Project
PRJEB1986: zebrafish developmental stages transcriptome
Navigation
Downloads
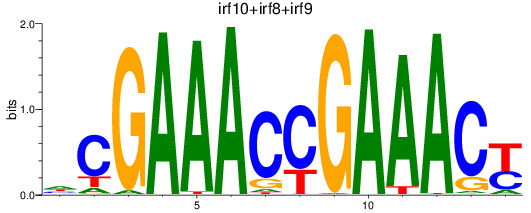
Results for irf10+irf8+irf9
Z-value: 0.87
Transcription factors associated with irf10+irf8+irf9
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
irf9
|
ENSDARG00000016457 | interferon regulatory factor 9 |
|
irf10
|
ENSDARG00000027658 | interferon regulatory factor 10 |
|
irf8
|
ENSDARG00000056407 | interferon regulatory factor 8 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| irf8 | dr11_v1_chr18_+_30567945_30567945 | 0.52 | 2.1e-02 | Click! |
| irf9 | dr11_v1_chr12_+_13282797_13282807 | 0.36 | 1.3e-01 | Click! |
| irf10 | dr11_v1_chr23_+_642001_642052 | 0.26 | 2.9e-01 | Click! |
Activity profile of irf10+irf8+irf9 motif
Sorted Z-values of irf10+irf8+irf9 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr5_+_38662125 | 1.31 |
ENSDART00000136949
|
si:dkey-58f10.13
|
si:dkey-58f10.13 |
| chr19_-_7115229 | 1.11 |
ENSDART00000001930
|
psmb13a
|
proteasome subunit beta 13a |
| chr15_-_17071328 | 1.02 |
ENSDART00000122617
|
si:ch211-24o10.6
|
si:ch211-24o10.6 |
| chr19_+_7115223 | 1.00 |
ENSDART00000001359
|
psmb12
|
proteasome subunit beta 12 |
| chr11_-_20096018 | 0.96 |
ENSDART00000030420
|
ogfrl2
|
opioid growth factor receptor-like 2 |
| chr4_-_12795030 | 0.87 |
ENSDART00000150427
|
b2m
|
beta-2-microglobulin |
| chr15_-_43625549 | 0.85 |
ENSDART00000168589
|
ctsc
|
cathepsin C |
| chr21_-_22648007 | 0.79 |
ENSDART00000121788
|
gig2l
|
grass carp reovirus (GCRV)-induced gene 2l |
| chr16_+_29509133 | 0.78 |
ENSDART00000112116
|
ctss2.1
|
cathepsin S, ortholog2, tandem duplicate 1 |
| chr18_+_26422124 | 0.76 |
ENSDART00000060245
|
ctsh
|
cathepsin H |
| chr1_-_7930679 | 0.67 |
ENSDART00000146090
|
si:dkey-79f11.10
|
si:dkey-79f11.10 |
| chr10_+_17776981 | 0.66 |
ENSDART00000141693
|
ccl19b
|
chemokine (C-C motif) ligand 19b |
| chr13_-_37642890 | 0.64 |
ENSDART00000146483
ENSDART00000136071 ENSDART00000111786 |
si:dkey-188i13.9
|
si:dkey-188i13.9 |
| chr9_-_18215644 | 0.64 |
ENSDART00000145876
|
epsti1
|
epithelial stromal interaction 1 |
| chr3_+_29941777 | 0.63 |
ENSDART00000113889
|
ifi35
|
interferon-induced protein 35 |
| chr19_-_7110617 | 0.62 |
ENSDART00000104838
|
psmb8a
|
proteasome subunit beta 8A |
| chr15_+_24588963 | 0.60 |
ENSDART00000155075
|
zgc:198241
|
zgc:198241 |
| chr4_-_17669881 | 0.59 |
ENSDART00000066997
|
dram1
|
DNA-damage regulated autophagy modulator 1 |
| chr15_+_2421729 | 0.57 |
ENSDART00000082294
ENSDART00000156428 |
hephl1a
|
hephaestin-like 1a |
| chr3_+_8826905 | 0.57 |
ENSDART00000167474
ENSDART00000168088 |
si:dkeyp-30d6.2
|
si:dkeyp-30d6.2 |
| chr16_+_48678655 | 0.55 |
ENSDART00000150156
|
tap1
|
transporter 1, ATP-binding cassette, sub-family B (MDR/TAP) |
| chr3_-_36127234 | 0.54 |
ENSDART00000130917
|
coil
|
coilin p80 |
| chr15_+_2421432 | 0.53 |
ENSDART00000193772
|
hephl1a
|
hephaestin-like 1a |
| chr15_-_15227541 | 0.52 |
ENSDART00000184787
|
rrp8
|
ribosomal RNA processing 8, methyltransferase, homolog (yeast) |
| chr22_-_17499513 | 0.52 |
ENSDART00000105460
|
si:ch211-197g15.6
|
si:ch211-197g15.6 |
| chr9_+_23765587 | 0.51 |
ENSDART00000145120
|
si:ch211-219a4.3
|
si:ch211-219a4.3 |
| chr15_-_29162193 | 0.51 |
ENSDART00000138449
ENSDART00000099885 |
xaf1
|
XIAP associated factor 1 |
| chr16_+_29514473 | 0.50 |
ENSDART00000034102
|
ctss2.2
|
cathepsin S, ortholog 2, tandem duplicate 2 |
| chr8_-_27656765 | 0.49 |
ENSDART00000078491
|
mov10b.2
|
Moloney leukemia virus 10b, tandem duplicate 2 |
| chr12_-_35936329 | 0.47 |
ENSDART00000166634
|
rnf213b
|
ring finger protein 213b |
| chr13_-_37631092 | 0.46 |
ENSDART00000108855
|
si:dkey-188i13.7
|
si:dkey-188i13.7 |
| chr3_+_52092599 | 0.45 |
ENSDART00000104648
|
zgc:165582
|
zgc:165582 |
| chr1_-_47161996 | 0.45 |
ENSDART00000053153
|
mhc1zba
|
