Project
PRJEB1986: zebrafish developmental stages transcriptome
Navigation
Downloads
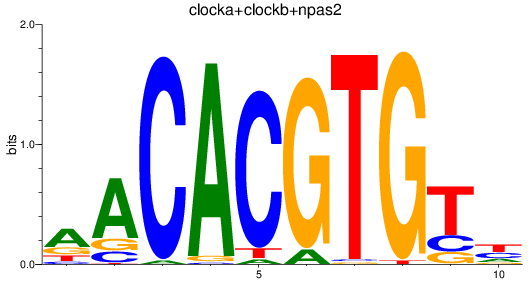
Results for clocka+clockb+npas2
Z-value: 2.30
Transcription factors associated with clocka+clockb+npas2
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
clockb
|
ENSDARG00000003631 | clock circadian regulator b |
|
clocka
|
ENSDARG00000011703 | clock circadian regulator a |
|
npas2
|
ENSDARG00000016536 | neuronal PAS domain protein 2 |
|
npas2
|
ENSDARG00000116993 | neuronal PAS domain protein 2 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| clocka | dr11_v1_chr20_+_22130284_22130284 | -0.57 | 1.2e-02 | Click! |
| clockb | dr11_v1_chr1_+_19433004_19433004 | -0.44 | 5.9e-02 | Click! |
| npas2 | dr11_v1_chr5_+_22791686_22791686 | -0.35 | 1.5e-01 | Click! |
Activity profile of clocka+clockb+npas2 motif
Sorted Z-values of clocka+clockb+npas2 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr10_+_22034477 | 8.82 |
ENSDART00000133304
ENSDART00000134189 ENSDART00000021240 ENSDART00000100526 |
npm1a
|
nucleophosmin 1a |
| chr3_+_53773256 | 5.30 |
ENSDART00000170461
|
col5a3a
|
collagen, type V, alpha 3a |
| chr23_+_25292147 | 5.01 |
ENSDART00000131486
|
pa2g4b
|
proliferation-associated 2G4, b |
| chr8_+_26059677 | 4.71 |
ENSDART00000009178
|
impdh2
|
IMP (inosine 5'-monophosphate) dehydrogenase 2 |
| chr9_-_11587070 | 4.69 |
ENSDART00000030995
|
umps
|
uridine monophosphate synthetase |
| chr19_+_16016038 | 4.54 |
ENSDART00000131319
|
ctps1a
|
CTP synthase 1a |
| chr16_-_34195002 | 4.49 |
ENSDART00000054026
|
rcc1
|
regulator of chromosome condensation 1 |
| chr13_+_13930263 | 4.49 |
ENSDART00000079154
|
rpia
|
ribose 5-phosphate isomerase A (ribose 5-phosphate epimerase) |
| chr7_+_29163762 | 4.21 |
ENSDART00000173762
|
slc38a8b
|
solute carrier family 38, member 8b |
| chr13_+_15701849 | 4.19 |
ENSDART00000003517
|
trmt61a
|
tRNA methyltransferase 61A |
| chr7_+_29167744 | 4.09 |
ENSDART00000076345
|
slc38a8b
|
solute carrier family 38, member 8b |
| chr1_-_44933094 | 3.87 |
ENSDART00000147527
|
si:dkey-9i23.14
|
si:dkey-9i23.14 |
| chr14_+_14836468 | 3.86 |
ENSDART00000166728
|
si:dkey-102m7.3
|
si:dkey-102m7.3 |
| chr4_+_47257854 | 3.82 |
ENSDART00000173868
|
crestin
|
crestin |
| chr5_-_32505109 | 3.82 |
ENSDART00000188219
|
ntmt1
|
N-terminal Xaa-Pro-Lys N-methyltransferase 1 |
| chr6_+_3717613 | 3.69 |
ENSDART00000184330
|
ssb
|
Sjogren syndrome antigen B (autoantigen La) |
| chr3_-_1339220 | 3.69 |
ENSDART00000172000
|
tmem106c
|
transmembrane protein 106C |
| chr21_+_43702016 | 3.66 |
ENSDART00000017176
|
dkc1
|
dyskeratosis congenita 1, dyskerin |
| chr5_-_32505276 | 3.56 |
ENSDART00000034705
ENSDART00000187597 |
ntmt1
|
N-terminal Xaa-Pro-Lys N-methyltransferase 1 |
| chr8_-_11229523 | 3.55 |
ENSDART00000002164
|
unc45b
|
unc-45 myosin chaperone B |
| chr20_-_25626693 | 3.40 |
ENSDART00000132247
|
paics
|
phosphoribosylaminoimidazole carboxylase, phosphoribosylaminoimidazole succinocarboxamide synthetase |
| chr13_-_24260609 | 3.37 |
ENSDART00000138747
|
urb2
|
URB2 ribosome biogenesis 2 homolog (S. cerevisiae) |
| chr5_-_24543526 | 3.36 |
ENSDART00000046384
|
trmt2a
|
tRNA methyltransferase 2 homolog A |
| chr5_+_68807170 | 3.30 |
ENSDART00000017849
|
her7
|
hairy and enhancer of split related-7 |
| chr25_+_19734038 | 3.21 |
ENSDART00000067354
|
zgc:101783
|
zgc:101783 |
| chr20_-_38827623 | 3.21 |
ENSDART00000153310
|
cad
|
carbamoyl-phosphate synthetase 2, aspartate transcarbamylase, and dihydroorotase |
| chr17_-_49407091 | 3.20 |
ENSDART00000021950
|
mthfd1b
|
methylenetetrahydrofolate dehydrogenase (NADP+ dependent) 1b |
| chr25_-_8625601 | 3.11 |
ENSDART00000155280
|
GDPGP1
|
zgc:153343 |
| chr2_+_25840463 | 3.09 |
ENSDART00000125178
|
eif5a2
|
eukaryotic translation initiation factor 5A2 |
| chr7_-_32021853 | 3.08 |
ENSDART00000134521
|
kif18a
|
kinesin family member 18A |
| chr7_+_41812636 | 3.08 |
ENSDART00000174333
|
orc6
|
origin recognition complex, subunit 6 |
| chr14_+_22076596 | 3.04 |
ENSDART00000106147
ENSDART00000100278 ENSDART00000131489 |
slc43a1a
|
solute carrier family 43 (amino acid system L transporter), member 1a |
| chr13_-_31370184 | 3.02 |
ENSDART00000034829
|
rrp12
|
ribosomal RNA processing 12 homolog |
| chr20_-_25626198 | 3.00 |
ENSDART00000126716
|
paics
|
phosphoribosylaminoimidazole carboxylase, phosphoribosylaminoimidazole succinocarboxamide synthetase |
| chr10_+_17714866 | 3.00 |
ENSDART00000039969
|
slc20a1b
|
solute carrier family 20 (phosphate transporter), member 1b |
| chr19_+_48359259 | 2.98 |
ENSDART00000167353
|
sgo1
|
shugoshin 1 |
| chr24_+_36317544 | 2.97 |
ENSDART00000048640
ENSDART00000156096 |
pus3
|
pseudouridylate synthase 3 |
| chr9_+_21358941 | 2.96 |
ENSDART00000147619
ENSDART00000059402 |
eef1akmt1
|
EEF1A lysine methyltransferase 1 |
| chr7_-_30174882 | 2.96 |
ENSDART00000110409
|
frmd5
|
FERM domain containing 5 |
| chr14_+_6423973 | 2.96 |
ENSDART00000051556
|
abca1b
|
ATP-binding cassette, sub-family A (ABC1), member 1B |
| chr20_+_36806398 | 2.95 |
ENSDART00000153317
|
abracl
|
ABRA C-terminal like |
| chr23_+_25291891 | 2.93 |
ENSDART00000016248
|
pa2g4b
|
proliferation-associated 2G4, b |
| chr11_+_3959495 | 2.86 |
ENSDART00000122953
|
gnl3
|
guanine nucleotide binding protein-like 3 (nucleolar) |
| chr13_+_24380132 | 2.85 |
ENSDART00000022682
|
pdcd2
|
programmed cell death 2 |
| chr5_+_12836913 | 2.83 |
ENSDART00000023101
|
pes
|
pescadillo |
| chr17_+_24821627 | 2.83 |
ENSDART00000112389
|
wdr43
|
WD repeat domain 43 |
| chr3_-_25377163 | 2.81 |
ENSDART00000055490
|
kpna2
|
karyopherin alpha 2 (RAG cohort 1, importin alpha 1) |
| chr4_+_25607743 | 2.79 |
ENSDART00000028297
|
acot14
|
acyl-CoA thioesterase 14 |
| chr21_-_21465111 | 2.79 |
ENSDART00000141487
|
nectin3b
|
nectin cell adhesion molecule 3b |
| chr20_-_25626428 | 2.78 |
ENSDART00000136475
|
paics
|
phosphoribosylaminoimidazole carboxylase, phosphoribosylaminoimidazole succinocarboxamide synthetase |
| chr19_-_18135724 | 2.77 |
ENSDART00000186609
|
cbx3a
|
chromobox homolog 3a (HP1 gamma homolog, Drosophila) |
| chr2_+_25839650 | 2.76 |
ENSDART00000134077
ENSDART00000140804 |
eif5a2
|
eukaryotic translation initiation factor 5A2 |
| chr10_-_11323134 | 2.71 |
ENSDART00000039145
|
inip
|
ints3 and nabp interacting protein |
| chr22_+_9922301 | 2.71 |
ENSDART00000105924
|
blf
|
bloody fingers |
| chr6_-_52348562 | 2.70 |
ENSDART00000142565
ENSDART00000145369 ENSDART00000016890 |
eif6
|
eukaryotic translation initiation factor 6 |
| chr9_+_29985010 | 2.69 |
ENSDART00000020743
|
cmss1
|
cms1 ribosomal small subunit homolog (yeast) |
| chr6_-_18531349 | 2.67 |
ENSDART00000160693
ENSDART00000169780 |
utp6
|
UTP6, small subunit (SSU) processome component, homolog (yeast) |
| chr15_-_14038227 | 2.63 |
ENSDART00000139068
|
zgc:114130
|
zgc:114130 |
| chr10_+_39283985 | 2.63 |
ENSDART00000016464
|
dcps
|
decapping enzyme, scavenger |
| chr7_+_30201611 | 2.60 |
ENSDART00000075588
|
wdr76
|
WD repeat domain 76 |
| chr1_-_52461322 | 2.60 |
ENSDART00000083836
|
si:ch211-217k17.7
|
si:ch211-217k17.7 |
| chr24_-_12938922 | 2.60 |
ENSDART00000024084
|
pck2
|
phosphoenolpyruvate carboxykinase 2 (mitochondrial) |
| chr19_+_16015881 | 2.60 |
ENSDART00000187135
|
ctps1a
|
CTP synthase 1a |
| chr1_+_5485799 | 2.58 |
ENSDART00000022307
|
atic
|
5-aminoimidazole-4-carboxamide ribonucleotide formyltransferase/IMP cyclohydrolase |
| chr1_-_51219965 | 2.56 |
ENSDART00000146612
|
esf1
|
ESF1, nucleolar pre-rRNA processing protein, homolog (S. cerevisiae) |
| chr4_-_4756349 | 2.55 |
ENSDART00000057507
|
ssbp1
|
single-stranded DNA binding protein 1 |
| chr13_+_15702142 | 2.54 |
ENSDART00000135960
|
trmt61a
|
tRNA methyltransferase 61A |
| chr13_-_4992395 | 2.53 |
ENSDART00000102651
|
nolc1
|
nucleolar and coiled-body phosphoprotein 1 |
| chr13_+_6188759 | 2.53 |
ENSDART00000161062
|
ppm1g
|
protein phosphatase, Mg2+/Mn2+ dependent, 1G |
| chr7_+_1505507 | 2.53 |
ENSDART00000161015
|
nop10
|
NOP10 ribonucleoprotein homolog (yeast) |
| chr20_+_35279968 | 2.52 |
ENSDART00000168216
ENSDART00000153332 |
fam49a
|
family with sequence similarity 49, member A |
| chr6_-_57476465 | 2.51 |
ENSDART00000128065
|
ddx27
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 27 |
| chr2_+_25839940 | 2.48 |
ENSDART00000139927
|
eif5a2
|
eukaryotic translation initiation factor 5A2 |
| chr9_+_29671553 | 2.47 |
ENSDART00000015377
|
rnaseh2b
|
ribonuclease H2, subunit B |
| chr7_+_41812817 | 2.47 |
ENSDART00000174165
|
orc6
|
origin recognition complex, subunit 6 |
| chr13_+_18311410 | 2.45 |
ENSDART00000036718
ENSDART00000132073 |
eif4e1c
|
eukaryotic translation initiation factor 4E family member 1c |
| chr13_+_31390313 | 2.41 |
ENSDART00000111477
ENSDART00000137291 |
METTL18
