Project
PRJEB1986: zebrafish developmental stages transcriptome
Navigation
Downloads
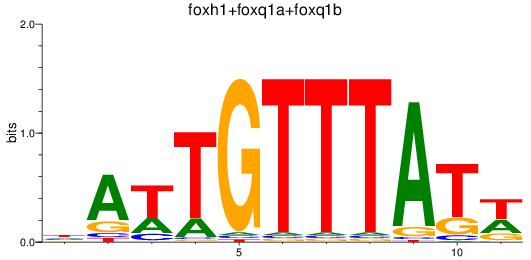
Results for foxh1+foxq1a+foxq1b
Z-value: 0.69
Transcription factors associated with foxh1+foxq1a+foxq1b
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
foxq1a
|
ENSDARG00000030896 | forkhead box Q1a |
|
foxq1b
|
ENSDARG00000055395 | forkhead box Q1b |
|
foxh1
|
ENSDARG00000055630 | forkhead box H1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| foxq1a | dr11_v1_chr2_-_722156_722170 | -0.40 | 9.1e-02 | Click! |
| foxh1 | dr11_v1_chr12_-_13729263_13729263 | -0.40 | 9.3e-02 | Click! |
| foxq1b | dr11_v1_chr20_+_26683933_26683940 | 0.05 | 8.5e-01 | Click! |
Activity profile of foxh1+foxq1a+foxq1b motif
Sorted Z-values of foxh1+foxq1a+foxq1b motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr17_+_15534815 | 2.21 |
ENSDART00000159426
|
marcksb
|
myristoylated alanine-rich protein kinase C substrate b |
| chr18_+_20034023 | 1.55 |
ENSDART00000139441
|
morf4l1
|
mortality factor 4 like 1 |
| chr22_+_15960005 | 1.55 |
ENSDART00000033617
|
stil
|
scl/tal1 interrupting locus |
| chr22_+_15960514 | 1.52 |
ENSDART00000181617
|
stil
|
scl/tal1 interrupting locus |
| chr22_+_15959844 | 1.44 |
ENSDART00000182201
|
stil
|
scl/tal1 interrupting locus |
| chr8_+_39634114 | 1.41 |
ENSDART00000144293
|
msi1
|
musashi RNA-binding protein 1 |
| chr4_+_21741228 | 1.41 |
ENSDART00000112035
ENSDART00000127664 |
myf5
|
myogenic factor 5 |
| chr14_+_34495216 | 1.38 |
ENSDART00000147756
|
wnt8a
|
wingless-type MMTV integration site family, member 8a |
| chr25_+_18556588 | 1.35 |
ENSDART00000073726
|
cav2
|
caveolin 2 |
| chr13_+_25286538 | 1.30 |
ENSDART00000180282
|
chuk
|
conserved helix-loop-helix ubiquitous kinase |
| chr1_+_26676758 | 1.26 |
ENSDART00000152299
|
si:dkey-25o16.4
|
si:dkey-25o16.4 |
| chr10_+_22775253 | 1.23 |
ENSDART00000190141
|
tmem88a
|
transmembrane protein 88 a |
| chr1_+_25696798 | 1.23 |
ENSDART00000054228
|
lrata
|
lecithin retinol acyltransferase a |
| chr16_+_35536075 | 1.22 |
ENSDART00000183618
|
cited4b
|
Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 4b |
| chr20_+_33294428 | 1.20 |
ENSDART00000024104
|
mycn
|
MYCN proto-oncogene, bHLH transcription factor |
| chr3_-_37148594 | 1.19 |
ENSDART00000140855
|
mlx
|
MLX, MAX dimerization protein |
| chr17_-_42213285 | 1.18 |
ENSDART00000140549
|
nkx2.2a
|
NK2 homeobox 2a |
| chr13_-_25198025 | 1.18 |
ENSDART00000159585
ENSDART00000144227 |
adka
|
adenosine kinase a |
| chr1_+_19538299 | 1.18 |
ENSDART00000109416
|
smc2
|
structural maintenance of chromosomes 2 |
| chr5_+_25762271 | 1.18 |
ENSDART00000181323
|
tmem2
|
transmembrane protein 2 |
| chr16_+_35535375 | 1.16 |
ENSDART00000171675
|
cited4b
|
Cbp/p300-interacting transactivator, with Glu/Asp-rich carboxy-terminal domain, 4b |
| chr2_+_36701322 | 1.13 |
ENSDART00000002510
|
golim4b
|
golgi integral membrane protein 4b |
| chr16_-_42461263 | 1.11 |
ENSDART00000109259
|
smarcc1a
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily c, member 1a |
| chr5_-_45958838 | 1.10 |
ENSDART00000135072
|
poc5
|
POC5 centriolar protein homolog (Chlamydomonas) |
| chr21_-_44772738 | 1.09 |
ENSDART00000026178
|
kif4
|
kinesin family member 4 |
| chr16_-_30564319 | 1.05 |
ENSDART00000145087
|
lmna
|
lamin A |
| chr7_-_26076970 | 1.05 |
ENSDART00000101120
|
zgc:92664
|
zgc:92664 |
| chr14_+_28486213 | 1.04 |
ENSDART00000161852
|
stag2b
|
stromal antigen 2b |
| chr7_-_51102479 | 1.04 |
ENSDART00000174023
|
col4a6
|
collagen, type IV, alpha 6 |
| chr13_+_23157053 | 1.03 |
ENSDART00000162359
|
sorbs1
|
sorbin and SH3 domain containing 1 |
| chr17_-_42218652 | 1.03 |
ENSDART00000081396
ENSDART00000190007 |
nkx2.2a
|
NK2 homeobox 2a |
| chr7_-_54679595 | 1.02 |
ENSDART00000165320
|
ccnd1
|
cyclin D1 |
| chr1_+_9290103 | 0.97 |
ENSDART00000055009
|
uncx4.1
|
Unc4.1 homeobox (C. elegans) |
| chr20_-_25748407 | 0.96 |
ENSDART00000063152
|
chst14
|
carbohydrate (N-acetylgalactosamine 4-0) sulfotransferase 14 |
| chr9_+_34397843 | 0.96 |
ENSDART00000146314
|
med14
|
mediator complex subunit 14 |
| chr19_+_25971000 | 0.93 |
ENSDART00000089836
|
jarid2b
|
jumonji, AT rich interactive domain 2b |
| chr2_+_7132292 | 0.92 |
ENSDART00000153404
ENSDART00000012119 |
zgc:110366
|
zgc:110366 |
| chr16_+_11818126 | 0.89 |
ENSDART00000145727
|
cxcr3.2
|
chemokine (C-X-C motif) receptor 3, tandem duplicate 2 |
| chr9_+_34950942 | 0.88 |
ENSDART00000077800
|
tfdp1a
|
transcription factor Dp-1, a |
| chr11_+_37612214 | 0.87 |
ENSDART00000172899
ENSDART00000077496 |
hp1bp3
|
heterochromatin protein 1, binding protein 3 |
| chr17_-_42213822 | 0.87 |
ENSDART00000187904
ENSDART00000180029 |
nkx2.2a
|
NK2 homeobox 2a |
| chr11_+_25257022 | 0.86 |
ENSDART00000156052
|
tp53inp2
|
tumor protein p53 inducible nuclear protein 2 |
| chr1_-_45580787 | 0.85 |
ENSDART00000135089
|
atf7ip
