Project
PRJEB1986: zebrafish developmental stages transcriptome
Navigation
Downloads
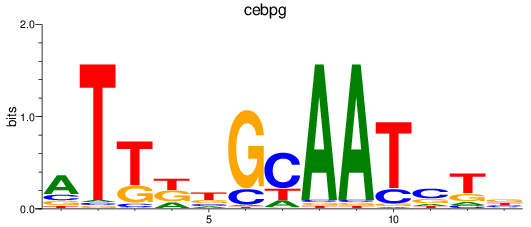
Results for cebpg
Z-value: 0.74
Transcription factors associated with cebpg
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
cebpg
|
ENSDARG00000036073 | CCAAT enhancer binding protein gamma |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| cebpg | dr11_v1_chr7_+_38089650_38089650 | 0.78 | 7.0e-05 | Click! |
Activity profile of cebpg motif
Sorted Z-values of cebpg motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr9_+_32301456 | 1.09 |
ENSDART00000078608
ENSDART00000185153 ENSDART00000144947 |
hspe1
|
heat shock 10 protein 1 |
| chr18_-_40905901 | 1.08 |
ENSDART00000064848
|
pglyrp5
|
peptidoglycan recognition protein 5 |
| chr3_+_23703704 | 1.00 |
ENSDART00000024256
|
hoxb6a
|
homeobox B6a |
| chr1_-_28885919 | 0.92 |
ENSDART00000152182
|
poglut1
|
protein O-glucosyltransferase 1 |
| chr19_+_16016038 | 0.89 |
ENSDART00000131319
|
ctps1a
|
CTP synthase 1a |
| chr19_-_25113660 | 0.89 |
ENSDART00000035538
|
ptp4a3
|
protein tyrosine phosphatase type IVA, member 3 |
| chr22_-_23668356 | 0.89 |
ENSDART00000167106
ENSDART00000159622 ENSDART00000163228 |
cfh
|
complement factor H |
| chr9_-_32300783 | 0.82 |
ENSDART00000078596
|
hspd1
|
heat shock 60 protein 1 |
| chr9_-_32300611 | 0.81 |
ENSDART00000127938
|
hspd1
|
heat shock 60 protein 1 |
| chr9_+_426392 | 0.79 |
ENSDART00000172515
|
bzw1b
|
basic leucine zipper and W2 domains 1b |
| chr25_-_3830272 | 0.78 |
ENSDART00000055843
|
cd151
|
CD151 molecule |
| chr11_-_34783938 | 0.76 |
ENSDART00000135725
ENSDART00000039847 |
chchd4a
|
coiled-coil-helix-coiled-coil-helix domain containing 4a |
| chr10_-_36808348 | 0.76 |
ENSDART00000099320
|
dhrs13a.1
|
dehydrogenase/reductase (SDR family) member 13a, tandem duplicate 1 |
| chr7_+_21887787 | 0.72 |
ENSDART00000162252
|
pop7
|
POP7 homolog, ribonuclease P/MRP subunit |
| chr25_+_20715950 | 0.69 |
ENSDART00000180223
|
ergic2
|
ERGIC and golgi 2 |
| chr3_+_54581987 | 0.68 |
ENSDART00000018071
|
eif3g
|
eukaryotic translation initiation factor 3, subunit G |
| chr9_+_32301017 | 0.68 |
ENSDART00000127916
ENSDART00000183298 ENSDART00000143103 |
hspe1
|
heat shock 10 protein 1 |
| chr10_-_22803740 | 0.67 |
ENSDART00000079469
ENSDART00000187968 ENSDART00000122543 |
pcolcea
|
procollagen C-endopeptidase enhancer a |
| chr2_-_40199780 | 0.67 |
ENSDART00000113901
|
ccl34a.4
|
chemokine (C-C motif) ligand 34a, duplicate 4 |
| chr15_-_3282220 | 0.67 |
ENSDART00000092942
|
slc25a15a
|
solute carrier family 25 (mitochondrial carrier; ornithine transporter) member 15a |
| chr17_+_51743908 | 0.66 |
ENSDART00000149039
ENSDART00000148869 |
odc1
|
ornithine decarboxylase 1 |
| chr10_-_23099809 | 0.65 |
ENSDART00000148333
ENSDART00000079703 ENSDART00000162444 |
nle1
|
notchless homolog 1 (Drosophila) |
| chr14_-_11507211 | 0.64 |
ENSDART00000186873
ENSDART00000109181 ENSDART00000186166 ENSDART00000186986 |
zgc:174917
|
zgc:174917 |
| chr17_+_12942634 | 0.63 |
ENSDART00000016597
|
nfkbiab
|
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha b |
| chr10_-_29827000 | 0.61 |
ENSDART00000131418
|
zpr1
|
ZPR1 zinc finger |
| chr8_-_50147948 | 0.61 |
ENSDART00000149010
|
hp
|
haptoglobin |
| chr3_+_34986837 | 0.61 |
ENSDART00000190341
|
smarce1
|
SWI/SNF related, matrix associated, actin dependent regulator of chromatin, subfamily e, member 1 |
| chr23_-_33986762 | 0.61 |
ENSDART00000144609
|
tmed4
|
transmembrane p24 trafficking protein 4 |
| chr17_+_51744450 | 0.61 |
ENSDART00000190955
ENSDART00000149807 |
odc1
|
ornithine decarboxylase 1 |
| chr18_-_26797723 | 0.60 |
ENSDART00000008013
|
sec11a
|
SEC11 homolog A, signal peptidase complex subunit |
| chr3_-_57666518 | 0.60 |
ENSDART00000102062
|
timp2b
|
TIMP metallopeptidase inhibitor 2b |
| chr4_+_42604252 | 0.60 |
ENSDART00000184850
|
CR925768.1
|
|
| chr14_-_38865800 | 0.59 |
ENSDART00000173047
|
gsr
|
glutathione reductase |
| chr4_+_30776883 | 0.59 |
ENSDART00000169519
|
si:dkey-11d20.1
|
si:dkey-11d20.1 |
| chr16_+_23984179 | 0.59 |
ENSDART00000175879
|
apoc2
|
apolipoprotein C-II |
| chr5_+_41510387 | 0.59 |
ENSDART00000023779
|
vcp
|
valosin containing protein |
| chr1_-_10071422 | 0.58 |
ENSDART00000135522
ENSDART00000033118 |
fga
|
fibrinogen alpha chain |
| chr17_-_10838434 | 0.57 |
ENSDART00000064597
|
lgals3b
|
lectin, galactoside binding soluble 3b |
| chr25_-_16146851 | 0.57 |
ENSDART00000104043
|
dkk3b
|
dickkopf WNT signaling pathway inhibitor 3b |
| chr12_-_19862912 | 0.57 |
ENSDART00000145788
|
shisa9a
|
shisa family member 9a |
| chr25_-_16076257 | 0.56 |
ENSDART00000140780
|
ovch2
|
ovochymase 2 |
| chr17_+_12942021 | 0.55 |
ENSDART00000192514
|
nfkbiab
|
nuclear factor of kappa light polypeptide gene enhancer in B-cells inhibitor, alpha b |
| chr20_-_30900947 | 0.55 |
ENSDART00000153419
ENSDART00000062536 |
hebp2
|
heme binding protein 2 |
| chr1_+_8694196 | 0.54 |
ENSDART00000025604
|
zgc:77849
|
zgc:77849 |
| chr11_-_31172276 | 0.53 |
ENSDART00000171520
|
si:dkey-238i5.3
|
si:dkey-238i5.3 |
| chr23_-_31763753 | 0.52 |
ENSDART00000053399
|
aldh8a1
|
aldehyde dehydrogenase 8 family, member A1 |
| chr23_-_16843649 | 0.51 |
ENSDART00000129971
|
si:dkey-63j12.4
|
si:dkey-63j12.4 |
| chr20_-_5369105 | 0.51 |
ENSDART00000114316
|
sptlc2b
|
serine palmitoyltransferase, long chain base subunit 2b |
| chr2_-_9696283 | 0.50 |
ENSDART00000165712
|
selenot1a
|
selenoprotein T, 1a |
| chr3_-_26190804 | 0.50 |
ENSDART00000136001
|
ypel3
|
yippee-like 3 |
| chr6_-_10708960 | 0.50 |
ENSDART00000157704
|
si:dkey-34m19.3
|
si:dkey-34m19.3 |
| chr5_-_15283509 | 0.49 |
ENSDART00000052712
|
gnb1l
|
guanine nucleotide binding protein (G protein), beta polypeptide 1-like |
| chr14_-_26392146 | 0.48 |
ENSDART00000037999
|
b4galt7
|
xylosylprotein beta 1,4-galactosyltransferase, polypeptide 7 (galactosyltransferase I) |
| chr17_-_12058171 | 0.48 |
ENSDART00000105236
|
smyd3
|
SET and MYND domain containing 3 |
| chr23_-_33944597 | 0.48 |
ENSDART00000133223
|
si:dkey-190g6.2
|
si:dkey-190g6.2 |
| chr25_-_26753196 | 0.48 |
ENSDART00000155698
|
usp3
|
ubiquitin specific peptidase 3 |
| chr1_+_41131481 | 0.48 |
ENSDART00000145272
|
lrpap1
|
low density lipoprotein receptor-related protein associated protein 1 |
| chr25_+_20716176 | 0.47 |
ENSDART00000073651
|
ergic2
|
ERGIC and golgi 2 |
| chr7_+_38089650 | 0.47 |
ENSDART00000052365
|
cebpg
|
CCAAT/enhancer binding protein (C/EBP), gamma |
| chr3_+_43086548 | 0.47 |
ENSDART00000163579
|
si:dkey-43p13.5
|
si:dkey-43p13.5 |
| chr1_+_17892944 | 0.46 |
ENSDART00000013021
|
tlr3
|
toll-like receptor 3 |
| chr13_-_24257631 | 0.46 |
ENSDART00000146524
|
urb2
|
URB2 ribosome biogenesis 2 homolog (S. cerevisiae) |
| chr13_-_50463938 | 0.46 |
ENSDART00000083857
|
ccnj
|
cyclin J |
| chr12_-_35949936 | 0.45 |
ENSDART00000192583
|
AL954682.1
|
|
| chr5_-_63509581 | 0.45 |
ENSDART00000097325
|
c5
|
complement component 5 |
| chr22_-_16317886 | 0.44 |
ENSDART00000163664
|
trmt13
|
tRNA methyltransferase 13 homolog (S. cerevisiae) |
| chr16_+_49601838 | 0.44 |
ENSDART00000168570
ENSDART00000159236 |
si:dkey-82o10.4
|
si:dkey-82o10.4 |
| chr24_+_16905188 | 0.41 |
ENSDART00000066760
|
cct5
|
chaperonin containing TCP1, subunit 5 (epsilon) |
| chr21_+_17051478 | 0.41 |
ENSDART00000047201
ENSDART00000161650 ENSDART00000167298 |
atp2a2a
|
ATPase sarcoplasmic/endoplasmic reticulum Ca2+ transporting 2a |
| chr3_-_34547000 | 0.40 |
ENSDART00000166623
|
sept9a
|
septin 9a |
| chr2_+_37971353 | 0.40 |
ENSDART00000165085
|
apol1
|
apolipoprotein L, 1 |
| chr19_-_18135724 | 0.40 |
ENSDART00000186609
|
cbx3a
|
chromobox homolog 3a (HP1 gamma homolog, Drosophila) |
| chr13_+_30903816 | 0.40 |
ENSDART00000191727
|
ercc6
|
excision repair cross-complementation group 6 |
| chr15_+_31296517 | 0.40 |
ENSDART00000132518
|
or116-1
|
odorant receptor, family F, subfamily 116, member 1 |
| chr22_-_24757785 | 0.39 |
ENSDART00000078225
|
vtg5
|
vitellogenin 5 |
| chr25_+_24616717 | 0.38 |
ENSDART00000089113
|
abtb2b
|
ankyrin repeat and BTB (POZ) domain containing 2b |
| chr4_-_73227710 | 0.38 |
ENSDART00000193165
|
LO018260.3
|
|
| chr13_+_11436130 | 0.37 |
ENSDART00000169895
|
zbtb18
|
zinc finger and BTB domain containing 18 |
| chr21_-_22709251 | 0.36 |
ENSDART00000140032
|
si:dkeyp-69c1.9
|
si:dkeyp-69c1.9 |
| chr17_+_12058509 | 0.35 |
ENSDART00000150209
|
tfb2m
|
transcription factor B2, mitochondrial |
| chr19_+_16015881 | 0.35 |
ENSDART00000187135
|
ctps1a
|
CTP synthase 1a |
| chr3_+_40809011 | 0.35 |
ENSDART00000033713
|
arpc1b
|
actin related protein 2/3 complex, subunit 1B |
| chr24_+_38671054 | 0.35 |
ENSDART00000154214
|
si:ch73-70c5.1
|
si:ch73-70c5.1 |
| chr20_+_40457599 | 0.35 |
ENSDART00000017553
|
serinc1
|
serine incorporator 1 |
| chr1_+_14253118 | 0.35 |
ENSDART00000161996
|
cxcl8a
|
chemokine (C-X-C motif) ligand 8a |
| chr9_+_38292947 | 0.34 |
ENSDART00000146663
|
tfcp2l1
|
transcription factor CP2-like 1 |
| chr5_+_25736979 | 0.34 |
ENSDART00000175959
|
abhd17b
|
abhydrolase domain containing 17B |
| chr18_-_25276932 | 0.34 |
ENSDART00000076183
|
plin1
|
perilipin 1 |
| chr13_-_25842074 | 0.34 |
ENSDART00000015154
|
papolg
|
poly(A) polymerase gamma |
| chr5_+_4298636 | 0.34 |
ENSDART00000100061
|
prdx4
|
peroxiredoxin 4 |
| chr10_+_6496185 | 0.33 |
ENSDART00000164770
|
reep5
|
receptor accessory protein 5 |
| chr24_-_25184553 | 0.33 |
ENSDART00000166917
|
plcxd2
|
phosphatidylinositol-specific phospholipase C, X domain containing 2 |
| chr14_-_38929885 | 0.33 |
ENSDART00000148737
|
btk
|
Bruton agammaglobulinemia tyrosine kinase |
| chr2_+_25378457 | 0.33 |
ENSDART00000089108
|
fndc3ba
|
fibronectin type III domain containing 3Ba |
| chr22_-_10051401 | 0.32 |
ENSDART00000106300
ENSDART00000175910 |
zgc:174564
BX324216.3
|
zgc:174564 |
| chr3_-_46818001 | 0.32 |
ENSDART00000166505
|
elavl3
|
ELAV like neuron-specific RNA binding protein 3 |
| chr21_+_21309164 | 0.31 |
ENSDART00000132174
|
si:ch211-191j22.8
|
si:ch211-191j22.8 |
| chr25_+_32474031 | 0.31 |
ENSDART00000152124
|
sqor
|
sulfide quinone oxidoreductase |
| chr8_-_2602572 | 0.30 |
ENSDART00000110482
|
zdhhc12a
|
zinc finger, DHHC-type containing 12a |
| chr15_+_28355023 | 0.30 |
ENSDART00000122159
|
si:dkey-118k5.3
|
si:dkey-118k5.3 |
| chr2_+_38055529 | 0.30 |
ENSDART00000145642
|
si:rp71-1g18.1
|
si:rp71-1g18.1 |
| chr4_-_20313810 | 0.29 |
ENSDART00000136350
|
dcp1b
|
decapping mRNA 1B |
| chr3_-_31254979 | 0.29 |
ENSDART00000130280
|
apnl
|
actinoporin-like protein |
| chr9_+_32073606 | 0.29 |
ENSDART00000184170
ENSDART00000180355 ENSDART00000110204 ENSDART00000123278 |
pikfyve
|
phosphoinositide kinase, FYVE finger containing |
| chr2_+_4207209 | 0.29 |
ENSDART00000157903
ENSDART00000166476 |
gata6
|
GATA binding protein 6 |
| chr5_-_27993972 | 0.29 |
ENSDART00000175819
|
ppp3cca
|
protein phosphatase 3, catalytic subunit, gamma isozyme, a |
| chr19_-_20403318 | 0.29 |
ENSDART00000136826
|
dazl
|
deleted in azoospermia-like |
| chr10_-_41980797 | 0.28 |
ENSDART00000076575
|
rhof
|
ras homolog family member F |
| chr10_-_20357013 | 0.28 |
ENSDART00000080143
|
sfrp1b
|
secreted frizzled-related protein 1b |
| chr13_-_30996072 | 0.28 |
ENSDART00000181661
|
wdfy4
|
WDFY family member 4 |
| chr7_-_13884610 | 0.27 |
ENSDART00000006897
|
rlbp1a
|
retinaldehyde binding protein 1a |
| chr2_+_42247560 | 0.27 |
ENSDART00000101230
ENSDART00000143094 |
ftr06
|
finTRIM family, member 6 |
| chr5_+_53482597 | 0.27 |
ENSDART00000180333
|
BX323994.1
|
|
| chr22_-_10570749 | 0.26 |
ENSDART00000140736
|
si:dkey-42i9.6
|
si:dkey-42i9.6 |
| chr4_-_5595237 | 0.26 |
ENSDART00000109854
|
vegfab
|
vascular endothelial growth factor Ab |
| chr13_-_24260609 | 0.26 |
ENSDART00000138747
|
urb2
|
URB2 ribosome biogenesis 2 homolog (S. cerevisiae) |
| chr6_+_13117598 | 0.25 |
ENSDART00000104744
|
casp8l1
|
caspase 8, apoptosis-related cysteine peptidase, like 1 |
| chr24_-_26485098 | 0.25 |
ENSDART00000135496
ENSDART00000009609 ENSDART00000133782 ENSDART00000141029 ENSDART00000113739 |
eif5a
|
eukaryotic translation initiation factor 5A |
| chr15_+_31307825 | 0.24 |
ENSDART00000173697
|
or117-1
|
odorant receptor, family F, subfamily 117, member 1 |
| chr4_-_20292821 | 0.24 |
ENSDART00000136069
ENSDART00000192504 |
cacna2d4a
|
calcium channel, voltage-dependent, alpha 2/delta subunit 4a |
| chr14_-_32089117 | 0.24 |
ENSDART00000158014
|
si:ch211-69b22.5
|
si:ch211-69b22.5 |
| chr4_+_8670662 | 0.23 |
ENSDART00000168768
|
adipor2
|
adiponectin receptor 2 |
| chr16_-_44945224 | 0.23 |
ENSDART00000156921
|
ncam3
|
neural cell adhesion molecule 3 |
| chr7_-_16596938 | 0.22 |
ENSDART00000134548
|
e2f8
|
E2F transcription factor 8 |
| chr9_-_33785093 | 0.22 |
ENSDART00000140779
ENSDART00000059837 |
fundc1
|
FUN14 domain containing 1 |
| chr17_+_18031899 | 0.22 |
ENSDART00000022758
|
setd3
|
SET domain containing 3 |
| chr22_-_10050856 | 0.22 |
ENSDART00000144811
|
zgc:174564
|
zgc:174564 |
| chr12_+_25775734 | 0.22 |
ENSDART00000024415
ENSDART00000149198 |
epas1a
|
endothelial PAS domain protein 1a |
| chr25_-_3469576 | 0.22 |
ENSDART00000186738
|
hbp1
|
HMG-box transcription factor 1 |
| chr8_-_18582922 | 0.22 |
ENSDART00000123917
|
tmem47
|
transmembrane protein 47 |
| chr3_-_16055432 | 0.21 |
ENSDART00000123621
ENSDART00000023859 |
atp6v0ca
|
ATPase H+ transporting V0 subunit ca |
| chr12_+_19362335 | 0.21 |
ENSDART00000041711
|
gspt1
|
G1 to S phase transition 1 |
| chr22_-_10487490 | 0.21 |
ENSDART00000064798
|
aspn
|
asporin (LRR class 1) |
| chr20_+_32118559 | 0.21 |
ENSDART00000026273
|
cd164
|
CD164 molecule, sialomucin |
| chr14_-_14705750 | 0.21 |
ENSDART00000168092
|
ogt.1
|
O-linked N-acetylglucosamine (GlcNAc) transferase, tandem duplicate 1 |
| chr12_-_33770299 | 0.21 |
ENSDART00000189849
|
llgl2
|
lethal giant larvae homolog 2 (Drosophila) |
| chr6_+_27452289 | 0.21 |
ENSDART00000186265
|
sned1
|
sushi, nidogen and EGF-like domains 1 |
| chr16_-_43356018 | 0.21 |
ENSDART00000181683
|
FO704821.1
|
|
| chr4_+_70563225 | 0.20 |
ENSDART00000159508
|
si:dkey-11d20.1
|
si:dkey-11d20.1 |
| chr6_+_10333920 | 0.19 |
ENSDART00000151667
ENSDART00000151477 |
cobll1a
|
cordon-bleu WH2 repeat protein-like 1a |
| chr10_-_13772847 | 0.19 |
ENSDART00000145103
ENSDART00000184491 |
cntfr
|
ciliary neurotrophic factor receptor |
| chr17_-_20218357 | 0.18 |
ENSDART00000155990
ENSDART00000155632 ENSDART00000156540 ENSDART00000063523 |
mgmt
|
O-6-methylguanine-DNA methyltransferase |
| chr15_-_24960730 | 0.18 |
ENSDART00000109990
ENSDART00000186706 |
abhd15a
|
abhydrolase domain containing 15a |
| chr9_+_50000504 | 0.18 |
ENSDART00000164409
ENSDART00000165451 ENSDART00000166509 |
slc38a11
|
solute carrier family 38, member 11 |
| chr11_+_25044082 | 0.18 |
ENSDART00000123263
|
phf20a
|
PHD finger protein 20, a |
| chr22_-_18491813 | 0.18 |
ENSDART00000105419
|
si:ch211-212d10.2
|
si:ch211-212d10.2 |
| chr24_-_33284945 | 0.18 |
ENSDART00000155429
ENSDART00000112845 |
zgc:195173
|
zgc:195173 |
| chr12_+_46386983 | 0.18 |
ENSDART00000183982
|
BX005305.3
|
Danio rerio legumain (LOC100005356), mRNA. |
| chr8_-_21988833 | 0.17 |
ENSDART00000167708
|
nphp4
|
nephronophthisis 4 |
| chr21_+_26535034 | 0.17 |
ENSDART00000180709
|
vegfbb
|
vascular endothelial growth factor Bb |
| chr14_+_7377552 | 0.17 |
ENSDART00000142158
ENSDART00000141471 |
hars
|
histidyl-tRNA synthetase |
| chr7_+_56577906 | 0.17 |
ENSDART00000184023
|
hp
|
haptoglobin |
| chr4_+_9508505 | 0.16 |
ENSDART00000080842
|
kitlgb
|
kit ligand b |
| chr6_+_28018390 | 0.16 |
ENSDART00000123324
ENSDART00000150915 |
sap130a
|
Sin3A-associated protein a |
| chr5_+_54400971 | 0.16 |
ENSDART00000169695
|
bspry
|
B-box and SPRY domain containing |
| chr20_+_53181017 | 0.15 |
ENSDART00000189692
ENSDART00000177109 |
FIG4
|
FIG4 phosphoinositide 5-phosphatase |
| chr12_+_46512881 | 0.15 |
ENSDART00000105454
|
CU855711.1
|
|
| chr9_-_21838045 | 0.15 |
ENSDART00000147471
|
acod1
|
aconitate decarboxylase 1 |
| chr18_+_32844439 | 0.15 |
ENSDART00000166038
|
olfcg1
|
olfactory receptor C family, g1 |
| chr17_-_11368662 | 0.15 |
ENSDART00000159061
ENSDART00000188694 ENSDART00000190932 |
si:ch211-185a18.2
|
si:ch211-185a18.2 |
| chr4_+_17279966 | 0.15 |
ENSDART00000067005
ENSDART00000137487 |
bcat1
|
branched chain amino-acid transaminase 1, cytosolic |
| chr9_-_9671244 | 0.14 |
ENSDART00000018228
|
gsk3b
|
glycogen synthase kinase 3 beta |
| chr9_-_12874774 | 0.13 |
ENSDART00000131385
|
ankzf1
|
ankyrin repeat and zinc finger domain containing 1 |
| chr15_-_29348212 | 0.13 |
ENSDART00000133117
|
tsku
|
tsukushi small leucine rich proteoglycan homolog (Xenopus laevis) |
| chr20_+_37300296 | 0.13 |
ENSDART00000180789
|
vta1
|
vesicle (multivesicular body) trafficking 1 |
| chr22_+_6587280 | 0.13 |
ENSDART00000106185
|
CR352226.7
|
|
| chr3_-_32873641 | 0.13 |
ENSDART00000075277
|
zgc:113090
|
zgc:113090 |
| chr12_+_46404307 | 0.13 |
ENSDART00000185011
|
BX005305.4
|
|
| chr12_+_46462090 | 0.12 |
ENSDART00000130748
|
BX005305.2
|
|
| chr15_-_23793641 | 0.12 |
ENSDART00000122891
|
tmem97
|
transmembrane protein 97 |
| chr19_-_34979837 | 0.12 |
ENSDART00000044838
|
ndrg1a
|
N-myc downstream regulated 1a |
| chr21_+_20237006 | 0.12 |
ENSDART00000132853
|
si:dkey-247m21.3
|
si:dkey-247m21.3 |
| chr9_-_40664923 | 0.12 |
ENSDART00000135134
|
bard1
|
BRCA1 associated RING domain 1 |
| chr10_-_7785930 | 0.12 |
ENSDART00000043961
ENSDART00000111058 |
mpx
|
myeloid-specific peroxidase |
| chr18_-_15551360 | 0.12 |
ENSDART00000159915
ENSDART00000172690 |
ppfibp1b
|
PTPRF interacting protein, binding protein 1b (liprin beta 1) |
| chr16_+_28601974 | 0.12 |
ENSDART00000018235
|
crot
|
carnitine O-octanoyltransferase |
| chr22_+_6711549 | 0.12 |
ENSDART00000132094
|
CT583625.4
|
|
| chr15_-_21014270 | 0.12 |
ENSDART00000154019
|
si:ch211-212c13.10
|
si:ch211-212c13.10 |
| chr20_+_29436601 | 0.11 |
ENSDART00000136804
|
fmn1
|
formin 1 |
| chr10_+_41945890 | 0.11 |
ENSDART00000063013
ENSDART00000128313 |
tmem120b
|
transmembrane protein 120B |
| chr12_+_46425800 | 0.11 |
ENSDART00000191965
|
BX005305.1
|
|
| chr7_+_33152723 | 0.11 |
ENSDART00000132658
|
si:ch211-194p6.10
|
si:ch211-194p6.10 |
| chr23_-_40814080 | 0.11 |
ENSDART00000135713
|
si:dkeyp-27c8.1
|
si:dkeyp-27c8.1 |
| chr9_-_9419704 | 0.10 |
ENSDART00000138996
|
si:ch211-214p13.9
|
si:ch211-214p13.9 |
| chr8_+_22955478 | 0.09 |
ENSDART00000099911
|
zgc:136605
|
zgc:136605 |
| chr1_+_30551777 | 0.09 |
ENSDART00000112778
|
gpr183b
|
G protein-coupled receptor 183b |
| chr14_-_43602968 | 0.09 |
ENSDART00000108825
|
adh5
|
alcohol dehydrogenase 5 |
| chr12_+_46443477 | 0.09 |
ENSDART00000191873
|
BX005305.1
|
|
| chr8_+_22955262 | 0.09 |
ENSDART00000193806
|
zgc:136605
|
zgc:136605 |
| chr23_-_40792128 | 0.08 |
ENSDART00000145360
|
si:dkey-194e6.2
|
si:dkey-194e6.2 |
| chr24_+_13925066 | 0.08 |
ENSDART00000134221
ENSDART00000012253 ENSDART00000081595 ENSDART00000136443 |
eya1
|
EYA transcriptional coactivator and phosphatase 1 |
| chr5_+_39504136 | 0.07 |
ENSDART00000121460
|
prdm8b
|
PR domain containing 8b |
| chr10_+_15310811 | 0.07 |
ENSDART00000136239
|
kcnv2a
|
potassium channel, subfamily V, member 2a |
| chr18_+_20869923 | 0.07 |
ENSDART00000138471
|
pgpep1l
|
pyroglutamyl-peptidase I-like |
| chr12_+_26632448 | 0.07 |
ENSDART00000185762
|
arhgap12b
|
Rho GTPase activating protein 12b |
| chr5_-_23783739 | 0.06 |
ENSDART00000139502
|
GBGT1 (1 of many)
|
si:ch211-287c22.1 |
| chr12_+_4712215 | 0.06 |
ENSDART00000152134
|
kansl1a
|
KAT8 regulatory NSL complex subunit 1a |
| chr14_-_5642371 | 0.06 |
ENSDART00000183859
ENSDART00000054876 |
npm1b
|
nucleophosmin 1b |
| chr19_+_7759354 | 0.05 |
ENSDART00000151400
|
ubap2l
|
ubiquitin associated protein 2-like |
| chr19_-_2822372 | 0.05 |
ENSDART00000109130
ENSDART00000122385 |
recql4
|
RecQ helicase-like 4 |
Network of associatons between targets according to the STRING database.
First level regulatory network of cebpg
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 2.4 | GO:0045041 | protein import into mitochondrial intermembrane space(GO:0045041) |
| 0.3 | 1.2 | GO:0044210 | 'de novo' CTP biosynthetic process(GO:0044210) |
| 0.2 | 0.7 | GO:0000066 | mitochondrial ornithine transport(GO:0000066) mitochondrial L-ornithine transmembrane transport(GO:1990575) |
| 0.2 | 0.6 | GO:0032510 | endosome to lysosome transport via multivesicular body sorting pathway(GO:0032510) mitotic spindle disassembly(GO:0051228) spindle disassembly(GO:0051230) |
| 0.2 | 0.5 | GO:0097053 | L-kynurenine metabolic process(GO:0097052) L-kynurenine catabolic process(GO:0097053) |
| 0.2 | 1.1 | GO:0000270 | peptidoglycan metabolic process(GO:0000270) peptidoglycan catabolic process(GO:0009253) |
| 0.2 | 0.5 | GO:0034138 | chemokine production(GO:0032602) toll-like receptor 3 signaling pathway(GO:0034138) |
| 0.1 | 1.3 | GO:0033387 | putrescine biosynthetic process from ornithine(GO:0033387) |
| 0.1 | 0.3 | GO:0002432 | granuloma formation(GO:0002432) chronic inflammatory response(GO:0002544) regulation of granuloma formation(GO:0002631) regulation of chronic inflammatory response(GO:0002676) |
| 0.1 | 0.6 | GO:0048245 | eosinophil chemotaxis(GO:0048245) eosinophil migration(GO:0072677) |
| 0.1 | 0.3 | GO:0070221 | sulfide oxidation(GO:0019418) sulfide oxidation, using sulfide:quinone oxidoreductase(GO:0070221) |
| 0.1 | 0.6 | GO:0051045 | negative regulation of membrane protein ectodomain proteolysis(GO:0051045) |
| 0.1 | 0.5 | GO:0034447 | very-low-density lipoprotein particle clearance(GO:0034447) |
| 0.1 | 0.6 | GO:0001561 | fatty acid alpha-oxidation(GO:0001561) |
| 0.1 | 0.4 | GO:0006391 | transcription initiation from mitochondrial promoter(GO:0006391) |
| 0.1 | 0.4 | GO:0032978 | protein insertion into membrane from inner side(GO:0032978) protein insertion into mitochondrial membrane from inner side(GO:0032979) protein insertion into mitochondrial membrane(GO:0051204) |
| 0.1 | 0.6 | GO:0006465 | signal peptide processing(GO:0006465) |
| 0.1 | 0.6 | GO:0048172 | regulation of short-term neuronal synaptic plasticity(GO:0048172) |
| 0.1 | 0.6 | GO:0003160 | endocardium morphogenesis(GO:0003160) |
| 0.1 | 0.2 | GO:0033301 | cell cycle comprising mitosis without cytokinesis(GO:0033301) |
| 0.1 | 0.3 | GO:0043011 | myeloid dendritic cell activation(GO:0001773) myeloid dendritic cell differentiation(GO:0043011) |
| 0.1 | 0.4 | GO:0006283 | transcription-coupled nucleotide-excision repair(GO:0006283) |
| 0.1 | 1.8 | GO:0051084 | 'de novo' protein folding(GO:0006458) 'de novo' posttranslational protein folding(GO:0051084) |
| 0.1 | 1.4 | GO:0045454 | cell redox homeostasis(GO:0045454) |
| 0.1 | 0.9 | GO:0045747 | positive regulation of Notch signaling pathway(GO:0045747) |
| 0.0 | 0.3 | GO:0031087 | nuclear-transcribed mRNA catabolic process, deadenylation-independent decay(GO:0031086) deadenylation-independent decapping of nuclear-transcribed mRNA(GO:0031087) |
| 0.0 | 0.3 | GO:0031640 | killing of cells of other organism(GO:0031640) disruption of cells of other organism(GO:0044364) disruption of cells of other organism involved in symbiotic interaction(GO:0051818) killing of cells in other organism involved in symbiotic interaction(GO:0051883) |
| 0.0 | 0.3 | GO:0060965 | negative regulation of posttranscriptional gene silencing(GO:0060149) negative regulation of gene silencing by miRNA(GO:0060965) negative regulation of gene silencing by RNA(GO:0060967) |
| 0.0 | 0.5 | GO:0046512 | diol biosynthetic process(GO:0034312) sphingosine biosynthetic process(GO:0046512) |
| 0.0 | 0.9 | GO:0042159 | lipoprotein catabolic process(GO:0042159) |
| 0.0 | 0.2 | GO:0008594 | photoreceptor cell morphogenesis(GO:0008594) negative regulation of epithelial to mesenchymal transition(GO:0010719) |
| 0.0 | 0.8 | GO:0042574 | retinal metabolic process(GO:0042574) |
| 0.0 | 0.4 | GO:0060753 | mast cell chemotaxis(GO:0002551) regulation of mast cell chemotaxis(GO:0060753) positive regulation of mast cell chemotaxis(GO:0060754) mast cell migration(GO:0097531) |
| 0.0 | 0.6 | GO:0072378 | blood coagulation, fibrin clot formation(GO:0072378) |
| 0.0 | 0.4 | GO:0030240 | skeletal muscle thin filament assembly(GO:0030240) |
| 0.0 | 0.1 | GO:0009099 | leucine biosynthetic process(GO:0009098) valine biosynthetic process(GO:0009099) |
| 0.0 | 0.2 | GO:0045901 | positive regulation of translational elongation(GO:0045901) positive regulation of translational termination(GO:0045905) |
| 0.0 | 0.2 | GO:2000096 | positive regulation of Wnt signaling pathway, planar cell polarity pathway(GO:2000096) |
| 0.0 | 0.3 | GO:0010890 | positive regulation of sequestering of triglyceride(GO:0010890) |
| 0.0 | 0.1 | GO:0003418 | chondrocyte differentiation involved in endochondral bone morphogenesis(GO:0003413) growth plate cartilage chondrocyte differentiation(GO:0003418) |
| 0.0 | 0.2 | GO:0002184 | cytoplasmic translational termination(GO:0002184) |
| 0.0 | 0.2 | GO:1903426 | regulation of reactive oxygen species biosynthetic process(GO:1903426) |
| 0.0 | 0.7 | GO:0090502 | RNA phosphodiester bond hydrolysis, endonucleolytic(GO:0090502) |
| 0.0 | 0.2 | GO:0018026 | peptidyl-lysine monomethylation(GO:0018026) |
| 0.0 | 0.2 | GO:0048680 | positive regulation of axon regeneration(GO:0048680) positive regulation of neuron projection regeneration(GO:0070572) |
| 0.0 | 0.6 | GO:0000027 | ribosomal large subunit assembly(GO:0000027) |
| 0.0 | 1.2 | GO:0006890 | retrograde vesicle-mediated transport, Golgi to ER(GO:0006890) |
| 0.0 | 0.7 | GO:0001732 | formation of cytoplasmic translation initiation complex(GO:0001732) |
| 0.0 | 0.3 | GO:0006658 | phosphatidylserine metabolic process(GO:0006658) |
| 0.0 | 0.1 | GO:2000377 | regulation of reactive oxygen species metabolic process(GO:2000377) |
| 0.0 | 0.5 | GO:1901663 | ubiquinone metabolic process(GO:0006743) ubiquinone biosynthetic process(GO:0006744) quinone biosynthetic process(GO:1901663) |
| 0.0 | 0.2 | GO:0060036 | notochord cell vacuolation(GO:0060036) |
| 0.0 | 0.2 | GO:0033211 | adiponectin-activated signaling pathway(GO:0033211) |
| 0.0 | 0.2 | GO:0071941 | nitrogen cycle metabolic process(GO:0071941) |
| 0.0 | 0.1 | GO:0032287 | peripheral nervous system myelin maintenance(GO:0032287) |
| 0.0 | 0.7 | GO:0002548 | monocyte chemotaxis(GO:0002548) |
| 0.0 | 0.4 | GO:0002574 | thrombocyte differentiation(GO:0002574) |
| 0.0 | 0.2 | GO:0061484 | hematopoietic stem cell homeostasis(GO:0061484) |
| 0.0 | 0.7 | GO:1902036 | regulation of hematopoietic stem cell differentiation(GO:1902036) |
| 0.0 | 0.0 | GO:0044038 | cell wall biogenesis(GO:0042546) cell wall macromolecule metabolic process(GO:0044036) cell wall macromolecule biosynthetic process(GO:0044038) cellular component macromolecule biosynthetic process(GO:0070589) cell wall organization or biogenesis(GO:0071554) |
| 0.0 | 0.2 | GO:0046549 | retinal cone cell development(GO:0046549) |
| 0.0 | 0.4 | GO:0035775 | pronephric glomerulus morphogenesis(GO:0035775) |
| 0.0 | 0.1 | GO:0046292 | formaldehyde metabolic process(GO:0046292) formaldehyde catabolic process(GO:0046294) |
| 0.0 | 0.4 | GO:0030488 | tRNA methylation(GO:0030488) |
| 0.0 | 0.3 | GO:0006378 | mRNA polyadenylation(GO:0006378) RNA polyadenylation(GO:0043631) |
| 0.0 | 0.9 | GO:0090090 | negative regulation of canonical Wnt signaling pathway(GO:0090090) |
| 0.0 | 0.1 | GO:0045444 | fat cell differentiation(GO:0045444) |
| 0.0 | 0.2 | GO:0071456 | cellular response to hypoxia(GO:0071456) |
| 0.0 | 0.2 | GO:0021508 | floor plate formation(GO:0021508) floor plate morphogenesis(GO:0033505) |
| 0.0 | 0.3 | GO:0007634 | optokinetic behavior(GO:0007634) |
| 0.0 | 0.2 | GO:0050994 | regulation of lipid catabolic process(GO:0050994) |
| 0.0 | 0.4 | GO:0032355 | response to estradiol(GO:0032355) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.2 | GO:0097268 | cytoophidium(GO:0097268) |
| 0.2 | 0.6 | GO:0005577 | fibrinogen complex(GO:0005577) |
| 0.1 | 0.6 | GO:0005787 | signal peptidase complex(GO:0005787) |
| 0.1 | 0.6 | GO:0034098 | VCP-NPL4-UFD1 AAA ATPase complex(GO:0034098) |
| 0.1 | 0.3 | GO:0044217 | other organism(GO:0044215) other organism cell(GO:0044216) other organism part(GO:0044217) other organism cell membrane(GO:0044218) other organism membrane(GO:0044279) |
| 0.1 | 0.6 | GO:0001772 | immunological synapse(GO:0001772) |
| 0.1 | 0.6 | GO:0034362 | low-density lipoprotein particle(GO:0034362) |
| 0.1 | 0.5 | GO:0031211 | serine C-palmitoyltransferase complex(GO:0017059) endoplasmic reticulum palmitoyltransferase complex(GO:0031211) |
| 0.0 | 0.3 | GO:0005955 | calcineurin complex(GO:0005955) |
| 0.0 | 1.2 | GO:0033116 | endoplasmic reticulum-Golgi intermediate compartment membrane(GO:0033116) |
| 0.0 | 0.4 | GO:0005832 | chaperonin-containing T-complex(GO:0005832) |
| 0.0 | 0.6 | GO:0032591 | dendritic spine membrane(GO:0032591) |
| 0.0 | 0.2 | GO:0018444 | translation release factor complex(GO:0018444) |
| 0.0 | 0.7 | GO:0016282 | eukaryotic 43S preinitiation complex(GO:0016282) |
| 0.0 | 1.1 | GO:0005793 | endoplasmic reticulum-Golgi intermediate compartment(GO:0005793) |
| 0.0 | 0.1 | GO:0019815 | B cell receptor complex(GO:0019815) |
| 0.0 | 3.7 | GO:0005759 | mitochondrial matrix(GO:0005759) |
| 0.0 | 0.8 | GO:0031970 | mitochondrial intermembrane space(GO:0005758) organelle envelope lumen(GO:0031970) |
| 0.0 | 0.4 | GO:0005885 | Arp2/3 protein complex(GO:0005885) |
| 0.0 | 0.4 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.0 | 0.2 | GO:0031307 | integral component of mitochondrial outer membrane(GO:0031307) |
| 0.0 | 0.6 | GO:0016514 | SWI/SNF complex(GO:0016514) |
| 0.0 | 0.2 | GO:0044545 | NSL complex(GO:0044545) |
| 0.0 | 0.2 | GO:0033179 | proton-transporting V-type ATPase, V0 domain(GO:0033179) |
| 0.0 | 0.1 | GO:0070531 | BRCA1-A complex(GO:0070531) |
| 0.0 | 0.0 | GO:0061702 | inflammasome complex(GO:0061702) |
| 0.0 | 0.4 | GO:0031105 | septin ring(GO:0005940) septin complex(GO:0031105) septin cytoskeleton(GO:0032156) |
| 0.0 | 0.5 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.2 | GO:0003883 | CTP synthase activity(GO:0003883) |
| 0.3 | 1.1 | GO:0008745 | N-acetylmuramoyl-L-alanine amidase activity(GO:0008745) |
| 0.1 | 1.3 | GO:0004586 | ornithine decarboxylase activity(GO:0004586) |
| 0.1 | 1.0 | GO:0048019 | receptor antagonist activity(GO:0048019) |
| 0.1 | 0.9 | GO:0035252 | UDP-xylosyltransferase activity(GO:0035252) |
| 0.1 | 0.6 | GO:0004362 | glutathione-disulfide reductase activity(GO:0004362) |
| 0.1 | 0.4 | GO:0034246 | core DNA-dependent RNA polymerase binding promoter specificity activity(GO:0000996) mitochondrial RNA polymerase binding promoter specificity activity(GO:0034246) |
| 0.1 | 0.6 | GO:0019863 | IgE binding(GO:0019863) immunoglobulin binding(GO:0019865) |
| 0.1 | 0.3 | GO:0070224 | sulfide:quinone oxidoreductase activity(GO:0070224) |
| 0.1 | 0.3 | GO:0005153 | interleukin-8 receptor binding(GO:0005153) |
| 0.1 | 3.4 | GO:0051087 | chaperone binding(GO:0051087) |
| 0.1 | 0.3 | GO:0004652 | polynucleotide adenylyltransferase activity(GO:0004652) |
| 0.1 | 0.8 | GO:0052650 | NADP-retinol dehydrogenase activity(GO:0052650) |
| 0.1 | 0.7 | GO:0000064 | L-ornithine transmembrane transporter activity(GO:0000064) |
| 0.1 | 0.5 | GO:0004758 | serine C-palmitoyltransferase activity(GO:0004758) C-palmitoyltransferase activity(GO:0016454) |
| 0.1 | 0.6 | GO:0016922 | ligand-dependent nuclear receptor binding(GO:0016922) |
| 0.1 | 0.3 | GO:0043183 | vascular endothelial growth factor receptor 1 binding(GO:0043183) |
| 0.1 | 0.7 | GO:0004526 | ribonuclease P activity(GO:0004526) |
| 0.1 | 0.5 | GO:0004791 | thioredoxin-disulfide reductase activity(GO:0004791) |
| 0.1 | 0.4 | GO:0045735 | nutrient reservoir activity(GO:0045735) |
| 0.1 | 0.5 | GO:0048039 | ubiquinone binding(GO:0048039) |
| 0.0 | 0.3 | GO:0008379 | thioredoxin peroxidase activity(GO:0008379) |
| 0.0 | 0.3 | GO:0004723 | calcium-dependent protein serine/threonine phosphatase activity(GO:0004723) calmodulin-dependent protein phosphatase activity(GO:0033192) |
| 0.0 | 0.6 | GO:0008191 | metalloendopeptidase inhibitor activity(GO:0008191) |
| 0.0 | 0.2 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 0.0 | 0.2 | GO:0005173 | stem cell factor receptor binding(GO:0005173) |
| 0.0 | 0.1 | GO:0052655 | branched-chain-amino-acid transaminase activity(GO:0004084) L-leucine transaminase activity(GO:0052654) L-valine transaminase activity(GO:0052655) L-isoleucine transaminase activity(GO:0052656) |
| 0.0 | 0.2 | GO:0097363 | protein N-acetylglucosaminyltransferase activity(GO:0016262) protein O-GlcNAc transferase activity(GO:0097363) |
| 0.0 | 0.5 | GO:0035250 | UDP-galactosyltransferase activity(GO:0035250) |
| 0.0 | 0.3 | GO:0008494 | translation activator activity(GO:0008494) |
| 0.0 | 0.2 | GO:0001217 | bacterial-type RNA polymerase transcription factor activity, sequence-specific DNA binding(GO:0001130) bacterial-type RNA polymerase transcriptional repressor activity, sequence-specific DNA binding(GO:0001217) |
| 0.0 | 0.8 | GO:0015035 | protein disulfide oxidoreductase activity(GO:0015035) |
| 0.0 | 0.2 | GO:0034338 | short-chain carboxylesterase activity(GO:0034338) |
| 0.0 | 0.2 | GO:0098809 | nitrite reductase activity(GO:0098809) |
| 0.0 | 0.2 | GO:0005172 | vascular endothelial growth factor receptor binding(GO:0005172) |
| 0.0 | 0.4 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.0 | 0.1 | GO:0016415 | octanoyltransferase activity(GO:0016415) |
| 0.0 | 0.1 | GO:0016920 | pyroglutamyl-peptidase activity(GO:0016920) |
| 0.0 | 0.2 | GO:0008172 | S-methyltransferase activity(GO:0008172) |
| 0.0 | 0.1 | GO:0051903 | alcohol dehydrogenase activity, zinc-dependent(GO:0004024) S-(hydroxymethyl)glutathione dehydrogenase activity(GO:0051903) |
| 0.0 | 0.2 | GO:0008079 | translation release factor activity(GO:0003747) translation termination factor activity(GO:0008079) |
| 0.0 | 0.3 | GO:0008474 | palmitoyl-(protein) hydrolase activity(GO:0008474) palmitoyl hydrolase activity(GO:0098599) |
| 0.0 | 0.7 | GO:0048020 | CCR chemokine receptor binding(GO:0048020) |
| 0.0 | 0.2 | GO:0046975 | histone methyltransferase activity (H3-K36 specific)(GO:0046975) |
| 0.0 | 0.7 | GO:0043022 | ribosome binding(GO:0043022) |
| 0.0 | 0.6 | GO:0031593 | polyubiquitin binding(GO:0031593) |
| 0.0 | 0.2 | GO:0045159 | myosin II binding(GO:0045159) |
| 0.0 | 0.5 | GO:0016620 | oxidoreductase activity, acting on the aldehyde or oxo group of donors, NAD or NADP as acceptor(GO:0016620) |
| 0.0 | 0.4 | GO:0008175 | tRNA methyltransferase activity(GO:0008175) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 2.1 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.0 | 0.6 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.0 | 0.6 | PID INTEGRIN2 PATHWAY | Beta2 integrin cell surface interactions |
| 0.0 | 0.9 | PID PRL SIGNALING EVENTS PATHWAY | Signaling events mediated by PRL |
| 0.0 | 0.6 | PID REELIN PATHWAY | Reelin signaling pathway |
| 0.0 | 1.4 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.0 | 0.3 | ST GA12 PATHWAY | G alpha 12 Pathway |
| 0.0 | 0.2 | PID MYC PATHWAY | C-MYC pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 0.9 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 0.1 | 0.6 | REACTOME CHYLOMICRON MEDIATED LIPID TRANSPORT | Genes involved in Chylomicron-mediated lipid transport |
| 0.1 | 0.6 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GLP1 | Genes involved in Synthesis, Secretion, and Inactivation of Glucagon-like Peptide-1 (GLP-1) |
| 0.1 | 0.6 | REACTOME COMMON PATHWAY | Genes involved in Common Pathway |
| 0.1 | 0.5 | REACTOME IKK COMPLEX RECRUITMENT MEDIATED BY RIP1 | Genes involved in IKK complex recruitment mediated by RIP1 |
| 0.0 | 0.3 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 0.0 | 0.4 | REACTOME SYNTHESIS OF PIPS AT THE LATE ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the late endosome membrane |
| 0.0 | 1.6 | REACTOME MITOCHONDRIAL PROTEIN IMPORT | Genes involved in Mitochondrial Protein Import |
| 0.0 | 1.3 | REACTOME REGULATION OF ORNITHINE DECARBOXYLASE ODC | Genes involved in Regulation of ornithine decarboxylase (ODC) |
| 0.0 | 0.6 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.0 | 0.5 | REACTOME A TETRASACCHARIDE LINKER SEQUENCE IS REQUIRED FOR GAG SYNTHESIS | Genes involved in A tetrasaccharide linker sequence is required for GAG synthesis |
| 0.0 | 0.4 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 0.0 | 0.4 | REACTOME FORMATION OF TUBULIN FOLDING INTERMEDIATES BY CCT TRIC | Genes involved in Formation of tubulin folding intermediates by CCT/TriC |
| 0.0 | 0.3 | REACTOME MRNA DECAY BY 5 TO 3 EXORIBONUCLEASE | Genes involved in mRNA Decay by 5' to 3' Exoribonuclease |
| 0.0 | 0.8 | REACTOME CELL JUNCTION ORGANIZATION | Genes involved in Cell junction organization |
| 0.0 | 0.4 | REACTOME RNA POL I TRANSCRIPTION INITIATION | Genes involved in RNA Polymerase I Transcription Initiation |
| 0.0 | 0.5 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |