Project
PRJDB7713: Age associated transcriptome analysis in 5 tissues of zebrafish
Navigation
Downloads
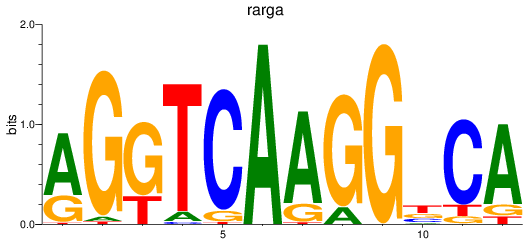
Results for rarga
Z-value: 1.99
Transcription factors associated with rarga
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
rarga
|
ENSDARG00000034117 | retinoic acid receptor gamma a |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| rarga | dr11_v1_chr23_+_35918530_35918530 | -0.88 | 1.4e-31 | Click! |
Activity profile of rarga motif
Sorted Z-values of rarga motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr2_-_42234484 | 42.91 |
ENSDART00000132617
ENSDART00000136690 ENSDART00000141358 |
apom
|
apolipoprotein M |
| chr13_-_12660318 | 31.21 |
ENSDART00000008498
|
adh8a
|
alcohol dehydrogenase 8a |
| chr5_+_28830388 | 30.77 |
ENSDART00000149150
|
serpina7
|
serpin peptidase inhibitor, clade A (alpha-1 antiproteinase, antitrypsin), member 7 |
| chr11_+_37201483 | 30.70 |
ENSDART00000160930
ENSDART00000173439 ENSDART00000171273 |
zgc:112265
|
zgc:112265 |
| chr16_-_17197546 | 29.83 |
ENSDART00000139939
ENSDART00000135146 ENSDART00000063800 ENSDART00000163606 |
gapdh
|
glyceraldehyde-3-phosphate dehydrogenase |
| chr4_-_17409533 | 29.54 |
ENSDART00000011943
|
pah
|
phenylalanine hydroxylase |
| chr5_-_41531629 | 27.42 |
ENSDART00000051082
|
akr1a1a
|
aldo-keto reductase family 1, member A1a (aldehyde reductase) |
| chr7_+_52122224 | 26.80 |
ENSDART00000174268
|
cyp2x12
|
cytochrome P450, family 2, subfamily X, polypeptide 12 |
| chr20_+_31269778 | 25.36 |
ENSDART00000133353
|
apobb.1
|
apolipoprotein Bb, tandem duplicate 1 |
| chr13_-_22862133 | 24.34 |
ENSDART00000138563
|
pbld2
|
phenazine biosynthesis-like protein domain containing 2 |
| chr8_+_1769475 | 24.07 |
ENSDART00000079073
|
serpind1
|
serpin peptidase inhibitor, clade D (heparin cofactor), member 1 |
| chr21_-_30658509 | 23.75 |
ENSDART00000139764
|
si:dkey-22f5.9
|
si:dkey-22f5.9 |
| chr13_-_22843562 | 23.40 |
ENSDART00000142738
|
pbld
|
phenazine biosynthesis like protein domain containing |
| chr25_-_26018424 | 22.87 |
ENSDART00000089332
|
acsbg1
|
acyl-CoA synthetase bubblegum family member 1 |
| chr22_-_23781083 | 22.81 |
ENSDART00000166563
ENSDART00000170458 ENSDART00000166158 ENSDART00000171246 |
cfhl3
|
complement factor H like 3 |
| chr3_-_15080226 | 22.61 |
ENSDART00000109818
ENSDART00000139835 |
nme4
|
NME/NM23 nucleoside diphosphate kinase 4 |
| chr12_-_6172154 | 21.31 |
ENSDART00000185434
|
a1cf
|
apobec1 complementation factor |
| chr20_+_6142433 | 21.26 |
ENSDART00000054084
ENSDART00000136986 |
ttr
|
transthyretin (prealbumin, amyloidosis type I) |
| chr8_+_19514294 | 18.08 |
ENSDART00000170622
|
si:ch73-281k2.5
|
si:ch73-281k2.5 |
| chr6_-_8498908 | 15.52 |
ENSDART00000149222
|
pglyrp2
|
peptidoglycan recognition protein 2 |
| chr16_+_25126935 | 15.50 |
ENSDART00000058945
|
zgc:92590
|
zgc:92590 |
| chr2_+_51028269 | 15.49 |
ENSDART00000161254
|
