Project
PRJEB1986: zebrafish developmental stages transcriptome
Navigation
Downloads
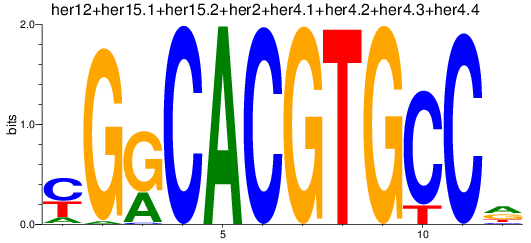
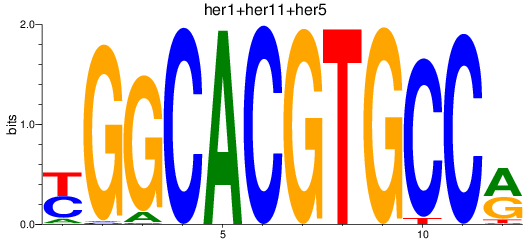
Results for her1+her11+her5_her12+her15.1+her15.2+her2+her4.1+her4.2+her4.3+her4.4
Z-value: 0.51
Transcription factors associated with her1+her11+her5_her12+her15.1+her15.2+her2+her4.1+her4.2+her4.3+her4.4
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
her11
|
ENSDARG00000002707 | hairy-related 11 |
|
her5
|
ENSDARG00000008796 | hairy-related 5 |
|
her1
|
ENSDARG00000014722 | hairy-related 1 |
|
her4.4
|
ENSDARG00000009822 | hairy-related 4, tandem duplicate 4 |
|
her12
|
ENSDARG00000032963 | hairy-related 12 |
|
her2
|
ENSDARG00000038205 | hairy-related 2 |
|
her15.2
|
ENSDARG00000054560 | hairy and enhancer of split-related 15, tandem duplicate 2 |
|
her15.1
|
ENSDARG00000054562 | hairy and enhancer of split-related 15, tandem duplicate 1 |
|
her4.2
|
ENSDARG00000056729 | hairy-related 4, tandem duplicate 2 |
|
her4.1
|
ENSDARG00000056732 | hairy-related 4, tandem duplicate 1 |
|
her4.3
|
ENSDARG00000070770 | hairy-related 4, tandem duplicate 3 |
|
her4.2
|
ENSDARG00000094426 | hairy-related 4, tandem duplicate 2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| her1 | dr11_v1_chr5_-_68795063_68795063 | -0.82 | 1.7e-05 | Click! |
| her5 | dr11_v1_chr14_+_30543008_30543008 | -0.67 | 1.6e-03 | Click! |
| her11 | dr11_v1_chr14_-_30540967_30540967 | -0.48 | 3.7e-02 | Click! |
| her4.1 | dr11_v1_chr23_-_21453614_21453614 | 0.45 | 5.2e-02 | Click! |
| her2 | dr11_v1_chr11_-_41966854_41966854 | -0.39 | 9.9e-02 | Click! |
| her4.2 | dr11_v1_chr23_+_21455152_21455154 | 0.35 | 1.5e-01 | Click! |
| her4.3 | dr11_v1_chr23_+_21459263_21459263 | 0.30 | 2.1e-01 | Click! |
| her4.4 | dr11_v1_chr23_-_21463788_21463788 | 0.30 | 2.1e-01 | Click! |
| her15.2 | dr11_v1_chr11_-_41996957_41996957 | 0.21 | 3.9e-01 | Click! |
| her15.1 | dr11_v1_chr11_+_41981959_41981959 | 0.08 | 7.3e-01 | Click! |
| her12 | dr11_v1_chr23_-_21446985_21446985 | -0.06 | 8.2e-01 | Click! |
Activity profile of her1+her11+her5_her12+her15.1+her15.2+her2+her4.1+her4.2+her4.3+her4.4 motif
Sorted Z-values of her1+her11+her5_her12+her15.1+her15.2+her2+her4.1+her4.2+her4.3+her4.4 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr19_+_31183495 | 2.27 |
ENSDART00000088618
|
meox2b
|
mesenchyme homeobox 2b |
| chr16_-_45069882 | 2.01 |
ENSDART00000058384
|
gapdhs
|
glyceraldehyde-3-phosphate dehydrogenase, spermatogenic |
| chr14_+_46313135 | 1.99 |
ENSDART00000172902
|
cryba1l1
|
crystallin, beta A1, like 1 |
| chr6_-_60147517 | 1.79 |
ENSDART00000083453
|
slc32a1
|
solute carrier family 32 (GABA vesicular transporter), member 1 |
| chr6_+_39184236 | 1.04 |
ENSDART00000156187
|
tac3b
|
tachykinin 3b |
| chr9_-_44295071 | 1.00 |
ENSDART00000011837
|
neurod1
|
neuronal differentiation 1 |
| chr14_+_46313396 | 0.86 |
ENSDART00000047525
|
cryba1l1
|
crystallin, beta A1, like 1 |
| chr23_+_21459263 | 0.67 |
ENSDART00000104209
|
her4.3
|
hairy-related 4, tandem duplicate 3 |
| chr23_-_21463788 | 0.64 |
ENSDART00000079265
|
her4.4
|
hairy-related 4, tandem duplicate 4 |
| chr20_-_27733683 | 0.50 |
ENSDART00000103317
ENSDART00000138139 |
zgc:153157
|
zgc:153157 |
| chr8_+_21146262 | 0.50 |
ENSDART00000045684
|
porcn
|
porcupine O-acyltransferase |
| chr19_+_42470396 | 0.42 |
ENSDART00000191679
|
si:dkey-166k12.1
|
si:dkey-166k12.1 |
| chr19_+_42469058 | 0.32 |
ENSDART00000076915
|
si:dkey-166k12.1
|
si:dkey-166k12.1 |
| chr3_+_23029934 | 0.28 |
ENSDART00000110343
|
nags
|
N-acetylglutamate synthase |
| chr5_-_45894802 | 0.27 |
ENSDART00000097648
|
crfb6
|
cytokine receptor family member b6 |
| chr10_-_27566481 | 0.26 |
ENSDART00000078920
|
auts2a
|
autism susceptibility candidate 2a |
| chr11_-_3959889 | 0.26 |
ENSDART00000159683
|
pbrm1
|
polybromo 1 |
| chr23_-_31969786 | 0.23 |
ENSDART00000134550
|
ormdl2
|
ORMDL sphingolipid biosynthesis regulator 2 |
| chr10_+_158590 | 0.23 |
ENSDART00000081982
|
KCNJ15
|
potassium voltage-gated channel subfamily J member 15 |
| chr7_+_32021669 | 0.21 |
ENSDART00000173976
|
mettl15
|
methyltransferase like 15 |
| chr11_-_3959477 | 0.20 |
ENSDART00000045971
|
pbrm1
|
polybromo 1 |
| chr18_+_6126506 | 0.19 |
ENSDART00000125725
|
si:ch1073-390k14.1
|
si:ch1073-390k14.1 |
| chr8_+_39760258 | 0.17 |
ENSDART00000037914
|
cox6a1
|
cytochrome c oxidase subunit VIa polypeptide 1 |
| chr12_+_33038757 | 0.16 |
ENSDART00000153146
|
rbfox3a
|
RNA binding fox-1 homolog 3a |
| chr5_-_46329880 | 0.15 |
ENSDART00000156577
|
si:ch211-130m23.5
|
si:ch211-130m23.5 |
| chr7_+_32021982 | 0.14 |
ENSDART00000173848
|
mettl15
|
methyltransferase like 15 |
| chr2_-_32386791 | 0.12 |
ENSDART00000056634
|
ubtfl
|
upstream binding transcription factor, like |
| chr9_+_45789887 | 0.11 |
ENSDART00000135202
|
si:dkey-34f9.3
|
si:dkey-34f9.3 |
| chr10_+_17026870 | 0.10 |
ENSDART00000184529
ENSDART00000157480 |
CR855996.2
|
|
| chr18_+_30370559 | 0.10 |
ENSDART00000165450
|
gse1
|
Gse1 coiled-coil protein |
| chr21_-_38730557 | 0.09 |
ENSDART00000150984
ENSDART00000111885 |
taf9
|
TAF9 RNA polymerase II, TATA box binding protein (TBP)-associated factor |
| chr18_+_30370339 | 0.09 |
ENSDART00000158871
|
gse1
|
Gse1 coiled-coil protein |
| chr3_+_32425202 | 0.09 |
ENSDART00000156464
|
prr12b
|
proline rich 12b |
| chr3_-_6441619 | 0.08 |
ENSDART00000157771
ENSDART00000166758 |
sumo2b
|
small ubiquitin-like modifier 2b |
| chr24_+_31459715 | 0.07 |
ENSDART00000181102
ENSDART00000189950 ENSDART00000192321 ENSDART00000126380 |
cnbd1
|
cyclic nucleotide binding domain containing 1 |
| chr19_-_35596207 | 0.07 |
ENSDART00000136811
|
col8a2
|
collagen, type VIII, alpha 2 |
| chr20_+_19212962 | 0.07 |
ENSDART00000063706
|
fndc4a
|
fibronectin type III domain containing 4a |
| chr19_+_42142381 | 0.04 |
ENSDART00000129915
|
kcnq4
|
potassium voltage-gated channel, KQT-like subfamily, member 4 |
| chr25_-_37284370 | 0.03 |
ENSDART00000103222
|
nudt7
|
nudix (nucleoside diphosphate linked moiety X)-type motif 7 |
| chr16_-_4026265 | 0.00 |
ENSDART00000149339
|
si:ch211-175f12.2
|
si:ch211-175f12.2 |
Network of associatons between targets according to the STRING database.
First level regulatory network of her1+her11+her5_her12+her15.1+her15.2+her2+her4.1+her4.2+her4.3+her4.4
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.0 | GO:0007217 | tachykinin receptor signaling pathway(GO:0007217) |
| 0.2 | 1.8 | GO:0098700 | neurotransmitter loading into synaptic vesicle(GO:0098700) |
| 0.2 | 0.5 | GO:0061355 | Wnt protein secretion(GO:0061355) |
| 0.1 | 0.3 | GO:0006526 | arginine biosynthetic process(GO:0006526) |
| 0.1 | 0.2 | GO:0090155 | regulation of sphingolipid biosynthetic process(GO:0090153) negative regulation of sphingolipid biosynthetic process(GO:0090155) negative regulation of ceramide biosynthetic process(GO:1900060) regulation of membrane lipid metabolic process(GO:1905038) regulation of ceramide biosynthetic process(GO:2000303) |
| 0.1 | 1.3 | GO:0048936 | peripheral nervous system neuron axonogenesis(GO:0048936) |
| 0.0 | 0.8 | GO:0048923 | posterior lateral line neuromast hair cell differentiation(GO:0048923) |
| 0.0 | 0.3 | GO:0070475 | rRNA base methylation(GO:0070475) |
| 0.0 | 2.0 | GO:0006096 | glycolytic process(GO:0006096) |
| 0.0 | 2.8 | GO:0002088 | lens development in camera-type eye(GO:0002088) |
| 0.0 | 0.2 | GO:0006123 | mitochondrial electron transport, cytochrome c to oxygen(GO:0006123) |
| 0.0 | 0.3 | GO:0021884 | forebrain neuron development(GO:0021884) |
| 0.0 | 2.3 | GO:0061053 | somite development(GO:0061053) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.8 | GO:0098563 | integral component of synaptic vesicle membrane(GO:0030285) intrinsic component of synaptic vesicle membrane(GO:0098563) |
| 0.1 | 0.2 | GO:0035339 | SPOTS complex(GO:0035339) |
| 0.1 | 0.5 | GO:0016586 | RSC complex(GO:0016586) |
| 0.0 | 0.2 | GO:0005751 | mitochondrial respiratory chain complex IV(GO:0005751) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 2.0 | GO:0043891 | glyceraldehyde-3-phosphate dehydrogenase (NAD+) (phosphorylating) activity(GO:0004365) glyceraldehyde-3-phosphate dehydrogenase (NAD(P)+) (phosphorylating) activity(GO:0043891) |
| 0.1 | 1.8 | GO:0015295 | solute:proton symporter activity(GO:0015295) |
| 0.1 | 0.3 | GO:0016434 | rRNA (cytosine) methyltransferase activity(GO:0016434) |
| 0.1 | 0.3 | GO:0034618 | arginine binding(GO:0034618) |
| 0.0 | 2.8 | GO:0005212 | structural constituent of eye lens(GO:0005212) |
| 0.0 | 0.5 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.0 | 0.5 | GO:0017147 | Wnt-protein binding(GO:0017147) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.0 | 0.2 | PID INTEGRIN A9B1 PATHWAY | Alpha9 beta1 integrin signaling events |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.8 | REACTOME GABA SYNTHESIS RELEASE REUPTAKE AND DEGRADATION | Genes involved in GABA synthesis, release, reuptake and degradation |
| 0.1 | 2.0 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.1 | 1.0 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.0 | 0.2 | REACTOME INHIBITION OF VOLTAGE GATED CA2 CHANNELS VIA GBETA GAMMA SUBUNITS | Genes involved in Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits |