Project
PRJEB1986: zebrafish developmental stages transcriptome
Navigation
Downloads
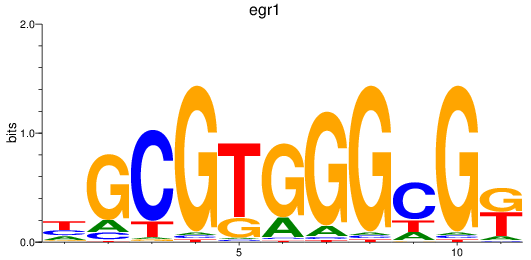
Results for egr1
Z-value: 0.95
Transcription factors associated with egr1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
egr1
|
ENSDARG00000037421 | early growth response 1 |
|
egr1
|
ENSDARG00000115563 | early growth response 1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| egr1 | dr11_v1_chr14_-_21336385_21336385 | 0.76 | 1.8e-04 | Click! |
Activity profile of egr1 motif
Sorted Z-values of egr1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr25_-_207214 | 4.09 |
ENSDART00000193448
|
FP236318.3
|
|
| chr17_+_27456804 | 3.75 |
ENSDART00000017756
ENSDART00000181461 ENSDART00000180178 |
ctsl.1
|
cathepsin L.1 |
| chr2_-_22535 | 3.75 |
ENSDART00000157877
|
CABZ01092282.1
|
|
| chr3_-_49504023 | 2.78 |
ENSDART00000168108
|
prkacaa
|
protein kinase, cAMP-dependent, catalytic, alpha, genome duplicate a |
| chr9_-_54001502 | 2.58 |
ENSDART00000085253
|
mid1
|
midline 1 |
| chr14_+_15155684 | 2.51 |
ENSDART00000167966
|
zgc:158852
|
zgc:158852 |
| chr19_-_703898 | 2.28 |
ENSDART00000181096
ENSDART00000121462 |
slc6a19a.2
|
solute carrier family 6 (neutral amino acid transporter), member 19a, tandem duplicate 2 |
| chr6_-_35446110 | 2.25 |
ENSDART00000058773
|
rgs16
|
regulator of G protein signaling 16 |
| chr24_+_24923166 | 2.22 |
ENSDART00000065288
|
pcyt1ba
|
phosphate cytidylyltransferase 1, choline, beta a |
| chr6_-_15604157 | 2.19 |
ENSDART00000141597
|
lrrfip1b
|
leucine rich repeat (in FLII) interacting protein 1b |
| chr19_+_342094 | 2.19 |
ENSDART00000151013
ENSDART00000187622 |
ensaa
|
endosulfine alpha a |
| chr1_-_7917062 | 2.08 |
ENSDART00000177068
|
mmd2b
|
monocyte to macrophage differentiation-associated 2b |
| chr3_-_61185746 | 2.05 |
ENSDART00000028219
|
pvalb4
|
parvalbumin 4 |
| chr3_+_16762483 | 1.97 |
ENSDART00000132732
|
tmem86b
|
transmembrane protein 86B |
| chr20_-_45661049 | 1.95 |
ENSDART00000124582
ENSDART00000131251 |
napbb
|
N-ethylmaleimide-sensitive factor attachment protein, beta b |
| chr6_-_15604417 | 1.95 |
ENSDART00000157817
|
lrrfip1b
|
leucine rich repeat (in FLII) interacting protein 1b |
| chr13_-_49666615 | 1.94 |
ENSDART00000148083
|
tomm20a
|
translocase of outer mitochondrial membrane 20 |
| chr11_+_2855430 | 1.94 |
ENSDART00000172837
|
kif21b
|
kinesin family member 21B |
| chr6_+_21740672 | 1.90 |
ENSDART00000193734
|
lhfpl4a
|
lipoma HMGIC fusion partner-like 4a |
| chr19_-_6193448 | 1.88 |
ENSDART00000151405
|
erf
|
Ets2 repressor factor |
| chr3_+_19299309 | 1.87 |
ENSDART00000046297
ENSDART00000146955 |
ldlra
|
low density lipoprotein receptor a |
| chr23_+_44665230 | 1.84 |
ENSDART00000144958
|
camta2
|
calmodulin binding transcription activator 2 |
| chr6_+_2271559 | 1.80 |
ENSDART00000165223
|
pbx1b
|
pre-B-cell leukemia homeobox 1b |
| chr2_+_47582488 | 1.74 |
ENSDART00000149967
|
scg2b
|
secretogranin II (chromogranin C), b |
| chr16_+_52105227 | 1.71 |
ENSDART00000150025
ENSDART00000097863 |
MAN1C1
|
si:ch73-373m9.1 |
| chr25_-_37501371 | 1.68 |
ENSDART00000160498
|
CABZ01088346.1
|
|
| chr2_-_24603325 | 1.62 |
ENSDART00000113356
|
crtc1a
|
CREB regulated transcription coactivator 1a |
| chr15_+_3284416 | 1.61 |
ENSDART00000187665
ENSDART00000171723 |
foxo1a
|
forkhead box O1 a |
| chr6_+_34511886 | 1.58 |
ENSDART00000179450
|
lrp8
|
low density lipoprotein receptor-related protein 8, apolipoprotein e receptor |
| chr9_+_51784922 | 1.57 |
ENSDART00000125850
|
cd302
|
CD302 molecule |
| chr17_+_15983247 | 1.56 |
ENSDART00000154275
|
clmn
|
calmin |
| chr13_+_9432501 | 1.55 |
ENSDART00000058064
|
zgc:123321
|
zgc:123321 |
| chr21_-_280769 | 1.52 |
ENSDART00000157753
|
plgrkt
|
plasminogen receptor, C-terminal lysine transmembrane protein |
| chr10_+_23553533 | 1.50 |
ENSDART00000159654
|
CR847844.2
|
|
| chr9_-_30363770 | 1.49 |
ENSDART00000147030
|
sytl5
|
synaptotagmin-like 5 |
| chr2_-_54639964 | 1.49 |
ENSDART00000100103
|
acss2l
|
acyl-CoA synthetase short chain family member 2 like |
| chr15_-_12319065 | 1.48 |
ENSDART00000162973
ENSDART00000170543 |
fxyd6
|
FXYD domain containing ion transport regulator 6 |
| chr18_+_402048 | 1.46 |
ENSDART00000166345
|
gpib
|
glucose-6-phosphate isomerase b |
| chr17_+_46387086 | 1.45 |
ENSDART00000157079
|
si:dkey-206p8.1
|
si:dkey-206p8.1 |
| chr21_+_28958471 | 1.43 |
ENSDART00000144331
ENSDART00000005929 |
ppp3ca
|
protein phosphatase 3, catalytic subunit, alpha isozyme |
| chr13_-_21739142 | 1.39 |
ENSDART00000078460
|
si:dkey-191g9.5
|
si:dkey-191g9.5 |
| chr25_+_34984333 | 1.36 |
ENSDART00000154760
|
ccdc136b
|
coiled-coil domain containing 136b |
| chr1_+_54673846 | 1.35 |
ENSDART00000145018
|
gprc5bb
|
G protein-coupled receptor, class C, group 5, member Bb |
| chr2_+_47582681 | 1.34 |
ENSDART00000187579
|
scg2b
|
secretogranin II (chromogranin C), b |
| chr1_-_8917902 | 1.33 |
ENSDART00000137900
|
grin2ab
|
glutamate receptor, ionotropic, N-methyl D-aspartate 2A, b |
| chr8_+_24854600 | 1.30 |
ENSDART00000156570
|
slc6a17
|
solute carrier family 6 (neutral amino acid transporter), member 17 |
| chr24_+_26379441 | 1.24 |
ENSDART00000137786
|
si:ch211-230g15.5
|
si:ch211-230g15.5 |
| chr9_+_45227028 | 1.23 |
ENSDART00000185579
|
adarb1b
|
adenosine deaminase, RNA-specific, B1b |
| chr5_-_21524251 | 1.18 |
ENSDART00000176793
ENSDART00000040184 |
tenm1
|
teneurin transmembrane protein 1 |
| chr4_-_9586713 | 1.17 |
ENSDART00000145613
|
shank3b
|
SH3 and multiple ankyrin repeat domains 3b |
| chr11_-_37880492 | 1.13 |
ENSDART00000102868
|
etnk2
|
ethanolamine kinase 2 |
| chr15_-_19250543 | 1.12 |
ENSDART00000092705
ENSDART00000138895 |
igsf9ba
|
immunoglobulin superfamily, member 9Ba |
| chr1_-_59422880 | 1.10 |
ENSDART00000167244
|
si:ch211-188p14.2
|
si:ch211-188p14.2 |
| chr11_+_77526 | 1.09 |
ENSDART00000193521
|
CABZ01072242.1
|
|
| chr16_-_22022456 | 1.07 |
ENSDART00000140175
|
si:dkey-71b5.3
|
si:dkey-71b5.3 |
| chr14_-_21218891 | 1.06 |
ENSDART00000158294
|
ppp2r2cb
|
protein phosphatase 2, regulatory subunit B, gamma b |
| chr23_+_37323962 | 1.05 |
ENSDART00000102881
|
fam43b
|
family with sequence similarity 43, member B |
| chr12_-_48943467 | 1.04 |
ENSDART00000191829
|
CABZ01092907.1
|
|
| chr20_-_54462551 | 1.03 |
ENSDART00000171769
ENSDART00000169692 |
evlb
|
Enah/Vasp-like b |
| chr14_-_7888748 | 1.03 |
ENSDART00000166293
|
ppp3cb
|
protein phosphatase 3, catalytic subunit, beta isozyme |
| chr14_+_2243 | 0.99 |
ENSDART00000191193
|
CYTL1
|
cytokine like 1 |
| chr25_+_21177326 | 0.99 |
ENSDART00000149142
|
erc1a
|
ELKS/RAB6-interacting/CAST family member 1a |
| chr10_+_252425 | 0.98 |
ENSDART00000059478
|
lrrc32
|
leucine rich repeat containing 32 |
| chr20_-_915234 | 0.98 |
ENSDART00000164816
|
cnr1
|
cannabinoid receptor 1 |
| chr6_+_34512313 | 0.97 |
ENSDART00000102554
|
lrp8
|
low density lipoprotein receptor-related protein 8, apolipoprotein e receptor |
| chr9_-_23990416 | 0.97 |
ENSDART00000113176
|
col6a3
|
collagen, type VI, alpha 3 |
| chr25_+_29688671 | 0.97 |
ENSDART00000073478
|
brd1b
|
bromodomain containing 1b |
| chr6_-_25181012 | 0.96 |
ENSDART00000161585
|
lrrc8db
|
leucine rich repeat containing 8 VRAC subunit Db |
| chr20_+_32523576 | 0.96 |
ENSDART00000147319
|
scml4
|
Scm polycomb group protein like 4 |
| chr2_+_38924975 | 0.95 |
ENSDART00000109219
|
rem2
|
RAS (RAD and GEM)-like GTP binding 2 |
| chr2_-_48966431 | 0.94 |
ENSDART00000147948
|
kcnj9
|
potassium inwardly-rectifying channel, subfamily J, member 9 |
| chr17_+_15983557 | 0.94 |
ENSDART00000190806
|
clmn
|
calmin |
| chr2_-_155270 | 0.93 |
ENSDART00000131177
|
adcy1b
|
adenylate cyclase 1b |
| chr25_+_21176937 | 0.93 |
ENSDART00000163751
ENSDART00000188930 ENSDART00000187155 |
erc1a
|
ELKS/RAB6-interacting/CAST family member 1a |
| chr25_+_35706493 | 0.92 |
ENSDART00000176741
|
kif21a
|
kinesin family member 21A |
| chr14_-_1565317 | 0.92 |
ENSDART00000169496
|
CABZ01064248.1
|
|
| chr16_+_10918252 | 0.92 |
ENSDART00000172949
|
pou2f2a
|
POU class 2 homeobox 2a |
| chr8_-_39903803 | 0.89 |
ENSDART00000012391
|
cabp1a
|
calcium binding protein 1a |
| chr18_+_15106518 | 0.89 |
ENSDART00000168639
|
cry1ab
|
cryptochrome circadian clock 1ab |
| chr19_-_2115040 | 0.88 |
ENSDART00000020497
|
snx13
|
sorting nexin 13 |
| chr23_-_1348933 | 0.88 |
ENSDART00000168981
|
CABZ01078120.1
|
|
| chr23_-_270847 | 0.87 |
ENSDART00000191867
|
anks1aa
|
ankyrin repeat and sterile alpha motif domain containing 1Aa |
| chr23_+_20422661 | 0.86 |
ENSDART00000144047
ENSDART00000104336 |
tnnc2
|
troponin C type 2 (fast) |
| chr10_+_11282883 | 0.86 |
ENSDART00000135355
|
si:ch211-126i22.5
|
si:ch211-126i22.5 |
| chr11_+_45461419 | 0.86 |
ENSDART00000173424
|
sos1
|
son of sevenless homolog 1 (Drosophila) |
| chr3_-_5228137 | 0.82 |
ENSDART00000137105
|
myh9b
|
myosin, heavy chain 9b, non-muscle |
| chr17_-_14726824 | 0.79 |
ENSDART00000162947
|
si:ch73-305o9.3
|
si:ch73-305o9.3 |
| chr19_+_58954 | 0.78 |
ENSDART00000162379
|
col14a1b
|
collagen, type XIV, alpha 1b |
| chr21_+_11415224 | 0.78 |
ENSDART00000049036
|
zgc:92275
|
zgc:92275 |
| chr13_+_44495063 | 0.76 |
ENSDART00000169591
|
CU694264.1
|
|
| chr4_+_2619132 | 0.76 |
ENSDART00000128807
|
gpr22a
|
G protein-coupled receptor 22a |
| chr10_+_23548674 | 0.76 |
ENSDART00000079686
|
zmp:0000001103
|
zmp:0000001103 |
| chr21_-_24865454 | 0.76 |
ENSDART00000142907
|
igsf9bb
|
immunoglobulin superfamily, member 9Bb |
| chr12_+_47448558 | 0.75 |
ENSDART00000185689
ENSDART00000105328 |
fmn2b
|
formin 2b |
| chr9_-_41401564 | 0.74 |
ENSDART00000059628
|
nab1b
|
NGFI-A binding protein 1b (EGR1 binding protein 1) |
| chr23_+_14269311 | 0.72 |
ENSDART00000170414
|
CR354537.1
|
|
| chr21_-_43398122 | 0.72 |
ENSDART00000050533
|
ccni2
|
cyclin I family, member 2 |
| chr3_+_16722014 | 0.72 |
ENSDART00000008711
|
gys1
|
glycogen synthase 1 (muscle) |
| chr23_-_32100106 | 0.70 |
ENSDART00000044658
|
letmd1
|
LETM1 domain containing 1 |
| chr19_+_935565 | 0.68 |
ENSDART00000113368
|
RNF5
|
ring finger protein 5 |
| chr14_+_909769 | 0.68 |
ENSDART00000021346
ENSDART00000172777 |
arl3l2
|
ADP-ribosylation factor-like 3, like 2 |
| chr1_+_176583 | 0.67 |
ENSDART00000168760
ENSDART00000160425 |
LAMP1
|
lysosomal associated membrane protein 1 |
| chr23_+_45822935 | 0.67 |
ENSDART00000161892
|
vdra
|
vitamin D receptor a |
| chr8_+_28724692 | 0.66 |
ENSDART00000140115
|
ccser1
|
coiled-coil serine-rich protein 1 |
| chr23_-_27547931 | 0.66 |
ENSDART00000144419
|
larp4aa
|
La ribonucleoprotein domain family, member 4Aa |
| chr21_-_43398457 | 0.65 |
ENSDART00000166530
|
ccni2
|
cyclin I family, member 2 |
| chr25_+_34888886 | 0.64 |
ENSDART00000035245
|
spire2
|
spire-type actin nucleation factor 2 |
| chr9_-_24413008 | 0.63 |
ENSDART00000135897
|
tmeff2a
|
transmembrane protein with EGF-like and two follistatin-like domains 2a |
| chr12_+_48511605 | 0.61 |
ENSDART00000168389
|
arhgap44
|
Rho GTPase activating protein 44 |
| chr24_+_3328354 | 0.61 |
ENSDART00000147468
|
bdh1
|
3-hydroxybutyrate dehydrogenase, type 1 |
| chr1_-_411331 | 0.61 |
ENSDART00000092524
|
rasa3
|
RAS p21 protein activator 3 |
| chr1_+_32528097 | 0.61 |
ENSDART00000128317
|
nlgn4a
|
neuroligin 4a |
| chr25_-_7999756 | 0.60 |
ENSDART00000159908
|
camk1db
|
calcium/calmodulin-dependent protein kinase 1Db |
| chr22_-_80533 | 0.60 |
ENSDART00000183124
|
CABZ01101813.1
|
|
| chr12_+_48851381 | 0.60 |
ENSDART00000187241
ENSDART00000187796 |
dlg5b.1
|
discs, large homolog 5b (Drosophila), tandem duplicate 1 |
| chr5_+_65492183 | 0.59 |
ENSDART00000162804
|
si:dkey-21e5.1
|
si:dkey-21e5.1 |
| chr11_+_37251825 | 0.59 |
ENSDART00000169804
|
il17rc
|
interleukin 17 receptor C |
| chr22_+_34784075 | 0.59 |
ENSDART00000167538
|
lcor
|
ligand dependent nuclear receptor corepressor |
| chr17_+_8323348 | 0.59 |
ENSDART00000169900
|
cdc42bpaa
|
CDC42 binding protein kinase alpha (DMPK-like) a |
| chr19_+_56351 | 0.59 |
ENSDART00000168334
|
col14a1b
|
collagen, type XIV, alpha 1b |
| chr10_-_15919839 | 0.59 |
ENSDART00000065032
|
pip5k1ba
|
phosphatidylinositol-4-phosphate 5-kinase, type I, beta a |
| chr15_-_17099560 | 0.58 |
ENSDART00000101724
|
mos
|
v-mos Moloney murine sarcoma viral oncogene homolog |
| chr18_-_49078428 | 0.58 |
ENSDART00000160702
ENSDART00000174103 |
BX663503.3
|
|
| chr25_+_37348730 | 0.57 |
ENSDART00000156639
|
pamr1
|
peptidase domain containing associated with muscle regeneration 1 |
| chr14_-_7245971 | 0.57 |
ENSDART00000108796
|
stox2b
|
storkhead box 2b |
| chr3_+_45687266 | 0.57 |
ENSDART00000131652
|
gpr146
|
G protein-coupled receptor 146 |
| chr23_+_9057999 | 0.55 |
ENSDART00000091899
|
ccm2l
|
cerebral cavernous malformation 2-like |
| chr1_-_48922273 | 0.55 |
ENSDART00000150254
|
si:ch211-112b1.2
|
si:ch211-112b1.2 |
| chr18_-_19103929 | 0.55 |
ENSDART00000188370
ENSDART00000177621 |
dennd4a
|
DENN/MADD domain containing 4A |
| chr6_-_7776612 | 0.55 |
ENSDART00000190269
|
myh9a
|
myosin, heavy chain 9a, non-muscle |
| chr16_-_54405976 | 0.55 |
ENSDART00000055395
|
osr2
|
odd-skipped related transciption factor 2 |
| chr13_+_12739283 | 0.55 |
ENSDART00000102279
|
lingo2b
|
leucine rich repeat and Ig domain containing 2b |
| chr19_-_3303995 | 0.53 |
ENSDART00000105150
|
si:ch211-133n4.9
|
si:ch211-133n4.9 |
| chr12_-_44016898 | 0.53 |
ENSDART00000175304
|
si:dkey-201i2.4
|
si:dkey-201i2.4 |
| chr11_+_45436703 | 0.53 |
ENSDART00000168295
ENSDART00000173293 |
sos1
|
son of sevenless homolog 1 (Drosophila) |
| chr6_+_42338309 | 0.52 |
ENSDART00000015277
|
gpx1b
|
glutathione peroxidase 1b |
| chr4_-_9579299 | 0.50 |
ENSDART00000183079
ENSDART00000192968 ENSDART00000091809 |
shank3b
|
SH3 and multiple ankyrin repeat domains 3b |
| chr24_-_42072886 | 0.50 |
ENSDART00000171389
|
CABZ01095370.1
|
|
| chr24_+_32472155 | 0.49 |
ENSDART00000098859
|
neurod6a
|
neuronal differentiation 6a |
| chr21_-_24865217 | 0.48 |
ENSDART00000101136
|
igsf9bb
|
immunoglobulin superfamily, member 9Bb |
| chr25_+_1304173 | 0.48 |
ENSDART00000155229
|
rxfp3.3b
|
relaxin/insulin-like family peptide receptor 3.3b |
| chr17_+_53311243 | 0.48 |
ENSDART00000160241
ENSDART00000160009 ENSDART00000162239 |
asb2b
|
ankyrin repeat and SOCS box containing 2b |
| chr25_+_34889061 | 0.47 |
ENSDART00000136226
|
spire2
|
spire-type actin nucleation factor 2 |
| chr5_-_67241633 | 0.47 |
ENSDART00000114783
|
clip1a
|
CAP-GLY domain containing linker protein 1a |
| chr16_-_26132122 | 0.46 |
ENSDART00000157787
|
lipeb
|
lipase, hormone-sensitive b |
| chr20_-_48172556 | 0.46 |
ENSDART00000097888
|
CABZ01059099.1
|
|
| chr12_+_41510492 | 0.46 |
ENSDART00000170976
ENSDART00000176164 |
kif5bb
|
kinesin family member 5B, b |
| chr5_-_1999417 | 0.46 |
ENSDART00000155437
ENSDART00000145781 |
si:ch211-160e1.5
|
si:ch211-160e1.5 |
| chr1_-_49434798 | 0.45 |
ENSDART00000150842
ENSDART00000150789 |
ATP5MD
|
si:dkeyp-80c12.10 |
| chr8_-_34762163 | 0.45 |
ENSDART00000114080
|
setd1bb
|
SET domain containing 1B, b |
| chr9_-_18877597 | 0.45 |
ENSDART00000099446
|
kctd4
|
potassium channel tetramerization domain containing 4 |
| chr6_-_57408576 | 0.44 |
ENSDART00000163930
|
znfx1
|
zinc finger, NFX1-type containing 1 |
| chr13_-_479129 | 0.44 |
ENSDART00000159803
ENSDART00000082127 |
heatr5b
|
HEAT repeat containing 5B |
| chr5_+_15667874 | 0.43 |
ENSDART00000127015
|
srrm4
|
serine/arginine repetitive matrix 4 |
| chr1_+_1599979 | 0.43 |
ENSDART00000097626
|
urp2
|
urotensin II-related peptide |
| chr14_-_4044545 | 0.43 |
ENSDART00000169527
|
snx25
|
sorting nexin 25 |
| chr16_-_6821927 | 0.43 |
ENSDART00000149070
ENSDART00000149570 |
mbpb
|
myelin basic protein b |
| chr4_+_50401133 | 0.43 |
ENSDART00000150644
|
si:dkey-156k2.7
|
si:dkey-156k2.7 |
| chr7_-_12596727 | 0.41 |
ENSDART00000186413
|
adamtsl3
|
ADAMTS-like 3 |
| chr12_-_15620090 | 0.41 |
ENSDART00000038032
|
acbd4
|
acyl-CoA binding domain containing 4 |
| chr5_+_66170479 | 0.38 |
ENSDART00000172117
|
gldc
|
glycine dehydrogenase (decarboxylating) |
| chr21_+_7900107 | 0.38 |
ENSDART00000056560
|
ch25hl2
|
cholesterol 25-hydroxylase like 2 |
| chr1_-_53259746 | 0.37 |
ENSDART00000134222
|
il15
|
interleukin 15 |
| chr5_+_337215 | 0.36 |
ENSDART00000167982
|
rnf170
|
ring finger protein 170 |
| chr8_-_14144707 | 0.35 |
ENSDART00000148061
|
si:dkey-6n6.1
|
si:dkey-6n6.1 |
| chr20_-_46808870 | 0.35 |
ENSDART00000190911
|
esrrgb
|
estrogen-related receptor gamma b |
| chr19_-_7225060 | 0.34 |
ENSDART00000133179
ENSDART00000151138 ENSDART00000104799 ENSDART00000128331 |
col11a2
|
collagen, type XI, alpha 2 |
| chr15_-_14486534 | 0.33 |
ENSDART00000179368
|
numbl
|
numb homolog (Drosophila)-like |
| chr25_-_36370292 | 0.33 |
ENSDART00000152766
|
CR354435.1
|
Histone H2B 1/2 |
| chr23_-_29556844 | 0.32 |
ENSDART00000138021
|
rbp7a
|
retinol binding protein 7a, cellular |
| chr22_-_16275236 | 0.30 |
ENSDART00000149051
|
cdc14ab
|
cell division cycle 14Ab |
| chr4_-_5019113 | 0.30 |
ENSDART00000189321
ENSDART00000081990 |
strip2
|
striatin interacting protein 2 |
| chr19_-_25427255 | 0.30 |
ENSDART00000036854
|
glcci1a
|
glucocorticoid induced 1a |
| chr9_-_52386733 | 0.29 |
ENSDART00000171721
|
dap1b
|
death associated protein 1b |
| chr3_-_22829710 | 0.29 |
ENSDART00000055659
|
cyb561
|
cytochrome b561 |
| chr6_+_11026354 | 0.29 |
ENSDART00000065433
|
abcc6a
|
ATP-binding cassette, sub-family C (CFTR/MRP), member 6a |
| chr1_-_56363108 | 0.29 |
ENSDART00000021878
|
gcgrb
|
glucagon receptor b |
| chr6_-_53391727 | 0.28 |
ENSDART00000155483
|
si:ch211-161c3.6
|
si:ch211-161c3.6 |
| chr9_+_38075851 | 0.28 |
ENSDART00000135314
|
cacnb4a
|
calcium channel, voltage-dependent, beta 4a subunit |
| chr14_-_14566417 | 0.27 |
ENSDART00000159056
|
si:dkey-27i16.2
|
si:dkey-27i16.2 |
| chr22_+_15979430 | 0.26 |
ENSDART00000189703
ENSDART00000192674 |
rc3h1a
|
ring finger and CCCH-type domains 1a |
| chr25_+_12012 | 0.25 |
ENSDART00000170348
|
cntn1a
|
contactin 1a |
| chr22_-_20289948 | 0.25 |
ENSDART00000132951
|
si:dkey-110c1.10
|
si:dkey-110c1.10 |
| chr14_+_12316581 | 0.25 |
ENSDART00000170115
ENSDART00000149757 |
si:ch211-125c23.3
cx31.7
|
si:ch211-125c23.3 connexin 31.7 |
| chr6_+_11026866 | 0.24 |
ENSDART00000189619
|
abcc6a
|
ATP-binding cassette, sub-family C (CFTR/MRP), member 6a |
| chr6_-_13498745 | 0.22 |
ENSDART00000027684
ENSDART00000189438 |
mylkb
|
myosin light chain kinase b |
| chr21_+_15295685 | 0.22 |
ENSDART00000135603
|
BX901878.1
|
|
| chr1_-_50438247 | 0.21 |
ENSDART00000114098
|
dkk2
|
dickkopf WNT signaling pathway inhibitor 2 |
| chr15_+_43906043 | 0.21 |
ENSDART00000010881
|
naalad2
|
N-acetylated alpha-linked acidic dipeptidase 2 |
| chr3_-_16039619 | 0.20 |
ENSDART00000143324
|
spsb3a
|
splA/ryanodine receptor domain and SOCS box containing 3a |
| chr14_+_164556 | 0.20 |
ENSDART00000185606
|
WDR1
|
WD repeat domain 1 |
| chr21_+_3941758 | 0.20 |
ENSDART00000181345
|
setx
|
senataxin |
| chr23_+_2136344 | 0.19 |
ENSDART00000085290
ENSDART00000092757 |
cpeb2
|
cytoplasmic polyadenylation element binding protein 2 |
| chr1_-_10473630 | 0.18 |
ENSDART00000040116
|
tnrc5
|
trinucleotide repeat containing 5 |
| chr23_+_41799748 | 0.18 |
ENSDART00000144257
|
pdyn
|
prodynorphin |
| chr4_+_39742119 | 0.17 |
ENSDART00000176004
|
CR749167.2
|
|
| chr8_+_2642384 | 0.17 |
ENSDART00000143242
|
naif1
|
nuclear apoptosis inducing factor 1 |
| chr5_+_37032038 | 0.16 |
ENSDART00000045036
|
nfkbib
|
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, beta |
| chr7_+_31051603 | 0.16 |
ENSDART00000108721
|
tjp1a
|
tight junction protein 1a |
| chr8_-_53490376 | 0.14 |
ENSDART00000158789
|
chdh
|
choline dehydrogenase |
Network of associatons between targets according to the STRING database.
First level regulatory network of egr1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 2.0 | GO:0010807 | regulation of synaptic vesicle priming(GO:0010807) |
| 0.5 | 1.5 | GO:0090004 | positive regulation of Golgi to plasma membrane protein transport(GO:0042998) positive regulation of establishment of protein localization to plasma membrane(GO:0090004) |
| 0.5 | 1.9 | GO:0097112 | gamma-aminobutyric acid receptor clustering(GO:0097112) |
| 0.4 | 1.3 | GO:0015824 | proline transport(GO:0015824) |
| 0.4 | 1.5 | GO:0010755 | regulation of plasminogen activation(GO:0010755) |
| 0.3 | 1.9 | GO:0032367 | intracellular cholesterol transport(GO:0032367) |
| 0.3 | 2.8 | GO:0071331 | cellular response to carbohydrate stimulus(GO:0071322) cellular response to monosaccharide stimulus(GO:0071326) cellular response to hexose stimulus(GO:0071331) cellular response to glucose stimulus(GO:0071333) |
| 0.3 | 0.9 | GO:1901187 | regulation of ephrin receptor signaling pathway(GO:1901187) |
| 0.2 | 1.5 | GO:0019427 | acetate metabolic process(GO:0006083) acetyl-CoA biosynthetic process from acetate(GO:0019427) |
| 0.2 | 1.7 | GO:0098815 | postsynaptic density assembly(GO:0097107) modulation of excitatory postsynaptic potential(GO:0098815) positive regulation of excitatory postsynaptic potential(GO:2000463) |
| 0.2 | 1.6 | GO:0032793 | positive regulation of CREB transcription factor activity(GO:0032793) |
| 0.2 | 1.4 | GO:0048745 | smooth muscle tissue development(GO:0048745) |
| 0.2 | 0.6 | GO:1902103 | meiotic cell cycle phase transition(GO:0044771) metaphase/anaphase transition of meiotic cell cycle(GO:0044785) regulation of meiotic cell cycle phase transition(GO:1901993) negative regulation of meiotic cell cycle phase transition(GO:1901994) regulation of metaphase/anaphase transition of meiotic cell cycle(GO:1902102) negative regulation of metaphase/anaphase transition of meiotic cell cycle(GO:1902103) regulation of meiotic chromosome separation(GO:1905132) negative regulation of meiotic chromosome separation(GO:1905133) |
| 0.2 | 1.9 | GO:0048790 | maintenance of presynaptic active zone structure(GO:0048790) maintenance of synapse structure(GO:0099558) |
| 0.2 | 2.5 | GO:0033173 | calcineurin-NFAT signaling cascade(GO:0033173) |
| 0.2 | 1.1 | GO:0033206 | meiotic cytokinesis(GO:0033206) polar body extrusion after meiotic divisions(GO:0040038) |
| 0.1 | 0.8 | GO:0070650 | actin filament bundle distribution(GO:0070650) |
| 0.1 | 1.0 | GO:2000253 | cannabinoid signaling pathway(GO:0038171) positive regulation of feeding behavior(GO:2000253) |
| 0.1 | 1.3 | GO:0060079 | excitatory postsynaptic potential(GO:0060079) |
| 0.1 | 0.5 | GO:0061439 | renal system vasculature morphogenesis(GO:0061438) kidney vasculature morphogenesis(GO:0061439) glomerulus vasculature morphogenesis(GO:0072103) glomerular capillary formation(GO:0072104) |
| 0.1 | 1.1 | GO:0070262 | peptidyl-serine dephosphorylation(GO:0070262) |
| 0.1 | 1.4 | GO:2000054 | negative regulation of Wnt signaling pathway involved in dorsal/ventral axis specification(GO:2000054) |
| 0.1 | 1.2 | GO:0006382 | adenosine to inosine editing(GO:0006382) |
| 0.1 | 0.4 | GO:0006546 | glycine catabolic process(GO:0006546) glycine decarboxylation via glycine cleavage system(GO:0019464) |
| 0.1 | 0.5 | GO:0010269 | response to selenium ion(GO:0010269) |
| 0.1 | 0.7 | GO:0071632 | optomotor response(GO:0071632) |
| 0.1 | 0.6 | GO:0045053 | protein retention in Golgi apparatus(GO:0045053) |
| 0.1 | 0.5 | GO:0031116 | positive regulation of microtubule polymerization(GO:0031116) |
| 0.1 | 2.2 | GO:0035308 | negative regulation of dephosphorylation(GO:0035305) negative regulation of protein dephosphorylation(GO:0035308) |
| 0.1 | 1.0 | GO:0044154 | histone H3-K14 acetylation(GO:0044154) |
| 0.1 | 0.6 | GO:0043363 | nucleate erythrocyte differentiation(GO:0043363) |
| 0.1 | 0.9 | GO:0016199 | axon midline choice point recognition(GO:0016199) |
| 0.1 | 1.5 | GO:0051156 | glucose 6-phosphate metabolic process(GO:0051156) |
| 0.1 | 0.4 | GO:0031048 | chromatin silencing by small RNA(GO:0031048) |
| 0.1 | 2.6 | GO:1903670 | regulation of sprouting angiogenesis(GO:1903670) |
| 0.1 | 1.6 | GO:0019835 | cytolysis(GO:0019835) |
| 0.1 | 0.4 | GO:0032475 | otolith formation(GO:0032475) |
| 0.0 | 0.1 | GO:0019695 | choline metabolic process(GO:0019695) |
| 0.0 | 0.9 | GO:0014823 | response to activity(GO:0014823) |
| 0.0 | 1.0 | GO:0051289 | protein homotetramerization(GO:0051289) |
| 0.0 | 1.0 | GO:0030168 | platelet activation(GO:0030168) |
| 0.0 | 1.0 | GO:0071542 | dopaminergic neuron differentiation(GO:0071542) |
| 0.0 | 0.6 | GO:0032483 | regulation of Rab protein signal transduction(GO:0032483) |
| 0.0 | 0.4 | GO:0021754 | facial nucleus development(GO:0021754) |
| 0.0 | 1.2 | GO:0007157 | heterophilic cell-cell adhesion via plasma membrane cell adhesion molecules(GO:0007157) |
| 0.0 | 0.3 | GO:0071850 | mitotic cell cycle arrest(GO:0071850) |
| 0.0 | 1.5 | GO:2000649 | regulation of sodium ion transmembrane transporter activity(GO:2000649) |
| 0.0 | 0.9 | GO:2001237 | negative regulation of extrinsic apoptotic signaling pathway(GO:2001237) |
| 0.0 | 0.2 | GO:0006369 | termination of RNA polymerase II transcription(GO:0006369) |
| 0.0 | 0.8 | GO:0071910 | determination of liver left/right asymmetry(GO:0071910) |
| 0.0 | 0.5 | GO:0019433 | triglyceride catabolic process(GO:0019433) |
| 0.0 | 0.4 | GO:0036230 | granulocyte activation(GO:0036230) neutrophil activation(GO:0042119) positive regulation of tyrosine phosphorylation of STAT protein(GO:0042531) |
| 0.0 | 2.3 | GO:0035725 | sodium ion transmembrane transport(GO:0035725) |
| 0.0 | 0.4 | GO:0051568 | histone H3-K4 methylation(GO:0051568) |
| 0.0 | 0.6 | GO:0055003 | cardiac myofibril assembly(GO:0055003) |
| 0.0 | 0.5 | GO:0030500 | regulation of bone mineralization(GO:0030500) regulation of biomineral tissue development(GO:0070167) |
| 0.0 | 0.4 | GO:0010811 | positive regulation of cell-substrate adhesion(GO:0010811) |
| 0.0 | 0.9 | GO:0010107 | potassium ion import(GO:0010107) potassium ion import across plasma membrane(GO:1990573) |
| 0.0 | 0.4 | GO:0030512 | negative regulation of transforming growth factor beta receptor signaling pathway(GO:0030512) negative regulation of cellular response to transforming growth factor beta stimulus(GO:1903845) |
| 0.0 | 0.3 | GO:0055117 | regulation of cardiac muscle contraction(GO:0055117) |
| 0.0 | 0.7 | GO:0005978 | glycogen biosynthetic process(GO:0005978) glucan biosynthetic process(GO:0009250) |
| 0.0 | 2.1 | GO:0008654 | phospholipid biosynthetic process(GO:0008654) |
| 0.0 | 0.3 | GO:0010675 | regulation of cellular carbohydrate metabolic process(GO:0010675) regulation of glucose metabolic process(GO:0010906) |
| 0.0 | 0.1 | GO:0035469 | determination of pancreatic left/right asymmetry(GO:0035469) |
| 0.0 | 1.0 | GO:0060216 | definitive hemopoiesis(GO:0060216) |
| 0.0 | 0.3 | GO:0010507 | negative regulation of autophagy(GO:0010507) |
| 0.0 | 0.7 | GO:0003146 | heart jogging(GO:0003146) |
| 0.0 | 0.6 | GO:0009948 | anterior/posterior axis specification(GO:0009948) |
| 0.0 | 0.5 | GO:0060349 | bone morphogenesis(GO:0060349) |
| 0.0 | 0.4 | GO:0016126 | sterol biosynthetic process(GO:0016126) |
| 0.0 | 2.9 | GO:0007018 | microtubule-based movement(GO:0007018) |
| 0.0 | 0.3 | GO:0060319 | primitive erythrocyte differentiation(GO:0060319) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 1.9 | GO:0060077 | inhibitory synapse(GO:0060077) |
| 0.5 | 2.0 | GO:0070044 | synaptobrevin 2-SNAP-25-syntaxin-1a complex(GO:0070044) |
| 0.4 | 2.5 | GO:0005955 | calcineurin complex(GO:0005955) |
| 0.3 | 2.8 | GO:0001669 | acrosomal vesicle(GO:0001669) |
| 0.2 | 1.4 | GO:0070062 | extracellular exosome(GO:0070062) |
| 0.2 | 1.9 | GO:0005742 | mitochondrial outer membrane translocase complex(GO:0005742) |
| 0.1 | 0.4 | GO:0005960 | glycine cleavage complex(GO:0005960) |
| 0.1 | 1.3 | GO:0017146 | NMDA selective glutamate receptor complex(GO:0017146) |
| 0.1 | 1.9 | GO:0048788 | cytoskeleton of presynaptic active zone(GO:0048788) |
| 0.1 | 1.0 | GO:0070775 | H3 histone acetyltransferase complex(GO:0070775) MOZ/MORF histone acetyltransferase complex(GO:0070776) |
| 0.1 | 0.3 | GO:1902636 | kinociliary basal body(GO:1902636) |
| 0.0 | 3.1 | GO:0030141 | secretory granule(GO:0030141) |
| 0.0 | 1.1 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.0 | 2.9 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.0 | 0.6 | GO:0043256 | basal lamina(GO:0005605) laminin complex(GO:0043256) |
| 0.0 | 0.4 | GO:0048188 | Set1C/COMPASS complex(GO:0048188) |
| 0.0 | 0.4 | GO:0043209 | myelin sheath(GO:0043209) |
| 0.0 | 1.7 | GO:0098878 | ionotropic glutamate receptor complex(GO:0008328) neurotransmitter receptor complex(GO:0098878) |
| 0.0 | 0.9 | GO:0005861 | troponin complex(GO:0005861) |
| 0.0 | 0.5 | GO:0035371 | microtubule plus-end(GO:0035371) |
| 0.0 | 3.8 | GO:0005764 | lysosome(GO:0005764) |
| 0.0 | 2.8 | GO:0099503 | secretory vesicle(GO:0099503) |
| 0.0 | 1.5 | GO:0005802 | trans-Golgi network(GO:0005802) |
| 0.0 | 1.2 | GO:0030027 | lamellipodium(GO:0030027) |
| 0.0 | 1.1 | GO:0005581 | collagen trimer(GO:0005581) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 2.8 | GO:0004679 | AMP-activated protein kinase activity(GO:0004679) |
| 0.5 | 1.9 | GO:0030228 | lipoprotein particle receptor activity(GO:0030228) |
| 0.4 | 2.2 | GO:0004105 | choline-phosphate cytidylyltransferase activity(GO:0004105) |
| 0.4 | 2.5 | GO:0033192 | calcium-dependent protein serine/threonine phosphatase activity(GO:0004723) calmodulin-dependent protein phosphatase activity(GO:0033192) |
| 0.4 | 1.5 | GO:0071253 | connexin binding(GO:0071253) |
| 0.3 | 2.0 | GO:0005483 | soluble NSF attachment protein activity(GO:0005483) |
| 0.2 | 1.5 | GO:0003987 | acetate-CoA ligase activity(GO:0003987) |
| 0.2 | 2.0 | GO:0016803 | ether hydrolase activity(GO:0016803) |
| 0.2 | 1.0 | GO:0004949 | cannabinoid receptor activity(GO:0004949) |
| 0.2 | 1.7 | GO:0030160 | GKAP/Homer scaffold activity(GO:0030160) |
| 0.2 | 1.6 | GO:0008140 | cAMP response element binding protein binding(GO:0008140) |
| 0.2 | 0.7 | GO:0005499 | vitamin D binding(GO:0005499) |
| 0.2 | 0.7 | GO:0004373 | glycogen (starch) synthase activity(GO:0004373) |
| 0.1 | 2.2 | GO:0051721 | protein phosphatase 2A binding(GO:0051721) |
| 0.1 | 1.2 | GO:0003726 | double-stranded RNA adenosine deaminase activity(GO:0003726) |
| 0.1 | 1.9 | GO:0098882 | structural constituent of presynaptic active zone(GO:0098882) |
| 0.1 | 0.9 | GO:0015467 | G-protein activated inward rectifier potassium channel activity(GO:0015467) |
| 0.1 | 1.5 | GO:0016861 | intramolecular oxidoreductase activity, interconverting aldoses and ketoses(GO:0016861) |
| 0.1 | 1.0 | GO:0005522 | profilin binding(GO:0005522) |
| 0.1 | 0.3 | GO:0000293 | ferric-chelate reductase activity(GO:0000293) |
| 0.1 | 1.9 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.1 | 0.4 | GO:0070052 | collagen V binding(GO:0070052) |
| 0.1 | 0.3 | GO:0070513 | death domain binding(GO:0070513) |
| 0.1 | 1.0 | GO:0043994 | H3 histone acetyltransferase activity(GO:0010484) histone acetyltransferase activity (H3-K23 specific)(GO:0043994) |
| 0.1 | 1.3 | GO:0004972 | NMDA glutamate receptor activity(GO:0004972) |
| 0.1 | 0.2 | GO:0001160 | transcription termination site sequence-specific DNA binding(GO:0001147) transcription termination site DNA binding(GO:0001160) |
| 0.1 | 0.5 | GO:0033878 | hormone-sensitive lipase activity(GO:0033878) |
| 0.1 | 0.3 | GO:0016519 | gastric inhibitory peptide receptor activity(GO:0016519) |
| 0.1 | 1.5 | GO:0042043 | neurexin family protein binding(GO:0042043) |
| 0.1 | 1.5 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.1 | 0.4 | GO:0000254 | C-4 methylsterol oxidase activity(GO:0000254) |
| 0.0 | 0.6 | GO:0030368 | interleukin-17 receptor activity(GO:0030368) |
| 0.0 | 0.9 | GO:0046875 | ephrin receptor binding(GO:0046875) |
| 0.0 | 0.3 | GO:0035613 | RNA stem-loop binding(GO:0035613) |
| 0.0 | 0.4 | GO:0016594 | glycine binding(GO:0016594) |
| 0.0 | 0.9 | GO:0004016 | adenylate cyclase activity(GO:0004016) |
| 0.0 | 0.9 | GO:0048306 | calcium-dependent protein binding(GO:0048306) |
| 0.0 | 0.4 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.0 | 2.9 | GO:0003777 | microtubule motor activity(GO:0003777) |
| 0.0 | 0.5 | GO:0004602 | glutathione peroxidase activity(GO:0004602) |
| 0.0 | 0.8 | GO:0051537 | 2 iron, 2 sulfur cluster binding(GO:0051537) |
| 0.0 | 2.8 | GO:0004197 | cysteine-type endopeptidase activity(GO:0004197) |
| 0.0 | 3.6 | GO:0015293 | symporter activity(GO:0015293) |
| 0.0 | 0.1 | GO:0004974 | leukotriene receptor activity(GO:0004974) |
| 0.0 | 0.6 | GO:0016307 | phosphatidylinositol phosphate kinase activity(GO:0016307) |
| 0.0 | 0.1 | GO:0008349 | MAP kinase kinase kinase kinase activity(GO:0008349) |
| 0.0 | 0.4 | GO:0000062 | fatty-acyl-CoA binding(GO:0000062) |
| 0.0 | 0.1 | GO:0072320 | volume-sensitive chloride channel activity(GO:0072320) |
| 0.0 | 0.5 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
| 0.0 | 0.1 | GO:0003836 | beta-galactoside (CMP) alpha-2,3-sialyltransferase activity(GO:0003836) |
| 0.0 | 0.3 | GO:0005246 | calcium channel regulator activity(GO:0005246) |
| 0.0 | 0.5 | GO:0019957 | C-C chemokine binding(GO:0019957) |
| 0.0 | 1.1 | GO:0019888 | protein phosphatase regulator activity(GO:0019888) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 2.5 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
| 0.1 | 2.6 | PID REELIN PATHWAY | Reelin signaling pathway |
| 0.1 | 0.9 | ST ADRENERGIC | Adrenergic Pathway |
| 0.1 | 1.3 | SA TRKA RECEPTOR | The TrkA receptor binds nerve growth factor to activate MAP kinase pathways and promote cell growth. |
| 0.1 | 0.9 | ST G ALPHA S PATHWAY | G alpha s Pathway |
| 0.0 | 0.7 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
| 0.0 | 1.0 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.0 | 0.6 | ST ERK1 ERK2 MAPK PATHWAY | ERK1/ERK2 MAPK Pathway |
| 0.0 | 1.1 | NABA COLLAGENS | Genes encoding collagen proteins |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 2.6 | REACTOME PLATELET SENSITIZATION BY LDL | Genes involved in Platelet sensitization by LDL |
| 0.2 | 2.5 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.1 | 1.3 | REACTOME SOS MEDIATED SIGNALLING | Genes involved in SOS-mediated signalling |
| 0.1 | 1.1 | REACTOME SYNTHESIS OF PE | Genes involved in Synthesis of PE |
| 0.0 | 1.7 | REACTOME TRANSPORT TO THE GOLGI AND SUBSEQUENT MODIFICATION | Genes involved in Transport to the Golgi and subsequent modification |
| 0.0 | 0.9 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.0 | 0.6 | REACTOME INHIBITION OF VOLTAGE GATED CA2 CHANNELS VIA GBETA GAMMA SUBUNITS | Genes involved in Inhibition of voltage gated Ca2+ channels via Gbeta/gamma subunits |
| 0.0 | 1.1 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.0 | 0.1 | REACTOME TERMINATION OF O GLYCAN BIOSYNTHESIS | Genes involved in Termination of O-glycan biosynthesis |
| 0.0 | 0.7 | REACTOME GLUCOSE METABOLISM | Genes involved in Glucose metabolism |