eef1da
|
eukaryotic translation elongation factor 1 delta a (guanine nucleotide exchange protein) |
| chr2_-_20120904 | 14.29 |
ENSDART00000186002
ENSDART00000124724 |
dpydb
|
dihydropyrimidine dehydrogenase b |
| chr2_-_24289641 | 13.38 |
ENSDART00000128784
ENSDART00000123565 ENSDART00000141922 ENSDART00000184550 ENSDART00000191469 |
myh7l
|
myosin heavy chain 7-like |
| chr6_-_8498676 | 13.23 |
ENSDART00000148627
|
pglyrp2
|
peptidoglycan recognition protein 2 |
| chr17_-_2584423 | 13.17 |
ENSDART00000013506
|
zp3.2
|
zona pellucida glycoprotein 3, tandem duplicate 2 |
| chr13_-_4707018 | 12.99 |
ENSDART00000128422
|
oit3
|
oncoprotein induced transcript 3 |
| chr25_+_3327071 | 12.87 |
ENSDART00000136131
ENSDART00000133243 |
ldhbb
|
lactate dehydrogenase Bb |
| chr14_-_36412473 | 12.77 |
ENSDART00000128244
ENSDART00000138376 |
asb5a
|
ankyrin repeat and SOCS box containing 5a |
| chr12_-_29301022 | 12.77 |
ENSDART00000187826
|
sh2d4bb
|
SH2 domain containing 4Bb |
| chr7_+_52135791 | 12.73 |
ENSDART00000098705
|
cyp2x12
|
cytochrome P450, family 2, subfamily X, polypeptide 12 |
| chr22_-_24967348 | 12.07 |
ENSDART00000153490
ENSDART00000084871 |
fam20cl
|
family with sequence similarity 20, member C like |
| chr25_-_29415369 | 11.03 |
ENSDART00000110774
ENSDART00000019183 |
ugt5a2
ugt5a1
|
UDP glucuronosyltransferase 5 family, polypeptide A2 UDP glucuronosyltransferase 5 family, polypeptide A1 |
| chr13_+_22480857 | 11.02 |
ENSDART00000078721
ENSDART00000044719 ENSDART00000130957 ENSDART00000078757 ENSDART00000130424 ENSDART00000078747 |
ldb3a
|
LIM domain binding 3a |
| chr25_+_3326885 | 10.61 |
ENSDART00000104866
|
ldhbb
|
lactate dehydrogenase Bb |
| chr5_+_19343880 | 10.53 |
ENSDART00000148130
|
acacb
|
acetyl-CoA carboxylase beta |
| chr9_-_43082945 | 10.48 |
ENSDART00000142257
|
ccdc141
|
coiled-coil domain containing 141 |
| chr6_-_49898881 | 10.47 |
ENSDART00000150204
|
atp5f1e
|
ATP synthase F1 subunit epsilon |
| chr9_+_24095677 | 10.16 |
ENSDART00000150443
|
lrrfip1a
|
leucine rich repeat (in FLII) interacting protein 1a |
| chr24_-_9989634 | 10.12 |
ENSDART00000115275
|
zgc:152652
|
zgc:152652 |
| chr6_-_15653494 | 10.11 |
ENSDART00000038133
|
trim63a
|
tripartite motif containing 63a |
| chr17_-_2590222 | 9.90 |
ENSDART00000185711
|
CR759892.1
|
|
| chr2_-_57469115 | 9.79 |
ENSDART00000192201
|
pias4b
|
protein inhibitor of activated STAT, 4b |
| chr22_-_17595310 | 9.70 |
ENSDART00000099056
|
gpx4a
|
glutathione peroxidase 4a |
| chr5_-_64168415 | 9.65 |
ENSDART00000048395
|
cmlc1
|
cardiac myosin light chain-1 |
| chr13_+_22480496 | 9.44 |
ENSDART00000136863
ENSDART00000131870 ENSDART00000078720 ENSDART00000078740 ENSDART00000139218 |
ldb3a
|
LIM domain binding 3a |
| chr2_-_48826707 | 9.38 |
ENSDART00000134711
|
svilb
|
supervillin b |
| chr10_+_28428222 | 9.36 |
ENSDART00000135003
|
si:ch211-222e20.4
|
si:ch211-222e20.4 |
| chr24_-_9979342 | 9.14 |
ENSDART00000138576
ENSDART00000191206 |
zgc:171977
|
zgc:171977 |
| chr8_+_8046086 | 9.05 |
ENSDART00000163232
ENSDART00000111392 |
abcd1
|
ATP-binding cassette, sub-family D (ALD), member 1 |
| chr9_-_1970071 | 8.85 |
ENSDART00000080608
|
hoxd10a
|
homeobox D10a |
| chr19_+_40856807 | 8.70 |
ENSDART00000139083
|
gngt1
|
guanine nucleotide binding protein (G protein), gamma transducing activity polypeptide 1 |
| chr22_-_18779232 | 8.58 |
ENSDART00000186726
|
atp5f1d
|
ATP synthase F1 subunit delta |
| chr6_-_20952187 | 8.32 |
ENSDART00000074327
|
igfbp2a
|
insulin-like growth factor binding protein 2a |
| chr6_-_43047774 | 8.08 |
ENSDART00000161722
|
glyctk
|
glycerate kinase |
| chr1_-_58913813 | 8.08 |
ENSDART00000056494
|
zgc:171687
|
zgc:171687 |
| chr17_-_15498275 | 7.97 |
ENSDART00000156905
ENSDART00000080661 |
si:ch211-266g18.10
|
si:ch211-266g18.10 |
| chr19_-_14155781 | 7.61 |
ENSDART00000169232
|
nr0b2b
|
nuclear receptor subfamily 0, group B, member 2b |
| chr6_+_28054639 | 7.50 |
ENSDART00000187478
ENSDART00000189194 |
si:ch73-194h10.2
|
si:ch73-194h10.2 |
| chr5_+_25585869 | 7.41 |
ENSDART00000138060
|
si:dkey-229d2.7
|
si:dkey-229d2.7 |
| chr10_-_16028082 | 7.38 |
ENSDART00000122540
|
aldh7a1
|
aldehyde dehydrogenase 7 family, member A1 |
| chr4_+_9836465 | 7.29 |
ENSDART00000004879
|
hsp90b1
|
heat shock protein 90, beta (grp94), member 1 |
| chr24_-_14711597 | 7.27 |
ENSDART00000131830
|
jph1a
|
junctophilin 1a |
| chr22_-_18778988 | 7.08 |
ENSDART00000019235
|
atp5f1d
|
ATP synthase F1 subunit delta |
| chr15_+_3284416 | 6.97 |
ENSDART00000187665
ENSDART00000171723 |
foxo1a
|
forkhead box O1 a |
| chr22_-_38258053 | 6.83 |
ENSDART00000132516
|
elavl2
|
ELAV like neuron-specific RNA binding protein 2 |
| chr21_-_43117327 | 6.82 |
ENSDART00000122352
|
p4ha2
|
procollagen-proline, 2-oxoglutarate 4-dioxygenase (proline 4-hydroxylase), alpha polypeptide 2 |
| chr15_+_46386261 | 6.72 |
ENSDART00000191793
|
igsf11
|
immunoglobulin superfamily member 11 |
| chr23_+_36087219 | 6.66 |
ENSDART00000154825
|
hoxc3a
|
homeobox C3a |
| chr19_+_40856534 | 6.59 |
ENSDART00000051950
|
gngt1
|
guanine nucleotide binding protein (G protein), gamma transducing activity polypeptide 1 |
| chr6_-_10835849 | 6.43 |
ENSDART00000005903
ENSDART00000135065 |
atp5mc3b
|
ATP synthase membrane subunit c locus 3b |
| chr6_-_40448416 | 6.42 |
ENSDART00000187019
ENSDART00000179469 |
tatdn2
|
TatD DNase domain containing 2 |
| chr17_+_6538733 | 6.41 |
ENSDART00000193712
|
slc5a6b
|
solute carrier family 5 (sodium/multivitamin and iodide cotransporter), member 6 |
| chr24_+_19986432 | 6.38 |
ENSDART00000184402
|
CR381686.5
|
|
| chr19_-_703898 | 6.13 |
ENSDART00000181096
ENSDART00000121462 |
slc6a19a.2
|
solute carrier family 6 (neutral amino acid transporter), member 19a, tandem duplicate 2 |
| chr4_-_9836342 | 6.10 |
ENSDART00000150519
|
ttc41
|
tetratricopeptide repeat domain 41 |
| chr7_-_16237861 | 6.08 |
ENSDART00000173647
|
btr07
|
bloodthirsty-related gene family, member 7 |
| chr7_-_27033080 | 5.94 |
ENSDART00000173516
|
nucb2a
|
nucleobindin 2a |
| chr12_+_46605600 | 5.92 |
ENSDART00000123533
|
FADS6
|
fatty acid desaturase 6 |
| chr3_-_3210640 | 5.89 |
ENSDART00000187699
|
si:ch211-229i14.2
|
si:ch211-229i14.2 |
| chr7_-_16205471 | 5.87 |
ENSDART00000173584
|
btr05
|
bloodthirsty-related gene family, member 5 |
| chr13_+_38817871 | 5.87 |
ENSDART00000187708
|
col19a1
|
collagen, type XIX, alpha 1 |
| chr21_-_308852 | 5.86 |
ENSDART00000151613
|
lhfpl2a
|
LHFPL tetraspan subfamily member 2a |
| chr22_+_28969071 | 5.79 |
ENSDART00000163427
|
pimr95
|
Pim proto-oncogene, serine/threonine kinase, related 95 |
| chr15_-_36727462 | 5.73 |
ENSDART00000085971
|
nphs1
|
nephrosis 1, congenital, Finnish type (nephrin) |
| chr19_+_30867845 | 5.69 |
ENSDART00000047461
|
mfsd2ab
|
major facilitator superfamily domain containing 2ab |
| chr25_-_37284370 | 5.63 |
ENSDART00000103222
|
nudt7
|
nudix (nucleoside diphosphate linked moiety X)-type motif 7 |
| chr9_-_22834860 | 5.59 |
ENSDART00000146486
|
neb
|
nebulin |
| chr25_-_17367578 | 5.57 |
ENSDART00000064591
ENSDART00000110692 |
cyp2x6
|
cytochrome P450, family 2, subfamily X, polypeptide 6 |
| chr1_-_52167509 | 5.56 |
ENSDART00000184935
|
BX901942.2
|
|
| chr1_+_56886214 | 5.50 |
ENSDART00000152718
ENSDART00000182408 |
si:ch211-1f22.8
|
si:ch211-1f22.8 |
| chr25_-_17368231 | 5.46 |
ENSDART00000189291
|
cyp2x6
|
cytochrome P450, family 2, subfamily X, polypeptide 6 |
| chr15_-_30853246 | 5.43 |
ENSDART00000112511
|
akap1b
|
A kinase (PRKA) anchor protein 1b |
| chr20_+_572037 | 5.37 |
ENSDART00000028062
ENSDART00000152736 ENSDART00000031759 ENSDART00000162198 |
smyd2b
|
SET and MYND domain containing 2b |
| chr22_-_5171829 | 5.34 |
ENSDART00000140313
|
tnfaip8l1
|
tumor necrosis factor, alpha-induced protein 8-like 1 |
| chr24_-_17047918 | 5.25 |
ENSDART00000020204
|
msrb2
|
methionine sulfoxide reductase B2 |
| chr16_+_16978424 | 5.25 |
ENSDART00000143128
|
rpl18
|
ribosomal protein L18 |
| chr14_+_32918484 | 5.20 |
ENSDART00000105721
|
lnx2b
|
ligand of numb-protein X 2b |
| chr17_+_26352372 | 5.10 |
ENSDART00000155177
|
grid1a
|
glutamate receptor, ionotropic, delta 1a |
| chr16_+_32736588 | 4.99 |
ENSDART00000075191
ENSDART00000168358 |
zgc:172323
|
zgc:172323 |
| chr10_+_26813710 | 4.99 |
ENSDART00000136122
|
si:ch211-263p13.7
|
si:ch211-263p13.7 |
| chr14_+_32918172 | 4.97 |
ENSDART00000182867
|
lnx2b
|
ligand of numb-protein X 2b |
| chr1_-_46632948 | 4.90 |
ENSDART00000148893
ENSDART00000053232 |
cdadc1
|
cytidine and dCMP deaminase domain containing 1 |
| chr13_+_22264914 | 4.86 |
ENSDART00000060576
|
myoz1a
|
myozenin 1a |
| chr8_-_13419049 | 4.82 |
ENSDART00000133656
|
pimr101
|
Pim proto-oncogene, serine/threonine kinase, related 101 |
| chr6_+_49021703 | 4.82 |
ENSDART00000149394
|
slc16a1a
|
solute carrier family 16 (monocarboxylate transporter), member 1a |
| chr19_-_48349002 | 4.72 |
ENSDART00000168468
|
si:ch73-359m17.8
|
si:ch73-359m17.8 |
| chr17_-_32370047 | 4.72 |
ENSDART00000145487
|
klf11b
|
Kruppel-like factor 11b |
| chr20_-_28956303 | 4.68 |
ENSDART00000062342
|
acot21
|
acyl-CoA thioesterase 21 |
| chr22_+_29009541 | 4.68 |
ENSDART00000169449
|
pimr97
|
Pim proto-oncogene, serine/threonine kinase, related 97 |
| chr21_-_551014 | 4.64 |
ENSDART00000099252
|
vimr1
|
vimentin-related 1 |
| chr1_+_54626491 | 4.61 |
ENSDART00000136063
|
si:ch211-202h22.9
|
si:ch211-202h22.9 |
| chr9_-_21459074 | 4.58 |
ENSDART00000136427
|
zmym2
|
zinc finger, MYM-type 2 |
| chr11_-_37720266 | 4.57 |
ENSDART00000142734
|
yod1
|
YOD1 deubiquitinase |
| chr8_-_13471916 | 4.55 |
ENSDART00000146558
|
pimr105
|
Pim proto-oncogene, serine/threonine kinase, related 105 |
| chr18_-_22753637 | 4.54 |
ENSDART00000181589
ENSDART00000009912 |
hsf4
|
heat shock transcription factor 4 |
| chr15_-_24883956 | 4.41 |
ENSDART00000113199
|
aipl1
|
aryl hydrocarbon receptor interacting protein-like 1 |
| chr21_-_5879897 | 4.39 |
ENSDART00000184034
|
rpl35
|
ribosomal protein L35 |
| chr7_-_63637644 | 4.37 |
ENSDART00000145168
|
si:ch211-208g1.1
|
si:ch211-208g1.1 |
| chr8_-_13428740 | 4.34 |
ENSDART00000131826
|
CR354547.3
|
|
| chr22_+_29067388 | 4.25 |
ENSDART00000133673
|
pimr100
|
Pim proto-oncogene, serine/threonine kinase, related 100 |
| chr7_-_23768234 | 4.23 |
ENSDART00000173981
|
si:ch211-200p22.4
|
si:ch211-200p22.4 |
| chr8_-_35736141 | 4.10 |
ENSDART00000190863
|
BX322622.1
|
|
| chr1_+_44767657 | 4.07 |
ENSDART00000073717
|
si:dkey-9i23.4
|
si:dkey-9i23.4 |
| chr13_+_24579108 | 4.07 |
ENSDART00000001830
|
echs1
|
enoyl CoA hydratase, short chain, 1, mitochondrial |
| chr25_+_34887999 | 4.03 |
ENSDART00000012287
ENSDART00000182506 ENSDART00000183561 |
spire2
|
spire-type actin nucleation factor 2 |
| chr20_+_23960525 | 3.93 |
ENSDART00000042123
|
cx52.6
|
connexin 52.6 |
| chr6_+_43390387 | 3.90 |
ENSDART00000122793
|
mitfa
|
microphthalmia-associated transcription factor a |
| chr6_+_52235441 | 3.85 |
ENSDART00000056319
|
cox6c
|
cytochrome c oxidase subunit VIc |
| chr15_-_19681870 | 3.85 |
ENSDART00000152266
|
si:dkey-4p15.4
|
si:dkey-4p15.4 |
| chr20_+_33534038 | 3.84 |
ENSDART00000029206
|
kcnf1a
|
potassium voltage-gated channel, subfamily F, member 1a |
| chr1_-_55118745 | 3.80 |
ENSDART00000133915
|
sertad2a
|
SERTA domain containing 2a |
| chr8_+_41712799 | 3.79 |
ENSDART00000150337
|
si:ch211-208g1.1
|
si:ch211-208g1.1 |
| chr5_-_3889047 | 3.78 |
ENSDART00000185817
|
mlxipl
|
MLX interacting protein like |
| chr19_-_15177324 | 3.76 |
ENSDART00000169883
|
phactr4a
|
phosphatase and actin regulator 4a |
| chr2_+_46009507 | 3.75 |
ENSDART00000193596
|
CR762395.1
|
|
| chr2_+_37110504 | 3.75 |
ENSDART00000132794
ENSDART00000042974 |
slc1a8b
|
solute carrier family 1 (glutamate transporter), member 8b |
| chr10_-_40964739 | 3.69 |
ENSDART00000076359
|
npffr1l1
|
neuropeptide FF receptor 1 like 1 |
| chr25_-_19661198 | 3.67 |
ENSDART00000149641
|
atp2b1b
|
ATPase plasma membrane Ca2+ transporting 1b |
| chr12_+_4573696 | 3.64 |
ENSDART00000152534
|
si:dkey-94f20.4
|
si:dkey-94f20.4 |
| chr1_+_57331813 | 3.59 |
ENSDART00000152440
ENSDART00000062841 |
epn3b
|
epsin 3b |
| chr7_+_4193869 | 3.56 |
ENSDART00000138277
ENSDART00000142539 |
si:dkey-28d5.5
|
si:dkey-28d5.5 |
| chr5_-_11971946 | 3.53 |
ENSDART00000166285
|
si:ch73-47f2.1
|
si:ch73-47f2.1 |
| chr10_-_4980150 | 3.53 |
ENSDART00000093228
|
mat2al
|
methionine adenosyltransferase II, alpha-like |
| chr16_-_19026414 | 3.52 |
ENSDART00000141208
|
pimr98
|
Pim proto-oncogene, serine/threonine kinase, related 98 |
| chr7_+_26100024 | 3.48 |
ENSDART00000173726
|
si:ch211-196f2.3
|
si:ch211-196f2.3 |
| chr17_-_51224800 | 3.48 |
ENSDART00000150089
|
psen1
|
presenilin 1 |
| chr20_-_14665002 | 3.45 |
ENSDART00000152816
|
scrn2
|
secernin 2 |
| chr18_-_21746421 | 3.45 |
ENSDART00000188809
|
pskh1
|
protein serine kinase H1 |
| chr6_+_11990733 | 3.41 |
ENSDART00000151075
|
baz2ba
|
bromodomain adjacent to zinc finger domain, 2Ba |
| chr17_-_12196865 | 3.39 |
ENSDART00000154694
|
kif28
|
kinesin family member 28 |
| chr7_-_6417339 | 3.39 |
ENSDART00000172795
|
si:ch73-368j24.12
|
si:ch73-368j24.12 |
| chr20_+_6543674 | 3.38 |
ENSDART00000134204
|
tns3.1
|
tensin 3, tandem duplicate 1 |
| chr22_+_28995391 | 3.33 |
ENSDART00000164486
|
pimr99
|
Pim proto-oncogene, serine/threonine kinase, related 99 |
| chr3_-_2613990 | 3.33 |
ENSDART00000137102
|
si:dkey-217f16.6
|
si:dkey-217f16.6 |
| chr23_-_10017245 | 3.32 |
ENSDART00000131669
|
plxnb1a
|
plexin b1a |
| chr15_+_1137337 | 3.27 |
ENSDART00000191508
|
p2ry13
|
purinergic receptor P2Y13 |
| chr15_-_43768776 | 3.25 |
ENSDART00000170398
|
grm5b
|
glutamate receptor, metabotropic 5b |
| chr15_+_19681718 | 3.21 |
ENSDART00000164803
|
si:dkey-4p15.5
|
si:dkey-4p15.5 |
| chr8_-_50287949 | 3.20 |
ENSDART00000023639
|
nkx2.7
|
NK2 transcription factor related 7 |
| chr1_-_9109699 | 3.14 |
ENSDART00000147833
|
vap
|
vascular associated protein |
| chr10_+_11265387 | 3.10 |
ENSDART00000038888
|
hsdl2
|
hydroxysteroid dehydrogenase like 2 |
| chr2_+_7818368 | 3.08 |
ENSDART00000007068
|
kcnmb2
|
potassium large conductance calcium-activated channel, subfamily M, beta member 2 |
| chr10_-_29120515 | 3.08 |
ENSDART00000162016
ENSDART00000149140 |
zpld1a
|
zona pellucida-like domain containing 1a |
| chr17_-_24364527 | 3.07 |
ENSDART00000155192
|
wdpcp
|
WD repeat containing planar cell polarity effector |
| chr3_-_18792492 | 3.04 |
ENSDART00000134208
ENSDART00000034373 |
hagh
|
hydroxyacylglutathione hydrolase |
| chr16_-_40727455 | 3.02 |
ENSDART00000162331
|
si:dkey-22o22.2
|
si:dkey-22o22.2 |
| chr8_+_20679759 | 2.99 |
ENSDART00000088668
|
nfic
|
nuclear factor I/C |
| chr23_+_16469530 | 2.98 |
ENSDART00000132898
|
ntsr1
|
neurotensin receptor 1 (high affinity) |
| chr5_-_71838520 | 2.86 |
ENSDART00000174396
|
CU927890.1
|
|
| chr8_-_13454281 | 2.84 |
ENSDART00000141959
|
CR354547.2
|
|
| chr2_-_27774783 | 2.83 |
ENSDART00000161864
|
XKR4
|
zgc:123035 |
| chr14_+_46216703 | 2.82 |
ENSDART00000136045
ENSDART00000142317 |
mgst2
|
microsomal glutathione S-transferase 2 |
| chr11_+_3578543 | 2.80 |
ENSDART00000191015
|
si:dkey-33m11.8
|
si:dkey-33m11.8 |
| chr25_-_16826219 | 2.80 |
ENSDART00000191299
ENSDART00000188504 |
dyrk4
|
dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 4 |
| chr9_+_27379193 | 2.78 |
ENSDART00000142656
|
tlr20.4
|
toll-like receptor 20, tandem duplicate 4 |
| chr13_-_9119867 | 2.73 |
ENSDART00000137255
|
si:dkey-112g5.15
|
si:dkey-112g5.15 |
| chr11_+_26403873 | 2.68 |
ENSDART00000109298
|
fer1l4
|
fer-1-like family member 4 |
| chr17_+_51224421 | 2.64 |
ENSDART00000025229
|
adi1
|
acireductone dioxygenase 1 |
| chr9_+_27371240 | 2.63 |
ENSDART00000060337
|
tlr20.3
|
toll-like receptor 20, tandem duplicate 3 |
| chr15_+_24644251 | 2.62 |
ENSDART00000181660
|
smtnl
|
smoothelin, like |
| chr19_+_17642356 | 2.62 |
ENSDART00000176431
|
CR382334.1
|
|
| chr14_+_6159162 | 2.61 |
ENSDART00000128638
|
bscl2l
|
Bernardinelli-Seip congenital lipodystrophy 2, like |
| chr7_-_3657284 | 2.59 |
ENSDART00000142591
|
si:ch211-282j17.3
|
si:ch211-282j17.3 |
| chr18_-_6862738 | 2.58 |
ENSDART00000192592
|
CT009745.1
|
|
| chr8_-_13486258 | 2.58 |
ENSDART00000137459
|
pimr104
|
Pim proto-oncogene, serine/threonine kinase, related 104 |
| chr15_+_19682013 | 2.58 |
ENSDART00000127368
|
si:dkey-4p15.5
|
si:dkey-4p15.5 |
| chr2_-_25553607 | 2.56 |
ENSDART00000150075
ENSDART00000078695 |
ghsra
|
growth hormone secretagogue receptor a |
| chr20_-_14259032 | 2.55 |
ENSDART00000169482
|
dlgap2b
|
discs, large (Drosophila) homolog-associated protein 2b |
| chr21_+_43702016 | 2.54 |
ENSDART00000017176
|
dkc1
|
dyskeratosis congenita 1, dyskerin |
| chr5_+_13243446 | 2.52 |
ENSDART00000186140
|
CR933528.1
|
|
| chr20_+_192170 | 2.51 |
ENSDART00000189675
|
cx28.8
|
connexin 28.8 |
| chr12_+_33433882 | 2.51 |
ENSDART00000185053
|
fasn
|
fatty acid synthase |
| chr6_+_27418541 | 2.49 |
ENSDART00000187410
|
crocc2
|
ciliary rootlet coiled-coil, rootletin family member 2 |
| chr8_-_13442871 | 2.48 |
ENSDART00000144887
|
pimr106
|
Pim proto-oncogene, serine/threonine kinase, related 106 |
| chr11_-_20956309 | 2.47 |
ENSDART00000188659
|
CABZ01008739.1
|
|
| chr16_+_6021908 | 2.46 |
ENSDART00000163786
|
BX511115.1
|
|
| chr13_-_37254777 | 2.46 |
ENSDART00000139734
|
syne2b
|
spectrin repeat containing, nuclear envelope 2b |
| chr18_-_16791331 | 2.44 |
ENSDART00000148222
|
ampd3b
|
adenosine monophosphate deaminase 3b |
| chr7_-_58729894 | 2.41 |
ENSDART00000149347
|
chchd7
|
coiled-coil-helix-coiled-coil-helix domain containing 7 |
Network of associatons between targets according to the STRING database.