|
si:dkey-15f17.8 |
| chr2_+_29976419 | 2.41 |
ENSDART00000056748
|
en2b
|
engrailed homeobox 2b |
| chr19_+_10831362 | 2.39 |
ENSDART00000053325
|
tomm40l
|
translocase of outer mitochondrial membrane 40 homolog, like |
| chr8_-_18203092 | 2.38 |
ENSDART00000140620
|
elovl8b
|
ELOVL fatty acid elongase 8b |
| chr20_-_37481180 | 2.37 |
ENSDART00000152959
|
si:ch211-202p1.5
|
si:ch211-202p1.5 |
| chr14_-_35414559 | 2.35 |
ENSDART00000145033
|
rnaseh2c
|
ribonuclease H2, subunit C |
| chr11_-_6974022 | 2.34 |
ENSDART00000172851
|
COMP
|
si:ch211-43f4.1 |
| chr19_+_43579786 | 2.28 |
ENSDART00000138404
|
si:ch211-199g17.2
|
si:ch211-199g17.2 |
| chr1_-_25438934 | 2.28 |
ENSDART00000111686
|
fhdc1
|
FH2 domain containing 1 |
| chr25_-_2081371 | 2.27 |
ENSDART00000104915
ENSDART00000156925 |
wnt7bb
|
wingless-type MMTV integration site family, member 7Bb |
| chr4_+_17642731 | 2.26 |
ENSDART00000026509
|
cwf19l1
|
CWF19-like 1, cell cycle control |
| chr21_+_38855551 | 2.25 |
ENSDART00000171977
|
ddx52
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 52 |
| chr12_+_19384615 | 2.25 |
ENSDART00000078266
|
rsl1d1
|
ribosomal L1 domain containing 1 |
| chr5_-_67499279 | 2.25 |
ENSDART00000128050
|
si:dkey-251i10.3
|
si:dkey-251i10.3 |
| chr25_+_19733704 | 2.24 |
ENSDART00000145334
ENSDART00000138946 ENSDART00000138851 |
zgc:101783
|
zgc:101783 |
| chr19_-_24555623 | 2.24 |
ENSDART00000176022
|
polr3gla
|
polymerase (RNA) III (DNA directed) polypeptide G like a |
| chr25_+_18964782 | 2.23 |
ENSDART00000017299
|
tdg.1
|
thymine DNA glycosylase, tandem duplicate 1 |
| chr1_-_19648227 | 2.21 |
ENSDART00000054574
|
polr1e
|
polymerase (RNA) I polypeptide E |
| chr9_+_27720428 | 2.21 |
ENSDART00000112415
|
lcmt2
|
leucine carboxyl methyltransferase 2 |
| chr17_-_114121 | 2.20 |
ENSDART00000172408
ENSDART00000157784 |
arhgap11a
|
Rho GTPase activating protein 11A |
| chr21_+_8341774 | 2.20 |
ENSDART00000129749
ENSDART00000055325 ENSDART00000133804 |
psmb7
|
proteasome subunit beta 7 |
| chr12_-_21684197 | 2.18 |
ENSDART00000152999
ENSDART00000153109 ENSDART00000148698 |
eme1
|
essential meiotic structure-specific endonuclease 1 |
| chr22_-_24984733 | 2.18 |
ENSDART00000142147
ENSDART00000187284 |
dnal4b
|
dynein, axonemal, light chain 4b |
| chr6_-_32093830 | 2.16 |
ENSDART00000017695
|
foxd3
|
forkhead box D3 |
| chr19_-_24555935 | 2.15 |
ENSDART00000132660
ENSDART00000162801 |
polr3gla
|
polymerase (RNA) III (DNA directed) polypeptide G like a |
| chr25_+_28555584 | 2.14 |
ENSDART00000157046
|
SLC15A5
|
si:ch211-190o6.3 |
| chr1_-_19764038 | 2.13 |
ENSDART00000005733
|
tma16
|
translation machinery associated 16 homolog |
| chr1_+_14454663 | 2.12 |
ENSDART00000005067
|
rbpja
|
recombination signal binding protein for immunoglobulin kappa J region a |
| chr25_+_19095231 | 2.12 |
ENSDART00000154066
|
isg20
|
interferon stimulated exonuclease gene |
| chr25_+_21098675 | 2.10 |
ENSDART00000079012
|
rad52
|
RAD52 homolog, DNA repair protein |
| chr25_+_21098990 | 2.10 |
ENSDART00000017488
|
rad52
|
RAD52 homolog, DNA repair protein |
| chr16_-_17251898 | 2.10 |
ENSDART00000114272
|
nop2
|
NOP2 nucleolar protein homolog (yeast) |
| chr19_-_47555956 | 2.08 |
ENSDART00000114549
|
actc1a
|
actin, alpha, cardiac muscle 1a |
| chr23_+_17417539 | 2.07 |
ENSDART00000182605
|
BX649300.2
|
|
| chr22_-_12726556 | 2.05 |
ENSDART00000105776
|
fam207a
|
family with sequence similarity 207, member A |
| chr7_+_41812190 | 2.05 |
ENSDART00000113732
ENSDART00000174137 |
orc6
|
origin recognition complex, subunit 6 |
| chr7_+_14536083 | 2.05 |
ENSDART00000027331
|
mrpl46
|
mitochondrial ribosomal protein L46 |
| chr14_+_16151636 | 2.05 |
ENSDART00000159352
|
polr1a
|
polymerase (RNA) I polypeptide A |
| chr16_-_31919568 | 2.04 |
ENSDART00000027364
|
rbfox1l
|
RNA binding fox-1 homolog 1, like |
| chr13_-_25199260 | 2.04 |
ENSDART00000057605
|
adka
|
adenosine kinase a |
| chr2_+_3044992 | 2.04 |
ENSDART00000020463
|
zgc:63882
|
zgc:63882 |
| chr2_+_25839193 | 2.03 |
ENSDART00000078634
|
eif5a2
|
eukaryotic translation initiation factor 5A2 |
| chr14_-_25985698 | 2.03 |
ENSDART00000172909
ENSDART00000123053 |
atox1
|
antioxidant 1 copper chaperone |
| chr7_-_29164818 | 2.03 |
ENSDART00000052348
|
exosc6
|
exosome component 6 |
| chr3_-_7296189 | 2.03 |
ENSDART00000148349
|
si:ch1073-186o8.3
|
si:ch1073-186o8.3 |
| chr4_-_4755622 | 2.00 |
ENSDART00000193929
|
ssbp1
|
single-stranded DNA binding protein 1 |
| chr4_+_18442941 | 2.00 |
ENSDART00000190819
|
ncaph2
|
non-SMC condensin II complex, subunit H2 |
| chr7_-_30779575 | 1.98 |
ENSDART00000004782
|
mphosph10
|
M-phase phosphoprotein 10 (U3 small nucleolar ribonucleoprotein) |
| chr21_+_21611867 | 1.98 |
ENSDART00000189148
|
b9d2
|
B9 domain containing 2 |
| chr21_-_30408775 | 1.98 |
ENSDART00000101037
|
nhp2
|
NHP2 ribonucleoprotein homolog (yeast) |
| chr23_-_3759345 | 1.96 |
ENSDART00000132205
ENSDART00000137707 ENSDART00000189382 |
hmga1a
|
high mobility group AT-hook 1a |
| chr2_+_20967673 | 1.94 |
ENSDART00000057174
|
arpc5a
|
actin related protein 2/3 complex, subunit 5A |
| chr9_+_24920677 | 1.92 |
ENSDART00000037025
|
slc39a10
|
solute carrier family 39 (zinc transporter), member 10 |
| chr20_+_25626479 | 1.91 |
ENSDART00000143883
|
ppat
|
phosphoribosyl pyrophosphate amidotransferase |
| chr12_-_28349026 | 1.90 |
ENSDART00000183768
ENSDART00000152998 |
zgc:195081
|
zgc:195081 |
| chr13_+_8892784 | 1.88 |
ENSDART00000075054
ENSDART00000143705 |
thada
|
thyroid adenoma associated |
| chr7_-_57637779 | 1.87 |
ENSDART00000028017
|
mad2l1
|
MAD2 mitotic arrest deficient-like 1 (yeast) |
| chr15_+_36309070 | 1.87 |
ENSDART00000157034
|
gmnc
|
geminin coiled-coil domain containing |
| chr25_-_14424406 | 1.87 |
ENSDART00000073609
|
prmt7
|
protein arginine methyltransferase 7 |
| chr11_-_6206520 | 1.86 |
ENSDART00000150199
ENSDART00000148246 ENSDART00000019440 |
pole4
|
polymerase (DNA-directed), epsilon 4, accessory subunit |
| chr9_-_29985390 | 1.86 |
ENSDART00000134157
|
il1rapl1a
|
interleukin 1 receptor accessory protein-like 1a |
| chr1_-_26675969 | 1.85 |
ENSDART00000054184
|
trmo
|
tRNA methyltransferase O |
| chr23_+_7548797 | 1.84 |
ENSDART00000006765
|
tm9sf4
|
transmembrane 9 superfamily protein member 4 |
| chr16_+_16968682 | 1.82 |
ENSDART00000111074
|
si:ch211-120k19.1
|
si:ch211-120k19.1 |
| chr5_+_38462121 | 1.81 |
ENSDART00000144425
|
gltpd2
|
glycolipid transfer protein domain containing 2 |
| chr1_+_43686251 | 1.81 |
ENSDART00000074604
ENSDART00000137791 |
cisd2
|
CDGSH iron sulfur domain 2 |
| chr3_+_54581987 | 1.80 |
ENSDART00000018071
|
eif3g
|
eukaryotic translation initiation factor 3, subunit G |
| chr16_-_14074594 | 1.80 |
ENSDART00000090234
|
trim109
|
tripartite motif containing 109 |
| chr17_-_15189397 | 1.80 |
ENSDART00000133710
ENSDART00000110507 |
wdhd1
|
WD repeat and HMG-box DNA binding protein 1 |
| chr5_-_45958838 | 1.79 |
ENSDART00000135072
|
poc5
|
POC5 centriolar protein homolog (Chlamydomonas) |
| chr5_+_60847823 | 1.79 |
ENSDART00000074426
|
lig3
|
ligase III, DNA, ATP-dependent |
| chr3_+_15893039 | 1.78 |
ENSDART00000055780
|
jpt2
|
Jupiter microtubule associated homolog 2 |
| chr10_-_15340362 | 1.78 |
ENSDART00000148119
ENSDART00000127277 ENSDART00000154037 ENSDART00000189109 |
pum3
|
pumilio RNA-binding family member 3 |
| chr13_+_22555342 | 1.77 |
ENSDART00000193633
|
bmpr1aa
|
bone morphogenetic protein receptor, type IAa |
| chr3_-_42086577 | 1.76 |
ENSDART00000083111
ENSDART00000187312 |
ttyh3a
|
tweety family member 3a |
| chr8_-_17184482 | 1.76 |
ENSDART00000025803
|
pola2
|
polymerase (DNA directed), alpha 2 |
| chr12_+_29240124 | 1.75 |
ENSDART00000053761
ENSDART00000130172 |
bms1
|
BMS1 ribosome biogenesis factor |
| chr1_-_157563 | 1.75 |
ENSDART00000018741
|
pcid2
|
PCI domain containing 2 |
| chr16_+_16969060 | 1.70 |
ENSDART00000182819
ENSDART00000191876 |
si:ch211-120k19.1
rpl18
|
si:ch211-120k19.1 ribosomal protein L18 |
| chr14_+_36896689 | 1.69 |
ENSDART00000168684
ENSDART00000187827 ENSDART00000182511 |
LECT2 (1 of many)
|
si:ch211-132p1.4 |
| chr3_-_3398383 | 1.69 |
ENSDART00000047865
|
si:dkey-46g23.2
|
si:dkey-46g23.2 |
| chr5_+_72194444 | 1.69 |
ENSDART00000165436
|
ddx54
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 54 |
| chr20_+_42537768 | 1.69 |
ENSDART00000134066
ENSDART00000153434 |
si:dkeyp-93d12.1
|
si:dkeyp-93d12.1 |
| chr15_-_34056733 | 1.68 |
ENSDART00000170130
ENSDART00000188272 |
VWDE
|
si:dkey-30e9.7 |
| chr15_-_2519640 | 1.68 |
ENSDART00000047013
|
srprb
|
signal recognition particle receptor, B subunit |
| chr19_+_20201254 | 1.68 |
ENSDART00000010140
|
igf2bp3
|
insulin-like growth factor 2 mRNA binding protein 3 |
| chr13_+_6189203 | 1.67 |
ENSDART00000109665
|
ppm1g
|
protein phosphatase, Mg2+/Mn2+ dependent, 1G |
| chr20_-_29483514 | 1.67 |
ENSDART00000062370
|
actc1a
|
actin, alpha, cardiac muscle 1a |
| chr3_-_7464250 | 1.67 |
ENSDART00000159873
|
znf1001
|
zinc finger protein 1001 |
| chr5_+_24543862 | 1.66 |
ENSDART00000029699
|
atp6v0a2b
|
ATPase H+ transporting V0 subunit a2b |
| chr3_+_6443992 | 1.65 |
ENSDART00000169325
ENSDART00000162255 |
nup85
|
nucleoporin 85 |
| chr25_+_30442389 | 1.64 |
ENSDART00000003346
|
pdcd2l
|
programmed cell death 2-like |
| chr5_+_65040228 | 1.64 |
ENSDART00000164278
|
pmpca
|
peptidase (mitochondrial processing) alpha |
| chr20_-_3310017 | 1.64 |
ENSDART00000099315
|
cebpz
|
CCAAT/enhancer binding protein (C/EBP), zeta |
| chr3_-_58165254 | 1.63 |
ENSDART00000093031
|
snu13a
|
SNU13 homolog, small nuclear ribonucleoprotein a (U4/U6.U5) |
| chr5_-_8817458 | 1.61 |
ENSDART00000191098
|
fgf10b
|
fibroblast growth factor 10b |
| chr19_+_2685779 | 1.60 |
ENSDART00000160533
ENSDART00000097531 |
tomm7
|
translocase of outer mitochondrial membrane 7 homolog (yeast) |
| chr7_-_14535862 | 1.59 |
ENSDART00000112980
ENSDART00000135446 |
mrps11
|
mitochondrial ribosomal protein S11 |
| chr5_+_29714786 | 1.58 |
ENSDART00000148314
|
ddx31
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 31 |
| chr15_+_5116179 | 1.58 |
ENSDART00000101937
|
pgm2l1
|
phosphoglucomutase 2-like 1 |
| chr7_+_72012397 | 1.58 |
ENSDART00000011303
|
ywhae2
|
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, epsilon polypeptide 2 |
| chr9_-_1483879 | 1.57 |
ENSDART00000093415
|
rbm45
|
RNA binding motif protein 45 |
| chr3_-_40162843 | 1.56 |
ENSDART00000129664
ENSDART00000025285 |
drg2
|
developmentally regulated GTP binding protein 2 |
| chr3_-_1387292 | 1.56 |
ENSDART00000163535
|
ddx47
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 47 |
| chr7_+_65296901 | 1.55 |
ENSDART00000162693
ENSDART00000161609 |
bola3
|
bolA family member 3 |
| chr17_+_53156530 | 1.55 |
ENSDART00000126277
ENSDART00000156774 |
dph6
|
diphthamine biosynthesis 6 |
| chr18_+_184746 | 1.54 |
ENSDART00000140897
|
larp6a
|
La ribonucleoprotein domain family, member 6a |
| chr4_-_5455506 | 1.53 |
ENSDART00000156593
ENSDART00000154676 |
si:dkey-14d8.22
|
si:dkey-14d8.22 |
| chr21_-_929448 | 1.52 |
ENSDART00000133976
|
txnl1
|
thioredoxin-like 1 |
| chr24_-_36175365 | 1.52 |
ENSDART00000065338
|
pak1ip1
|
PAK1 interacting protein 1 |
| chr6_+_9893554 | 1.51 |
ENSDART00000064979
|
pimr74
|
Pim proto-oncogene, serine/threonine kinase, related 74 |
| chr6_-_18531760 | 1.51 |
ENSDART00000167167
|
utp6
|
UTP6, small subunit (SSU) processome component, homolog (yeast) |
| chr25_-_6292560 | 1.51 |
ENSDART00000153496
|
wdr61
|
WD repeat domain 61 |
| chr9_-_27719998 | 1.51 |
ENSDART00000161068
ENSDART00000148195 ENSDART00000138386 |
gtf2e1
|
general transcription factor IIE, polypeptide 1, alpha |
| chr16_+_25761101 | 1.49 |
ENSDART00000110619
|
cblc
|
Cbl proto-oncogene C, E3 ubiquitin protein ligase |
| chr21_+_18274825 | 1.48 |
ENSDART00000144322
ENSDART00000147768 |
wdr5
|
WD repeat domain 5 |
| chr13_+_39277178 | 1.48 |
ENSDART00000113259
|
si:dkey-85a20.4
|
si:dkey-85a20.4 |
| chr7_+_30202104 | 1.47 |
ENSDART00000173525
|
wdr76
|
WD repeat domain 76 |
| chr15_+_7086327 | 1.47 |
ENSDART00000114560
|
pik3cb
|
phosphatidylinositol-4,5-bisphosphate 3-kinase, catalytic subunit beta |
| chr21_+_3796620 | 1.47 |
ENSDART00000099535
ENSDART00000144515 |
spout1
|
SPOUT domain containing methyltransferase 1 |
| chr14_+_16034447 | 1.46 |
ENSDART00000161348
|
prelid1a
|
PRELI domain containing 1a |
| chr25_+_245438 | 1.46 |
ENSDART00000004689
|
zgc:92481
|
zgc:92481 |
| chr8_+_17184602 | 1.45 |
ENSDART00000050228
ENSDART00000140531 |
dimt1l
|
DIM1 dimethyladenosine transferase 1-like (S. cerevisiae) |
| chr12_-_34827477 | 1.45 |
ENSDART00000153026
|
NDUFAF8
|
si:dkey-21c1.6 |
Network of associatons between targets according to the STRING database.
First level regulatory network of clocka+clockb+npas2
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.7 | 8.2 | GO:0031120 | snRNA pseudouridine synthesis(GO:0031120) |
| 2.5 | 7.4 | GO:0006480 | N-terminal protein amino acid methylation(GO:0006480) |
| 1.9 | 7.5 | GO:0044210 | 'de novo' CTP biosynthetic process(GO:0044210) |
| 1.6 | 7.9 | GO:0044205 | 'de novo' UMP biosynthetic process(GO:0044205) |
| 1.5 | 4.5 | GO:0051095 | regulation of helicase activity(GO:0051095) positive regulation of helicase activity(GO:0051096) |
| 1.5 | 10.4 | GO:0045901 | positive regulation of translational elongation(GO:0045901) positive regulation of translational termination(GO:0045905) |
| 1.4 | 4.2 | GO:0045002 | double-strand break repair via single-strand annealing(GO:0045002) |
| 1.4 | 13.7 | GO:0006189 | 'de novo' IMP biosynthetic process(GO:0006189) |
| 1.2 | 10.9 | GO:0000056 | ribosomal small subunit export from nucleus(GO:0000056) |
| 1.1 | 6.5 | GO:0045721 | negative regulation of gluconeogenesis(GO:0045721) |
| 0.9 | 2.8 | GO:0001113 | DNA-templated transcriptional open complex formation(GO:0001112) transcriptional open complex formation at RNA polymerase II promoter(GO:0001113) protein-DNA complex remodeling(GO:0001120) |
| 0.9 | 2.7 | GO:1902626 | mature ribosome assembly(GO:0042256) assembly of large subunit precursor of preribosome(GO:1902626) |
| 0.9 | 4.5 | GO:0009052 | pentose-phosphate shunt, non-oxidative branch(GO:0009052) |
| 0.8 | 4.1 | GO:2000001 | regulation of DNA damage checkpoint(GO:2000001) |
| 0.7 | 3.0 | GO:0071962 | mitotic sister chromatid cohesion, centromeric(GO:0071962) |
| 0.7 | 3.0 | GO:1990481 | mRNA pseudouridine synthesis(GO:1990481) |
| 0.7 | 2.2 | GO:0031591 | wybutosine metabolic process(GO:0031590) wybutosine biosynthetic process(GO:0031591) |
| 0.7 | 2.0 | GO:0071051 | U4 snRNA 3'-end processing(GO:0034475) polyadenylation-dependent snoRNA 3'-end processing(GO:0071051) |
| 0.7 | 6.8 | GO:0046037 | GMP biosynthetic process(GO:0006177) GMP metabolic process(GO:0046037) |
| 0.7 | 2.6 | GO:0019401 | glycerol biosynthetic process(GO:0006114) alditol biosynthetic process(GO:0019401) short-chain fatty acid catabolic process(GO:0019626) response to dexamethasone(GO:0071548) |
| 0.6 | 4.5 | GO:1901673 | regulation of mitotic spindle assembly(GO:1901673) |
| 0.6 | 1.9 | GO:0050861 | positive regulation of B cell receptor signaling pathway(GO:0050861) |
| 0.6 | 3.7 | GO:0003272 | endocardial cushion formation(GO:0003272) |
| 0.6 | 1.9 | GO:0071514 | genetic imprinting(GO:0071514) |
| 0.6 | 1.8 | GO:0043504 | mitochondrial DNA repair(GO:0043504) |
| 0.6 | 1.8 | GO:0060912 | cardiac cell fate specification(GO:0060912) |
| 0.6 | 1.7 | GO:0000973 | posttranscriptional tethering of RNA polymerase II gene DNA at nuclear periphery(GO:0000973) |
| 0.5 | 2.1 | GO:0072314 | glomerular visceral epithelial cell fate commitment(GO:0072149) glomerular epithelial cell fate commitment(GO:0072314) |
| 0.5 | 1.1 | GO:0000469 | cleavage involved in rRNA processing(GO:0000469) |
| 0.5 | 6.7 | GO:0009303 | rRNA transcription(GO:0009303) |
| 0.5 | 2.0 | GO:0010032 | meiotic chromosome condensation(GO:0010032) |
| 0.5 | 2.4 | GO:0003100 | regulation of systemic arterial blood pressure by endothelin(GO:0003100) vein smooth muscle contraction(GO:0014826) regulation of prostaglandin-endoperoxide synthase activity(GO:0060584) positive regulation of prostaglandin-endoperoxide synthase activity(GO:0060585) |
| 0.5 | 2.8 | GO:0045943 | regulation of transcription from RNA polymerase I promoter(GO:0006356) positive regulation of transcription from RNA polymerase I promoter(GO:0045943) |
| 0.5 | 1.8 | GO:0010259 | multicellular organism aging(GO:0010259) |
| 0.4 | 1.3 | GO:0019408 | dolichol biosynthetic process(GO:0019408) |
| 0.4 | 2.2 | GO:0097066 | response to thyroid hormone(GO:0097066) |
| 0.4 | 1.3 | GO:0070914 | UV-damage excision repair(GO:0070914) |
| 0.4 | 1.3 | GO:0002188 | translation reinitiation(GO:0002188) |
| 0.4 | 1.6 | GO:0060437 | lung growth(GO:0060437) secretion by lung epithelial cell involved in lung growth(GO:0061033) |
| 0.4 | 13.7 | GO:0006270 | DNA replication initiation(GO:0006270) |
| 0.4 | 7.5 | GO:0000466 | maturation of 5.8S rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000466) |
| 0.4 | 0.8 | GO:0033314 | mitotic DNA replication checkpoint(GO:0033314) |
| 0.4 | 3.1 | GO:0036372 | opsin transport(GO:0036372) |
| 0.4 | 2.6 | GO:0000290 | deadenylation-dependent decapping of nuclear-transcribed mRNA(GO:0000290) |
| 0.4 | 1.1 | GO:0098581 | detection of biotic stimulus(GO:0009595) detection of bacterium(GO:0016045) detection of other organism(GO:0098543) detection of external biotic stimulus(GO:0098581) |
| 0.4 | 1.5 | GO:0051661 | maintenance of centrosome location(GO:0051661) |
| 0.4 | 1.1 | GO:0000390 | spliceosomal complex disassembly(GO:0000390) |
| 0.4 | 1.4 | GO:1903292 | protein localization to Golgi membrane(GO:1903292) |
| 0.4 | 6.4 | GO:0030150 | protein import into mitochondrial matrix(GO:0030150) |
| 0.4 | 1.1 | GO:0033690 | positive regulation of osteoblast proliferation(GO:0033690) |
| 0.3 | 2.0 | GO:0044209 | AMP salvage(GO:0044209) |
| 0.3 | 10.5 | GO:0000462 | maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000462) |
| 0.3 | 3.0 | GO:0070475 | rRNA base methylation(GO:0070475) |
| 0.3 | 1.3 | GO:0048014 | Tie signaling pathway(GO:0048014) |
| 0.3 | 10.5 | GO:0030488 | tRNA methylation(GO:0030488) |
| 0.3 | 3.0 | GO:0033700 | phospholipid efflux(GO:0033700) |
| 0.3 | 2.0 | GO:0048739 | cardiac muscle fiber development(GO:0048739) |
| 0.3 | 0.9 | GO:0042823 | pyridoxal phosphate metabolic process(GO:0042822) pyridoxal phosphate biosynthetic process(GO:0042823) |
| 0.3 | 1.4 | GO:0030576 | nuclear body organization(GO:0030575) Cajal body organization(GO:0030576) |
| 0.3 | 1.4 | GO:0032776 | DNA methylation on cytosine(GO:0032776) |
| 0.3 | 1.1 | GO:0009957 | epidermal cell fate specification(GO:0009957) |
| 0.3 | 1.6 | GO:0017183 | peptidyl-diphthamide metabolic process(GO:0017182) peptidyl-diphthamide biosynthetic process from peptidyl-histidine(GO:0017183) |
| 0.3 | 2.6 | GO:0055070 | copper ion homeostasis(GO:0055070) |
| 0.3 | 1.3 | GO:0006107 | oxaloacetate metabolic process(GO:0006107) |
| 0.2 | 1.5 | GO:0007175 | negative regulation of epidermal growth factor-activated receptor activity(GO:0007175) |
| 0.2 | 1.7 | GO:0070572 | positive regulation of axon regeneration(GO:0048680) positive regulation of neuron projection regeneration(GO:0070572) |
| 0.2 | 2.8 | GO:0006607 | NLS-bearing protein import into nucleus(GO:0006607) |
| 0.2 | 3.9 | GO:0030490 | maturation of SSU-rRNA(GO:0030490) |
| 0.2 | 0.9 | GO:2000659 | regulation of interleukin-1-mediated signaling pathway(GO:2000659) negative regulation of interleukin-1-mediated signaling pathway(GO:2000660) |
| 0.2 | 1.4 | GO:0033183 | negative regulation of histone ubiquitination(GO:0033183) regulation of protein K63-linked ubiquitination(GO:1900044) negative regulation of protein K63-linked ubiquitination(GO:1900045) regulation of histone H2A K63-linked ubiquitination(GO:1901314) negative regulation of histone H2A K63-linked ubiquitination(GO:1901315) regulation of protein polyubiquitination(GO:1902914) negative regulation of protein polyubiquitination(GO:1902915) |
| 0.2 | 1.6 | GO:0006627 | protein processing involved in protein targeting to mitochondrion(GO:0006627) |
| 0.2 | 21.8 | GO:0006364 | rRNA processing(GO:0006364) |
| 0.2 | 3.2 | GO:0035999 | tetrahydrofolate interconversion(GO:0035999) |
| 0.2 | 0.4 | GO:0045117 | azole transport(GO:0045117) |
| 0.2 | 1.6 | GO:1902262 | apoptotic process involved in patterning of blood vessels(GO:1902262) |
| 0.2 | 0.8 | GO:1902946 | protein localization to early endosome(GO:1902946) |
| 0.2 | 2.2 | GO:0006285 | base-excision repair, AP site formation(GO:0006285) |
| 0.2 | 1.9 | GO:0045920 | negative regulation of exocytosis(GO:0045920) |
| 0.2 | 2.2 | GO:0031573 | intra-S DNA damage checkpoint(GO:0031573) |
| 0.2 | 1.1 | GO:0019184 | nonribosomal peptide biosynthetic process(GO:0019184) |
| 0.2 | 2.5 | GO:0045580 | regulation of T cell differentiation(GO:0045580) |
| 0.2 | 5.1 | GO:0006383 | transcription from RNA polymerase III promoter(GO:0006383) |
| 0.2 | 0.6 | GO:0045938 | positive regulation of circadian sleep/wake cycle, sleep(GO:0045938) |
| 0.2 | 1.3 | GO:0070933 | histone H4 deacetylation(GO:0070933) |
| 0.2 | 0.6 | GO:0089709 | L-histidine transmembrane transport(GO:0089709) L-histidine transport(GO:1902024) |
| 0.2 | 3.5 | GO:0035435 | phosphate ion transmembrane transport(GO:0035435) |
| 0.2 | 1.1 | GO:0045110 | intermediate filament bundle assembly(GO:0045110) |
| 0.1 | 12.3 | GO:0042254 | ribosome biogenesis(GO:0042254) |
| 0.1 | 0.4 | GO:0002407 | dendritic cell chemotaxis(GO:0002407) dendritic cell migration(GO:0036336) |
| 0.1 | 0.7 | GO:1903361 | protein localization to basolateral plasma membrane(GO:1903361) |
| 0.1 | 5.3 | GO:0030199 | collagen fibril organization(GO:0030199) |
| 0.1 | 0.7 | GO:0060561 | apoptotic process involved in morphogenesis(GO:0060561) |
| 0.1 | 0.9 | GO:0006283 | transcription-coupled nucleotide-excision repair(GO:0006283) |
| 0.1 | 1.2 | GO:0042987 | beta-amyloid formation(GO:0034205) amyloid precursor protein catabolic process(GO:0042987) |
| 0.1 | 1.4 | GO:0021754 | facial nucleus development(GO:0021754) |
| 0.1 | 1.1 | GO:1901998 | toxin transport(GO:1901998) |
| 0.1 | 1.0 | GO:0006348 | chromatin silencing at telomere(GO:0006348) |
| 0.1 | 3.2 | GO:0001732 | formation of cytoplasmic translation initiation complex(GO:0001732) |
| 0.1 | 0.5 | GO:0061469 | regulation of type B pancreatic cell proliferation(GO:0061469) |
| 0.1 | 3.3 | GO:0001757 | somite specification(GO:0001757) |
| 0.1 | 0.4 | GO:0036076 | ligamentous ossification(GO:0036076) |
| 0.1 | 1.7 | GO:0051452 | vacuolar acidification(GO:0007035) pH reduction(GO:0045851) intracellular pH reduction(GO:0051452) |
| 0.1 | 0.7 | GO:0006556 | S-adenosylmethionine biosynthetic process(GO:0006556) |
| 0.1 | 1.3 | GO:0038007 | netrin-activated signaling pathway(GO:0038007) |
| 0.1 | 1.0 | GO:0006337 | nucleosome disassembly(GO:0006337) chromatin disassembly(GO:0031498) protein-DNA complex disassembly(GO:0032986) |
| 0.1 | 0.3 | GO:1990575 | mitochondrial ornithine transport(GO:0000066) mitochondrial L-ornithine transmembrane transport(GO:1990575) |
| 0.1 | 1.4 | GO:0001678 | cellular glucose homeostasis(GO:0001678) |
| 0.1 | 0.7 | GO:0006542 | glutamine biosynthetic process(GO:0006542) |
| 0.1 | 3.2 | GO:0007286 | spermatid development(GO:0007286) |
| 0.1 | 0.8 | GO:0090520 | sphingolipid mediated signaling pathway(GO:0090520) |
| 0.1 | 5.2 | GO:0003333 | amino acid transmembrane transport(GO:0003333) |
| 0.1 | 0.7 | GO:0034389 | lipid particle organization(GO:0034389) |
| 0.1 | 0.2 | GO:0051973 | positive regulation of telomerase activity(GO:0051973) |
| 0.1 | 2.1 | GO:0034625 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.1 | 0.3 | GO:0035971 | peptidyl-histidine dephosphorylation(GO:0035971) |
| 0.1 | 0.4 | GO:0042754 | negative regulation of circadian rhythm(GO:0042754) |
| 0.1 | 1.2 | GO:0061026 | cardiac muscle tissue regeneration(GO:0061026) |
| 0.1 | 1.8 | GO:0008354 | germ cell migration(GO:0008354) |
| 0.1 | 3.5 | GO:0061077 | chaperone-mediated protein folding(GO:0061077) |
| 0.1 | 0.3 | GO:0071939 | vitamin A transport(GO:0071938) vitamin A import(GO:0071939) |
| 0.1 | 0.7 | GO:0044528 | regulation of mitochondrial mRNA stability(GO:0044528) |
| 0.1 | 0.4 | GO:0036498 | IRE1-mediated unfolded protein response(GO:0036498) |
| 0.1 | 5.8 | GO:0051028 | mRNA transport(GO:0051028) |
| 0.1 | 1.4 | GO:0032392 | DNA geometric change(GO:0032392) DNA duplex unwinding(GO:0032508) |
| 0.1 | 2.2 | GO:0010499 | proteasomal ubiquitin-independent protein catabolic process(GO:0010499) |
| 0.1 | 0.2 | GO:0090244 | Wnt signaling pathway involved in somitogenesis(GO:0090244) |
| 0.1 | 1.5 | GO:0036092 | phosphatidylinositol-3-phosphate biosynthetic process(GO:0036092) |
| 0.1 | 0.4 | GO:0071596 | ubiquitin-dependent protein catabolic process via the N-end rule pathway(GO:0071596) |
| 0.1 | 0.5 | GO:2000576 | positive regulation of microtubule motor activity(GO:2000576) regulation of ATP-dependent microtubule motor activity, plus-end-directed(GO:2000580) positive regulation of ATP-dependent microtubule motor activity, plus-end-directed(GO:2000582) |
| 0.1 | 1.5 | GO:0051568 | histone H3-K4 methylation(GO:0051568) |
| 0.1 | 0.6 | GO:0021588 | cerebellum formation(GO:0021588) |
| 0.1 | 1.3 | GO:0021555 | midbrain-hindbrain boundary morphogenesis(GO:0021555) |
| 0.1 | 1.7 | GO:0072599 | protein targeting to ER(GO:0045047) establishment of protein localization to endoplasmic reticulum(GO:0072599) |
| 0.1 | 1.5 | GO:0032968 | positive regulation of transcription elongation from RNA polymerase II promoter(GO:0032968) |
| 0.1 | 1.1 | GO:0033173 | calcineurin-NFAT signaling cascade(GO:0033173) |
| 0.1 | 0.6 | GO:0006787 | porphyrin-containing compound catabolic process(GO:0006787) tetrapyrrole catabolic process(GO:0033015) heme catabolic process(GO:0042167) pigment catabolic process(GO:0046149) |
| 0.1 | 1.2 | GO:0043001 | Golgi to plasma membrane protein transport(GO:0043001) |
| 0.1 | 0.3 | GO:1904105 | positive regulation of convergent extension involved in gastrulation(GO:1904105) |
| 0.1 | 0.9 | GO:0032094 | response to food(GO:0032094) |
| 0.1 | 2.7 | GO:0010212 | response to ionizing radiation(GO:0010212) |
| 0.1 | 3.1 | GO:0007019 | microtubule depolymerization(GO:0007019) |
| 0.1 | 0.3 | GO:0046952 | ketone body catabolic process(GO:0046952) |
| 0.1 | 1.8 | GO:0007632 | visual behavior(GO:0007632) |
| 0.1 | 0.4 | GO:0032447 | tRNA wobble position uridine thiolation(GO:0002143) protein urmylation(GO:0032447) |
| 0.1 | 2.9 | GO:0002181 | cytoplasmic translation(GO:0002181) |
| 0.1 | 4.2 | GO:0030433 | ER-associated ubiquitin-dependent protein catabolic process(GO:0030433) |
| 0.1 | 1.0 | GO:0038061 | NIK/NF-kappaB signaling(GO:0038061) |
| 0.1 | 1.3 | GO:0090557 | establishment of endothelial intestinal barrier(GO:0090557) |
| 0.1 | 0.2 | GO:2000303 | regulation of sphingolipid biosynthetic process(GO:0090153) negative regulation of sphingolipid biosynthetic process(GO:0090155) cellular sphingolipid homeostasis(GO:0090156) negative regulation of ceramide biosynthetic process(GO:1900060) regulation of membrane lipid metabolic process(GO:1905038) regulation of ceramide biosynthetic process(GO:2000303) |
| 0.1 | 0.2 | GO:0019287 | isopentenyl diphosphate biosynthetic process, mevalonate pathway(GO:0019287) |
| 0.1 | 0.9 | GO:0033617 | mitochondrial respiratory chain complex IV assembly(GO:0033617) mitochondrial respiratory chain complex IV biogenesis(GO:0097034) |
| 0.1 | 1.1 | GO:0043535 | regulation of blood vessel endothelial cell migration(GO:0043535) |
| 0.1 | 2.3 | GO:0046330 | positive regulation of JNK cascade(GO:0046330) |
| 0.1 | 0.7 | GO:0060729 | maintenance of gastrointestinal epithelium(GO:0030277) intestinal epithelial structure maintenance(GO:0060729) |
| 0.1 | 2.4 | GO:0030901 | midbrain development(GO:0030901) |
| 0.1 | 5.6 | GO:0006401 | RNA catabolic process(GO:0006401) |
| 0.1 | 0.2 | GO:0006295 | nucleotide-excision repair, DNA incision, 3'-to lesion(GO:0006295) |
| 0.1 | 0.2 | GO:0006116 | NADH oxidation(GO:0006116) glycerol-3-phosphate catabolic process(GO:0046168) |
| 0.0 | 3.5 | GO:0007088 | regulation of mitotic nuclear division(GO:0007088) |
| 0.0 | 1.1 | GO:0043049 | otic placode formation(GO:0043049) |
| 0.0 | 0.4 | GO:0046477 | glycosylceramide catabolic process(GO:0046477) |
| 0.0 | 0.7 | GO:0045747 | positive regulation of Notch signaling pathway(GO:0045747) |
| 0.0 | 0.8 | GO:0034472 | snRNA 3'-end processing(GO:0034472) |
| 0.0 | 1.4 | GO:0032543 | mitochondrial translation(GO:0032543) |
| 0.0 | 1.0 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.0 | 1.1 | GO:0060059 | embryonic retina morphogenesis in camera-type eye(GO:0060059) |
| 0.0 | 1.4 | GO:0006367 | transcription initiation from RNA polymerase II promoter(GO:0006367) |
| 0.0 | 2.9 | GO:0006413 | translational initiation(GO:0006413) |
| 0.0 | 0.7 | GO:0070536 | protein K63-linked deubiquitination(GO:0070536) |
| 0.0 | 0.4 | GO:0001522 | pseudouridine synthesis(GO:0001522) |
| 0.0 | 1.4 | GO:0035987 | endodermal cell differentiation(GO:0035987) |
| 0.0 | 0.8 | GO:0002377 | immunoglobulin production(GO:0002377) |
| 0.0 | 1.8 | GO:0015914 | phospholipid transport(GO:0015914) |
| 0.0 | 0.9 | GO:0014866 | skeletal myofibril assembly(GO:0014866) |
| 0.0 | 0.3 | GO:0050957 | equilibrioception(GO:0050957) |
| 0.0 | 0.9 | GO:0000387 | spliceosomal snRNP assembly(GO:0000387) |
| 0.0 | 0.3 | GO:0021694 | cerebellar Purkinje cell layer formation(GO:0021694) cerebellar Purkinje cell differentiation(GO:0021702) |
| 0.0 | 0.8 | GO:0006509 | membrane protein ectodomain proteolysis(GO:0006509) |
| 0.0 | 1.8 | GO:0006261 | DNA-dependent DNA replication(GO:0006261) |
| 0.0 | 0.5 | GO:0006003 | fructose 2,6-bisphosphate metabolic process(GO:0006003) |
| 0.0 | 0.7 | GO:0030032 | lamellipodium assembly(GO:0030032) |
| 0.0 | 0.7 | GO:0045494 | photoreceptor cell maintenance(GO:0045494) |
| 0.0 | 0.1 | GO:0051012 | microtubule sliding(GO:0051012) |
| 0.0 | 0.1 | GO:0021680 | cerebellar Purkinje cell layer development(GO:0021680) |
| 0.0 | 0.5 | GO:0060287 | epithelial cilium movement involved in determination of left/right asymmetry(GO:0060287) |
| 0.0 | 0.7 | GO:0035329 | hippo signaling(GO:0035329) |
| 0.0 | 0.3 | GO:0060385 | axonogenesis involved in innervation(GO:0060385) |
| 0.0 | 0.7 | GO:0006099 | tricarboxylic acid cycle(GO:0006099) |
| 0.0 | 0.6 | GO:0045454 | cell redox homeostasis(GO:0045454) |
| 0.0 | 0.8 | GO:0016126 | sterol biosynthetic process(GO:0016126) |
| 0.0 | 0.1 | GO:0006369 | termination of RNA polymerase II transcription(GO:0006369) |
| 0.0 | 2.3 | GO:0010466 | negative regulation of peptidase activity(GO:0010466) |
| 0.0 | 12.6 | GO:0006396 | RNA processing(GO:0006396) |
| 0.0 | 0.6 | GO:0035305 | negative regulation of dephosphorylation(GO:0035305) negative regulation of protein dephosphorylation(GO:0035308) |
| 0.0 | 0.4 | GO:0000077 | DNA damage checkpoint(GO:0000077) DNA integrity checkpoint(GO:0031570) |
| 0.0 | 7.7 | GO:0043066 | negative regulation of apoptotic process(GO:0043066) |
| 0.0 | 0.9 | GO:0005978 | glycogen biosynthetic process(GO:0005978) glucan biosynthetic process(GO:0009250) |
| 0.0 | 0.3 | GO:0043486 | histone exchange(GO:0043486) |
| 0.0 | 0.4 | GO:0006491 | N-glycan processing(GO:0006491) |
| 0.0 | 0.6 | GO:0060972 | left/right pattern formation(GO:0060972) |
| 0.0 | 0.2 | GO:0006265 | DNA topological change(GO:0006265) |
| 0.0 | 0.6 | GO:0043123 | positive regulation of I-kappaB kinase/NF-kappaB signaling(GO:0043123) |
| 0.0 | 0.1 | GO:0015862 | uridine transport(GO:0015862) pyrimidine nucleoside transport(GO:0015864) |
| 0.0 | 0.7 | GO:0002028 | regulation of sodium ion transport(GO:0002028) |
| 0.0 | 2.6 | GO:0006412 | translation(GO:0006412) |
| 0.0 | 0.4 | GO:0072112 | renal filtration cell differentiation(GO:0061318) glomerular epithelium development(GO:0072010) glomerular visceral epithelial cell development(GO:0072015) glomerular visceral epithelial cell differentiation(GO:0072112) glomerular epithelial cell development(GO:0072310) glomerular epithelial cell differentiation(GO:0072311) |
| 0.0 | 1.8 | GO:0001755 | neural crest cell migration(GO:0001755) |
| 0.0 | 0.6 | GO:0018107 | peptidyl-threonine phosphorylation(GO:0018107) |
| 0.0 | 0.3 | GO:0019835 | cytolysis(GO:0019835) |
| 0.0 | 0.5 | GO:0030050 | vesicle transport along actin filament(GO:0030050) actin filament-based transport(GO:0099515) |
| 0.0 | 2.6 | GO:0006470 | protein dephosphorylation(GO:0006470) |
| 0.0 | 0.1 | GO:0015740 | C4-dicarboxylate transport(GO:0015740) acidic amino acid transport(GO:0015800) aspartate transport(GO:0015810) L-glutamate transport(GO:0015813) malate-aspartate shuttle(GO:0043490) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.0 | 8.2 | GO:0072588 | box H/ACA snoRNP complex(GO:0031429) box H/ACA RNP complex(GO:0072588) |
| 1.9 | 7.5 | GO:0097268 | cytoophidium(GO:0097268) |
| 1.3 | 6.7 | GO:0031515 | tRNA (m1A) methyltransferase complex(GO:0031515) |
| 1.2 | 4.8 | GO:0032299 | ribonuclease H2 complex(GO:0032299) |
| 0.9 | 2.8 | GO:0070545 | PeBoW complex(GO:0070545) |
| 0.8 | 3.1 | GO:0061673 | mitotic spindle astral microtubule(GO:0061673) |
| 0.7 | 9.0 | GO:0000808 | origin recognition complex(GO:0000808) nuclear origin of replication recognition complex(GO:0005664) |
| 0.7 | 2.0 | GO:0034457 | Mpp10 complex(GO:0034457) |
| 0.6 | 4.2 | GO:0034388 | Pwp2p-containing subcomplex of 90S preribosome(GO:0034388) |
| 0.6 | 4.5 | GO:0009295 | nucleoid(GO:0009295) mitochondrial nucleoid(GO:0042645) |
| 0.6 | 2.8 | GO:0005673 | transcription factor TFIIE complex(GO:0005673) |
| 0.5 | 1.9 | GO:0008622 | epsilon DNA polymerase complex(GO:0008622) |
| 0.5 | 5.4 | GO:0005736 | DNA-directed RNA polymerase I complex(GO:0005736) |
| 0.5 | 2.7 | GO:0070876 | SOSS complex(GO:0070876) |
| 0.4 | 7.5 | GO:0030686 | 90S preribosome(GO:0030686) |
| 0.4 | 5.7 | GO:0005666 | DNA-directed RNA polymerase III complex(GO:0005666) |
| 0.4 | 1.5 | GO:0035327 | transcriptionally active chromatin(GO:0035327) |
| 0.4 | 2.2 | GO:0048476 | Holliday junction resolvase complex(GO:0048476) |
| 0.4 | 4.0 | GO:0005742 | mitochondrial outer membrane translocase complex(GO:0005742) |
| 0.4 | 1.8 | GO:0070724 | BMP receptor complex(GO:0070724) |
| 0.4 | 1.8 | GO:0005658 | alpha DNA polymerase:primase complex(GO:0005658) |
| 0.3 | 1.7 | GO:0070390 | transcription export complex 2(GO:0070390) |
| 0.3 | 1.7 | GO:0000839 | Hrd1p ubiquitin ligase ERAD-L complex(GO:0000839) |
| 0.3 | 1.4 | GO:0005672 | transcription factor TFIIA complex(GO:0005672) |
| 0.3 | 1.4 | GO:0005655 | nucleolar ribonuclease P complex(GO:0005655) |
| 0.3 | 1.4 | GO:0001405 | presequence translocase-associated import motor(GO:0001405) |
| 0.3 | 1.3 | GO:0000110 | nucleotide-excision repair factor 1 complex(GO:0000110) |
| 0.3 | 1.6 | GO:0031428 | box C/D snoRNP complex(GO:0031428) |
| 0.2 | 2.0 | GO:0000796 | condensin complex(GO:0000796) |
| 0.2 | 0.7 | GO:0042709 | succinate-CoA ligase complex(GO:0042709) |
| 0.2 | 3.5 | GO:0071014 | post-mRNA release spliceosomal complex(GO:0071014) |
| 0.2 | 3.7 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.2 | 2.0 | GO:0030688 | preribosome, small subunit precursor(GO:0030688) |
| 0.2 | 2.0 | GO:0000177 | cytoplasmic exosome (RNase complex)(GO:0000177) |
| 0.2 | 12.0 | GO:0005643 | nuclear pore(GO:0005643) |
| 0.2 | 3.1 | GO:0036038 | MKS complex(GO:0036038) |
| 0.2 | 1.1 | GO:0071012 | U2-type catalytic step 1 spliceosome(GO:0071006) catalytic step 1 spliceosome(GO:0071012) |
| 0.2 | 49.3 | GO:0005730 | nucleolus(GO:0005730) |
| 0.2 | 1.5 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.2 | 0.7 | GO:0048269 | methionine adenosyltransferase complex(GO:0048269) |
| 0.2 | 0.7 | GO:0097025 | MPP7-DLG1-LIN7 complex(GO:0097025) |
| 0.2 | 1.7 | GO:0000220 | vacuolar proton-transporting V-type ATPase, V0 domain(GO:0000220) |
| 0.2 | 3.7 | GO:0005852 | eukaryotic translation initiation factor 3 complex(GO:0005852) |
| 0.2 | 1.1 | GO:0005744 | mitochondrial inner membrane presequence translocase complex(GO:0005744) |
| 0.1 | 2.1 | GO:0015030 | Cajal body(GO:0015030) |
| 0.1 | 2.5 | GO:0042555 | MCM complex(GO:0042555) |
| 0.1 | 0.4 | GO:1990604 | IRE1-TRAF2-ASK1 complex(GO:1990604) |
| 0.1 | 6.0 | GO:0000315 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.1 | 0.9 | GO:0032797 | SMN complex(GO:0032797) |
| 0.1 | 0.4 | GO:0033391 | chromatoid body(GO:0033391) |
| 0.1 | 1.8 | GO:0032391 | photoreceptor connecting cilium(GO:0032391) |
| 0.1 | 2.5 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.1 | 0.5 | GO:0043540 | 6-phosphofructo-2-kinase/fructose-2,6-biphosphatase complex(GO:0043540) |
| 0.1 | 1.9 | GO:0005885 | Arp2/3 protein complex(GO:0005885) |
| 0.1 | 3.4 | GO:0005791 | rough endoplasmic reticulum(GO:0005791) |
| 0.1 | 1.1 | GO:0045095 | keratin filament(GO:0045095) |
| 0.1 | 1.5 | GO:0048188 | Set1C/COMPASS complex(GO:0048188) |
| 0.1 | 1.8 | GO:0043596 | nuclear replication fork(GO:0043596) |
| 0.1 | 0.9 | GO:0005832 | chaperonin-containing T-complex(GO:0005832) |
| 0.1 | 1.1 | GO:0008250 | oligosaccharyltransferase complex(GO:0008250) |
| 0.1 | 2.2 | GO:0005839 | proteasome core complex(GO:0005839) |
| 0.1 | 2.2 | GO:0031672 | A band(GO:0031672) |
| 0.1 | 1.7 | GO:0071565 | nBAF complex(GO:0071565) |
| 0.1 | 1.4 | GO:0005669 | transcription factor TFIID complex(GO:0005669) |
| 0.1 | 2.1 | GO:0017053 | transcriptional repressor complex(GO:0017053) |
| 0.1 | 0.2 | GO:0035339 | SPOTS complex(GO:0035339) |
| 0.1 | 0.9 | GO:0016514 | SWI/SNF complex(GO:0016514) |
| 0.1 | 0.8 | GO:0071339 | MLL1/2 complex(GO:0044665) MLL1 complex(GO:0071339) |
| 0.1 | 0.6 | GO:0005689 | U12-type spliceosomal complex(GO:0005689) |
| 0.1 | 0.8 | GO:0033202 | Ino80 complex(GO:0031011) DNA helicase complex(GO:0033202) |
| 0.0 | 2.7 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.0 | 1.9 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.0 | 0.8 | GO:0032039 | integrator complex(GO:0032039) |
| 0.0 | 3.9 | GO:0000776 | kinetochore(GO:0000776) |
| 0.0 | 0.2 | GO:0030877 | beta-catenin destruction complex(GO:0030877) |
| 0.0 | 3.5 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.0 | 0.3 | GO:0097346 | Swr1 complex(GO:0000812) INO80-type complex(GO:0097346) |
| 0.0 | 0.9 | GO:0000313 | organellar ribosome(GO:0000313) mitochondrial ribosome(GO:0005761) |
| 0.0 | 0.2 | GO:0009331 | glycerol-3-phosphate dehydrogenase complex(GO:0009331) |
| 0.0 | 0.2 | GO:0005686 | U2 snRNP(GO:0005686) |
| 0.0 | 1.1 | GO:0071013 | catalytic step 2 spliceosome(GO:0071013) |
| 0.0 | 3.9 | GO:0043296 | apical junction complex(GO:0043296) |
| 0.0 | 0.6 | GO:0030131 | clathrin adaptor complex(GO:0030131) |
| 0.0 | 2.6 | GO:0005802 | trans-Golgi network(GO:0005802) |
| 0.0 | 0.7 | GO:0016605 | PML body(GO:0016605) |
| 0.0 | 0.3 | GO:0035869 | ciliary transition zone(GO:0035869) |
| 0.0 | 0.3 | GO:0031680 | G-protein beta/gamma-subunit complex(GO:0031680) |
| 0.0 | 1.8 | GO:0005840 | ribosome(GO:0005840) |
| 0.0 | 2.2 | GO:0030176 | integral component of endoplasmic reticulum membrane(GO:0030176) |
| 0.0 | 0.9 | GO:0005793 | endoplasmic reticulum-Golgi intermediate compartment(GO:0005793) |
| 0.0 | 0.2 | GO:0044545 | NSL complex(GO:0044545) |
| 0.0 | 0.6 | GO:0030864 | cortical actin cytoskeleton(GO:0030864) |
| 0.0 | 0.9 | GO:0005942 | phosphatidylinositol 3-kinase complex(GO:0005942) |
| 0.0 | 0.5 | GO:0005902 | microvillus(GO:0005902) |
| 0.0 | 0.9 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.0 | 13.5 | GO:0005829 | cytosol(GO:0005829) |
| 0.0 | 3.8 | GO:1990904 | ribonucleoprotein complex(GO:1990904) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.5 | 7.4 | GO:0071885 | N-terminal protein N-methyltransferase activity(GO:0071885) |
| 2.3 | 9.2 | GO:0004639 | phosphoribosylaminoimidazolesuccinocarboxamide synthase activity(GO:0004639) |
| 1.9 | 7.5 | GO:0003883 | CTP synthase activity(GO:0003883) |
| 1.5 | 4.5 | GO:0034513 | box H/ACA snoRNA binding(GO:0034513) |
| 1.4 | 9.7 | GO:0016426 | tRNA (adenine) methyltransferase activity(GO:0016426) |
| 1.1 | 3.2 | GO:0016743 | carboxyl- or carbamoyltransferase activity(GO:0016743) |
| 1.0 | 3.0 | GO:0016434 | rRNA (cytosine) methyltransferase activity(GO:0016434) |
| 0.9 | 5.7 | GO:0003938 | IMP dehydrogenase activity(GO:0003938) |
| 0.8 | 2.3 | GO:0072591 | N-acetyl-L-aspartate-L-glutamate ligase activity(GO:0072590) citrate-L-glutamate ligase activity(GO:0072591) |
| 0.7 | 2.7 | GO:0034511 | U3 snoRNA binding(GO:0034511) |
| 0.6 | 1.9 | GO:0035243 | protein-arginine omega-N symmetric methyltransferase activity(GO:0035243) |
| 0.6 | 1.9 | GO:0050135 | NAD(P)+ nucleosidase activity(GO:0050135) |
| 0.6 | 1.8 | GO:0003910 | DNA ligase activity(GO:0003909) DNA ligase (ATP) activity(GO:0003910) |
| 0.6 | 3.0 | GO:0090556 | phospholipid-translocating ATPase activity(GO:0004012) phosphatidylcholine-translocating ATPase activity(GO:0090554) phosphatidylserine-translocating ATPase activity(GO:0090556) |
| 0.5 | 1.5 | GO:0047690 | aspartyltransferase activity(GO:0047690) |
| 0.5 | 5.8 | GO:0019238 | cyclohydrolase activity(GO:0019238) |
| 0.5 | 2.4 | GO:0031708 | endothelin B receptor binding(GO:0031708) |
| 0.4 | 2.7 | GO:0043023 | ribosomal large subunit binding(GO:0043023) |
| 0.4 | 5.6 | GO:0030515 | snoRNA binding(GO:0030515) |
| 0.4 | 2.0 | GO:0004001 | adenosine kinase activity(GO:0004001) |
| 0.4 | 2.0 | GO:0016531 | copper chaperone activity(GO:0016531) |
| 0.4 | 2.6 | GO:0050072 | m7G(5')pppN diphosphatase activity(GO:0050072) |
| 0.4 | 1.1 | GO:0016019 | peptidoglycan receptor activity(GO:0016019) |
| 0.4 | 7.0 | GO:0009982 | pseudouridine synthase activity(GO:0009982) |
| 0.4 | 1.1 | GO:0004422 | hypoxanthine phosphoribosyltransferase activity(GO:0004422) |
| 0.3 | 1.4 | GO:0033204 | ribonuclease P RNA binding(GO:0033204) |
| 0.3 | 1.3 | GO:0050211 | procollagen galactosyltransferase activity(GO:0050211) |
| 0.3 | 1.3 | GO:0003865 | 3-oxo-5-alpha-steroid 4-dehydrogenase activity(GO:0003865) steroid dehydrogenase activity, acting on the CH-CH group of donors(GO:0033765) enone reductase activity(GO:0035671) cholestenone 5-alpha-reductase activity(GO:0047751) |
| 0.3 | 4.5 | GO:0016861 | intramolecular oxidoreductase activity, interconverting aldoses and ketoses(GO:0016861) |
| 0.3 | 1.5 | GO:0000179 | rRNA (adenine-N6,N6-)-dimethyltransferase activity(GO:0000179) |
| 0.3 | 1.2 | GO:0034416 | bisphosphoglycerate phosphatase activity(GO:0034416) |
| 0.3 | 4.6 | GO:0004724 | magnesium-dependent protein serine/threonine phosphatase activity(GO:0004724) |
| 0.3 | 1.4 | GO:0004565 | beta-galactosidase activity(GO:0004565) |
| 0.3 | 0.8 | GO:0017050 | D-erythro-sphingosine kinase activity(GO:0017050) |
| 0.3 | 1.1 | GO:0033745 | L-methionine-(R)-S-oxide reductase activity(GO:0033745) |
| 0.3 | 0.8 | GO:0003913 | DNA photolyase activity(GO:0003913) |
| 0.3 | 3.4 | GO:0008320 | protein transmembrane transporter activity(GO:0008320) |
| 0.3 | 0.8 | GO:1902945 | metalloendopeptidase activity involved in amyloid precursor protein catabolic process(GO:1902945) |
| 0.3 | 12.3 | GO:0043022 | ribosome binding(GO:0043022) |
| 0.2 | 2.2 | GO:0008263 | mismatch base pair DNA N-glycosylase activity(GO:0000700) pyrimidine-specific mismatch base pair DNA N-glycosylase activity(GO:0008263) |
| 0.2 | 0.7 | GO:0004775 | succinate-CoA ligase activity(GO:0004774) succinate-CoA ligase (ADP-forming) activity(GO:0004775) succinate-CoA ligase (GDP-forming) activity(GO:0004776) |
| 0.2 | 3.5 | GO:0051879 | Hsp90 protein binding(GO:0051879) |
| 0.2 | 1.4 | GO:0019158 | glucokinase activity(GO:0004340) hexokinase activity(GO:0004396) glucose binding(GO:0005536) fructokinase activity(GO:0008865) mannokinase activity(GO:0019158) |
| 0.2 | 3.5 | GO:0005315 | inorganic phosphate transmembrane transporter activity(GO:0005315) |
| 0.2 | 1.6 | GO:0070181 | small ribosomal subunit rRNA binding(GO:0070181) |
| 0.2 | 1.1 | GO:0043140 | ATP-dependent 3'-5' DNA helicase activity(GO:0043140) single-stranded DNA-dependent ATP-dependent 3'-5' DNA helicase activity(GO:1990518) |
| 0.2 | 0.9 | GO:0004373 | glycogen (starch) synthase activity(GO:0004373) |
| 0.2 | 0.4 | GO:1901474 | azole transporter activity(GO:0045118) azole transmembrane transporter activity(GO:1901474) |
| 0.2 | 2.1 | GO:0005049 | nuclear export signal receptor activity(GO:0005049) |
| 0.2 | 1.5 | GO:1990050 | phosphatidic acid transporter activity(GO:1990050) |
| 0.2 | 1.0 | GO:0004385 | guanylate kinase activity(GO:0004385) |
| 0.2 | 3.1 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 0.2 | 1.7 | GO:0072320 | volume-sensitive chloride channel activity(GO:0072320) |
| 0.2 | 1.1 | GO:0019215 | intermediate filament binding(GO:0019215) |
| 0.2 | 0.7 | GO:0004478 | methionine adenosyltransferase activity(GO:0004478) |
| 0.2 | 0.7 | GO:0097016 | L27 domain binding(GO:0097016) |
| 0.2 | 7.3 | GO:0016831 | carboxy-lyase activity(GO:0016831) |
| 0.2 | 4.7 | GO:0034062 | DNA-directed RNA polymerase activity(GO:0003899) RNA polymerase activity(GO:0034062) |
| 0.2 | 1.5 | GO:0004791 | thioredoxin-disulfide reductase activity(GO:0004791) |
| 0.2 | 2.8 | GO:0061608 | nuclear import signal receptor activity(GO:0061608) |
| 0.2 | 13.3 | GO:0003724 | RNA helicase activity(GO:0003724) |
| 0.1 | 1.8 | GO:0005025 | transforming growth factor beta receptor activity, type I(GO:0005025) |
| 0.1 | 2.8 | GO:0003688 | DNA replication origin binding(GO:0003688) |
| 0.1 | 4.1 | GO:0008175 | tRNA methyltransferase activity(GO:0008175) |
| 0.1 | 2.0 | GO:0000340 | RNA 7-methylguanosine cap binding(GO:0000340) |
| 0.1 | 1.4 | GO:0000774 | adenyl-nucleotide exchange factor activity(GO:0000774) |
| 0.1 | 1.6 | GO:0005111 | type 2 fibroblast growth factor receptor binding(GO:0005111) |
| 0.1 | 1.5 | GO:0052812 | phosphatidylinositol-4,5-bisphosphate 3-kinase activity(GO:0046934) phosphatidylinositol-3,4-bisphosphate 5-kinase activity(GO:0052812) |
| 0.1 | 1.6 | GO:0016868 | intramolecular transferase activity, phosphotransferases(GO:0016868) |
| 0.1 | 3.4 | GO:0008173 | RNA methyltransferase activity(GO:0008173) |
| 0.1 | 1.3 | GO:0005042 | netrin receptor activity(GO:0005042) |
| 0.1 | 0.8 | GO:0070180 | large ribosomal subunit rRNA binding(GO:0070180) |
| 0.1 | 0.7 | GO:0097027 | ubiquitin-protein transferase activator activity(GO:0097027) |
| 0.1 | 0.6 | GO:0003980 | UDP-glucose:glycoprotein glucosyltransferase activity(GO:0003980) |
| 0.1 | 2.2 | GO:0070003 | threonine-type endopeptidase activity(GO:0004298) threonine-type peptidase activity(GO:0070003) |
| 0.1 | 2.8 | GO:0043021 | ribonucleoprotein complex binding(GO:0043021) |
| 0.1 | 0.8 | GO:0000254 | C-4 methylsterol oxidase activity(GO:0000254) |
| 0.1 | 1.7 | GO:0051117 | ATPase binding(GO:0051117) |
| 0.1 | 2.4 | GO:0016628 | oxidoreductase activity, acting on the CH-CH group of donors, NAD or NADP as acceptor(GO:0016628) |
| 0.1 | 0.7 | GO:0004356 | glutamate-ammonia ligase activity(GO:0004356) ammonia ligase activity(GO:0016211) acid-ammonia (or amide) ligase activity(GO:0016880) |
| 0.1 | 1.4 | GO:0003886 | DNA (cytosine-5-)-methyltransferase activity(GO:0003886) |
| 0.1 | 1.3 | GO:0004687 | myosin light chain kinase activity(GO:0004687) |
| 0.1 | 1.9 | GO:0070122 | isopeptidase activity(GO:0070122) |
| 0.1 | 1.0 | GO:0010485 | H4 histone acetyltransferase activity(GO:0010485) |
| 0.1 | 2.1 | GO:0102336 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.1 | 0.7 | GO:0035173 | histone kinase activity(GO:0035173) |
| 0.1 | 0.6 | GO:0008597 | calcium-dependent protein serine/threonine phosphatase regulator activity(GO:0008597) |
| 0.1 | 3.1 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.1 | 1.2 | GO:0047631 | ADP-ribose diphosphatase activity(GO:0047631) |
| 0.1 | 0.9 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.1 | 6.0 | GO:0003697 | single-stranded DNA binding(GO:0003697) |
| 0.1 | 8.8 | GO:0015171 | amino acid transmembrane transporter activity(GO:0015171) |
| 0.1 | 3.0 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.1 | 0.3 | GO:0101006 | protein histidine phosphatase activity(GO:0101006) |
| 0.1 | 0.4 | GO:0008142 | oxysterol binding(GO:0008142) |
| 0.1 | 0.2 | GO:0046964 | 3'-phosphoadenosine 5'-phosphosulfate transmembrane transporter activity(GO:0046964) |
| 0.1 | 0.8 | GO:0003689 | DNA clamp loader activity(GO:0003689) protein-DNA loading ATPase activity(GO:0033170) |
| 0.1 | 2.1 | GO:0000175 | 3'-5'-exoribonuclease activity(GO:0000175) |
| 0.1 | 1.2 | GO:0016846 | carbon-sulfur lyase activity(GO:0016846) |
| 0.1 | 1.6 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.1 | 0.3 | GO:0008260 | 3-oxoacid CoA-transferase activity(GO:0008260) |
| 0.1 | 13.2 | GO:0042393 | histone binding(GO:0042393) |
| 0.1 | 0.5 | GO:0072518 | Rho-dependent protein serine/threonine kinase activity(GO:0072518) |
| 0.1 | 1.6 | GO:0016881 | acid-amino acid ligase activity(GO:0016881) |
| 0.1 | 0.8 | GO:0004017 | adenylate kinase activity(GO:0004017) |
| 0.1 | 1.8 | GO:0051537 | 2 iron, 2 sulfur cluster binding(GO:0051537) |
| 0.1 | 0.6 | GO:0030544 | Hsp70 protein binding(GO:0030544) |
| 0.1 | 0.9 | GO:0016879 | ligase activity, forming carbon-nitrogen bonds(GO:0016879) |
| 0.1 | 0.8 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.1 | 0.2 | GO:0004367 | glycerol-3-phosphate dehydrogenase [NAD+] activity(GO:0004367) |
| 0.0 | 9.1 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.0 | 0.4 | GO:0003828 | alpha-N-acetylneuraminate alpha-2,8-sialyltransferase activity(GO:0003828) |
| 0.0 | 1.0 | GO:0098748 | clathrin adaptor activity(GO:0035615) endocytic adaptor activity(GO:0098748) |
| 0.0 | 5.8 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.0 | 0.6 | GO:0070530 | K63-linked polyubiquitin binding(GO:0070530) |
| 0.0 | 1.9 | GO:0051536 | iron-sulfur cluster binding(GO:0051536) metal cluster binding(GO:0051540) |
| 0.0 | 3.0 | GO:0016279 | lysine N-methyltransferase activity(GO:0016278) protein-lysine N-methyltransferase activity(GO:0016279) |
| 0.0 | 0.3 | GO:0004300 | enoyl-CoA hydratase activity(GO:0004300) |
| 0.0 | 0.4 | GO:0004028 | 3-chloroallyl aldehyde dehydrogenase activity(GO:0004028) |
| 0.0 | 1.2 | GO:0005540 | hyaluronic acid binding(GO:0005540) |
| 0.0 | 0.9 | GO:0001103 | RNA polymerase II repressing transcription factor binding(GO:0001103) |
| 0.0 | 0.3 | GO:0034632 | retinol transporter activity(GO:0034632) |
| 0.0 | 0.8 | GO:0008143 | poly(A) binding(GO:0008143) |
| 0.0 | 2.7 | GO:0004386 | helicase activity(GO:0004386) |
| 0.0 | 0.9 | GO:0008094 | DNA-dependent ATPase activity(GO:0008094) |
| 0.0 | 0.4 | GO:0016783 | sulfurtransferase activity(GO:0016783) |
| 0.0 | 2.8 | GO:0001227 | transcriptional repressor activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001227) |
| 0.0 | 2.1 | GO:0043130 | ubiquitin binding(GO:0043130) |
| 0.0 | 0.5 | GO:0003873 | 6-phosphofructo-2-kinase activity(GO:0003873) fructose-2,6-bisphosphate 2-phosphatase activity(GO:0004331) |
| 0.0 | 5.0 | GO:0001228 | transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001228) |
| 0.0 | 2.2 | GO:0003730 | mRNA 3'-UTR binding(GO:0003730) |
| 0.0 | 1.1 | GO:0050840 | extracellular matrix binding(GO:0050840) |
| 0.0 | 0.3 | GO:0019843 | rRNA binding(GO:0019843) |
| 0.0 | 0.2 | GO:0008118 | N-acetyllactosaminide alpha-2,3-sialyltransferase activity(GO:0008118) |
| 0.0 | 1.8 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.0 | 0.4 | GO:0031078 | histone deacetylase activity (H3-K14 specific)(GO:0031078) NAD-dependent histone deacetylase activity (H3-K14 specific)(GO:0032041) |
| 0.0 | 1.4 | GO:0004521 | endoribonuclease activity(GO:0004521) |
| 0.0 | 1.9 | GO:0004518 | nuclease activity(GO:0004518) |
| 0.0 | 0.5 | GO:0016251 | obsolete general RNA polymerase II transcription factor activity(GO:0016251) |
| 0.0 | 1.4 | GO:0005507 | copper ion binding(GO:0005507) |
| 0.0 | 0.2 | GO:0035035 | histone acetyltransferase binding(GO:0035035) |
| 0.0 | 0.6 | GO:0051020 | GTPase binding(GO:0051020) |
| 0.0 | 16.8 | GO:0003723 | RNA binding(GO:0003723) |
| 0.0 | 2.1 | GO:0042803 | protein homodimerization activity(GO:0042803) |
| 0.0 | 0.3 | GO:0031726 | CCR1 chemokine receptor binding(GO:0031726) CCR4 chemokine receptor binding(GO:0031729) CCR5 chemokine receptor binding(GO:0031730) |
| 0.0 | 10.2 | GO:0005525 | GTP binding(GO:0005525) |
| 0.0 | 0.7 | GO:0004864 | protein phosphatase inhibitor activity(GO:0004864) |
| 0.0 | 0.2 | GO:0003917 | DNA topoisomerase activity(GO:0003916) DNA topoisomerase type I activity(GO:0003917) |
| 0.0 | 0.4 | GO:0016854 | racemase and epimerase activity(GO:0016854) |
| 0.0 | 0.8 | GO:0003755 | peptidyl-prolyl cis-trans isomerase activity(GO:0003755) |
| 0.0 | 0.6 | GO:0005347 | ATP transmembrane transporter activity(GO:0005347) |
| 0.0 | 1.0 | GO:0031625 | ubiquitin protein ligase binding(GO:0031625) |
| 0.0 | 0.3 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.0 | 0.5 | GO:0045505 | dynein intermediate chain binding(GO:0045505) |
| 0.0 | 5.2 | GO:0061630 | ubiquitin protein ligase activity(GO:0061630) |
| 0.0 | 0.9 | GO:0051082 | unfolded protein binding(GO:0051082) |
| 0.0 | 0.4 | GO:0042826 | histone deacetylase binding(GO:0042826) |
| 0.0 | 0.4 | GO:0017147 | Wnt-protein binding(GO:0017147) |
| 0.0 | 0.8 | GO:0004722 | protein serine/threonine phosphatase activity(GO:0004722) |
| 0.0 | 0.5 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 2.7 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.2 | 10.2 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.1 | 1.7 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.1 | 1.3 | PID RETINOIC ACID PATHWAY | Retinoic acid receptors-mediated signaling |
| 0.1 | 1.3 | PID WNT NONCANONICAL PATHWAY | Noncanonical Wnt signaling pathway |
| 0.1 | 1.5 | PID TRAIL PATHWAY | TRAIL signaling pathway |
| 0.1 | 0.9 | PID CIRCADIAN PATHWAY | Circadian rhythm pathway |
| 0.0 | 1.2 | PID ATM PATHWAY | ATM pathway |
| 0.0 | 1.3 | PID ANGIOPOIETIN RECEPTOR PATHWAY | Angiopoietin receptor Tie2-mediated signaling |
| 0.0 | 1.4 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.0 | 2.8 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.0 | 1.3 | PID NETRIN PATHWAY | Netrin-mediated signaling events |
| 0.0 | 1.0 | PID ERBB2 ERBB3 PATHWAY | ErbB2/ErbB3 signaling events |
| 0.0 | 0.8 | PID CERAMIDE PATHWAY | Ceramide signaling pathway |
| 0.0 | 2.1 | PID REG GR PATHWAY | Glucocorticoid receptor regulatory network |
| 0.0 | 0.9 | PID AJDISS 2PATHWAY | Posttranslational regulation of adherens junction stability and dissassembly |
| 0.0 | 0.6 | PID ECADHERIN NASCENT AJ PATHWAY | E-cadherin signaling in the nascent adherens junction |
| 0.0 | 0.6 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.0 | 0.7 | PID HEDGEHOG GLI PATHWAY | Hedgehog signaling events mediated by Gli proteins |
| 0.0 | 0.3 | PID WNT CANONICAL PATHWAY | Canonical Wnt signaling pathway |
| 0.0 | 3.7 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.0 | 1.6 | PID P53 DOWNSTREAM PATHWAY | Direct p53 effectors |
| 0.0 | 0.8 | PID SMAD2 3NUCLEAR PATHWAY | Regulation of nuclear SMAD2/3 signaling |
| 0.0 | 0.2 | PID FRA PATHWAY | Validated transcriptional targets of AP1 family members Fra1 and Fra2 |
| 0.0 | 0.4 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.0 | 0.2 | PID PRL SIGNALING EVENTS PATHWAY | Signaling events mediated by PRL |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.3 | 18.4 | REACTOME PURINE RIBONUCLEOSIDE MONOPHOSPHATE BIOSYNTHESIS | Genes involved in Purine ribonucleoside monophosphate biosynthesis |
| 0.6 | 9.0 | REACTOME E2F ENABLED INHIBITION OF PRE REPLICATION COMPLEX FORMATION | Genes involved in E2F-enabled inhibition of pre-replication complex formation |
| 0.4 | 4.7 | REACTOME MRNA DECAY BY 3 TO 5 EXORIBONUCLEASE | Genes involved in mRNA Decay by 3' to 5' Exoribonuclease |
| 0.4 | 11.8 | REACTOME PTM GAMMA CARBOXYLATION HYPUSINE FORMATION AND ARYLSULFATASE ACTIVATION | Genes involved in PTM: gamma carboxylation, hypusine formation and arylsulfatase activation |
| 0.4 | 7.9 | REACTOME PYRIMIDINE METABOLISM | Genes involved in Pyrimidine metabolism |
| 0.3 | 5.0 | REACTOME RNA POL III TRANSCRIPTION TERMINATION | Genes involved in RNA Polymerase III Transcription Termination |
| 0.3 | 4.6 | REACTOME HOMOLOGOUS RECOMBINATION REPAIR OF REPLICATION INDEPENDENT DOUBLE STRAND BREAKS | Genes involved in Homologous recombination repair of replication-independent double-strand breaks |
| 0.2 | 7.4 | REACTOME EXTENSION OF TELOMERES | Genes involved in Extension of Telomeres |
| 0.2 | 2.4 | REACTOME TRANSCRIPTION COUPLED NER TC NER | Genes involved in Transcription-coupled NER (TC-NER) |
| 0.2 | 5.6 | REACTOME RNA POL II TRANSCRIPTION PRE INITIATION AND PROMOTER OPENING | Genes involved in RNA Polymerase II Transcription Pre-Initiation And Promoter Opening |
| 0.2 | 1.4 | REACTOME GLUCOSE TRANSPORT | Genes involved in Glucose transport |
| 0.2 | 3.2 | REACTOME RNA POL I TRANSCRIPTION TERMINATION | Genes involved in RNA Polymerase I Transcription Termination |
| 0.1 | 3.1 | REACTOME KINESINS | Genes involved in Kinesins |
| 0.1 | 1.3 | REACTOME ANDROGEN BIOSYNTHESIS | Genes involved in Androgen biosynthesis |
| 0.1 | 1.9 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.1 | 1.2 | REACTOME HYALURONAN UPTAKE AND DEGRADATION | Genes involved in Hyaluronan uptake and degradation |
| 0.1 | 1.3 | REACTOME ROLE OF DCC IN REGULATING APOPTOSIS | Genes involved in Role of DCC in regulating apoptosis |
| 0.1 | 1.8 | REACTOME RESOLUTION OF AP SITES VIA THE SINGLE NUCLEOTIDE REPLACEMENT PATHWAY | Genes involved in Resolution of AP sites via the single-nucleotide replacement pathway |
| 0.1 | 1.1 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.1 | 5.7 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
| 0.1 | 7.7 | REACTOME MITOTIC PROMETAPHASE | Genes involved in Mitotic Prometaphase |
| 0.1 | 2.8 | REACTOME TIE2 SIGNALING | Genes involved in Tie2 Signaling |
| 0.1 | 1.2 | REACTOME REVERSIBLE HYDRATION OF CARBON DIOXIDE | Genes involved in Reversible Hydration of Carbon Dioxide |
| 0.1 | 2.8 | REACTOME ANTIVIRAL MECHANISM BY IFN STIMULATED GENES | Genes involved in Antiviral mechanism by IFN-stimulated genes |
| 0.1 | 1.3 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.1 | 1.1 | REACTOME UNWINDING OF DNA | Genes involved in Unwinding of DNA |
| 0.1 | 2.1 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 0.1 | 1.0 | REACTOME NEF MEDIATED DOWNREGULATION OF MHC CLASS I COMPLEX CELL SURFACE EXPRESSION | Genes involved in Nef mediated downregulation of MHC class I complex cell surface expression |
| 0.1 | 4.5 | REACTOME LATE PHASE OF HIV LIFE CYCLE | Genes involved in Late Phase of HIV Life Cycle |
| 0.1 | 1.4 | REACTOME GLYCOSPHINGOLIPID METABOLISM | Genes involved in Glycosphingolipid metabolism |
| 0.1 | 3.0 | REACTOME DESTABILIZATION OF MRNA BY AUF1 HNRNP D0 | Genes involved in Destabilization of mRNA by AUF1 (hnRNP D0) |
| 0.1 | 1.3 | REACTOME CIRCADIAN REPRESSION OF EXPRESSION BY REV ERBA | Genes involved in Circadian Repression of Expression by REV-ERBA |
| 0.1 | 1.1 | REACTOME TRANSPORT OF MATURE TRANSCRIPT TO CYTOPLASM | Genes involved in Transport of Mature Transcript to Cytoplasm |
| 0.1 | 1.0 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.1 | 0.7 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.1 | 0.9 | REACTOME SIGNALING BY FGFR1 FUSION MUTANTS | Genes involved in Signaling by FGFR1 fusion mutants |
| 0.1 | 0.8 | REACTOME ACTIVATION OF ATR IN RESPONSE TO REPLICATION STRESS | Genes involved in Activation of ATR in response to replication stress |
| 0.1 | 2.6 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.0 | 1.4 | REACTOME CYTOSOLIC TRNA AMINOACYLATION | Genes involved in Cytosolic tRNA aminoacylation |
| 0.0 | 3.5 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 0.0 | 0.6 | REACTOME IRAK2 MEDIATED ACTIVATION OF TAK1 COMPLEX UPON TLR7 8 OR 9 STIMULATION | Genes involved in IRAK2 mediated activation of TAK1 complex upon TLR7/8 or 9 stimulation |
| 0.0 | 0.7 | REACTOME MITOCHONDRIAL FATTY ACID BETA OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.0 | 1.3 | REACTOME SRP DEPENDENT COTRANSLATIONAL PROTEIN TARGETING TO MEMBRANE | Genes involved in SRP-dependent cotranslational protein targeting to membrane |
| 0.0 | 0.9 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.0 | 0.8 | REACTOME CYCLIN E ASSOCIATED EVENTS DURING G1 S TRANSITION | Genes involved in Cyclin E associated events during G1/S transition |
| 0.0 | 0.2 | REACTOME CTNNB1 PHOSPHORYLATION CASCADE | Genes involved in Beta-catenin phosphorylation cascade |
| 0.0 | 0.7 | REACTOME FORMATION OF TUBULIN FOLDING INTERMEDIATES BY CCT TRIC | Genes involved in Formation of tubulin folding intermediates by CCT/TriC |
| 0.0 | 0.4 | REACTOME SYNTHESIS OF PIPS AT THE LATE ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the late endosome membrane |
| 0.0 | 0.3 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.0 | 0.8 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.0 | 2.2 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 0.9 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.0 | 0.3 | REACTOME G ALPHA1213 SIGNALLING EVENTS | Genes involved in G alpha (12/13) signalling events |
| 0.0 | 0.8 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.0 | 0.2 | REACTOME HS GAG DEGRADATION | Genes involved in HS-GAG degradation |