|
activating transcription factor 7 interacting protein |
| chr5_-_67365750 | 0.85 |
ENSDART00000062359
|
unga
|
uracil DNA glycosylase a |
| chr18_-_45761868 | 0.85 |
ENSDART00000025423
|
cstf3
|
cleavage stimulation factor, 3' pre-RNA, subunit 3 |
| chr19_+_46240171 | 0.84 |
ENSDART00000162785
|
mapk15
|
mitogen-activated protein kinase 15 |
| chr16_-_30563129 | 0.84 |
ENSDART00000191716
|
lmna
|
lamin A |
| chr9_+_34397516 | 0.83 |
ENSDART00000011304
ENSDART00000192973 |
med14
|
mediator complex subunit 14 |
| chr1_+_14658801 | 0.83 |
ENSDART00000192194
|
CR847936.1
|
|
| chr14_-_24410673 | 0.82 |
ENSDART00000125923
|
cxcl14
|
chemokine (C-X-C motif) ligand 14 |
| chr18_-_44316920 | 0.81 |
ENSDART00000098599
|
si:ch211-151h10.2
|
si:ch211-151h10.2 |
| chr17_+_24590177 | 0.80 |
ENSDART00000092941
|
rlf
|
rearranged L-myc fusion |
| chr5_+_49744713 | 0.79 |
ENSDART00000133384
|
nr2f1a
|
nuclear receptor subfamily 2, group F, member 1a |
| chr19_+_19777437 | 0.78 |
ENSDART00000170662
|
hoxa3a
|
homeobox A3a |
| chr11_-_25257045 | 0.77 |
ENSDART00000130477
|
snai1a
|
snail family zinc finger 1a |
| chr17_+_34244345 | 0.77 |
ENSDART00000006058
|
eif2s1a
|
eukaryotic translation initiation factor 2, subunit 1 alpha a |
| chr5_+_32835219 | 0.75 |
ENSDART00000140832
ENSDART00000186055 |
si:ch211-208h16.4
|
si:ch211-208h16.4 |
| chr3_+_22442445 | 0.75 |
ENSDART00000190921
|
wnk4b
|
WNK lysine deficient protein kinase 4b |
| chr10_-_6587066 | 0.75 |
ENSDART00000171833
|
chd1
|
chromodomain helicase DNA binding protein 1 |
| chr1_-_23370395 | 0.75 |
ENSDART00000143014
ENSDART00000126785 ENSDART00000159138 |
pds5a
|
PDS5 cohesin associated factor A |
| chr16_-_39267185 | 0.70 |
ENSDART00000058550
ENSDART00000133642 |
gpd1l
|
glycerol-3-phosphate dehydrogenase 1 like |
| chr8_-_11324143 | 0.70 |
ENSDART00000008215
|
pip5k1bb
|
phosphatidylinositol-4-phosphate 5-kinase, type I, beta b |
| chr3_-_43356082 | 0.69 |
ENSDART00000171213
|
uncx
|
UNC homeobox |
| chr11_+_22134540 | 0.68 |
ENSDART00000190502
|
MDFI
|
si:dkey-91m3.1 |
| chr2_-_37956768 | 0.67 |
ENSDART00000034595
|
cbln10
|
cerebellin 10 |
| chr16_-_42186093 | 0.67 |
ENSDART00000076030
|
fbl
|
fibrillarin |
| chr2_+_15128418 | 0.67 |
ENSDART00000141921
|
arhgap29b
|
Rho GTPase activating protein 29b |
| chr24_+_26329018 | 0.67 |
ENSDART00000145752
|
mynn
|
myoneurin |
| chr7_+_9290929 | 0.65 |
ENSDART00000128530
|
snrpa1
|
small nuclear ribonucleoprotein polypeptide A' |
| chr7_+_22801465 | 0.64 |
ENSDART00000052862
ENSDART00000173633 |
rbm4.1
|
RNA binding motif protein 4.1 |
| chr7_+_24494299 | 0.64 |
ENSDART00000087568
|
nelfa
|
negative elongation factor complex member A |
| chr11_+_24046179 | 0.64 |
ENSDART00000006703
|
maf1
|
MAF1 homolog, negative regulator of RNA polymerase III |
| chr9_-_27398369 | 0.64 |
ENSDART00000186499
|
tex30
|
testis expressed 30 |
| chr11_-_25257595 | 0.63 |
ENSDART00000123567
|
snai1a
|
snail family zinc finger 1a |
| chr10_+_28428222 | 0.62 |
ENSDART00000135003
|
si:ch211-222e20.4
|
si:ch211-222e20.4 |
| chr8_+_32719930 | 0.62 |
ENSDART00000145362
|
hmcn2
|
hemicentin 2 |
| chr12_-_34887943 | 0.62 |
ENSDART00000027379
|
bicral
|
BRD4 interacting chromatin remodeling complex associated protein like |
| chr19_+_5143128 | 0.62 |
ENSDART00000138011
ENSDART00000108541 |
si:dkey-89b17.4
|
si:dkey-89b17.4 |
| chr5_-_32074534 | 0.61 |
ENSDART00000112342
ENSDART00000135855 |
arpc5lb
|
actin related protein 2/3 complex, subunit 5-like, b |
| chr1_+_32054159 | 0.60 |
ENSDART00000181442
|
sts
|
steroid sulfatase (microsomal), isozyme S |
| chr22_-_29906764 | 0.60 |
ENSDART00000019786
|
smc3
|
structural maintenance of chromosomes 3 |
| chr16_-_21692024 | 0.59 |
ENSDART00000123597
|
si:ch211-154o6.2
|
si:ch211-154o6.2 |
| chr3_+_40289418 | 0.59 |
ENSDART00000017304
|
cpsf4
|
cleavage and polyadenylation specific factor 4 |
| chr17_-_18888959 | 0.59 |
ENSDART00000080029
|
ak7b
|
adenylate kinase 7b |
| chr8_-_20243389 | 0.59 |
ENSDART00000184904
|
acer1
|
alkaline ceramidase 1 |
| chr23_+_28077953 | 0.58 |
ENSDART00000186122
ENSDART00000111570 |
slc26a10
|
solute carrier family 26, member 10 |
| chr19_+_20177887 | 0.57 |
ENSDART00000008595
|
tra2a
|
transformer 2 alpha homolog |
| chr4_+_23125689 | 0.57 |
ENSDART00000077854
|
mdm2
|
MDM2 oncogene, E3 ubiquitin protein ligase |
| chr9_-_53666031 | 0.57 |
ENSDART00000126314
|
pcdh8
|
protocadherin 8 |
| chr3_-_39695856 | 0.56 |
ENSDART00000148247
|
b9d1
|
B9 protein domain 1 |
| chr5_-_41531629 | 0.56 |
ENSDART00000051082
|
akr1a1a
|
aldo-keto reductase family 1, member A1a (aldehyde reductase) |
| chr5_-_36328688 | 0.56 |
ENSDART00000011399
|
efnb1
|
ephrin-B1 |
| chr13_+_18507592 | 0.55 |
ENSDART00000142622
|
si:ch211-198a12.6
|
si:ch211-198a12.6 |
| chr24_+_6107901 | 0.55 |
ENSDART00000156419
|
si:ch211-37e10.2
|
si:ch211-37e10.2 |
| chr3_+_23710839 | 0.55 |
ENSDART00000151584
|
hoxb4a
|
homeobox B4a |
| chr4_+_11723852 | 0.53 |
ENSDART00000028820
|
mkln1
|
muskelin 1, intracellular mediator containing kelch motifs |
| chr18_+_19648275 | 0.53 |
ENSDART00000100569
|
smad6b
|
SMAD family member 6b |
| chr7_-_18547420 | 0.53 |
ENSDART00000173969
|
rgs12a
|
regulator of G protein signaling 12a |
| chr15_+_16185118 | 0.50 |
ENSDART00000147900
|
mrm3a
|
mitochondrial rRNA methyltransferase 3a |
| chr11_+_7580079 | 0.50 |
ENSDART00000091550
ENSDART00000193223 ENSDART00000193386 |
adgrl2a
|
adhesion G protein-coupled receptor L2a |
| chr8_-_15150131 | 0.50 |
ENSDART00000138253
|
bcar3
|
BCAR3, NSP family adaptor protein |
| chr19_+_19756425 | 0.49 |
ENSDART00000167606
|
hoxa3a
|
homeobox A3a |
| chr12_+_27156943 | 0.49 |
ENSDART00000153030
ENSDART00000001737 |
skap1
|
src kinase associated phosphoprotein 1 |
| chr18_-_5850683 | 0.49 |
ENSDART00000082087
|
nip7
|
NIP7, nucleolar pre-rRNA processing protein |
| chr7_+_30240791 | 0.48 |
ENSDART00000109243
|
sema4bb
|
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4Bb |
| chr25_+_2776865 | 0.47 |
ENSDART00000156567
|
neo1b
|
neogenin 1b |
| chr6_-_19341184 | 0.47 |
ENSDART00000168236
ENSDART00000167674 |
mif4gda
|
MIF4G domain containing a |
| chr3_-_39696066 | 0.47 |
ENSDART00000015393
|
b9d1
|
B9 protein domain 1 |
| chr7_+_38808027 | 0.46 |
ENSDART00000052323
|
harbi1
|
harbinger transposase derived 1 |
| chr15_-_7337537 | 0.46 |
ENSDART00000161613
|
SLC7A1 (1 of many)
|
high affinity cationic amino acid transporter 1 |
| chr17_+_44441042 | 0.46 |
ENSDART00000142123
|
ap5m1
|
adaptor-related protein complex 5, mu 1 subunit |
| chr9_-_30494727 | 0.46 |
ENSDART00000148729
|
si:dkey-229b18.3
|
si:dkey-229b18.3 |
| chr19_+_43004408 | 0.45 |
ENSDART00000038230
|
snrnp40
|
small nuclear ribonucleoprotein 40 (U5) |
| chr11_+_2172335 | 0.45 |
ENSDART00000170593
|
hoxc12b
|
homeobox C12b |
| chr20_+_37300296 | 0.45 |
ENSDART00000180789
|
vta1
|
vesicle (multivesicular body) trafficking 1 |
| chr3_+_40284598 | 0.44 |
ENSDART00000009411
|
bud31
|
BUD31 homolog (S. cerevisiae) |
| chr16_-_9675982 | 0.44 |
ENSDART00000113724
|
mal2
|
mal, T cell differentiation protein 2 (gene/pseudogene) |
| chr16_+_9580699 | 0.44 |
ENSDART00000165565
|
taf2
|
TAF2 RNA polymerase II, TATA box binding protein (TBP)-associated factor |
| chr5_+_29726428 | 0.44 |
ENSDART00000143183
|
ddx31
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 31 |
| chr9_-_21067971 | 0.44 |
ENSDART00000004333
|
tbx15
|
T-box 15 |
| chr18_+_30421528 | 0.43 |
ENSDART00000140908
|
gse1
|
Gse1 coiled-coil protein |
| chr19_-_46018152 | 0.42 |
ENSDART00000159206
|
krit1
|
KRIT1, ankyrin repeat containing |
| chr5_+_34623107 | 0.42 |
ENSDART00000184126
|
enc1
|
ectodermal-neural cortex 1 |
| chr14_+_34966598 | 0.42 |
ENSDART00000004550
|
rnf145a
|
ring finger protein 145a |
| chr8_+_47726919 | 0.42 |
ENSDART00000160166
|
si:ch211-243p7.3
|
si:ch211-243p7.3 |
| chr16_+_25116827 | 0.41 |
ENSDART00000163244
|
si:ch211-261d7.6
|
si:ch211-261d7.6 |
| chr23_+_38159715 | 0.41 |
ENSDART00000137969
|
zgc:112994
|
zgc:112994 |
| chr2_-_10631767 | 0.41 |
ENSDART00000190033
|
mtf2
|
metal response element binding transcription factor 2 |
| chr5_-_28767573 | 0.40 |
ENSDART00000158299
ENSDART00000043466 |
traf2a
|
Tnf receptor-associated factor 2a |
| chr2_+_8649293 | 0.39 |
ENSDART00000081442
ENSDART00000186024 |
fubp1
|
far upstream element (FUSE) binding protein 1 |
| chr8_+_11425048 | 0.39 |
ENSDART00000018739
|
tjp2b
|
tight junction protein 2b (zona occludens 2) |
| chr18_+_40354998 | 0.39 |
ENSDART00000098791
ENSDART00000049171 |
sema6dl
|
sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6D, like |
| chr17_-_12764360 | 0.37 |
ENSDART00000003418
|
brms1la
|
breast cancer metastasis-suppressor 1-like a |
| chr8_+_48484455 | 0.37 |
ENSDART00000122737
|
PRDM16
|
si:ch211-263k4.2 |
| chr15_-_36727462 | 0.37 |
ENSDART00000085971
|
nphs1
|
nephrosis 1, congenital, Finnish type (nephrin) |
| chr20_-_16156419 | 0.37 |
ENSDART00000037420
|
ralgps2
|
Ral GEF with PH domain and SH3 binding motif 2 |
| chr9_+_31282161 | 0.36 |
ENSDART00000010774
|
zic2a
|
zic family member 2 (odd-paired homolog, Drosophila), a |
| chr5_-_63509581 | 0.36 |
ENSDART00000097325
|
c5
|
complement component 5 |
| chr14_-_12391506 | 0.36 |
ENSDART00000080864
|
magt1
|
magnesium transporter 1 |
| chr1_+_35790082 | 0.36 |
ENSDART00000085051
|
hhip
|
hedgehog interacting protein |
| chr15_-_37875601 | 0.36 |
ENSDART00000122439
|
si:dkey-238d18.4
|
si:dkey-238d18.4 |
| chr19_+_32553874 | 0.36 |
ENSDART00000078197
|
heyl
|
hes-related family bHLH transcription factor with YRPW motif-like |
| chr7_+_30254652 | 0.35 |
ENSDART00000173711
|
sema4bb
|
sema domain, immunoglobulin domain (Ig), transmembrane domain (TM) and short cytoplasmic domain, (semaphorin) 4Bb |
| chr24_-_26854032 | 0.35 |
ENSDART00000087991
|
fndc3bb
|
fibronectin type III domain containing 3Bb |
| chr16_+_44906324 | 0.34 |
ENSDART00000074960
|
cd22
|
cd22 molecule |
| chr6_+_54680730 | 0.34 |
ENSDART00000074602
|
smpd2b
|
sphingomyelin phosphodiesterase 2b, neutral membrane (neutral sphingomyelinase) |
| chr18_+_25225524 | 0.33 |
ENSDART00000055563
|
si:dkeyp-59c12.1
|
si:dkeyp-59c12.1 |
| chr24_+_37338169 | 0.33 |
ENSDART00000141771
|
gnptg
|
N-acetylglucosamine-1-phosphate transferase, gamma subunit |
| chr17_+_26803470 | 0.33 |
ENSDART00000023470
|
pgrmc2
|
progesterone receptor membrane component 2 |
| chr7_+_29080684 | 0.32 |
ENSDART00000173709
ENSDART00000173576 |
acd
|
ACD, shelterin complex subunit and telomerase recruitment factor |
| chr3_+_34234029 | 0.32 |
ENSDART00000044859
|
znf207a
|
zinc finger protein 207, a |
| chr21_+_17542473 | 0.32 |
ENSDART00000005750
ENSDART00000141326 |
stom
|
stomatin |
| chr13_-_3155243 | 0.32 |
ENSDART00000139183
ENSDART00000050934 |
pkdcca
|
protein kinase domain containing, cytoplasmic a |
| chr14_-_15990361 | 0.32 |
ENSDART00000168075
|
trim105
|
tripartite motif containing 105 |
| chr15_-_7337148 | 0.32 |
ENSDART00000182568
|
SLC7A1 (1 of many)
|
high affinity cationic amino acid transporter 1 |
| chr23_+_25354856 | 0.31 |
ENSDART00000109023
ENSDART00000147440 |
fmnl3
|
formin-like 3 |
| chr7_-_19999152 | 0.31 |
ENSDART00000173881
ENSDART00000100798 |
trip6
|
thyroid hormone receptor interactor 6 |
| chr8_-_44223473 | 0.31 |
ENSDART00000098525
|
stx2b
|
syntaxin 2b |
| chr21_-_13225402 | 0.31 |
ENSDART00000080347
|
wdr34
|
WD repeat domain 34 |
| chr23_+_2191773 | 0.31 |
ENSDART00000190876
ENSDART00000126768 |
cc2d2a
|
coiled-coil and C2 domain containing 2A |
| chr1_+_46194333 | 0.31 |
ENSDART00000010894
|
sox1b
|
SRY (sex determining region Y)-box 1b |
| chr15_+_23550890 | 0.30 |
ENSDART00000009796
ENSDART00000152720 |
MARK4
|
si:dkey-31m14.7 |
| chr4_+_2637947 | 0.30 |
ENSDART00000130623
|
dus4l
|
dihydrouridine synthase 4-like (S. cerevisiae) |
| chr13_+_39182099 | 0.29 |
ENSDART00000131434
|
fam135a
|
family with sequence similarity 135, member A |
| chr3_-_42086577 | 0.29 |
ENSDART00000083111
ENSDART00000187312 |
ttyh3a
|
tweety family member 3a |
| chr11_+_27347076 | 0.29 |
ENSDART00000173383
|
fbln2
|
fibulin 2 |
| chr9_+_6802641 | 0.29 |
ENSDART00000187278
|
FO203514.1
|
|
| chr7_+_24153070 | 0.28 |
ENSDART00000076735
|
lrp10
|
low density lipoprotein receptor-related protein 10 |
| chr3_+_18062871 | 0.28 |
ENSDART00000112384
|
ankrd40
|
ankyrin repeat domain 40 |
| chr15_+_21712328 | 0.28 |
ENSDART00000192553
|
NKAPD1
|
zgc:162339 |
| chr9_+_11293830 | 0.28 |
ENSDART00000144440
|
wnt6b
|
wingless-type MMTV integration site family, member 6b |
| chr3_-_50139860 | 0.28 |
ENSDART00000101563
|
btr02
|
bloodthirsty-related gene family, member 2 |
| chr1_+_19083501 | 0.26 |
ENSDART00000002886
ENSDART00000131948 |
exosc9
|
exosome component 9 |
| chr16_+_24733741 | 0.26 |
ENSDART00000155217
|
si:dkey-79d12.4
|
si:dkey-79d12.4 |
| chr12_-_34716037 | 0.26 |
ENSDART00000152876
|
bahcc1b
|
BAH domain and coiled-coil containing 1b |
| chr19_+_33476557 | 0.26 |
ENSDART00000181800
|
triqk
|
triple QxxK/R motif containing |
| chr24_+_26328787 | 0.25 |
ENSDART00000003884
|
mynn
|
myoneurin |
| chr7_+_52761841 | 0.25 |
ENSDART00000111444
|
ppip5k1a
|
diphosphoinositol pentakisphosphate kinase 1a |
| chr14_-_40389699 | 0.24 |
ENSDART00000181581
ENSDART00000173398 |
pcdh19
|
protocadherin 19 |
| chr21_-_38153824 | 0.23 |
ENSDART00000151226
|
klf5l
|
Kruppel-like factor 5 like |
| chr18_-_34156765 | 0.23 |
ENSDART00000059400
|
c18h3orf33
|
c18h3orf33 homolog (H. sapiens) |
| chr6_+_33881372 | 0.23 |
ENSDART00000165710
|
gpbp1l1
|
GC-rich promoter binding protein 1-like 1 |
| chr4_+_25181572 | 0.22 |
ENSDART00000078529
ENSDART00000136643 |
kin
|
Kin17 DNA and RNA binding protein |
| chr18_+_6039141 | 0.22 |
ENSDART00000138972
|
si:ch73-386h18.1
|
si:ch73-386h18.1 |
| chr23_+_36118738 | 0.22 |
ENSDART00000139319
|
hoxc5a
|
homeobox C5a |
| chr11_+_24348425 | 0.21 |
ENSDART00000089747
|
nfs1
|
NFS1 cysteine desulfurase |
| chr10_-_43117607 | 0.21 |
ENSDART00000148293
ENSDART00000089965 |
tmem167a
|
transmembrane protein 167A |
| chr9_+_8894788 | 0.21 |
ENSDART00000132068
|
naxd
|
NAD(P)HX dehydratase |
| chr1_+_24076243 | 0.21 |
ENSDART00000014608
|
mab21l2
|
mab-21-like 2 |
| chr10_+_43117661 | 0.20 |
ENSDART00000024644
ENSDART00000186932 |
xrcc4
|
X-ray repair complementing defective repair in Chinese hamster cells 4 |
| chr2_-_44282796 | 0.20 |
ENSDART00000163040
ENSDART00000166923 ENSDART00000056372 ENSDART00000109251 ENSDART00000132682 |
mpz
|
myelin protein zero |
| chr25_+_18711804 | 0.20 |
ENSDART00000011149
|
fam185a
|
family with sequence similarity 185, member A |
| chr15_-_19334448 | 0.20 |
ENSDART00000062576
|
thyn1
|
thymocyte nuclear protein 1 |
| chr6_-_8466717 | 0.19 |
ENSDART00000151577
ENSDART00000151800 ENSDART00000151227 |
si:dkey-217d24.6
|
si:dkey-217d24.6 |
| chr5_-_37997774 | 0.19 |
ENSDART00000139616
ENSDART00000167694 |
si:dkey-111e8.1
|
si:dkey-111e8.1 |
| chr22_+_15624371 | 0.19 |
ENSDART00000124868
|
lpl
|
lipoprotein lipase |
| chr11_-_16215143 | 0.19 |
ENSDART00000027014
|
rab7
|
RAB7, member RAS oncogene family |
| chr7_+_52950449 | 0.19 |
ENSDART00000187559
|
cdkn2aip
|
CDKN2A interacting protein |
| chr17_+_33453689 | 0.17 |
ENSDART00000156894
|
rin3
|
Ras and Rab interactor 3 |
| chr15_+_42285643 | 0.17 |
ENSDART00000152731
|
scaf4b
|
SR-related CTD-associated factor 4b |
Network of associatons between targets according to the STRING database.
First level regulatory network of foxh1+foxq1a+foxq1b
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 3.1 | GO:0021530 | spinal cord oligodendrocyte cell fate specification(GO:0021530) |
| 0.5 | 4.5 | GO:0071539 | protein localization to centrosome(GO:0071539) |
| 0.5 | 1.4 | GO:0002074 | extraocular skeletal muscle development(GO:0002074) |
| 0.5 | 1.4 | GO:0072111 | spinal cord anterior/posterior patterning(GO:0021512) cell proliferation involved in pronephros development(GO:0039015) cell proliferation involved in kidney development(GO:0072111) |
| 0.4 | 0.4 | GO:0001886 | endothelial cell morphogenesis(GO:0001886) |
| 0.4 | 1.2 | GO:1901295 | negative regulation of striated muscle cell differentiation(GO:0051154) negative regulation of cardiac muscle tissue development(GO:0055026) cardiac muscle cell fate commitment(GO:0060923) canonical Wnt signaling pathway involved in cardiac muscle cell fate commitment(GO:0061317) regulation of canonical Wnt signaling pathway involved in cardiac muscle cell fate commitment(GO:1901295) negative regulation of canonical Wnt signaling pathway involved in cardiac muscle cell fate commitment(GO:1901296) negative regulation of cardiocyte differentiation(GO:1905208) negative regulation of cardiac muscle cell differentiation(GO:2000726) |
| 0.4 | 1.6 | GO:0043968 | histone H2A acetylation(GO:0043968) |
| 0.3 | 1.9 | GO:0090342 | regulation of cell aging(GO:0090342) |
| 0.3 | 0.9 | GO:0051574 | positive regulation of histone H3-K9 methylation(GO:0051574) |
| 0.3 | 0.9 | GO:0002522 | leukocyte migration involved in immune response(GO:0002522) |
| 0.3 | 1.3 | GO:0007252 | I-kappaB phosphorylation(GO:0007252) |
| 0.3 | 1.3 | GO:0051988 | regulation of attachment of spindle microtubules to kinetochore(GO:0051988) |
| 0.2 | 1.0 | GO:0031571 | mitotic G1 DNA damage checkpoint(GO:0031571) |
| 0.2 | 1.2 | GO:0044209 | AMP salvage(GO:0044209) |
| 0.2 | 1.4 | GO:0001937 | negative regulation of endothelial cell proliferation(GO:0001937) |
| 0.2 | 2.2 | GO:0003382 | epithelial cell morphogenesis(GO:0003382) |
| 0.2 | 1.3 | GO:0060017 | parathyroid gland development(GO:0060017) |
| 0.2 | 0.7 | GO:0046168 | NADH oxidation(GO:0006116) glycerol-3-phosphate catabolic process(GO:0046168) |
| 0.2 | 1.0 | GO:0038065 | collagen-activated tyrosine kinase receptor signaling pathway(GO:0038063) collagen-activated signaling pathway(GO:0038065) |
| 0.2 | 0.9 | GO:0097510 | base-excision repair, AP site formation via deaminated base removal(GO:0097510) |
| 0.2 | 0.3 | GO:0032211 | negative regulation of telomere maintenance via telomerase(GO:0032211) negative regulation of telomere maintenance via telomere lengthening(GO:1904357) |
| 0.2 | 1.0 | GO:0050655 | dermatan sulfate proteoglycan metabolic process(GO:0050655) |
| 0.2 | 0.8 | GO:0003096 | renal sodium ion transport(GO:0003096) renal sodium ion absorption(GO:0070294) |
| 0.1 | 0.6 | GO:1904036 | negative regulation of epithelial cell apoptotic process(GO:1904036) |
| 0.1 | 1.2 | GO:0042572 | retinol metabolic process(GO:0042572) |
| 0.1 | 0.6 | GO:0090497 | mesenchymal cell migration(GO:0090497) |
| 0.1 | 0.3 | GO:0071042 | U1 snRNA 3'-end processing(GO:0034473) U5 snRNA 3'-end processing(GO:0034476) nuclear polyadenylation-dependent mRNA catabolic process(GO:0071042) polyadenylation-dependent mRNA catabolic process(GO:0071047) |
| 0.1 | 0.8 | GO:0055011 | atrial cardiac muscle cell differentiation(GO:0055011) |
| 0.1 | 0.4 | GO:0045056 | transcytosis(GO:0045056) |
| 0.1 | 0.1 | GO:0010718 | positive regulation of epithelial to mesenchymal transition(GO:0010718) |
| 0.1 | 0.6 | GO:0003404 | optic vesicle morphogenesis(GO:0003404) |
| 0.1 | 0.7 | GO:0031126 | snoRNA 3'-end processing(GO:0031126) |
| 0.1 | 0.9 | GO:0070828 | heterochromatin organization(GO:0070828) |
| 0.1 | 1.4 | GO:0008078 | mesodermal cell migration(GO:0008078) |
| 0.1 | 1.8 | GO:0019827 | stem cell population maintenance(GO:0019827) |
| 0.1 | 0.6 | GO:0098787 | mRNA cleavage involved in mRNA processing(GO:0098787) pre-mRNA cleavage required for polyadenylation(GO:0098789) |
| 0.1 | 0.9 | GO:0031937 | methylation-dependent chromatin silencing(GO:0006346) positive regulation of chromatin silencing(GO:0031937) regulation of methylation-dependent chromatin silencing(GO:0090308) positive regulation of methylation-dependent chromatin silencing(GO:0090309) |
| 0.1 | 0.4 | GO:0021846 | cell proliferation in forebrain(GO:0021846) |
| 0.1 | 0.4 | GO:0048635 | negative regulation of muscle organ development(GO:0048635) |
| 0.1 | 0.6 | GO:0046512 | diol biosynthetic process(GO:0034312) sphingosine biosynthetic process(GO:0046512) |
| 0.1 | 0.2 | GO:0030238 | male sex determination(GO:0030238) |
| 0.1 | 1.3 | GO:0007064 | mitotic sister chromatid cohesion(GO:0007064) |
| 0.0 | 0.6 | GO:0034244 | negative regulation of DNA-templated transcription, elongation(GO:0032785) negative regulation of transcription elongation from RNA polymerase II promoter(GO:0034244) |
| 0.0 | 0.3 | GO:0030728 | ovulation(GO:0030728) |
| 0.0 | 0.2 | GO:2000382 | positive regulation of mesoderm development(GO:2000382) |
| 0.0 | 0.2 | GO:0051103 | DNA ligation(GO:0006266) immunoglobulin V(D)J recombination(GO:0033152) DNA ligation involved in DNA repair(GO:0051103) |
| 0.0 | 0.3 | GO:1901224 | positive regulation of NIK/NF-kappaB signaling(GO:1901224) |
| 0.0 | 0.1 | GO:0065001 | specification of axis polarity(GO:0065001) |
| 0.0 | 1.1 | GO:0007062 | sister chromatid cohesion(GO:0007062) |
| 0.0 | 0.1 | GO:0048210 | Golgi vesicle fusion to target membrane(GO:0048210) |
| 0.0 | 0.1 | GO:0034405 | response to fluid shear stress(GO:0034405) |
| 0.0 | 1.1 | GO:0007632 | visual behavior(GO:0007632) |
| 0.0 | 0.3 | GO:0008608 | attachment of spindle microtubules to kinetochore(GO:0008608) |
| 0.0 | 1.2 | GO:0030261 | chromosome condensation(GO:0030261) |
| 0.0 | 0.6 | GO:0048026 | positive regulation of mRNA splicing, via spliceosome(GO:0048026) |
| 0.0 | 0.9 | GO:0010508 | positive regulation of autophagy(GO:0010508) |
| 0.0 | 0.4 | GO:0033209 | tumor necrosis factor-mediated signaling pathway(GO:0033209) |
| 0.0 | 0.2 | GO:0000455 | enzyme-directed rRNA pseudouridine synthesis(GO:0000455) |
| 0.0 | 0.2 | GO:0061303 | cornea development in camera-type eye(GO:0061303) |
| 0.0 | 0.1 | GO:0019388 | galactose catabolic process(GO:0019388) galactose catabolic process via UDP-galactose(GO:0033499) |
| 0.0 | 0.5 | GO:0033138 | positive regulation of peptidyl-serine phosphorylation(GO:0033138) |
| 0.0 | 0.1 | GO:1901162 | serotonin biosynthetic process(GO:0042427) primary amino compound biosynthetic process(GO:1901162) |
| 0.0 | 0.4 | GO:0015693 | magnesium ion transport(GO:0015693) |
| 0.0 | 0.2 | GO:0006020 | inositol metabolic process(GO:0006020) |
| 0.0 | 0.4 | GO:0032511 | late endosome to vacuole transport via multivesicular body sorting pathway(GO:0032511) |
| 0.0 | 1.1 | GO:0043044 | ATP-dependent chromatin remodeling(GO:0043044) |
| 0.0 | 0.2 | GO:0001682 | tRNA 5'-leader removal(GO:0001682) |
| 0.0 | 0.5 | GO:0006446 | regulation of translational initiation(GO:0006446) |
| 0.0 | 0.4 | GO:0090557 | establishment of endothelial intestinal barrier(GO:0090557) |
| 0.0 | 1.1 | GO:0007052 | mitotic spindle organization(GO:0007052) |
| 0.0 | 0.3 | GO:0006684 | sphingomyelin metabolic process(GO:0006684) |
| 0.0 | 0.5 | GO:0030513 | positive regulation of BMP signaling pathway(GO:0030513) |
| 0.0 | 0.6 | GO:0034314 | Arp2/3 complex-mediated actin nucleation(GO:0034314) |
| 0.0 | 0.9 | GO:1902668 | negative regulation of axon extension involved in axon guidance(GO:0048843) negative regulation of axon guidance(GO:1902668) |
| 0.0 | 0.1 | GO:0036072 | intramembranous ossification(GO:0001957) direct ossification(GO:0036072) |
| 0.0 | 0.1 | GO:0060855 | venous endothelial cell migration involved in lymph vessel development(GO:0060855) |
| 0.0 | 0.4 | GO:0007340 | acrosome reaction(GO:0007340) |
| 0.0 | 0.2 | GO:0019433 | triglyceride catabolic process(GO:0019433) |
| 0.0 | 0.8 | GO:0031123 | RNA 3'-end processing(GO:0031123) |
| 0.0 | 0.1 | GO:1902915 | negative regulation of histone ubiquitination(GO:0033183) histone H2A K63-linked ubiquitination(GO:0070535) regulation of protein K63-linked ubiquitination(GO:1900044) negative regulation of protein K63-linked ubiquitination(GO:1900045) regulation of histone H2A K63-linked ubiquitination(GO:1901314) negative regulation of histone H2A K63-linked ubiquitination(GO:1901315) regulation of protein polyubiquitination(GO:1902914) negative regulation of protein polyubiquitination(GO:1902915) |
| 0.0 | 0.4 | GO:0003094 | glomerular filtration(GO:0003094) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.4 | GO:0097189 | apoptotic body(GO:0097189) |
| 0.3 | 0.8 | GO:0043614 | multi-eIF complex(GO:0043614) |
| 0.3 | 1.8 | GO:0070847 | core mediator complex(GO:0070847) |
| 0.2 | 1.3 | GO:0008385 | IkappaB kinase complex(GO:0008385) |
| 0.2 | 1.0 | GO:0016589 | NURF complex(GO:0016589) |
| 0.2 | 1.0 | GO:0098651 | collagen type IV trimer(GO:0005587) network-forming collagen trimer(GO:0098642) collagen network(GO:0098645) basement membrane collagen trimer(GO:0098651) |
| 0.1 | 1.6 | GO:0016580 | Sin3 complex(GO:0016580) |
| 0.1 | 0.7 | GO:0009331 | glycerol-3-phosphate dehydrogenase complex(GO:0009331) |
| 0.1 | 0.7 | GO:0031428 | box C/D snoRNP complex(GO:0031428) |
| 0.1 | 1.3 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.1 | 1.4 | GO:0032391 | photoreceptor connecting cilium(GO:0032391) |
| 0.1 | 0.6 | GO:0032021 | NELF complex(GO:0032021) |
| 0.1 | 1.1 | GO:0071564 | npBAF complex(GO:0071564) |
| 0.1 | 1.6 | GO:0008278 | cohesin complex(GO:0008278) |
| 0.1 | 0.2 | GO:0005958 | DNA-dependent protein kinase-DNA ligase 4 complex(GO:0005958) |
| 0.1 | 1.0 | GO:0036038 | MKS complex(GO:0036038) |
| 0.1 | 4.6 | GO:0005814 | centriole(GO:0005814) |
| 0.0 | 0.1 | GO:0031466 | Cul5-RING ubiquitin ligase complex(GO:0031466) |
| 0.0 | 0.2 | GO:0005655 | nucleolar ribonuclease P complex(GO:0005655) |
| 0.0 | 1.7 | GO:0044853 | caveola(GO:0005901) plasma membrane raft(GO:0044853) |
| 0.0 | 2.5 | GO:0032432 | actin filament bundle(GO:0032432) |
| 0.0 | 0.4 | GO:0070822 | Sin3-type complex(GO:0070822) |
| 0.0 | 0.6 | GO:0005885 | Arp2/3 protein complex(GO:0005885) |
| 0.0 | 1.0 | GO:0017053 | transcriptional repressor complex(GO:0017053) |
| 0.0 | 0.6 | GO:0005686 | U2 snRNP(GO:0005686) |
| 0.0 | 0.6 | GO:0016514 | SWI/SNF complex(GO:0016514) |
| 0.0 | 0.1 | GO:0070724 | BMP receptor complex(GO:0070724) |
| 0.0 | 0.4 | GO:0008250 | oligosaccharyltransferase complex(GO:0008250) |
| 0.0 | 0.3 | GO:0000177 | nuclear exosome (RNase complex)(GO:0000176) cytoplasmic exosome (RNase complex)(GO:0000177) |
| 0.0 | 0.4 | GO:0005669 | transcription factor TFIID complex(GO:0005669) |
| 0.0 | 0.3 | GO:0070187 | telosome(GO:0070187) |
| 0.0 | 0.5 | GO:0030687 | preribosome, large subunit precursor(GO:0030687) |
| 0.0 | 0.4 | GO:0005771 | multivesicular body(GO:0005771) |
| 0.0 | 1.9 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.0 | 0.4 | GO:0048787 | presynaptic active zone membrane(GO:0048787) |
| 0.0 | 0.6 | GO:0016605 | PML body(GO:0016605) |
| 0.0 | 0.2 | GO:0042627 | chylomicron(GO:0042627) |
| 0.0 | 0.1 | GO:0030688 | preribosome, small subunit precursor(GO:0030688) |
| 0.0 | 0.5 | GO:0030119 | AP-type membrane coat adaptor complex(GO:0030119) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.2 | GO:0047173 | phosphatidylcholine-retinol O-acyltransferase activity(GO:0047173) |
| 0.4 | 1.4 | GO:0051059 | NF-kappaB binding(GO:0051059) |
| 0.3 | 1.3 | GO:0008384 | IkappaB kinase activity(GO:0008384) |
| 0.3 | 1.5 | GO:0004032 | alditol:NADP+ 1-oxidoreductase activity(GO:0004032) |
| 0.3 | 0.9 | GO:0043739 | G/U mismatch-specific uracil-DNA glycosylase activity(GO:0043739) |
| 0.2 | 1.2 | GO:0004001 | adenosine kinase activity(GO:0004001) |
| 0.2 | 0.7 | GO:0004367 | glycerol-3-phosphate dehydrogenase [NAD+] activity(GO:0004367) |
| 0.2 | 1.0 | GO:0008158 | hedgehog receptor activity(GO:0008158) |
| 0.1 | 0.6 | GO:0030620 | U2 snRNA binding(GO:0030620) |
| 0.1 | 0.6 | GO:0004127 | cytidylate kinase activity(GO:0004127) |
| 0.1 | 0.6 | GO:0017040 | ceramidase activity(GO:0017040) |
| 0.1 | 0.9 | GO:0034647 | histone demethylase activity (H3-trimethyl-K4 specific)(GO:0034647) |
| 0.1 | 0.2 | GO:0047453 | ATP-dependent NAD(P)H-hydrate dehydratase activity(GO:0047453) ADP-dependent NAD(P)H-hydrate dehydratase activity(GO:0052855) |
| 0.1 | 0.2 | GO:0034185 | apolipoprotein binding(GO:0034185) |
| 0.1 | 0.2 | GO:0033857 | inositol heptakisphosphate kinase activity(GO:0000829) diphosphoinositol-pentakisphosphate kinase activity(GO:0033857) |
| 0.1 | 0.5 | GO:0008494 | translation activator activity(GO:0008494) |
| 0.1 | 0.3 | GO:0017150 | tRNA dihydrouridine synthase activity(GO:0017150) |
| 0.0 | 0.8 | GO:0019870 | chloride channel inhibitor activity(GO:0019869) potassium channel inhibitor activity(GO:0019870) |
| 0.0 | 1.2 | GO:0030165 | PDZ domain binding(GO:0030165) |
| 0.0 | 1.7 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.0 | 0.7 | GO:0008649 | rRNA methyltransferase activity(GO:0008649) |
| 0.0 | 0.3 | GO:0072320 | volume-sensitive chloride channel activity(GO:0072320) |
| 0.0 | 0.4 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.0 | 0.6 | GO:0004065 | arylsulfatase activity(GO:0004065) |
| 0.0 | 0.1 | GO:0004510 | tryptophan 5-monooxygenase activity(GO:0004510) |
| 0.0 | 0.6 | GO:0046875 | ephrin receptor binding(GO:0046875) |
| 0.0 | 0.3 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.0 | 0.2 | GO:0070700 | BMP receptor binding(GO:0070700) |
| 0.0 | 0.4 | GO:0016251 | obsolete general RNA polymerase II transcription factor activity(GO:0016251) |
| 0.0 | 0.1 | GO:0030504 | inorganic diphosphate transmembrane transporter activity(GO:0030504) |
| 0.0 | 0.7 | GO:0016307 | phosphatidylinositol phosphate kinase activity(GO:0016307) |
| 0.0 | 0.7 | GO:0019957 | C-C chemokine binding(GO:0019957) |
| 0.0 | 0.4 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.0 | 0.8 | GO:0043022 | ribosome binding(GO:0043022) |
| 0.0 | 0.2 | GO:0016783 | sulfurtransferase activity(GO:0016783) |
| 0.0 | 0.2 | GO:0015924 | mannosyl-oligosaccharide 1,2-alpha-mannosidase activity(GO:0004571) mannosyl-oligosaccharide mannosidase activity(GO:0015924) |
| 0.0 | 0.2 | GO:1904047 | S-adenosyl-L-methionine binding(GO:1904047) |
| 0.0 | 0.9 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.0 | 0.8 | GO:0008009 | chemokine activity(GO:0008009) |
| 0.0 | 2.2 | GO:0003713 | transcription coactivator activity(GO:0003713) |
| 0.0 | 0.2 | GO:0004526 | ribonuclease P activity(GO:0004526) |
| 0.0 | 0.1 | GO:0004962 | endothelin receptor activity(GO:0004962) |
| 0.0 | 0.1 | GO:0019784 | NEDD8-specific protease activity(GO:0019784) |
| 0.0 | 0.4 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.0 | 0.4 | GO:0005164 | tumor necrosis factor receptor binding(GO:0005164) |
| 0.0 | 0.1 | GO:0005025 | transforming growth factor beta receptor activity, type I(GO:0005025) |
| 0.0 | 0.1 | GO:0070569 | uridylyltransferase activity(GO:0070569) |
| 0.0 | 0.5 | GO:0008173 | RNA methyltransferase activity(GO:0008173) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 4.9 | PID HEDGEHOG 2PATHWAY | Signaling events mediated by the Hedgehog family |
| 0.1 | 1.0 | PID P38 GAMMA DELTA PATHWAY | Signaling mediated by p38-gamma and p38-delta |
| 0.1 | 1.4 | PID SYNDECAN 2 PATHWAY | Syndecan-2-mediated signaling events |
| 0.1 | 1.3 | PID AVB3 OPN PATHWAY | Osteopontin-mediated events |
| 0.0 | 1.9 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.0 | 0.6 | PID EPHRINB REV PATHWAY | Ephrin B reverse signaling |
| 0.0 | 1.4 | PID ATM PATHWAY | ATM pathway |
| 0.0 | 1.0 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
| 0.0 | 1.2 | PID AURORA B PATHWAY | Aurora B signaling |
| 0.0 | 1.0 | SIG INSULIN RECEPTOR PATHWAY IN CARDIAC MYOCYTES | Genes related to the insulin receptor pathway |
| 0.0 | 0.4 | PID NEPHRIN NEPH1 PATHWAY | Nephrin/Neph1 signaling in the kidney podocyte |
| 0.0 | 1.4 | WNT SIGNALING | Genes related to Wnt-mediated signal transduction |
| 0.0 | 0.5 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.0 | 0.3 | SIG BCR SIGNALING PATHWAY | Members of the BCR signaling pathway |
| 0.0 | 0.1 | PID ALK1 PATHWAY | ALK1 signaling events |
| 0.0 | 0.3 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.3 | REACTOME IKK COMPLEX RECRUITMENT MEDIATED BY RIP1 | Genes involved in IKK complex recruitment mediated by RIP1 |
| 0.1 | 2.8 | REACTOME MEIOTIC SYNAPSIS | Genes involved in Meiotic Synapsis |
| 0.0 | 0.6 | REACTOME THE ACTIVATION OF ARYLSULFATASES | Genes involved in The activation of arylsulfatases |
| 0.0 | 1.0 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.0 | 0.6 | REACTOME AKT PHOSPHORYLATES TARGETS IN THE CYTOSOL | Genes involved in AKT phosphorylates targets in the cytosol |
| 0.0 | 1.0 | REACTOME PRE NOTCH TRANSCRIPTION AND TRANSLATION | Genes involved in Pre-NOTCH Transcription and Translation |
| 0.0 | 0.8 | REACTOME PROCESSING OF INTRONLESS PRE MRNAS | Genes involved in Processing of Intronless Pre-mRNAs |
| 0.0 | 1.4 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.0 | 0.2 | REACTOME INTEGRATION OF PROVIRUS | Genes involved in Integration of provirus |
| 0.0 | 0.7 | REACTOME SYNTHESIS OF PA | Genes involved in Synthesis of PA |
| 0.0 | 2.0 | REACTOME TRANSCRIPTIONAL REGULATION OF WHITE ADIPOCYTE DIFFERENTIATION | Genes involved in Transcriptional Regulation of White Adipocyte Differentiation |
| 0.0 | 0.3 | REACTOME MRNA DECAY BY 3 TO 5 EXORIBONUCLEASE | Genes involved in mRNA Decay by 3' to 5' Exoribonuclease |
| 0.0 | 0.3 | REACTOME TRAFFICKING AND PROCESSING OF ENDOSOMAL TLR | Genes involved in Trafficking and processing of endosomal TLR |
| 0.0 | 0.9 | REACTOME CHONDROITIN SULFATE DERMATAN SULFATE METABOLISM | Genes involved in Chondroitin sulfate/dermatan sulfate metabolism |
| 0.0 | 1.0 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.0 | 0.6 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.0 | 0.4 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.0 | 0.4 | REACTOME ENDOSOMAL SORTING COMPLEX REQUIRED FOR TRANSPORT ESCRT | Genes involved in Endosomal Sorting Complex Required For Transport (ESCRT) |
| 0.0 | 0.1 | REACTOME ACTIVATION OF CHAPERONE GENES BY ATF6 ALPHA | Genes involved in Activation of Chaperone Genes by ATF6-alpha |
| 0.0 | 0.4 | REACTOME NOTCH1 INTRACELLULAR DOMAIN REGULATES TRANSCRIPTION | Genes involved in NOTCH1 Intracellular Domain Regulates Transcription |
| 0.0 | 0.4 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 0.0 | 0.5 | REACTOME MRNA SPLICING MINOR PATHWAY | Genes involved in mRNA Splicing - Minor Pathway |
| 0.0 | 0.6 | REACTOME MRNA SPLICING | Genes involved in mRNA Splicing |