Project
PRJEB1986: zebrafish developmental stages transcriptome
Navigation
Downloads
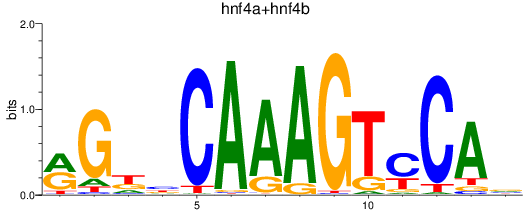
Results for hnf4a+hnf4b
Z-value: 2.56
Transcription factors associated with hnf4a+hnf4b
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
hnf4a
|
ENSDARG00000021494 | hepatocyte nuclear factor 4, alpha |
|
hnf4b
|
ENSDARG00000104742 | hepatic nuclear factor 4, beta |
|
hnf4a
|
ENSDARG00000112985 | hepatocyte nuclear factor 4, alpha |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| hnf4a | dr11_v1_chr23_+_25832689_25832689 | 0.91 | 4.2e-08 | Click! |
| CABZ01057488.1 | dr11_v1_chr7_+_69019851_69019851 | -0.25 | 3.0e-01 | Click! |
Activity profile of hnf4a+hnf4b motif
Sorted Z-values of hnf4a+hnf4b motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr10_-_11385155 | 15.03 |
ENSDART00000064214
|
plac8.1
|
placenta-specific 8, tandem duplicate 1 |
| chr7_-_26457208 | 10.29 |
ENSDART00000173519
|
zgc:172079
|
zgc:172079 |
| chr12_-_20362041 | 9.33 |
ENSDART00000184145
ENSDART00000105952 |
aqp8a.2
|
aquaporin 8a, tandem duplicate 2 |
| chr13_-_303137 | 8.95 |
ENSDART00000099131
|
chs1
|
chitin synthase 1 |
| chr13_+_50375800 | 8.00 |
ENSDART00000099537
|
cox5b2
|
cytochrome c oxidase subunit Vb 2 |
| chr25_-_26758253 | 7.59 |
ENSDART00000123004
|
si:dkeyp-73b11.8
|
si:dkeyp-73b11.8 |
| chr17_-_14613711 | 7.55 |
ENSDART00000157345
|
sdsl
|
serine dehydratase-like |
| chr6_+_30091811 | 7.48 |
ENSDART00000088403
|
meltf
|
melanotransferrin |
| chr9_-_9977827 | 7.23 |
ENSDART00000187315
ENSDART00000010246 |
ugt1ab
|
UDP glucuronosyltransferase 1 family a, b |
| chr7_+_73444325 | 7.08 |
ENSDART00000123016
|
si:ch211-142d6.2
|
si:ch211-142d6.2 |
| chr8_-_37263524 | 6.95 |
ENSDART00000061327
|
rh50
|
Rh50-like protein |
| chr7_-_24520866 | 6.80 |
ENSDART00000077039
|
faah2b
|
fatty acid amide hydrolase 2b |
| chr8_-_20838342 | 6.80 |
ENSDART00000141345
|
si:ch211-133l5.7
|
si:ch211-133l5.7 |
| chr20_+_1412193 | 6.66 |
ENSDART00000064419
|
leg1.1
|
liver-enriched gene 1, tandem duplicate 1 |
| chr23_-_45504991 | 6.49 |
ENSDART00000148761
|
col24a1
|
collagen type XXIV alpha 1 |
| chr9_-_9982696 | 6.46 |
ENSDART00000192548
ENSDART00000125852 |
ugt1ab
|
UDP glucuronosyltransferase 1 family a, b |
| chr12_+_22607761 | 6.41 |
ENSDART00000153112
|
si:dkey-219e21.2
|
si:dkey-219e21.2 |
| chr2_-_37960688 | 5.85 |
ENSDART00000055565
|
cbln14
|
cerebellin 14 |
| chr18_+_26829086 | 5.34 |
ENSDART00000098356
|
slc28a1
|
solute carrier family 28 (concentrative nucleoside transporter), member 1 |
| chr1_+_53374454 | 5.28 |
ENSDART00000038807
|
ucp1
|
uncoupling protein 1 |
| chr10_+_17026870 | 5.21 |
ENSDART00000184529
ENSDART00000157480 |
CR855996.2
|
|
| chr11_+_33284837 | 5.13 |
ENSDART00000166239
ENSDART00000111412 |
nkpd1
|
NTPase, KAP family P-loop domain containing 1 |
| chr3_-_50443607 | 5.10 |
ENSDART00000074036
|
rcvrna
|
recoverin a |
| chr8_-_33154677 | 4.98 |
ENSDART00000133300
|
zbtb34
|
zinc finger and BTB domain containing 34 |
| chr20_-_40754794 | 4.81 |
ENSDART00000187251
|
cx32.3
|
connexin 32.3 |
| chr15_+_19797918 | 4.78 |
ENSDART00000113314
|
si:ch211-229d2.5
|
si:ch211-229d2.5 |
| chr3_+_16762483 | 4.70 |
ENSDART00000132732
|
tmem86b
|
transmembrane protein 86B |
| chr6_-_49873020 | 4.66 |
ENSDART00000148511
|
gnas
|
GNAS complex locus |
| chr16_+_22345513 | 4.61 |
ENSDART00000078000
|
zgc:123238
|
zgc:123238 |
| chr12_-_31457301 | 4.55 |
ENSDART00000043887
ENSDART00000148603 |
acsl5
|
acyl-CoA synthetase long chain family member 5 |
| chr5_-_69940868 | 4.50 |
ENSDART00000185924
ENSDART00000097357 |
ugt2a4
|
UDP glucuronosyltransferase 2 family, polypeptide A4 |
| chr1_-_9641845 | 4.46 |
ENSDART00000121490
ENSDART00000159411 |
ugt5b2
ugt5b3
|
UDP glucuronosyltransferase 5 family, polypeptide B2 UDP glucuronosyltransferase 5 family, polypeptide B3 |
| chr8_-_25329967 | 4.41 |
ENSDART00000139682
|
eps8l3b
|
EPS8-like 3b |
| chr12_-_33706726 | 4.10 |
ENSDART00000153135
|
myo15b
|
myosin XVB |
| chr22_-_31517300 | 4.06 |
ENSDART00000164799
|
slc6a6b
|
solute carrier family 6 (neurotransmitter transporter), member 6b |
| chr18_+_26829362 | 3.95 |
ENSDART00000132728
|
slc28a1
|
solute carrier family 28 (concentrative nucleoside transporter), member 1 |
| chr6_+_27624023 | 3.81 |
ENSDART00000147789
|
slco2a1
|
solute carrier organic anion transporter family, member 2A1 |
| chr3_+_30257582 | 3.61 |
ENSDART00000159497
ENSDART00000103457 ENSDART00000121883 |
mybpc2a
|
myosin binding protein C, fast type a |
| chr18_+_36037223 | 3.55 |
ENSDART00000144410
|
tmem91
|
transmembrane protein 91 |
| chr7_-_33868903 | 3.53 |
ENSDART00000173500
ENSDART00000178746 |
uacab
|
uveal autoantigen with coiled-coil domains and ankyrin repeats b |
| chr21_-_27443995 | 3.51 |
ENSDART00000003508
|
bfb
|
complement component bfb |
| chr23_-_24493768 | 3.45 |
