Project
PRJEB1986: zebrafish developmental stages transcriptome
Navigation
Downloads
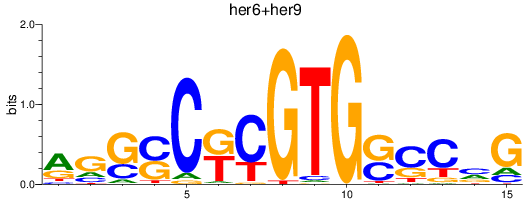
Results for her6+her9
Z-value: 0.31
Transcription factors associated with her6+her9
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
her6
|
ENSDARG00000006514 | hairy-related 6 |
|
her9
|
ENSDARG00000056438 | hairy-related 9 |
|
her6
|
ENSDARG00000111099 | hairy-related 6 |
|
her9
|
ENSDARG00000115775 | hairy-related 9 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| her6 | dr11_v1_chr6_-_36552844_36552844 | -0.29 | 2.2e-01 | Click! |
| her9 | dr11_v1_chr23_-_23401305_23401305 | -0.28 | 2.4e-01 | Click! |
Activity profile of her6+her9 motif
Sorted Z-values of her6+her9 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr19_-_6385594 | 0.58 |
ENSDART00000104950
|
atp1a3a
|
ATPase Na+/K+ transporting subunit alpha 3a |
| chr7_+_529522 | 0.51 |
ENSDART00000190811
|
nrxn2b
|
neurexin 2b |
| chr5_-_23429228 | 0.44 |
ENSDART00000049291
|
gria3a
|
glutamate receptor, ionotropic, AMPA 3a |
| chr23_+_22656477 | 0.39 |
ENSDART00000009337
ENSDART00000133322 |
eno1a
|
enolase 1a, (alpha) |
| chr20_+_39283849 | 0.32 |
ENSDART00000002481
ENSDART00000146683 |
scara3
|
scavenger receptor class A, member 3 |
| chr17_-_3986236 | 0.31 |
ENSDART00000188794
ENSDART00000160830 |
PLCB1
|
si:ch1073-140o9.2 |
| chr4_+_9669717 | 0.31 |
ENSDART00000004604
|
si:dkey-153k10.9
|
si:dkey-153k10.9 |
| chr23_+_40452157 | 0.29 |
ENSDART00000113106
ENSDART00000140136 |
soga3b
|
SOGA family member 3b |
| chr25_+_19954576 | 0.28 |
ENSDART00000149335
|
kcna1a
|
potassium voltage-gated channel, shaker-related subfamily, member 1a |
| chr14_+_15064870 | 0.28 |
ENSDART00000167075
|
nkx1.2lb
|
NK1 transcription factor related 2-like,b |
| chr1_+_31942961 | 0.28 |
ENSDART00000007522
|
anos1a
|
anosmin 1a |
| chr15_-_12319065 | 0.28 |
ENSDART00000162973
ENSDART00000170543 |
fxyd6
|
FXYD domain containing ion transport regulator 6 |
| chr14_-_32520295 | 0.27 |
ENSDART00000165463
|
atp11c
|
ATPase phospholipid transporting 11C |
| chr14_-_2189889 | 0.25 |
ENSDART00000181557
ENSDART00000106707 |
pcdh2ab9
pcdh2ab11
|
protocadherin 2 alpha b 9 protocadherin 2 alpha b 11 |
| chr22_-_10502780 | 0.25 |
ENSDART00000136961
|
ecm2
|
extracellular matrix protein 2, female organ and adipocyte specific |
| chr24_+_23791758 | 0.24 |
ENSDART00000066655
ENSDART00000146580 |
mybl1
|
v-myb avian myeloblastosis viral oncogene homolog-like 1 |
| chr23_+_44732863 | 0.24 |
ENSDART00000160044
ENSDART00000172268 |
atp1b2a
|
ATPase Na+/K+ transporting subunit beta 2a |
| chr18_-_14274803 | 0.22 |
ENSDART00000166643
|
mlycd
|
malonyl-CoA decarboxylase |
| chr20_-_5328637 | 0.20 |
ENSDART00000075974
|
ism2b
|
isthmin 2b |
| chr8_-_912821 | 0.20 |
ENSDART00000082296
|
ptger4a
|
prostaglandin E receptor 4 (subtype EP4) a |
| chr7_-_6438544 | 0.19 |
ENSDART00000173180
|
zgc:165555
|
zgc:165555 |
| chr16_+_48729947 | 0.18 |
ENSDART00000049051
|
brd2b
|
bromodomain containing 2b |
| chr22_+_19311411 | 0.18 |
ENSDART00000133234
ENSDART00000138284 |
si:dkey-21e2.16
|
si:dkey-21e2.16 |
| chr8_+_36509885 | 0.18 |
ENSDART00000109530
|
slc7a4
|
solute carrier family 7, member 4 |
| chr2_+_13069168 | 0.17 |
ENSDART00000192832
|
prkag2b
|
protein kinase, AMP-activated, gamma 2 non-catalytic subunit b |
| chr6_-_40195510 | 0.16 |
ENSDART00000156156
|
col7a1
|
collagen, type VII, alpha 1 |
| chr19_-_15397946 | 0.15 |
ENSDART00000143480
|
hivep3a
|
human immunodeficiency virus type I enhancer binding protein 3a |
| chr23_-_6765653 | 0.14 |
ENSDART00000192310
|
FP102169.1
|
|
| chr3_-_8765165 | 0.14 |
ENSDART00000191131
|
CABZ01058333.1
|
|
| chr4_+_332709 | 0.14 |
ENSDART00000067486
|
tulp4a
|
tubby like protein 4a |
| chr21_-_32680638 | 0.14 |
ENSDART00000147203
|
adamts2
|
ADAM metallopeptidase with thrombospondin type 1 motif, 2 |
| chr1_-_22756898 | 0.13 |
ENSDART00000158915
|
prom1b
|
prominin 1 b |
| chr17_-_20666616 | 0.13 |
ENSDART00000020167
|
slc16a9a
|
solute carrier family 16, member 9a |
| chr18_+_41772474 | 0.12 |
ENSDART00000097547
|
trpc6b
|
transient receptor potential cation channel, subfamily C, member 6b |
| chr10_+_29138021 | 0.11 |
ENSDART00000025227
ENSDART00000123033 ENSDART00000034242 |
picalma
|
phosphatidylinositol binding clathrin assembly protein a |
| chr2_-_58257624 | 0.11 |
ENSDART00000098940
|
foxl2b
|
forkhead box L2b |
| chr20_-_8304271 | 0.11 |
ENSDART00000179057
|
dab1a
|
Dab, reelin signal transducer, homolog 1a (Drosophila) |
| chr20_-_40766387 | 0.11 |
ENSDART00000061173
|
hsdl1
|
hydroxysteroid dehydrogenase like 1 |
| chr5_+_1965296 | 0.10 |
ENSDART00000156224
|
dhx33
|
DEAH (Asp-Glu-Ala-His) box polypeptide 33 |
| chr5_-_12560569 | 0.10 |
ENSDART00000133587
|
wsb2
|
WD repeat and SOCS box containing 2 |
| chr24_-_33291784 | 0.10 |
ENSDART00000124938
|
si:ch1073-406l10.2
|
si:ch1073-406l10.2 |
| chr18_-_494606 | 0.10 |
ENSDART00000157564
|
wtip
|
WT1 interacting protein |
| chr4_+_68562464 | 0.10 |
ENSDART00000192954
|
BX548011.4
|
|
| chr9_-_43538328 | 0.09 |
ENSDART00000140526
|
znf385b
|
zinc finger protein 385B |
| chr9_-_6624663 | 0.08 |
ENSDART00000092537
|
GPR45
|
si:dkeyp-118h3.5 |
| chr2_+_68789 | 0.07 |
ENSDART00000058569
|
cldn1
|
claudin 1 |
| chr25_+_388258 | 0.07 |
ENSDART00000166834
|
rfx7b
|
regulatory factor X7b |
| chr7_+_17938128 | 0.07 |
ENSDART00000141044
|
mta2
|
metastasis associated 1 family, member 2 |
| chr18_+_16125852 | 0.07 |
ENSDART00000061106
|
bhlhe41
|
basic helix-loop-helix family, member e41 |
| chr5_-_29122615 | 0.06 |
ENSDART00000144802
|
whrnb
|
whirlin b |
| chr5_-_29122834 | 0.06 |
ENSDART00000087197
|
whrnb
|
whirlin b |
| chr11_-_12800945 | 0.06 |
ENSDART00000191178
|
txlng
|
taxilin gamma |
| chr15_+_27387555 | 0.06 |
ENSDART00000018603
|
tbx4
|
T-box 4 |
| chr11_-_12801157 | 0.06 |
ENSDART00000103449
|
txlng
|
taxilin gamma |
| chr14_+_12110020 | 0.05 |
ENSDART00000192462
|
kctd12b
|
potassium channel tetramerisation domain containing 12b |
| chr1_+_37196106 | 0.05 |
ENSDART00000008756
ENSDART00000157503 ENSDART00000162971 ENSDART00000191004 ENSDART00000078206 ENSDART00000045111 |
dclk2a
|
doublecortin-like kinase 2a |
| chr1_-_38908197 | 0.04 |
ENSDART00000101489
|
vegfc
|
vascular endothelial growth factor c |
| chr24_+_39537565 | 0.04 |
ENSDART00000140868
ENSDART00000145358 |
si:dkey-161j23.5
|
si:dkey-161j23.5 |
| chr14_+_12109535 | 0.04 |
ENSDART00000054620
ENSDART00000121913 |
kctd12b
|
potassium channel tetramerisation domain containing 12b |
| chr12_+_19362547 | 0.03 |
ENSDART00000180148
|
gspt1
|
G1 to S phase transition 1 |
| chr9_+_9112159 | 0.03 |
ENSDART00000165932
|
sik1
|
salt-inducible kinase 1 |
| chr1_+_37195919 | 0.03 |
ENSDART00000159684
ENSDART00000172742 ENSDART00000158395 |
dclk2a
|
doublecortin-like kinase 2a |
| chr12_+_19362335 | 0.03 |
ENSDART00000041711
|
gspt1
|
G1 to S phase transition 1 |
| chr13_+_35925490 | 0.02 |
ENSDART00000046115
|
mfsd2aa
|
major facilitator superfamily domain containing 2aa |
| chr23_-_15088316 | 0.01 |
ENSDART00000082060
|
si:ch211-218g4.2
|
si:ch211-218g4.2 |
| chr3_+_431208 | 0.00 |
ENSDART00000154296
ENSDART00000048733 |
si:ch73-308m11.1
si:dkey-167k11.5
|
si:ch73-308m11.1 si:dkey-167k11.5 |
| chr14_-_41075262 | 0.00 |
ENSDART00000180518
|
drp2
|
dystrophin related protein 2 |
Network of associatons between targets according to the STRING database.
First level regulatory network of her6+her9
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | GO:0032889 | regulation of vacuole fusion, non-autophagic(GO:0032889) |
| 0.0 | 0.2 | GO:2001293 | malonyl-CoA metabolic process(GO:2001293) |
| 0.0 | 0.8 | GO:0071436 | cellular potassium ion homeostasis(GO:0030007) sodium ion export from cell(GO:0036376) sodium ion export(GO:0071436) |
| 0.0 | 0.1 | GO:0048496 | maintenance of organ identity(GO:0048496) |
| 0.0 | 0.1 | GO:0060148 | negative regulation of hippo signaling(GO:0035331) positive regulation of posttranscriptional gene silencing(GO:0060148) positive regulation of gene silencing by miRNA(GO:2000637) |
| 0.0 | 0.2 | GO:1990402 | embryonic liver development(GO:1990402) |
| 0.0 | 0.3 | GO:0021772 | olfactory bulb development(GO:0021772) olfactory lobe development(GO:0021988) |
| 0.0 | 0.1 | GO:0006972 | hyperosmotic response(GO:0006972) |
| 0.0 | 0.1 | GO:0045943 | positive regulation of transcription from RNA polymerase I promoter(GO:0045943) |
| 0.0 | 0.1 | GO:0097477 | spinal cord motor neuron migration(GO:0097476) lateral motor column neuron migration(GO:0097477) |
| 0.0 | 0.1 | GO:0002184 | cytoplasmic translational termination(GO:0002184) |
| 0.0 | 0.2 | GO:0010811 | positive regulation of cell-substrate adhesion(GO:0010811) |
| 0.0 | 0.1 | GO:0030326 | embryonic limb morphogenesis(GO:0030326) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.4 | GO:0000015 | phosphopyruvate hydratase complex(GO:0000015) |
| 0.0 | 0.2 | GO:0005782 | peroxisomal matrix(GO:0005782) microbody lumen(GO:0031907) |
| 0.0 | 0.2 | GO:0005890 | sodium:potassium-exchanging ATPase complex(GO:0005890) cation-transporting ATPase complex(GO:0090533) |
| 0.0 | 0.3 | GO:0044224 | paranode region of axon(GO:0033270) juxtaparanode region of axon(GO:0044224) |
| 0.0 | 0.1 | GO:0071914 | prominosome(GO:0071914) |
| 0.0 | 0.2 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.0 | 0.1 | GO:0002139 | stereocilia coupling link(GO:0002139) stereocilia ankle link(GO:0002141) stereocilia ankle link complex(GO:0002142) stereocilium tip(GO:0032426) |
| 0.0 | 0.1 | GO:0018444 | translation release factor complex(GO:0018444) |
| 0.0 | 0.1 | GO:0098835 | presynaptic endocytic zone(GO:0098833) presynaptic endocytic zone membrane(GO:0098835) extrinsic component of presynaptic membrane(GO:0098888) extrinsic component of presynaptic endocytic zone(GO:0098894) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.2 | GO:0070052 | collagen V binding(GO:0070052) |
| 0.1 | 0.6 | GO:0005391 | sodium:potassium-exchanging ATPase activity(GO:0005391) |
| 0.0 | 0.4 | GO:0004634 | phosphopyruvate hydratase activity(GO:0004634) |
| 0.0 | 0.4 | GO:0004971 | AMPA glutamate receptor activity(GO:0004971) |
| 0.0 | 0.0 | GO:0043185 | vascular endothelial growth factor receptor 3 binding(GO:0043185) |
| 0.0 | 0.2 | GO:0001671 | ATPase activator activity(GO:0001671) |
| 0.0 | 0.3 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | PID IL8 CXCR1 PATHWAY | IL8- and CXCR1-mediated signaling events |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.3 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |