO:0005178 | integrin binding(GO:0005178) |
| 0.0 | 0.3 | GO:0008378 | galactosyltransferase activity(GO:0008378) |
| 0.0 | 0.7 | GO:0046332 | SMAD binding(GO:0046332) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.0 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.1 | 0.6 | PID THROMBIN PAR4 PATHWAY | PAR4-mediated thrombin signaling events |
| 0.0 | 0.6 | PID LPA4 PATHWAY | LPA4-mediated signaling events |
| 0.0 | 1.1 | PID EPHA FWDPATHWAY | EPHA forward signaling |
| 0.0 | 0.8 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.0 | 0.2 | PID RHOA PATHWAY | RhoA signaling pathway |
| 0.0 | 0.4 | PID AR NONGENOMIC PATHWAY | Nongenotropic Androgen signaling |
| 0.0 | 2.2 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.0 | 1.2 | PID TRKR PATHWAY | Neurotrophic factor-mediated Trk receptor signaling |
| 0.0 | 0.6 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.0 | 0.3 | ST GA12 PATHWAY | G alpha 12 Pathway |
| 0.0 | 0.3 | PID PI3K PLC TRK PATHWAY | Trk receptor signaling mediated by PI3K and PLC-gamma |
| 0.0 | 0.3 | SA REG CASCADE OF CYCLIN EXPR | Expression of cyclins regulates progression through the cell cycle by activating cyclin-dependent kinases. |
| 0.0 | 0.7 | PID RET PATHWAY | Signaling events regulated by Ret tyrosine kinase |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.7 | REACTOME RECYCLING OF BILE ACIDS AND SALTS | Genes involved in Recycling of bile acids and salts |
| 0.1 | 1.4 | REACTOME TRAFFICKING AND PROCESSING OF ENDOSOMAL TLR | Genes involved in Trafficking and processing of endosomal TLR |
| 0.1 | 1.1 | REACTOME ALPHA LINOLENIC ACID ALA METABOLISM | Genes involved in alpha-linolenic acid (ALA) metabolism |
| 0.1 | 2.2 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.1 | 0.3 | REACTOME ROLE OF SECOND MESSENGERS IN NETRIN1 SIGNALING | Genes involved in Role of second messengers in netrin-1 signaling |
| 0.1 | 1.1 | REACTOME GABA A RECEPTOR ACTIVATION | Genes involved in GABA A receptor activation |
| 0.1 | 1.1 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.1 | 1.0 | REACTOME SIGNAL ATTENUATION | Genes involved in Signal attenuation |
| 0.1 | 0.6 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.1 | 0.8 | REACTOME MITOCHONDRIAL FATTY ACID BETA OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.1 | 0.6 | REACTOME ADENYLATE CYCLASE ACTIVATING PATHWAY | Genes involved in Adenylate cyclase activating pathway |
| 0.0 | 0.9 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.0 | 0.6 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.0 | 0.6 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.0 | 0.7 | REACTOME GLUCONEOGENESIS | Genes involved in Gluconeogenesis |
| 0.0 | 0.4 | REACTOME PLATELET CALCIUM HOMEOSTASIS | Genes involved in Platelet calcium homeostasis |
| 0.0 | 0.3 | REACTOME AMINE LIGAND BINDING RECEPTORS | Genes involved in Amine ligand-binding receptors |
| 0.0 | 0.4 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.0 | 0.4 | REACTOME ANTIGEN ACTIVATES B CELL RECEPTOR LEADING TO GENERATION OF SECOND MESSENGERS | Genes involved in Antigen Activates B Cell Receptor Leading to Generation of Second Messengers |
| 0.0 | 0.3 | REACTOME CYCLIN A B1 ASSOCIATED EVENTS DURING G2 M TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 0.0 | 0.8 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.0 | 0.2 | REACTOME HYALURONAN METABOLISM | Genes involved in Hyaluronan metabolism |
| 0.0 | 0.2 | REACTOME SEMA4D INDUCED CELL MIGRATION AND GROWTH CONE COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.0 | 1.0 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.0 | 0.3 | REACTOME METAL ION SLC TRANSPORTERS | Genes involved in Metal ion SLC transporters |
| 0.0 | 0.4 | REACTOME AMINO ACID AND OLIGOPEPTIDE SLC TRANSPORTERS | Genes involved in Amino acid and oligopeptide SLC transporters |
| 0.0 | 0.2 | REACTOME IONOTROPIC ACTIVITY OF KAINATE RECEPTORS | Genes involved in Ionotropic activity of Kainate Receptors |
| 0.0 | 0.4 | REACTOME REGULATION OF GLUCOKINASE BY GLUCOKINASE REGULATORY PROTEIN | Genes involved in Regulation of Glucokinase by Glucokinase Regulatory Protein |
| 0.0 | 0.4 | REACTOME VOLTAGE GATED POTASSIUM CHANNELS | Genes involved in Voltage gated Potassium channels |
| 0.0 | 0.3 | REACTOME KERATAN SULFATE BIOSYNTHESIS | Genes involved in Keratan sulfate biosynthesis |
| 0.0 | 0.1 | REACTOME CLASS C 3 METABOTROPIC GLUTAMATE PHEROMONE RECEPTORS | Genes involved in Class C/3 (Metabotropic glutamate/pheromone receptors) |