ENSDART00000135559
ENSDART00000046933 |
sult1st5
|
sulfotransferase family 1, cytosolic sulfotransferase 5 |
| chr10_+_10738880 | 3.44 |
ENSDART00000004181
|
slc27a4
|
solute carrier family 27 (fatty acid transporter), member 4 |
| chr15_-_24178893 | 3.40 |
ENSDART00000077980
|
pipox
|
pipecolic acid oxidase |
| chr10_+_45128375 | 3.37 |
ENSDART00000164805
|
camk2b2
|
calcium/calmodulin-dependent protein kinase (CaM kinase) II beta 2 |
| chr23_+_2778813 | 3.36 |
ENSDART00000142621
|
top1
|
DNA topoisomerase I |
| chr13_-_37254777 | 3.35 |
ENSDART00000139734
|
syne2b
|
spectrin repeat containing, nuclear envelope 2b |
| chr21_-_22910520 | 3.34 |
ENSDART00000065567
ENSDART00000191792 |
guca1d
|
guanylate cyclase activator 1d |
| chr7_+_25036188 | 3.29 |
ENSDART00000163957
ENSDART00000169749 |
sb:cb1058
|
sb:cb1058 |
| chr17_+_20173882 | 3.27 |
ENSDART00000155379
|
si:ch211-248a14.8
|
si:ch211-248a14.8 |
| chr23_-_43595956 | 3.20 |
ENSDART00000162186
|
itchb
|
itchy E3 ubiquitin protein ligase b |
| chr7_+_26100024 | 3.17 |
ENSDART00000173726
|
si:ch211-196f2.3
|
si:ch211-196f2.3 |
| chr1_-_46862190 | 3.17 |
ENSDART00000145167
|
agpat3
|
1-acylglycerol-3-phosphate O-acyltransferase 3 |
| chr24_-_32150276 | 3.15 |
ENSDART00000166212
|
cubn
|
cubilin (intrinsic factor-cobalamin receptor) |
| chr2_-_22927958 | 3.15 |
ENSDART00000141621
|
myo7bb
|
myosin VIIBb |
| chr15_-_1885247 | 3.13 |
ENSDART00000149703
|
porb
|
P450 (cytochrome) oxidoreductase b |
| chr18_-_370286 | 3.09 |
ENSDART00000162633
|
si:ch211-79l17.1
|
si:ch211-79l17.1 |
| chr1_+_17900306 | 3.06 |
ENSDART00000089480
|
cyp4v8
|
cytochrome P450, family 4, subfamily V, polypeptide 8 |
| chr17_+_20174044 | 2.99 |
ENSDART00000156028
|
si:ch211-248a14.8
|
si:ch211-248a14.8 |
| chr9_-_470038 | 2.99 |
ENSDART00000170338
ENSDART00000162054 |
si:dkey-11f4.16
|
si:dkey-11f4.16 |
| chr17_-_30702411 | 2.98 |
ENSDART00000114358
|
zgc:194392
|
zgc:194392 |
| chr14_+_9009600 | 2.90 |
ENSDART00000133904
|
si:ch211-274f20.2
|
si:ch211-274f20.2 |
| chr4_+_11375894 | 2.88 |
ENSDART00000190471
ENSDART00000143963 |
pcloa
|
piccolo presynaptic cytomatrix protein a |
| chr5_-_13076779 | 2.83 |
ENSDART00000192826
|
ypel1
|
yippee-like 1 |
| chr4_+_1283068 | 2.78 |
ENSDART00000167233
|
chrm2a
|
cholinergic receptor, muscarinic 2a |
| chr11_-_36957127 | 2.77 |
ENSDART00000168528
|
cacna1da
|
calcium channel, voltage-dependent, L type, alpha 1D subunit, a |
| chr19_-_25005609 | 2.76 |
ENSDART00000151129
|
xkr8.2
|
XK, Kell blood group complex subunit-related family, member 8, tandem duplicate 2 |
| chr2_-_22927581 | 2.72 |
ENSDART00000109515
|
myo7bb
|
myosin VIIBb |
| chr3_+_12744083 | 2.70 |
ENSDART00000158554
ENSDART00000169545 |
cyp2k21
|
cytochrome P450, family 2, subfamily k, polypeptide 21 |
| chr14_+_6429399 | 2.69 |
ENSDART00000149783
ENSDART00000148461 |
abca1b
|
ATP-binding cassette, sub-family A (ABC1), member 1B |
| chr6_-_46474483 | 2.64 |
ENSDART00000155761
|
rdh20
|
retinol dehydrogenase 20 |
| chr5_-_23317477 | 2.58 |
ENSDART00000090171
|
nlgn3b
|
neuroligin 3b |
| chr5_-_55981288 | 2.57 |
ENSDART00000146616
|
si:dkey-189h5.6
|
si:dkey-189h5.6 |
| chr7_+_24520518 | 2.50 |
ENSDART00000173604
|
btr09
|
bloodthirsty-related gene family, member 9 |
| chr20_+_1398564 | 2.49 |
ENSDART00000002242
|
leg1.2
|
liver-enriched gene 1, tandem duplicate 2 |
| chr3_+_46628885 | 2.48 |
ENSDART00000006602
|
pde4a
|
phosphodiesterase 4A, cAMP-specific |
| chr2_-_23778180 | 2.47 |
ENSDART00000136782
|
si:dkey-24c2.7
|
si:dkey-24c2.7 |
| chr6_+_28051978 | 2.47 |
ENSDART00000143218
|
si:ch73-194h10.2
|
si:ch73-194h10.2 |
| chr21_-_36571804 | 2.46 |
ENSDART00000138129
|
wwc1
|
WW and C2 domain containing 1 |
| chr22_+_15323930 | 2.46 |
ENSDART00000142416
|
si:dkey-236e20.3
|
si:dkey-236e20.3 |
| chr23_+_46183410 | 2.46 |
ENSDART00000167596
ENSDART00000151149 ENSDART00000150896 |
btr31
|
bloodthirsty-related gene family, member 31 |
| chr8_+_6576940 | 2.43 |
ENSDART00000138135
|
vsig8b
|
V-set and immunoglobulin domain containing 8b |
| chr22_-_16400484 | 2.43 |
ENSDART00000135987
|
lama3
|
laminin, alpha 3 |
| chr17_+_450956 | 2.37 |
ENSDART00000183022
ENSDART00000171386 |
zgc:194887
|
zgc:194887 |
| chr12_-_36260532 | 2.33 |
ENSDART00000022533
|
kcnj2a
|
potassium inwardly-rectifying channel, subfamily J, member 2a |
| chr12_-_6172154 | 2.27 |
ENSDART00000185434
|
a1cf
|
apobec1 complementation factor |
| chr6_+_36807861 | 2.25 |
ENSDART00000161708
|
si:ch73-29l19.1
|
si:ch73-29l19.1 |
| chr2_+_47249179 | 2.24 |
ENSDART00000144438
|
si:ch211-284d12.3
|
si:ch211-284d12.3 |
| chr5_-_41638039 | 2.18 |
ENSDART00000144525
|
epg5
|
ectopic P-granules autophagy protein 5 homolog (C. elegans) |
| chr13_-_22843562 | 2.18 |
ENSDART00000142738
|
pbld
|
phenazine biosynthesis like protein domain containing |
| chr16_+_32029090 | 2.16 |
ENSDART00000041054
|
tmc4
|
transmembrane channel-like 4 |
| chr12_-_17897134 | 2.14 |
ENSDART00000066407
|
nptx2b
|
neuronal pentraxin IIb |
| chr8_+_7778770 | 2.12 |
ENSDART00000171325
|
tfe3a
|
transcription factor binding to IGHM enhancer 3a |
| chr22_-_11648094 | 2.10 |
ENSDART00000191791
|
dpp4
|
dipeptidyl-peptidase 4 |
| chr20_-_52338782 | 2.10 |
ENSDART00000109735
ENSDART00000132941 |
si:ch1073-287p18.1
|
si:ch1073-287p18.1 |
| chr23_+_9522781 | 2.08 |
ENSDART00000136486
|
osbpl2b
|
oxysterol binding protein-like 2b |
| chr23_-_27702561 | 2.07 |
ENSDART00000053876
|
dnajc22
|
DnaJ (Hsp40) homolog, subfamily C, member 22 |
| chr6_-_1768724 | 2.02 |
ENSDART00000162488
ENSDART00000163613 |
zgc:158417
|
zgc:158417 |
| chr18_-_12957451 | 2.02 |
ENSDART00000140403
|
srgap1a
|
SLIT-ROBO Rho GTPase activating protein 1a |
| chr20_+_43379029 | 1.98 |
ENSDART00000142486
ENSDART00000186486 |
unc93a
|
unc-93 homolog A |
| chr19_+_31532043 | 1.98 |
ENSDART00000136289
|
tmem64
|
transmembrane protein 64 |
| chr8_-_65189 | 1.97 |
ENSDART00000168412
|
hsd17b4
|
hydroxysteroid (17-beta) dehydrogenase 4 |
| chr16_+_2840775 | 1.96 |
ENSDART00000108735
|
clec3ba
|
C-type lectin domain family 3, member Ba |
| chr7_-_60351876 | 1.95 |
ENSDART00000098563
|
plcb3
|
phospholipase C, beta 3 (phosphatidylinositol-specific) |
| chr6_-_40429411 | 1.94 |
ENSDART00000156005
ENSDART00000156357 |
si:dkey-28n18.9
|
si:dkey-28n18.9 |
| chr2_-_38261272 | 1.89 |
ENSDART00000143743
|
si:ch211-14a17.10
|
si:ch211-14a17.10 |
| chr18_+_27515640 | 1.88 |
ENSDART00000181593
|
tp53i11b
|
tumor protein p53 inducible protein 11b |
| chr15_+_26603395 | 1.84 |
ENSDART00000188667
|
slc47a3
|
solute carrier family 47 (multidrug and toxin extrusion), member 3 |
| chr2_-_24407933 | 1.83 |
ENSDART00000088584
|
si:dkey-208k22.6
|
si:dkey-208k22.6 |
| chr22_+_29994093 | 1.80 |
ENSDART00000104778
|
si:dkey-286j15.3
|
si:dkey-286j15.3 |
| chr7_-_5487593 | 1.77 |
ENSDART00000136594
|
arhgef11
|
Rho guanine nucleotide exchange factor (GEF) 11 |
| chr10_-_322769 | 1.75 |
ENSDART00000165244
|
akt2l
|
v-akt murine thymoma viral oncogene homolog 2, like |
| chr24_-_1341543 | 1.73 |
ENSDART00000169341
|
nrp1a
|
neuropilin 1a |
| chr4_-_67799941 | 1.72 |
ENSDART00000185830
|
si:ch211-66c13.1
|
si:ch211-66c13.1 |
| chr8_+_36803415 | 1.72 |
ENSDART00000111680
|
iqsec2b
|
IQ motif and Sec7 domain 2b |
| chr4_+_69619 | 1.71 |
ENSDART00000164425
|
mansc1
|
MANSC domain containing 1 |
| chr24_-_26369185 | 1.71 |
ENSDART00000080039
|
lrrc31
|
leucine rich repeat containing 31 |
| chr2_+_11670270 | 1.70 |
ENSDART00000100524
|
ftr01
|
finTRIM family, member 1 |
| chr2_-_37967368 | 1.69 |
ENSDART00000050345
|
cbln9
|
cerebellin 9 |
| chr22_+_37888249 | 1.67 |
ENSDART00000076082
|
fetub
|
fetuin B |
| chr22_+_15331214 | 1.66 |
ENSDART00000136566
|
sult3st4
|
sulfotransferase family 3, cytosolic sulfotransferase 4 |
| chr1_-_37383741 | 1.63 |
ENSDART00000193155
ENSDART00000191887 ENSDART00000189077 |
scpp1
|
secretory calcium-binding phosphoprotein 1 |
| chr18_-_7400075 | 1.55 |
ENSDART00000101250
|
si:dkey-30c15.13
|
si:dkey-30c15.13 |
| chr18_+_18861359 | 1.55 |
ENSDART00000144605
|
pllp
|
plasmolipin |
| chr2_+_3533458 | 1.50 |
ENSDART00000133007
|
gpt2l
|
glutamic pyruvate transaminase (alanine aminotransferase) 2, like |
| chr8_+_30699429 | 1.50 |
ENSDART00000005345
|
upb1
|
ureidopropionase, beta |
| chr2_-_47870649 | 1.49 |
ENSDART00000142854
|
ftr05
|
finTRIM family, member 5 |
| chr14_+_24934736 | 1.48 |
ENSDART00000191821
|
ppargc1b
|
peroxisome proliferator-activated receptor gamma, coactivator 1 beta |
| chr15_-_20412286 | 1.46 |
ENSDART00000008589
|
chp2
|
calcineurin-like EF-hand protein 2 |
| chr1_-_37383539 | 1.45 |
ENSDART00000127579
|
scpp1
|
secretory calcium-binding phosphoprotein 1 |
| chr11_+_6764207 | 1.44 |
ENSDART00000104292
|
pde4cb
|
phosphodiesterase 4C, cAMP-specific b |
| chr11_+_39672874 | 1.44 |
ENSDART00000046663
ENSDART00000157659 |
camta1b
|
calmodulin binding transcription activator 1b |
| chr3_-_45848257 | 1.37 |
ENSDART00000147198
|
igfals
|
insulin-like growth factor binding protein, acid labile subunit |
| chr24_-_17047918 | 1.32 |
ENSDART00000020204
|
msrb2
|
methionine sulfoxide reductase B2 |
| chr12_-_44199316 | 1.31 |
ENSDART00000170378
|
si:ch73-329n5.1
|
si:ch73-329n5.1 |
| chr6_+_18367388 | 1.31 |
ENSDART00000163394
|
dgke
|
diacylglycerol kinase, epsilon |
| chr5_-_3885727 | 1.29 |
ENSDART00000143250
|
mlxipl
|
MLX interacting protein like |
| chr19_-_30524952 | 1.27 |
ENSDART00000103506
|
hpcal4
|
hippocalcin like 4 |
| chr15_-_26552652 | 1.27 |
ENSDART00000152336
|
serpinf2b
|
serpin peptidase inhibitor, clade F (alpha-2 antiplasmin, pigment epithelium derived factor), member 2b |
| chr19_-_8604429 | 1.27 |
ENSDART00000151165
|
trim46b
|
tripartite motif containing 46b |
| chr7_+_52135791 | 1.21 |
ENSDART00000098705
|
cyp2x12
|
cytochrome P450, family 2, subfamily X, polypeptide 12 |
| chr18_+_31410652 | 1.19 |
ENSDART00000098504
|
def8
|
differentially expressed in FDCP 8 homolog (mouse) |
| chr5_+_8919698 | 1.18 |
ENSDART00000046440
|
agpat9l
|
1-acylglycerol-3-phosphate O-acyltransferase 9, like |
| chr5_-_24112175 | 1.17 |
ENSDART00000131595
|
si:ch211-114c12.5
|
si:ch211-114c12.5 |
| chr2_-_24407732 | 1.17 |
ENSDART00000180612
ENSDART00000153688 ENSDART00000179913 |
si:dkey-208k22.6
|
si:dkey-208k22.6 |
| chr3_-_36440705 | 1.15 |
ENSDART00000162875
|
rogdi
|
rogdi homolog (Drosophila) |
| chr22_-_20126230 | 1.15 |
ENSDART00000138688
|
creb3l3a
|
cAMP responsive element binding protein 3-like 3a |
| chr2_+_22659787 | 1.12 |
ENSDART00000043956
|
zgc:161973
|
zgc:161973 |
| chr11_+_11152214 | 1.11 |
ENSDART00000148030
|
ly75
|
lymphocyte antigen 75 |
| chr22_+_24645325 | 1.11 |
ENSDART00000159531
|
lpar3
|
lysophosphatidic acid receptor 3 |
| chr1_-_47431453 | 1.10 |
ENSDART00000101104
|
gja5b
|
gap junction protein, alpha 5b |
| chr20_-_3087162 | 1.09 |
ENSDART00000152495
|
map3k5
|
mitogen-activated protein kinase kinase kinase 5 |
| chr18_+_27926839 | 1.03 |
ENSDART00000191835
|
hipk3b
|
homeodomain interacting protein kinase 3b |
| chr16_-_25233515 | 1.03 |
ENSDART00000058943
|
zgc:110182
|
zgc:110182 |
| chr22_+_30502555 | 1.02 |
ENSDART00000139128
ENSDART00000104747 |
zgc:171679
|
zgc:171679 |
| chr5_-_62317496 | 1.01 |
ENSDART00000180089
|
zgc:85789
|
zgc:85789 |
| chr15_+_15403560 | 1.00 |
ENSDART00000049831
|
dhrs11b
|
dehydrogenase/reductase (SDR family) member 11b |
| chr21_+_41743493 | 0.99 |
ENSDART00000192669
|
ppp2r2bb
|
protein phosphatase 2, regulatory subunit B, beta b |
| chr23_+_9522942 | 0.99 |
ENSDART00000137751
|
osbpl2b
|
oxysterol binding protein-like 2b |
| chr6_+_27339962 | 0.98 |
ENSDART00000193726
|
klhl30
|
kelch-like family member 30 |
| chr5_+_52039067 | 0.98 |
ENSDART00000143276
|
setbp1
|
SET binding protein 1 |
| chr13_+_8696825 | 0.97 |
ENSDART00000109059
|
ttc7a
|
tetratricopeptide repeat domain 7A |
| chr15_+_40074923 | 0.95 |
ENSDART00000111018
|
ngef
|
neuronal guanine nucleotide exchange factor |
| chr3_-_3496738 | 0.95 |
ENSDART00000186849
|
CABZ01040998.1
|
|
| chr16_-_25606889 | 0.95 |
ENSDART00000077447
ENSDART00000131528 |
zgc:110410
|
zgc:110410 |
| chr10_-_15919839 | 0.94 |
ENSDART00000065032
|
pip5k1ba
|
phosphatidylinositol-4-phosphate 5-kinase, type I, beta a |
| chr14_-_16379843 | 0.92 |
ENSDART00000090711
|
ltc4s
|
leukotriene C4 synthase |
| chr8_+_1766206 | 0.90 |
ENSDART00000021820
|
serpind1
|
serpin peptidase inhibitor, clade D (heparin cofactor), member 1 |
| chr22_-_10774735 | 0.89 |
ENSDART00000081156
|
timm13
|
translocase of inner mitochondrial membrane 13 homolog (yeast) |
| chr23_-_21758253 | 0.89 |
ENSDART00000046613
|
vps13d
|
vacuolar protein sorting 13 homolog D (S. cerevisiae) |
| chr7_+_24115082 | 0.87 |
ENSDART00000182718
|
mrpl52
|
mitochondrial ribosomal protein L52 |
| chr19_+_9277327 | 0.86 |
ENSDART00000104623
ENSDART00000151164 |
si:rp71-15k1.1
|
si:rp71-15k1.1 |
| chr6_-_40722480 | 0.85 |
ENSDART00000188187
|
kbtbd12
|
kelch repeat and BTB (POZ) domain containing 12 |
| chr23_+_33718602 | 0.84 |
ENSDART00000024695
|
dazap2
|
DAZ associated protein 2 |
| chr7_+_24114694 | 0.78 |
ENSDART00000127177
|
mrpl52
|
mitochondrial ribosomal protein L52 |
| chr16_-_25607266 | 0.75 |
ENSDART00000192602
|
zgc:110410
|
zgc:110410 |
| chr12_+_2455429 | 0.74 |
ENSDART00000182929
|
ARHGAP22
|
si:dkey-191m6.4 |
| chr7_+_65398161 | 0.73 |
ENSDART00000166109
ENSDART00000157399 |
usp47
|
ubiquitin specific peptidase 47 |
| chr20_-_25669813 | 0.73 |
ENSDART00000153118
|
si:dkeyp-117h8.2
|
si:dkeyp-117h8.2 |
| chr17_-_17948587 | 0.72 |
ENSDART00000090447
|
hhipl1
|
HHIP-like 1 |
| chr2_-_5728843 | 0.71 |
ENSDART00000014020
|
sst2
|
somatostatin 2 |
| chr2_-_56655769 | 0.70 |
ENSDART00000113589
|
gpx4b
|
glutathione peroxidase 4b |
| chr6_+_40952031 | 0.70 |
ENSDART00000189219
|
patz1
|
POZ (BTB) and AT hook containing zinc finger 1 |
| chr10_-_21953643 | 0.69 |
ENSDART00000188921
ENSDART00000193569 |
FO744833.2
|
|
| chr22_-_21392748 | 0.66 |
ENSDART00000144648
|
ankrd24
|
ankyrin repeat domain 24 |
| chr23_-_44786844 | 0.65 |
ENSDART00000148669
|
si:ch73-269m23.5
|
si:ch73-269m23.5 |
| chr2_+_30916188 | 0.64 |
ENSDART00000137012
|
myom1a
|
myomesin 1a (skelemin) |
| chr8_+_47099033 | 0.63 |
ENSDART00000142979
|
arhgef16
|
Rho guanine nucleotide exchange factor (GEF) 16 |
| chr15_-_30857350 | 0.58 |
ENSDART00000138988
|
akap1b
|
A kinase (PRKA) anchor protein 1b |
| chr18_+_32615075 | 0.58 |
ENSDART00000166937
|
CABZ01012885.1
|
|
| chr14_+_33329420 | 0.54 |
ENSDART00000171090
ENSDART00000164062 |
sowahd
|
sosondowah ankyrin repeat domain family d |
| chr2_-_44199722 | 0.52 |
ENSDART00000140633
ENSDART00000145728 |
sdhc
|
succinate dehydrogenase complex, subunit C, integral membrane protein |
| chr5_-_30984010 | 0.52 |
ENSDART00000182367
|
spns3
|
spinster homolog 3 (Drosophila) |
| chr6_-_49078702 | 0.51 |
ENSDART00000135185
|
slc5a8l
|
solute carrier family 5 (iodide transporter), member 8-like |
| chr19_-_27564980 | 0.50 |
ENSDART00000171967
|
si:dkeyp-46h3.8
|
si:dkeyp-46h3.8 |
| chr12_+_2446837 | 0.47 |
ENSDART00000112032
|
ARHGAP22
|
si:dkey-191m6.4 |
| chr18_+_45200043 | 0.46 |
ENSDART00000015786
|
large2
|
LARGE xylosyl- and glucuronyltransferase 2 |
| chr16_-_32013913 | 0.45 |
ENSDART00000030282
ENSDART00000138701 |
gstk1
|
glutathione S-transferase kappa 1 |
| chr11_+_11201096 | 0.44 |
ENSDART00000171916
ENSDART00000171521 ENSDART00000087105 ENSDART00000159603 |
myom2a
|
myomesin 2a |
| chr10_+_26747755 | 0.44 |
ENSDART00000100329
|
f9b
|
coagulation factor IXb |
Network of associatons between targets according to the STRING database.
First level regulatory network of hnf4a+hnf4b
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.9 | 7.6 | GO:0006549 | isoleucine metabolic process(GO:0006549) isoleucine biosynthetic process(GO:0009097) |
| 1.8 | 9.0 | GO:0006031 | chitin biosynthetic process(GO:0006031) glucosamine-containing compound biosynthetic process(GO:1901073) |
| 1.5 | 4.6 | GO:0010747 | regulation of plasma membrane long-chain fatty acid transport(GO:0010746) positive regulation of plasma membrane long-chain fatty acid transport(GO:0010747) plasma membrane long-chain fatty acid transport(GO:0015911) fatty acid transmembrane transport(GO:1902001) positive regulation of anion transmembrane transport(GO:1903961) regulation of fatty acid transport(GO:2000191) positive regulation of fatty acid transport(GO:2000193) |
| 1.2 | 3.5 | GO:1901223 | negative regulation of NIK/NF-kappaB signaling(GO:1901223) |
| 1.1 | 6.5 | GO:0038065 | collagen-activated tyrosine kinase receptor signaling pathway(GO:0038063) collagen-activated signaling pathway(GO:0038065) |
| 1.0 | 3.1 | GO:0010430 | fatty acid omega-oxidation(GO:0010430) |
| 0.9 | 4.7 | GO:0007191 | adenylate cyclase-activating dopamine receptor signaling pathway(GO:0007191) |
| 0.9 | 7.0 | GO:0072488 | ammonium transmembrane transport(GO:0072488) |
| 0.9 | 3.4 | GO:0006554 | lysine catabolic process(GO:0006554) L-lysine catabolic process to acetyl-CoA(GO:0019474) L-lysine catabolic process(GO:0019477) L-lysine metabolic process(GO:0046440) |
| 0.7 | 4.4 | GO:1900029 | positive regulation of ruffle assembly(GO:1900029) |
| 0.7 | 2.8 | GO:0010459 | cardiac conduction system development(GO:0003161) negative regulation of heart rate(GO:0010459) |
| 0.7 | 6.7 | GO:1990402 | embryonic liver development(GO:1990402) |
| 0.7 | 5.3 | GO:1990845 | adaptive thermogenesis(GO:1990845) |
| 0.6 | 9.3 | GO:1901642 | nucleoside transport(GO:0015858) nucleoside transmembrane transport(GO:1901642) |
| 0.6 | 8.0 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
| 0.5 | 3.4 | GO:0044539 | long-chain fatty acid import(GO:0044539) |
| 0.5 | 5.1 | GO:0036368 | cone photoresponse recovery(GO:0036368) |
| 0.4 | 1.1 | GO:0045648 | positive regulation of erythrocyte differentiation(GO:0045648) |
| 0.4 | 2.5 | GO:0032475 | otolith formation(GO:0032475) |
| 0.3 | 1.7 | GO:0002164 | larval development(GO:0002164) larval heart development(GO:0007508) angiogenesis involved in wound healing(GO:0060055) |
| 0.3 | 1.3 | GO:0030091 | protein repair(GO:0030091) |
| 0.3 | 1.5 | GO:0019483 | beta-alanine metabolic process(GO:0019482) beta-alanine biosynthetic process(GO:0019483) |
| 0.3 | 2.9 | GO:1904071 | presynaptic active zone assembly(GO:1904071) |
| 0.3 | 2.7 | GO:0033700 | phospholipid efflux(GO:0033700) |
| 0.3 | 2.8 | GO:0070782 | phosphatidylserine exposure on apoptotic cell surface(GO:0070782) |
| 0.2 | 2.6 | GO:0097090 | presynaptic membrane organization(GO:0097090) postsynaptic membrane assembly(GO:0097104) presynaptic membrane assembly(GO:0097105) |
| 0.2 | 7.5 | GO:0006826 | iron ion transport(GO:0006826) |
| 0.2 | 2.2 | GO:0060575 | intestinal epithelial cell differentiation(GO:0060575) |
| 0.2 | 3.4 | GO:0006265 | DNA topological change(GO:0006265) |
| 0.2 | 3.9 | GO:0006198 | cAMP catabolic process(GO:0006198) |
| 0.2 | 0.9 | GO:0019370 | leukotriene metabolic process(GO:0006691) leukotriene biosynthetic process(GO:0019370) |
| 0.2 | 5.1 | GO:0051923 | sulfation(GO:0051923) |
| 0.2 | 0.9 | GO:0045039 | protein import into mitochondrial inner membrane(GO:0045039) |
| 0.2 | 2.8 | GO:0050910 | detection of mechanical stimulus involved in sensory perception of sound(GO:0050910) |
| 0.2 | 2.1 | GO:0045670 | regulation of osteoclast differentiation(GO:0045670) |
| 0.2 | 2.3 | GO:0016556 | mRNA modification(GO:0016556) |
| 0.2 | 2.9 | GO:0043252 | sodium-independent organic anion transport(GO:0043252) |
| 0.1 | 1.2 | GO:0046341 | CDP-diacylglycerol biosynthetic process(GO:0016024) CDP-diacylglycerol metabolic process(GO:0046341) |
| 0.1 | 0.9 | GO:0045053 | protein retention in Golgi apparatus(GO:0045053) |
| 0.1 | 2.0 | GO:0090279 | regulation of calcium ion import(GO:0090279) |
| 0.1 | 4.2 | GO:0072348 | sulfur compound transport(GO:0072348) |
| 0.1 | 1.1 | GO:0000186 | activation of MAPKK activity(GO:0000186) |
| 0.1 | 0.5 | GO:0006121 | mitochondrial electron transport, succinate to ubiquinone(GO:0006121) |
| 0.1 | 1.0 | GO:0070262 | peptidyl-serine dephosphorylation(GO:0070262) |
| 0.1 | 0.4 | GO:0046166 | methylglyoxal biosynthetic process(GO:0019242) glyceraldehyde-3-phosphate biosynthetic process(GO:0046166) |
| 0.1 | 3.2 | GO:0070936 | protein K48-linked ubiquitination(GO:0070936) |
| 0.1 | 4.0 | GO:0030968 | endoplasmic reticulum unfolded protein response(GO:0030968) |
| 0.1 | 3.4 | GO:0035118 | embryonic pectoral fin morphogenesis(GO:0035118) |
| 0.1 | 2.1 | GO:0048168 | regulation of neuronal synaptic plasticity(GO:0048168) |
| 0.1 | 0.4 | GO:0019254 | carnitine metabolic process, CoA-linked(GO:0019254) |
| 0.1 | 1.1 | GO:0030534 | adult behavior(GO:0030534) |
| 0.1 | 5.5 | GO:0007605 | sensory perception of sound(GO:0007605) |
| 0.1 | 0.4 | GO:0055005 | ventricular cardiac myofibril assembly(GO:0055005) |
| 0.1 | 0.4 | GO:0016572 | histone phosphorylation(GO:0016572) |
| 0.1 | 1.3 | GO:0007205 | protein kinase C-activating G-protein coupled receptor signaling pathway(GO:0007205) |
| 0.1 | 3.5 | GO:0006956 | complement activation(GO:0006956) |
| 0.0 | 1.8 | GO:0030038 | contractile actin filament bundle assembly(GO:0030038) stress fiber assembly(GO:0043149) |
| 0.0 | 1.6 | GO:0032543 | mitochondrial translation(GO:0032543) |
| 0.0 | 3.6 | GO:0010951 | negative regulation of endopeptidase activity(GO:0010951) |
| 0.0 | 0.2 | GO:0048755 | branching morphogenesis of a nerve(GO:0048755) |
| 0.0 | 1.7 | GO:0032012 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.0 | 2.0 | GO:0030282 | bone mineralization(GO:0030282) |
| 0.0 | 0.6 | GO:0010738 | regulation of protein kinase A signaling(GO:0010738) |
| 0.0 | 1.5 | GO:0048546 | digestive tract morphogenesis(GO:0048546) |
| 0.0 | 0.6 | GO:0032958 | inositol phosphate biosynthetic process(GO:0032958) |
| 0.0 | 0.9 | GO:0016601 | Rac protein signal transduction(GO:0016601) |
| 0.0 | 2.3 | GO:1990573 | potassium ion import across plasma membrane(GO:1990573) |
| 0.0 | 0.5 | GO:0035269 | protein O-linked mannosylation(GO:0035269) |
| 0.0 | 2.0 | GO:0030336 | negative regulation of cell migration(GO:0030336) |
| 0.0 | 1.5 | GO:0051091 | positive regulation of sequence-specific DNA binding transcription factor activity(GO:0051091) |
| 0.0 | 2.3 | GO:0016358 | dendrite development(GO:0016358) |
| 0.0 | 1.2 | GO:0042737 | drug metabolic process(GO:0017144) drug catabolic process(GO:0042737) exogenous drug catabolic process(GO:0042738) |
| 0.0 | 2.5 | GO:0055123 | digestive system development(GO:0055123) |
| 0.0 | 10.3 | GO:0045944 | positive regulation of transcription from RNA polymerase II promoter(GO:0045944) |
| 0.0 | 1.6 | GO:0090630 | activation of GTPase activity(GO:0090630) |
| 0.0 | 8.4 | GO:0016567 | protein ubiquitination(GO:0016567) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.2 | 6.5 | GO:0005592 | collagen type XI trimer(GO:0005592) |
| 1.1 | 9.0 | GO:0030428 | cell septum(GO:0030428) |
| 0.7 | 3.0 | GO:0061689 | tricellular tight junction(GO:0061689) |
| 0.4 | 2.1 | GO:0031258 | lamellipodium membrane(GO:0031258) |
| 0.3 | 0.9 | GO:0042719 | mitochondrial intermembrane space protein transporter complex(GO:0042719) |
| 0.3 | 1.2 | GO:0043291 | RAVE complex(GO:0043291) |
| 0.3 | 2.9 | GO:0098982 | GABA-ergic synapse(GO:0098982) |
| 0.1 | 2.8 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.1 | 5.8 | GO:0005902 | microvillus(GO:0005902) |
| 0.1 | 4.4 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.1 | 0.5 | GO:0045257 | mitochondrial respiratory chain complex II, succinate dehydrogenase complex (ubiquinone)(GO:0005749) succinate dehydrogenase complex (ubiquinone)(GO:0045257) respiratory chain complex II(GO:0045273) succinate dehydrogenase complex(GO:0045281) fumarate reductase complex(GO:0045283) |
| 0.1 | 1.1 | GO:0005922 | connexon complex(GO:0005922) |
| 0.1 | 7.5 | GO:0055037 | recycling endosome(GO:0055037) |
| 0.1 | 4.7 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.1 | 5.9 | GO:0016459 | myosin complex(GO:0016459) |
| 0.1 | 1.1 | GO:0031430 | M band(GO:0031430) |
| 0.0 | 2.6 | GO:0005811 | lipid particle(GO:0005811) |
| 0.0 | 1.6 | GO:0000315 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.0 | 1.0 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.0 | 2.3 | GO:0042579 | peroxisome(GO:0005777) microbody(GO:0042579) |
| 0.0 | 1.0 | GO:0016605 | PML body(GO:0016605) |
| 0.0 | 2.3 | GO:0005604 | basement membrane(GO:0005604) |
| 0.0 | 0.4 | GO:0043209 | myelin sheath(GO:0043209) |
| 0.0 | 3.8 | GO:0048471 | perinuclear region of cytoplasm(GO:0048471) |
| 0.0 | 7.7 | GO:0031966 | mitochondrial membrane(GO:0031966) |
| 0.0 | 6.7 | GO:0009986 | cell surface(GO:0009986) |
| 0.0 | 10.6 | GO:0005789 | endoplasmic reticulum membrane(GO:0005789) |
| 0.0 | 0.1 | GO:0016272 | prefoldin complex(GO:0016272) |
| 0.0 | 93.1 | GO:0016021 | integral component of membrane(GO:0016021) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.5 | 7.6 | GO:0016841 | ammonia-lyase activity(GO:0016841) |
| 1.3 | 5.3 | GO:0017077 | oxidative phosphorylation uncoupler activity(GO:0017077) |
| 1.1 | 9.0 | GO:0004100 | chitin synthase activity(GO:0004100) |
| 1.0 | 9.3 | GO:0015250 | water channel activity(GO:0015250) |
| 0.9 | 4.7 | GO:0031852 | mu-type opioid receptor binding(GO:0031852) dopamine receptor binding(GO:0050780) corticotropin-releasing hormone receptor 1 binding(GO:0051430) |
| 0.8 | 3.1 | GO:0003958 | NADPH-hemoprotein reductase activity(GO:0003958) |
| 0.6 | 4.7 | GO:0016803 | ether hydrolase activity(GO:0016803) |
| 0.6 | 3.4 | GO:0016647 | oxidoreductase activity, acting on the CH-NH group of donors, oxygen as acceptor(GO:0016647) |
| 0.5 | 2.7 | GO:0090556 | phospholipid-translocating ATPase activity(GO:0004012) phosphatidylcholine-translocating ATPase activity(GO:0090554) phosphatidylserine-translocating ATPase activity(GO:0090556) |
| 0.5 | 3.4 | GO:0031957 | very long-chain fatty acid-CoA ligase activity(GO:0031957) |
| 0.4 | 3.3 | GO:0008048 | calcium sensitive guanylate cyclase activator activity(GO:0008048) |
| 0.4 | 4.6 | GO:0047676 | arachidonate-CoA ligase activity(GO:0047676) |
| 0.4 | 3.5 | GO:0004062 | aryl sulfotransferase activity(GO:0004062) |
| 0.3 | 1.3 | GO:0033745 | L-methionine-(R)-S-oxide reductase activity(GO:0033745) |
| 0.3 | 4.3 | GO:0003841 | 1-acylglycerol-3-phosphate O-acyltransferase activity(GO:0003841) |
| 0.3 | 7.0 | GO:0008519 | ammonium transmembrane transporter activity(GO:0008519) |
| 0.3 | 17.2 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.3 | 8.0 | GO:0016675 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors(GO:0016675) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.3 | 3.4 | GO:0003917 | DNA topoisomerase type I activity(GO:0003917) |
| 0.3 | 1.5 | GO:0030331 | estrogen receptor binding(GO:0030331) |
| 0.3 | 12.9 | GO:0015370 | solute:sodium symporter activity(GO:0015370) |
| 0.3 | 2.8 | GO:0016907 | G-protein coupled acetylcholine receptor activity(GO:0016907) |
| 0.2 | 0.9 | GO:0004464 | leukotriene-C4 synthase activity(GO:0004464) |
| 0.2 | 1.1 | GO:0055077 | gap junction hemi-channel activity(GO:0055077) |
| 0.2 | 1.2 | GO:0003857 | 3-hydroxyacyl-CoA dehydrogenase activity(GO:0003857) |
| 0.2 | 0.5 | GO:0000104 | succinate dehydrogenase activity(GO:0000104) |
| 0.2 | 3.9 | GO:0004115 | 3',5'-cyclic-AMP phosphodiesterase activity(GO:0004115) |
| 0.2 | 2.9 | GO:0098882 | structural constituent of presynaptic active zone(GO:0098882) |
| 0.2 | 2.9 | GO:0015347 | sodium-independent organic anion transmembrane transporter activity(GO:0015347) |
| 0.1 | 1.7 | GO:0005021 | vascular endothelial growth factor-activated receptor activity(GO:0005021) |
| 0.1 | 1.9 | GO:0042910 | xenobiotic transporter activity(GO:0042910) |
| 0.1 | 1.8 | GO:0001965 | G-protein alpha-subunit binding(GO:0001965) |
| 0.1 | 0.5 | GO:0015355 | secondary active monocarboxylate transmembrane transporter activity(GO:0015355) |
| 0.1 | 2.1 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.1 | 0.4 | GO:0004092 | carnitine O-acetyltransferase activity(GO:0004092) |
| 0.1 | 0.3 | GO:0047291 | lactosylceramide alpha-2,3-sialyltransferase activity(GO:0047291) |
| 0.1 | 2.8 | GO:0015248 | sterol transporter activity(GO:0015248) |
| 0.1 | 1.5 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.1 | 2.6 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 0.1 | 0.4 | GO:0004807 | triose-phosphate isomerase activity(GO:0004807) methylglyoxal synthase activity(GO:0008929) |
| 0.1 | 8.3 | GO:0016811 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amides(GO:0016811) |
| 0.1 | 2.8 | GO:0008331 | high voltage-gated calcium channel activity(GO:0008331) |
| 0.1 | 11.3 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.1 | 1.1 | GO:0070915 | lysophosphatidic acid receptor activity(GO:0070915) |
| 0.1 | 8.3 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.1 | 0.2 | GO:0030273 | melanin-concentrating hormone receptor activity(GO:0030273) |
| 0.1 | 3.4 | GO:0004683 | calmodulin-dependent protein kinase activity(GO:0004683) |
| 0.1 | 2.2 | GO:0008381 | mechanically-gated ion channel activity(GO:0008381) mechanically gated channel activity(GO:0022833) |
| 0.1 | 2.3 | GO:0005242 | inward rectifier potassium channel activity(GO:0005242) |
| 0.1 | 3.9 | GO:0016712 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced flavin or flavoprotein as one donor, and incorporation of one atom of oxygen(GO:0016712) |
| 0.1 | 5.7 | GO:0001227 | transcriptional repressor activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001227) |
| 0.1 | 1.3 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.0 | 0.4 | GO:0004332 | fructose-bisphosphate aldolase activity(GO:0004332) |
| 0.0 | 1.1 | GO:0004709 | MAP kinase kinase kinase activity(GO:0004709) |
| 0.0 | 1.3 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.0 | 2.4 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.0 | 1.7 | GO:0004869 | cysteine-type endopeptidase inhibitor activity(GO:0004869) |
| 0.0 | 0.6 | GO:0000828 | inositol hexakisphosphate kinase activity(GO:0000828) |
| 0.0 | 3.0 | GO:0051082 | unfolded protein binding(GO:0051082) |
| 0.0 | 5.9 | GO:0003774 | motor activity(GO:0003774) |
| 0.0 | 0.4 | GO:0035497 | cAMP response element binding(GO:0035497) |
| 0.0 | 2.7 | GO:0004497 | monooxygenase activity(GO:0004497) |
| 0.0 | 8.8 | GO:0061630 | ubiquitin protein ligase activity(GO:0061630) |
| 0.0 | 4.4 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.0 | 2.6 | GO:0016616 | oxidoreductase activity, acting on the CH-OH group of donors, NAD or NADP as acceptor(GO:0016616) |
| 0.0 | 0.3 | GO:0004983 | neuropeptide Y receptor activity(GO:0004983) |
| 0.0 | 0.7 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.0 | 0.4 | GO:0015035 | protein disulfide oxidoreductase activity(GO:0015035) |
| 0.0 | 0.9 | GO:0008146 | sulfotransferase activity(GO:0008146) |
| 0.0 | 2.2 | GO:0004842 | ubiquitin-protein transferase activity(GO:0004842) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 6.5 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.1 | 2.4 | PID SYNDECAN 2 PATHWAY | Syndecan-2-mediated signaling events |
| 0.1 | 2.0 | PID IL8 CXCR1 PATHWAY | IL8- and CXCR1-mediated signaling events |
| 0.1 | 2.9 | ST GA13 PATHWAY | G alpha 13 Pathway |
| 0.1 | 0.3 | ST PAC1 RECEPTOR PATHWAY | PAC1 Receptor Pathway |
| 0.0 | 1.7 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.0 | 1.1 | PID LYSOPHOSPHOLIPID PATHWAY | LPA receptor mediated events |
| 0.0 | 0.1 | PID S1P S1P1 PATHWAY | S1P1 pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 16.5 | REACTOME TRANSPORT OF VITAMINS NUCLEOSIDES AND RELATED MOLECULES | Genes involved in Transport of vitamins, nucleosides, and related molecules |
| 0.6 | 4.6 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.5 | 2.0 | REACTOME G BETA GAMMA SIGNALLING THROUGH PLC BETA | Genes involved in G beta:gamma signalling through PLC beta |
| 0.3 | 2.0 | REACTOME ALPHA LINOLENIC ACID ALA METABOLISM | Genes involved in alpha-linolenic acid (ALA) metabolism |
| 0.3 | 4.7 | REACTOME PROSTACYCLIN SIGNALLING THROUGH PROSTACYCLIN RECEPTOR | Genes involved in Prostacyclin signalling through prostacyclin receptor |
| 0.2 | 3.0 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.2 | 2.1 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GLP1 | Genes involved in Synthesis, Secretion, and Inactivation of Glucagon-like Peptide-1 (GLP-1) |
| 0.2 | 1.5 | REACTOME PYRIMIDINE CATABOLISM | Genes involved in Pyrimidine catabolism |
| 0.2 | 1.4 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.1 | 6.5 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.1 | 2.5 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.1 | 1.8 | REACTOME SEMA4D INDUCED CELL MIGRATION AND GROWTH CONE COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.1 | 3.2 | REACTOME TRIGLYCERIDE BIOSYNTHESIS | Genes involved in Triglyceride Biosynthesis |
| 0.1 | 5.8 | REACTOME RESPIRATORY ELECTRON TRANSPORT ATP SYNTHESIS BY CHEMIOSMOTIC COUPLING AND HEAT PRODUCTION BY UNCOUPLING PROTEINS | Genes involved in Respiratory electron transport, ATP synthesis by chemiosmotic coupling, and heat production by uncoupling proteins. |
| 0.0 | 1.6 | REACTOME NRAGE SIGNALS DEATH THROUGH JNK | Genes involved in NRAGE signals death through JNK |
| 0.0 | 1.6 | REACTOME CELL JUNCTION ORGANIZATION | Genes involved in Cell junction organization |
| 0.0 | 0.2 | REACTOME SEMA3A PLEXIN REPULSION SIGNALING BY INHIBITING INTEGRIN ADHESION | Genes involved in SEMA3A-Plexin repulsion signaling by inhibiting Integrin adhesion |
| 0.0 | 0.9 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
| 0.0 | 0.9 | REACTOME INTEGRATION OF ENERGY METABOLISM | Genes involved in Integration of energy metabolism |
| 0.0 | 0.4 | REACTOME RECYCLING PATHWAY OF L1 | Genes involved in Recycling pathway of L1 |
| 0.0 | 0.4 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.0 | 1.4 | REACTOME G ALPHA I SIGNALLING EVENTS | Genes involved in G alpha (i) signalling events |