major histocompatibility complex class I ZBA |
| chr1_+_58424507 | 0.44 |
ENSDART00000114197
|
si:ch73-236c18.3
|
si:ch73-236c18.3 |
| chr11_-_19775182 | 0.44 |
ENSDART00000037894
|
namptb
|
nicotinamide phosphoribosyltransferase b |
| chr25_+_14092871 | 0.43 |
ENSDART00000067239
|
guca1g
|
guanylate cyclase activator 1g |
| chr5_+_38619813 | 0.42 |
ENSDART00000133314
|
si:ch211-271e10.3
|
si:ch211-271e10.3 |
| chr15_-_36369743 | 0.42 |
ENSDART00000155942
|
si:dkey-23k10.3
|
si:dkey-23k10.3 |
| chr4_-_26107841 | 0.41 |
ENSDART00000172012
|
si:ch211-244b2.4
|
si:ch211-244b2.4 |
| chr2_-_42035250 | 0.41 |
ENSDART00000056460
ENSDART00000140788 |
gbp1
|
guanylate binding protein 1 |
| chr4_-_76383223 | 0.40 |
ENSDART00000161866
ENSDART00000168456 ENSDART00000174287 ENSDART00000174048 |
zgc:123107
|
zgc:123107 |
| chr22_-_5958066 | 0.40 |
ENSDART00000145821
|
si:rp71-36a1.3
|
si:rp71-36a1.3 |
| chr1_-_7951002 | 0.40 |
ENSDART00000138187
|
si:dkey-79f11.8
|
si:dkey-79f11.8 |
| chr10_+_31809226 | 0.40 |
ENSDART00000087898
|
foxo1b
|
forkhead box O1 b |
| chr4_-_26108053 | 0.39 |
ENSDART00000066951
|
si:ch211-244b2.4
|
si:ch211-244b2.4 |
| chr24_+_12945803 | 0.39 |
ENSDART00000005105
|
psme1
|
proteasome activator subunit 1 |
| chr10_+_26689262 | 0.38 |
ENSDART00000123195
ENSDART00000132946 |
tmtops3b
|
teleost multiple tissue opsin 3b |
| chr2_-_38363017 | 0.38 |
ENSDART00000088026
|
prmt5
|
protein arginine methyltransferase 5 |
| chr15_+_24572926 | 0.37 |
ENSDART00000155636
ENSDART00000187800 |
dhrs13b
|
dehydrogenase/reductase (SDR family) member 13b |
| chr6_+_52947699 | 0.37 |
ENSDART00000180913
|
uba7
|
ubiquitin-like modifier activating enzyme 7 |
| chr2_-_47904043 | 0.36 |
ENSDART00000185328
ENSDART00000126740 |
ftr22
|
finTRIM family, member 22 |
| chr13_-_37608441 | 0.36 |
ENSDART00000140230
|
zgc:123068
|
zgc:123068 |
| chr10_+_26645953 | 0.35 |
ENSDART00000131482
|
adgrg4b
|
adhesion G protein-coupled receptor G4b |
| chr21_-_22676323 | 0.34 |
ENSDART00000167392
|
gig2h
|
grass carp reovirus (GCRV)-induced gene 2h |
| chr22_+_36902352 | 0.34 |
ENSDART00000036273
|
trim35-20
|
tripartite motif containing 35-20 |
| chr22_+_8800432 | 0.33 |
ENSDART00000139940
|
si:dkey-182g1.6
|
si:dkey-182g1.6 |
| chr7_+_19615056 | 0.33 |
ENSDART00000124752
ENSDART00000190297 |
si:ch211-212k18.15
|
si:ch211-212k18.15 |
| chr7_-_19614916 | 0.33 |
ENSDART00000169029
|
zgc:194655
|
zgc:194655 |
| chr3_+_1942219 | 0.33 |
ENSDART00000114520
|
zgc:165583
|
zgc:165583 |
| chr8_+_43016714 | 0.32 |
ENSDART00000142671
|
rassf2a
|
Ras association (RalGDS/AF-6) domain family member 2a |
| chr6_+_58569808 | 0.32 |
ENSDART00000154393
|
helz2
|
helicase with zinc finger 2, transcriptional coactivator |
| chr24_-_28419444 | 0.32 |
ENSDART00000105749
|
nrros
|
negative regulator of reactive oxygen species |
| chr2_+_42005475 | 0.31 |
ENSDART00000056461
|
gbp2
|
guanylate binding protein 2 |
| chr23_+_43668756 | 0.31 |
ENSDART00000112598
|
otud4
|
OTU deubiquitinase 4 |
| chr4_+_18707174 | 0.31 |
ENSDART00000142343
|
wnt7ba
|
wingless-type MMTV integration site family, member 7Ba |
| chr6_+_52947186 | 0.31 |
ENSDART00000155831
|
uba7
|
ubiquitin-like modifier activating enzyme 7 |
| chr12_-_19151708 | 0.31 |
ENSDART00000057124
|
tefa
|
thyrotrophic embryonic factor a |
| chr18_-_38270430 | 0.30 |
ENSDART00000139519
|
caprin1b
|
cell cycle associated protein 1b |
| chr11_+_11974708 | 0.30 |
ENSDART00000125060
|
zgc:64002
|
zgc:64002 |
| chr4_-_31993240 | 0.30 |
ENSDART00000167842
|
si:dkey-72l17.5
|
si:dkey-72l17.5 |
| chr25_-_25550938 | 0.29 |
ENSDART00000150412
ENSDART00000103622 |
irf7
|
interferon regulatory factor 7 |
| chr12_-_290782 | 0.28 |
ENSDART00000152527
ENSDART00000105694 |
adprm
|
ADP-ribose/CDP-alcohol diphosphatase, manganese-dependent |
| chr9_-_43001898 | 0.28 |
ENSDART00000138515
|
ttn.2
|
titin, tandem duplicate 2 |
| chr22_-_6941098 | 0.28 |
ENSDART00000105864
|
zgc:171500
|
zgc:171500 |
| chr16_+_13855039 | 0.28 |
ENSDART00000113764
ENSDART00000143983 |
zgc:174888
|
zgc:174888 |
| chr10_+_7719796 | 0.28 |
ENSDART00000191795
|
ggcx
|
gamma-glutamyl carboxylase |
| chr22_+_34616151 | 0.28 |
ENSDART00000155399
ENSDART00000104705 |
si:ch1073-214b20.2
|
si:ch1073-214b20.2 |
| chr9_+_51147764 | 0.28 |
ENSDART00000188480
|
ifih1
|
interferon induced with helicase C domain 1 |
| chr5_+_8919698 | 0.27 |
ENSDART00000046440
|
agpat9l
|
1-acylglycerol-3-phosphate O-acyltransferase 9, like |
| chr3_+_36127287 | 0.27 |
ENSDART00000058605
ENSDART00000182500 |
scpep1
|
serine carboxypeptidase 1 |
| chr6_+_29217392 | 0.27 |
ENSDART00000006386
|
atp1b1a
|
ATPase Na+/K+ transporting subunit beta 1a |
| chr3_-_1938588 | 0.27 |
ENSDART00000013001
ENSDART00000186405 |
zgc:152753
|
zgc:152753 |
| chr3_-_49015203 | 0.27 |
ENSDART00000186138
|
BX769173.2
|
|
| chr10_+_35152928 | 0.27 |
ENSDART00000063418
|
nsun5
|
NOP2/Sun domain family, member 5 |
| chr25_-_27665978 | 0.27 |
ENSDART00000123590
|
si:ch211-91p5.3
|
si:ch211-91p5.3 |
| chr15_-_24588649 | 0.26 |
ENSDART00000060469
|
gemin4
|
gem (nuclear organelle) associated protein 4 |
| chr15_-_47929455 | 0.26 |
ENSDART00000064462
|
psma6l
|
proteasome subunit alpha 6, like |
| chr17_+_45395846 | 0.26 |
ENSDART00000058793
|
nenf
|
neudesin neurotrophic factor |
| chr22_-_8536482 | 0.26 |
ENSDART00000148134
|
si:ch73-27e22.2
|
si:ch73-27e22.2 |
| chr25_+_35942867 | 0.26 |
ENSDART00000066985
|
hsd17b2
|
hydroxysteroid (17-beta) dehydrogenase 2 |
| chr17_+_35362851 | 0.26 |
ENSDART00000137659
|
cmpk2
|
cytidine monophosphate (UMP-CMP) kinase 2, mitochondrial |
| chr5_+_38684651 | 0.25 |
ENSDART00000137852
|
si:dkey-58f10.10
|
si:dkey-58f10.10 |
| chr1_-_49530615 | 0.25 |
ENSDART00000113315
|
si:ch211-281g13.5
|
si:ch211-281g13.5 |
| chr4_-_71177920 | 0.25 |
ENSDART00000158287
|
si:dkey-193i10.1
|
si:dkey-193i10.1 |
| chr22_-_8692305 | 0.25 |
ENSDART00000181602
|
CR450686.4
|
|
| chr11_+_1584747 | 0.25 |
ENSDART00000154583
|
si:dkey-40c23.2
|
si:dkey-40c23.2 |
| chr22_-_10580194 | 0.24 |
ENSDART00000105848
|
SHISA4 (1 of many)
|
si:dkey-42i9.7 |
| chr23_-_37536127 | 0.24 |
ENSDART00000078253
|
tor1l1
|
torsin family 1 like 1 |
| chr14_-_16810401 | 0.24 |
ENSDART00000158396
ENSDART00000170758 |
tcirg1b
|
T cell immune regulator 1, ATPase H+ transporting V0 subunit a3b |
| chr4_-_51693601 | 0.23 |
ENSDART00000189025
ENSDART00000185516 ENSDART00000189576 |
si:dkeyp-90h9.1
|
si:dkeyp-90h9.1 |
| chr21_-_22673758 | 0.23 |
ENSDART00000164910
|
gig2i
|
grass carp reovirus (GCRV)-induced gene 2i |
| chr4_+_70791887 | 0.23 |
ENSDART00000164234
|
si:ch211-127b6.2
|
si:ch211-127b6.2 |
| chr15_-_43873005 | 0.23 |
ENSDART00000190326
|
NOX4
|
NADPH oxidase 4 |
| chr19_-_325584 | 0.23 |
ENSDART00000134266
|
gpd1c
|
glycerol-3-phosphate dehydrogenase 1c |
| chr4_+_34417403 | 0.23 |
ENSDART00000186032
ENSDART00000167909 |
si:ch211-246b8.2
|
si:ch211-246b8.2 |
| chr5_-_57723929 | 0.23 |
ENSDART00000144237
|
gig2p
|
grass carp reovirus (GCRV)-induced gene 2p |
| chr2_-_43135700 | 0.23 |
ENSDART00000098284
|
ftr14
|
finTRIM family, member 14 |
| chr22_+_9060699 | 0.23 |
ENSDART00000133993
|
BX248395.1
|
|
| chr21_-_30994577 | 0.22 |
ENSDART00000065503
|
pgap2
|
post-GPI attachment to proteins 2 |
| chr4_+_68476490 | 0.22 |
ENSDART00000168543
|
si:dkey-237g15.2
|
si:dkey-237g15.2 |
| chr24_+_10310577 | 0.22 |
ENSDART00000141718
|
otulina
|
OTU deubiquitinase with linear linkage specificity a |
| chr4_+_70782916 | 0.22 |
ENSDART00000187960
|
si:dkey-16p6.1
|
si:dkey-16p6.1 |
| chr3_-_4455951 | 0.22 |
ENSDART00000193908
ENSDART00000074077 |
trim35-3
|
tripartite motif containing 35-3 |
| chr9_+_42269059 | 0.22 |
ENSDART00000113435
|
si:dkey-10c21.1
|
si:dkey-10c21.1 |
| chr21_+_5192016 | 0.21 |
ENSDART00000139288
|
si:dkey-121h17.7
|
si:dkey-121h17.7 |
| chr4_-_58945951 | 0.21 |
ENSDART00000159145
|
si:dkey-28i19.4
|
si:dkey-28i19.4 |
| chr8_+_47188154 | 0.21 |
ENSDART00000137319
|
si:dkeyp-100a1.6
|
si:dkeyp-100a1.6 |
| chr4_-_31699842 | 0.21 |
ENSDART00000166400
|
si:dkey-265e15.5
|
si:dkey-265e15.5 |
| chr5_-_64103863 | 0.21 |
ENSDART00000135014
ENSDART00000083684 |
pappab
|
pregnancy-associated plasma protein A, pappalysin 1b |
| chr9_-_35633827 | 0.20 |
ENSDART00000077745
|
zp2l1
|
zona pellucida glycoprotein 2, like 1 |
| chr17_-_23631400 | 0.20 |
ENSDART00000079563
|
fas
|
Fas cell surface death receptor |
| chr16_+_26439939 | 0.20 |
ENSDART00000143073
|
trim35-28
|
tripartite motif containing 35-28 |
| chr4_-_40008579 | 0.20 |
ENSDART00000167236
|
si:ch211-215p11.1
|
si:ch211-215p11.1 |
| chr23_-_37575030 | 0.20 |
ENSDART00000031875
|
tor1l3
|
torsin family 1 like 3 |
| chr15_-_16384184 | 0.20 |
ENSDART00000154504
|
fam222bb
|
family with sequence similarity 222, member Bb |
| chr2_+_20539402 | 0.20 |
ENSDART00000129585
|
si:ch73-14h1.2
|
si:ch73-14h1.2 |
| chr5_+_32845759 | 0.20 |
ENSDART00000136552
|
si:ch211-208h16.4
|
si:ch211-208h16.4 |
| chr21_+_20901505 | 0.20 |
ENSDART00000132741
|
c7b
|
complement component 7b |
| chr5_+_38743644 | 0.20 |
ENSDART00000137658
|
si:dkey-58f10.6
|
si:dkey-58f10.6 |
| chr4_-_46948863 | 0.19 |
ENSDART00000168835
|
si:dkey-16p6.1
|
si:dkey-16p6.1 |
| chr6_-_52234796 | 0.19 |
ENSDART00000001803
|
tomm34
|
translocase of outer mitochondrial membrane 34 |
| chr5_-_24869213 | 0.19 |
ENSDART00000112287
ENSDART00000144635 |
gas2l1
|
growth arrest-specific 2 like 1 |
| chr12_-_44018667 | 0.19 |
ENSDART00000170692
|
si:dkey-201i2.4
|
si:dkey-201i2.4 |
| chr18_+_34667486 | 0.19 |
ENSDART00000163583
|
CR626874.1
|
|
| chr4_+_34417798 | 0.19 |
ENSDART00000157757
|
si:ch211-246b8.2
|
si:ch211-246b8.2 |
| chr10_-_42147318 | 0.19 |
ENSDART00000134890
|
dusp11
|
dual specificity phosphatase 11 (RNA/RNP complex 1-interacting) |
| chr4_-_75616197 | 0.19 |
ENSDART00000157778
|
si:dkey-71l4.3
|
si:dkey-71l4.3 |
| chr22_-_36750589 | 0.19 |
ENSDART00000010824
|
acy1
|
aminoacylase 1 |
| chr4_-_75511162 | 0.19 |
ENSDART00000157651
|
si:dkey-71l4.5
|
si:dkey-71l4.5 |
| chr19_+_4066449 | 0.19 |
ENSDART00000162461
|
btr26
|
bloodthirsty-related gene family, member 26 |
| chr6_-_51811095 | 0.18 |
ENSDART00000161048
|
psmf1
|
proteasome inhibitor subunit 1 |
| chr4_-_76079075 | 0.18 |
ENSDART00000158469
|
si:ch211-232d10.3
|
si:ch211-232d10.3 |
| chr4_+_58240778 | 0.18 |
ENSDART00000181184
ENSDART00000192348 ENSDART00000189444 ENSDART00000164424 |
si:dkey-16p6.1
|
si:dkey-16p6.1 |
| chr23_-_37835794 | 0.18 |
ENSDART00000137358
|
cd40
|
CD40 molecule, TNF receptor superfamily member 5 |
| chr23_+_642395 | 0.18 |
ENSDART00000186995
|
irf10
|
interferon regulatory factor 10 |
| chr4_-_45043716 | 0.18 |
ENSDART00000181920
|
si:ch211-227e10.6
|
si:ch211-227e10.6 |
| chr2_+_20868286 | 0.17 |
ENSDART00000062591
|
odr4
|
odr-4 GPCR localization factor homolog |
| chr4_-_76206413 | 0.17 |
ENSDART00000170738
|
si:ch211-106j21.5
|
si:ch211-106j21.5 |
| chr4_+_35130622 | 0.17 |
ENSDART00000165372
|
si:dkey-279j5.1
|
si:dkey-279j5.1 |
| chr13_-_25548733 | 0.17 |
ENSDART00000168099
ENSDART00000135788 ENSDART00000077655 |
mcmbp
|
minichromosome maintenance complex binding protein |
| chr4_+_70911498 | 0.17 |
ENSDART00000157674
|
si:dkey-16p6.1
|
si:dkey-16p6.1 |
| chr15_-_37867995 | 0.16 |
ENSDART00000192698
|
si:dkey-238d18.4
|
si:dkey-238d18.4 |
| chr3_-_7997887 | 0.16 |
ENSDART00000172256
|
TRIM35 (1 of many)
|
si:ch211-175l6.2 |
| chr4_-_45028479 | 0.16 |
ENSDART00000143150
ENSDART00000076870 |
si:dkey-51d8.1
|
si:dkey-51d8.1 |
| chr4_+_5842433 | 0.16 |
ENSDART00000124085
ENSDART00000179848 |
usp18
|
ubiquitin specific peptidase 18 |
| chr6_-_19271210 | 0.16 |
ENSDART00000163628
ENSDART00000159124 |
zgc:174863
|
zgc:174863 |
| chr18_+_13248956 | 0.16 |
ENSDART00000080709
|
plcg2
|
phospholipase C, gamma 2 |
| chr4_-_42244412 | 0.16 |
ENSDART00000172686
|
si:ch211-129p6.2
|
si:ch211-129p6.2 |
| chr12_+_38556462 | 0.16 |
ENSDART00000021069
ENSDART00000145377 |
rpl38
|
ribosomal protein L38 |
| chr6_+_32834760 | 0.16 |
ENSDART00000121562
|
cyldl
|
cylindromatosis (turban tumor syndrome), like |
| chr7_+_17908235 | 0.15 |
ENSDART00000077113
|
mta2
|
metastasis associated 1 family, member 2 |
| chr3_-_22366032 | 0.15 |
ENSDART00000029849
|
ifnphi1
|
interferon phi 1 |
| chr4_+_35198656 | 0.15 |
ENSDART00000187990
|
si:dkey-269p2.1
|
si:dkey-269p2.1 |
| chr15_+_13984879 | 0.15 |
ENSDART00000159438
|
zgc:162730
|
zgc:162730 |
| chr10_+_2899108 | 0.15 |
ENSDART00000147031
|
erap1a
|
endoplasmic reticulum aminopeptidase 1a |
| chr1_-_25177086 | 0.15 |
ENSDART00000144711
ENSDART00000177225 |
tmem154
|
transmembrane protein 154 |
| chr1_-_30510839 | 0.15 |
ENSDART00000168189
ENSDART00000174868 |
igf2bp2b
|
insulin-like growth factor 2 mRNA binding protein 2b |
| chr1_+_58332000 | 0.15 |
ENSDART00000145234
|
ggt1l2.1
|
gamma-glutamyltransferase 1 like 2.1 |
| chr3_+_40164129 | 0.15 |
ENSDART00000102526
|
gfer
|
growth factor, augmenter of liver regeneration (ERV1 homolog, S. cerevisiae) |
| chr2_-_32688905 | 0.15 |
ENSDART00000041146
|
nrbp2a
|
nuclear receptor binding protein 2a |
| chr15_-_17169935 | 0.15 |
ENSDART00000110111
|
cul5a
|
cullin 5a |
| chr16_+_25806400 | 0.15 |
ENSDART00000154652
|
irgq1
|
immunity-related GTPase family, q1 |
| chr5_+_27421639 | 0.15 |
ENSDART00000146285
|
cyb561a3a
|
cytochrome b561 family, member A3a |
| chr5_-_39171302 | 0.14 |
ENSDART00000020808
|
paqr3a
|
progestin and adipoQ receptor family member IIIa |
| chr4_-_56673431 | 0.14 |
ENSDART00000161464
ENSDART00000191926 |
si:ch211-227p7.5
|
si:ch211-227p7.5 |
| chr4_+_40329524 | 0.14 |
ENSDART00000165722
|
znf993
|
zinc finger protein 993 |
| chr23_+_642001 | 0.14 |
ENSDART00000030643
ENSDART00000124850 |
irf10
|
interferon regulatory factor 10 |
| chr4_-_55227527 | 0.14 |
ENSDART00000184323
ENSDART00000148091 |
si:dkey-61p9.9
|
si:dkey-61p9.9 |
| chr2_+_42708674 | 0.14 |
ENSDART00000141989
ENSDART00000005767 |
ftr11
|
finTRIM family, member 11 |
| chr22_+_2512154 | 0.14 |
ENSDART00000097363
|
zgc:173726
|
zgc:173726 |
| chr4_+_53115165 | 0.14 |
ENSDART00000157470
|
si:dkey-8o9.2
|
si:dkey-8o9.2 |
| chr4_-_26095755 | 0.14 |
ENSDART00000100611
ENSDART00000191266 |
si:ch211-244b2.3
|
si:ch211-244b2.3 |
| chr13_-_45155792 | 0.14 |
ENSDART00000163556
|
runx3
|
runt-related transcription factor 3 |
| chr19_-_27448606 | 0.14 |
ENSDART00000079251
|
vars2
|
valyl-tRNA synthetase 2, mitochondrial (putative) |
| chr10_-_36808348 | 0.14 |
ENSDART00000099320
|
dhrs13a.1
|
dehydrogenase/reductase (SDR family) member 13a, tandem duplicate 1 |
| chr14_-_45967712 | 0.14 |
ENSDART00000043751
ENSDART00000141357 |
macrod1
|
MACRO domain containing 1 |
| chr10_+_26646104 | 0.14 |
ENSDART00000187685
|
adgrg4b
|
adhesion G protein-coupled receptor G4b |
| chr4_-_76129431 | 0.14 |
ENSDART00000162809
|
si:ch211-106j21.2
|
si:ch211-106j21.2 |
| chr2_-_24554416 | 0.13 |
ENSDART00000052061
|
cnn2
|
calponin 2 |
| chr4_-_76239517 | 0.13 |
ENSDART00000172599
|
si:ch211-106j21.6
|
si:ch211-106j21.6 |
| chr3_-_22366562 | 0.13 |
ENSDART00000129447
|
ifnphi1
|
interferon phi 1 |
| chr4_-_51692811 | 0.13 |
ENSDART00000159625
|
si:dkeyp-90h9.1
|
si:dkeyp-90h9.1 |
| chr9_-_40935934 | 0.13 |
ENSDART00000155604
|
C2orf88
|
Small membrane A-kinase anchor protein |
| chr4_-_32466676 | 0.13 |
ENSDART00000193347
ENSDART00000169396 ENSDART00000145784 |
si:dkey-16p6.1
|
si:dkey-16p6.1 |
| chr23_+_14269311 | 0.13 |
ENSDART00000170414
|
CR354537.1
|
|
| chr22_+_9009496 | 0.13 |
ENSDART00000132667
|
si:ch211-213a13.2
|
si:ch211-213a13.2 |
| chr15_-_42760110 | 0.13 |
ENSDART00000152490
|
si:ch211-181d7.3
|
si:ch211-181d7.3 |
| chr4_-_40008099 | 0.13 |
ENSDART00000165574
ENSDART00000171888 |
si:ch211-215p11.1
|
si:ch211-215p11.1 |
| chr4_+_43522945 | 0.12 |
ENSDART00000183921
ENSDART00000181832 |
si:dkeyp-53e4.4
|
si:dkeyp-53e4.4 |
| chr3_-_49925313 | 0.12 |
ENSDART00000164361
|
gcgra
|
glucagon receptor a |
Network of associatons between targets according to the STRING database.
First level regulatory network of irf10+irf8+irf9
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.6 | GO:0042590 | antigen processing and presentation of exogenous peptide antigen(GO:0002478) antigen processing and presentation of exogenous antigen(GO:0019884) antigen processing and presentation of exogenous peptide antigen via MHC class I(GO:0042590) |
| 0.1 | 0.5 | GO:0030576 | nuclear body organization(GO:0030575) Cajal body organization(GO:0030576) |
| 0.1 | 0.3 | GO:0036363 | transforming growth factor beta activation(GO:0036363) |
| 0.1 | 0.5 | GO:0098795 | mRNA cleavage involved in gene silencing by miRNA(GO:0035279) mRNA cleavage involved in gene silencing(GO:0098795) |
| 0.1 | 0.3 | GO:1901533 | negative regulation of hematopoietic progenitor cell differentiation(GO:1901533) |
| 0.1 | 0.4 | GO:0034969 | histone arginine methylation(GO:0034969) |
| 0.1 | 0.5 | GO:2000051 | negative regulation of non-canonical Wnt signaling pathway(GO:2000051) |
| 0.1 | 0.4 | GO:0071763 | nuclear membrane organization(GO:0071763) |
| 0.1 | 2.8 | GO:0010499 | proteasomal ubiquitin-independent protein catabolic process(GO:0010499) |
| 0.1 | 0.5 | GO:0000183 | chromatin silencing at rDNA(GO:0000183) |
| 0.1 | 0.3 | GO:0009211 | pyrimidine deoxyribonucleoside triphosphate metabolic process(GO:0009211) |
| 0.1 | 1.2 | GO:0034122 | negative regulation of toll-like receptor signaling pathway(GO:0034122) |
| 0.1 | 0.3 | GO:0032608 | interferon-beta production(GO:0032608) regulation of interferon-beta production(GO:0032648) positive regulation of interferon-beta production(GO:0032728) |
| 0.1 | 0.6 | GO:0031179 | peptide modification(GO:0031179) |
| 0.1 | 0.4 | GO:0002504 | antigen processing and presentation of peptide or polysaccharide antigen via MHC class II(GO:0002504) |
| 0.1 | 0.2 | GO:0042832 | response to protozoan(GO:0001562) TRIF-dependent toll-like receptor signaling pathway(GO:0035666) defense response to protozoan(GO:0042832) regulation of isotype switching(GO:0045191) positive regulation of isotype switching(GO:0045830) |
| 0.1 | 0.2 | GO:0006116 | NADH oxidation(GO:0006116) glycerol-3-phosphate catabolic process(GO:0046168) |
| 0.1 | 1.1 | GO:0006825 | copper ion transport(GO:0006825) |
| 0.1 | 0.2 | GO:0070227 | lymphocyte apoptotic process(GO:0070227) |
| 0.0 | 0.3 | GO:0017187 | peptidyl-glutamic acid carboxylation(GO:0017187) protein carboxylation(GO:0018214) |
| 0.0 | 0.3 | GO:0098773 | skin epidermis development(GO:0098773) |
| 0.0 | 0.2 | GO:1990108 | protein linear deubiquitination(GO:1990108) |
| 0.0 | 0.3 | GO:0046685 | response to arsenic-containing substance(GO:0046685) |
| 0.0 | 0.3 | GO:0046341 | CDP-diacylglycerol biosynthetic process(GO:0016024) CDP-diacylglycerol metabolic process(GO:0046341) |
| 0.0 | 0.4 | GO:1901222 | regulation of NIK/NF-kappaB signaling(GO:1901222) |
| 0.0 | 0.3 | GO:0070475 | rRNA base methylation(GO:0070475) |
| 0.0 | 0.4 | GO:0036065 | fucosylation(GO:0036065) |
| 0.0 | 0.1 | GO:0014005 | microglia development(GO:0014005) |
| 0.0 | 0.1 | GO:1901492 | positive regulation of lymphangiogenesis(GO:1901492) |
| 0.0 | 0.1 | GO:0006370 | 7-methylguanosine mRNA capping(GO:0006370) |
| 0.0 | 0.1 | GO:0071480 | response to gamma radiation(GO:0010332) cellular response to gamma radiation(GO:0071480) |
| 0.0 | 0.3 | GO:0048769 | sarcomerogenesis(GO:0048769) |
| 0.0 | 0.2 | GO:0007035 | vacuolar acidification(GO:0007035) pH reduction(GO:0045851) intracellular pH reduction(GO:0051452) |
| 0.0 | 0.1 | GO:0006686 | sphingomyelin biosynthetic process(GO:0006686) |
| 0.0 | 0.0 | GO:0005997 | xylulose metabolic process(GO:0005997) |
| 0.0 | 0.1 | GO:0071340 | skeletal muscle acetylcholine-gated channel clustering(GO:0071340) |
| 0.0 | 0.6 | GO:0048247 | lymphocyte chemotaxis(GO:0048247) |
| 0.0 | 0.3 | GO:0043171 | peptide catabolic process(GO:0043171) |
| 0.0 | 0.1 | GO:0006390 | transcription from mitochondrial promoter(GO:0006390) |
| 0.0 | 0.1 | GO:2000096 | positive regulation of Wnt signaling pathway, planar cell polarity pathway(GO:2000096) |
| 0.0 | 0.1 | GO:0015886 | heme transport(GO:0015886) iron coordination entity transport(GO:1901678) |
| 0.0 | 0.2 | GO:0051764 | actin crosslink formation(GO:0051764) |
| 0.0 | 0.1 | GO:0030826 | regulation of cGMP metabolic process(GO:0030823) positive regulation of cGMP metabolic process(GO:0030825) regulation of cGMP biosynthetic process(GO:0030826) positive regulation of cGMP biosynthetic process(GO:0030828) regulation of guanylate cyclase activity(GO:0031282) positive regulation of guanylate cyclase activity(GO:0031284) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.9 | GO:0042612 | MHC class I protein complex(GO:0042612) |
| 0.1 | 0.6 | GO:0042824 | MHC class I peptide loading complex(GO:0042824) |
| 0.1 | 0.5 | GO:0033553 | rDNA heterochromatin(GO:0033553) |
| 0.1 | 2.8 | GO:0005839 | proteasome core complex(GO:0005839) |
| 0.1 | 0.4 | GO:0008537 | proteasome activator complex(GO:0008537) |
| 0.1 | 0.4 | GO:0042611 | MHC protein complex(GO:0042611) MHC class II protein complex(GO:0042613) |
| 0.0 | 0.1 | GO:0031466 | Cul5-RING ubiquitin ligase complex(GO:0031466) |
| 0.0 | 0.2 | GO:0009331 | glycerol-3-phosphate dehydrogenase complex(GO:0009331) |
| 0.0 | 0.3 | GO:0032797 | SMN complex(GO:0032797) |
| 0.0 | 0.5 | GO:0015030 | Cajal body(GO:0015030) |
| 0.0 | 0.3 | GO:0005890 | sodium:potassium-exchanging ATPase complex(GO:0005890) cation-transporting ATPase complex(GO:0090533) |
| 0.0 | 0.2 | GO:0071797 | LUBAC complex(GO:0071797) |
| 0.0 | 0.2 | GO:0000220 | vacuolar proton-transporting V-type ATPase, V0 domain(GO:0000220) |
| 0.0 | 0.5 | GO:0043186 | P granule(GO:0043186) |
| 0.0 | 3.7 | GO:0005764 | lysosome(GO:0005764) |
| 0.0 | 0.2 | GO:0005579 | membrane attack complex(GO:0005579) |
| 0.0 | 0.1 | GO:0042645 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.5 | GO:0032574 | 5'-3' RNA helicase activity(GO:0032574) |
| 0.1 | 2.6 | GO:0070003 | threonine-type endopeptidase activity(GO:0004298) threonine-type peptidase activity(GO:0070003) |
| 0.1 | 0.4 | GO:0035243 | protein-arginine omega-N symmetric methyltransferase activity(GO:0035243) |
| 0.1 | 0.6 | GO:0015433 | peptide antigen-transporting ATPase activity(GO:0015433) peptide-transporting ATPase activity(GO:0015440) TAP binding(GO:0046977) |
| 0.1 | 0.5 | GO:0030620 | U2 snRNA binding(GO:0030620) |
| 0.1 | 0.7 | GO:0004839 | ubiquitin activating enzyme activity(GO:0004839) |
| 0.1 | 0.3 | GO:0005132 | type I interferon receptor binding(GO:0005132) |
| 0.1 | 0.3 | GO:0009041 | uridylate kinase activity(GO:0009041) |
| 0.1 | 1.1 | GO:0016724 | ferroxidase activity(GO:0004322) oxidoreductase activity, oxidizing metal ions, oxygen as acceptor(GO:0016724) |
| 0.1 | 0.3 | GO:0008488 | gamma-glutamyl carboxylase activity(GO:0008488) |
| 0.1 | 0.6 | GO:0036374 | peptidyltransferase activity(GO:0000048) glutathione hydrolase activity(GO:0036374) |
| 0.1 | 0.3 | GO:0047035 | testosterone dehydrogenase (NAD+) activity(GO:0047035) |
| 0.1 | 0.4 | GO:0061133 | endopeptidase activator activity(GO:0061133) |
| 0.1 | 0.2 | GO:0004367 | glycerol-3-phosphate dehydrogenase [NAD+] activity(GO:0004367) |
| 0.1 | 0.3 | GO:0004185 | serine-type carboxypeptidase activity(GO:0004185) |
| 0.1 | 0.4 | GO:0008048 | calcium sensitive guanylate cyclase activator activity(GO:0008048) |
| 0.0 | 0.1 | GO:0016971 | flavin-linked sulfhydryl oxidase activity(GO:0016971) |
| 0.0 | 0.2 | GO:0005031 | tumor necrosis factor-activated receptor activity(GO:0005031) |
| 0.0 | 0.2 | GO:0098518 | polynucleotide phosphatase activity(GO:0098518) |
| 0.0 | 0.6 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.0 | 0.3 | GO:0004366 | glycerol-3-phosphate O-acyltransferase activity(GO:0004366) |
| 0.0 | 0.4 | GO:0046920 | alpha-(1->3)-fucosyltransferase activity(GO:0046920) |
| 0.0 | 0.1 | GO:0072590 | N-acetyl-L-aspartate-L-glutamate ligase activity(GO:0072590) citrate-L-glutamate ligase activity(GO:0072591) |
| 0.0 | 1.4 | GO:0003950 | NAD+ ADP-ribosyltransferase activity(GO:0003950) |
| 0.0 | 0.1 | GO:0016519 | gastric inhibitory peptide receptor activity(GO:0016519) |
| 0.0 | 0.1 | GO:0003980 | UDP-glucose:glycoprotein glucosyltransferase activity(GO:0003980) |
| 0.0 | 0.3 | GO:0031386 | protein tag(GO:0031386) |
| 0.0 | 0.1 | GO:0047493 | sphingomyelin synthase activity(GO:0033188) ceramide cholinephosphotransferase activity(GO:0047493) |
| 0.0 | 0.4 | GO:0019894 | kinesin binding(GO:0019894) |
| 0.0 | 2.9 | GO:0004197 | cysteine-type endopeptidase activity(GO:0004197) |
| 0.0 | 0.1 | GO:0052650 | NADP-retinol dehydrogenase activity(GO:0052650) |
| 0.0 | 0.2 | GO:0004046 | aminoacylase activity(GO:0004046) |
| 0.0 | 0.2 | GO:0051117 | ATPase binding(GO:0051117) |
| 0.0 | 0.1 | GO:0015232 | heme transporter activity(GO:0015232) |
| 0.0 | 0.1 | GO:0002161 | aminoacyl-tRNA editing activity(GO:0002161) |
| 0.0 | 0.5 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.0 | 0.3 | GO:0001671 | ATPase activator activity(GO:0001671) |
| 0.0 | 0.6 | GO:0048020 | CCR chemokine receptor binding(GO:0048020) |
| 0.0 | 0.2 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 0.0 | 0.1 | GO:0008174 | mRNA methyltransferase activity(GO:0008174) |
| 0.0 | 0.1 | GO:0050700 | CARD domain binding(GO:0050700) |
| 0.0 | 0.3 | GO:0008307 | structural constituent of muscle(GO:0008307) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | SA FAS SIGNALING | The TNF-type receptor Fas induces apoptosis on ligand binding. |
| 0.0 | 0.9 | PID CD8 TCR DOWNSTREAM PATHWAY | Downstream signaling in naïve CD8+ T cells |
| 0.0 | 1.6 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 0.3 | PID CD40 PATHWAY | CD40/CD40L signaling |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.9 | REACTOME ENDOSOMAL VACUOLAR PATHWAY | Genes involved in Endosomal/Vacuolar pathway |
| 0.1 | 2.0 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 0.0 | 0.4 | REACTOME INTERFERON GAMMA SIGNALING | Genes involved in Interferon gamma signaling |
| 0.0 | 1.0 | REACTOME NEGATIVE REGULATORS OF RIG I MDA5 SIGNALING | Genes involved in Negative regulators of RIG-I/MDA5 signaling |
| 0.0 | 0.6 | REACTOME ANTIGEN PRESENTATION FOLDING ASSEMBLY AND PEPTIDE LOADING OF CLASS I MHC | Genes involved in Antigen Presentation: Folding, assembly and peptide loading of class I MHC |
| 0.0 | 0.3 | REACTOME GAMMA CARBOXYLATION TRANSPORT AND AMINO TERMINAL CLEAVAGE OF PROTEINS | Genes involved in Gamma-carboxylation, transport, and amino-terminal cleavage of proteins |
| 0.0 | 1.6 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.0 | 0.2 | REACTOME EXTRINSIC PATHWAY FOR APOPTOSIS | Genes involved in Extrinsic Pathway for Apoptosis |
| 0.0 | 0.2 | REACTOME IMMUNOREGULATORY INTERACTIONS BETWEEN A LYMPHOID AND A NON LYMPHOID CELL | Genes involved in Immunoregulatory interactions between a Lymphoid and a non-Lymphoid cell |
| 0.0 | 0.1 | REACTOME NFKB IS ACTIVATED AND SIGNALS SURVIVAL | Genes involved in NF-kB is activated and signals survival |
| 0.0 | 0.7 | REACTOME METABOLISM OF NON CODING RNA | Genes involved in Metabolism of non-coding RNA |