First level regulatory network of rarga
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 9.6 | 28.8 | GO:0098543 | detection of biotic stimulus(GO:0009595) detection of bacterium(GO:0016045) detection of other organism(GO:0098543) detection of external biotic stimulus(GO:0098581) |
| 5.1 | 25.4 | GO:0006642 | triglyceride mobilization(GO:0006642) |
| 4.8 | 14.3 | GO:0046125 | deoxyribonucleoside metabolic process(GO:0009120) thymidine metabolic process(GO:0046104) pyrimidine deoxyribonucleoside metabolic process(GO:0046125) |
| 4.5 | 31.2 | GO:0046294 | formaldehyde metabolic process(GO:0046292) formaldehyde catabolic process(GO:0046294) |
| 3.0 | 12.1 | GO:0070166 | enamel mineralization(GO:0070166) |
| 2.8 | 27.7 | GO:0072337 | modified amino acid transport(GO:0072337) |
| 2.6 | 10.5 | GO:2001295 | malonyl-CoA biosynthetic process(GO:2001295) |
| 2.5 | 29.5 | GO:1902221 | L-phenylalanine metabolic process(GO:0006558) L-phenylalanine catabolic process(GO:0006559) erythrose 4-phosphate/phosphoenolpyruvate family amino acid metabolic process(GO:1902221) erythrose 4-phosphate/phosphoenolpyruvate family amino acid catabolic process(GO:1902222) |
| 1.7 | 23.8 | GO:0016556 | mRNA modification(GO:0016556) |
| 1.6 | 9.6 | GO:0055005 | ventricular cardiac myofibril assembly(GO:0055005) |
| 1.5 | 4.5 | GO:1902746 | regulation of lens fiber cell differentiation(GO:1902746) |
| 1.4 | 22.6 | GO:0006228 | UTP biosynthetic process(GO:0006228) UTP metabolic process(GO:0046051) |
| 1.3 | 5.2 | GO:0030091 | protein repair(GO:0030091) |
| 1.3 | 32.6 | GO:0015986 | energy coupled proton transport, down electrochemical gradient(GO:0015985) ATP synthesis coupled proton transport(GO:0015986) |
| 1.2 | 7.3 | GO:0035889 | otolith tethering(GO:0035889) |
| 1.1 | 4.6 | GO:1990167 | protein K27-linked deubiquitination(GO:1990167) |
| 1.1 | 5.7 | GO:0045056 | transcytosis(GO:0045056) |
| 1.1 | 9.1 | GO:0031643 | positive regulation of myelination(GO:0031643) |
| 1.1 | 3.2 | GO:0099552 | trans-synaptic signaling by lipid, modulating synaptic transmission(GO:0099552) trans-synaptic signaling by endocannabinoid, modulating synaptic transmission(GO:0099553) |
| 1.0 | 22.9 | GO:0042759 | long-chain fatty acid biosynthetic process(GO:0042759) |
| 1.0 | 3.0 | GO:0006089 | lactate metabolic process(GO:0006089) |
| 1.0 | 7.0 | GO:0048745 | smooth muscle tissue development(GO:0048745) |
| 0.9 | 5.6 | GO:0071691 | cardiac muscle thin filament assembly(GO:0071691) |
| 0.9 | 45.4 | GO:0042737 | drug metabolic process(GO:0017144) drug catabolic process(GO:0042737) exogenous drug catabolic process(GO:0042738) |
| 0.9 | 29.8 | GO:0050821 | protein stabilization(GO:0050821) |
| 0.8 | 8.3 | GO:0040014 | regulation of multicellular organism growth(GO:0040014) |
| 0.8 | 13.2 | GO:2000344 | binding of sperm to zona pellucida(GO:0007339) egg coat formation(GO:0035803) regulation of acrosome reaction(GO:0060046) positive regulation of acrosome reaction(GO:2000344) |
| 0.7 | 5.4 | GO:0018026 | peptidyl-lysine monomethylation(GO:0018026) |
| 0.7 | 56.8 | GO:0010951 | negative regulation of endopeptidase activity(GO:0010951) |
| 0.7 | 5.9 | GO:0046546 | development of primary male sexual characteristics(GO:0046546) |
| 0.6 | 5.6 | GO:0009109 | coenzyme catabolic process(GO:0009109) |
| 0.6 | 5.9 | GO:0032099 | negative regulation of response to food(GO:0032096) negative regulation of appetite(GO:0032099) |
| 0.6 | 2.8 | GO:0019370 | leukotriene metabolic process(GO:0006691) leukotriene biosynthetic process(GO:0019370) |
| 0.5 | 3.8 | GO:2001045 | negative regulation of integrin-mediated signaling pathway(GO:2001045) |
| 0.5 | 2.1 | GO:0016554 | cytidine to uridine editing(GO:0016554) |
| 0.5 | 2.6 | GO:0019509 | L-methionine biosynthetic process from methylthioadenosine(GO:0019509) amino acid salvage(GO:0043102) L-methionine biosynthetic process(GO:0071265) L-methionine salvage(GO:0071267) |
| 0.5 | 3.1 | GO:0005513 | detection of calcium ion(GO:0005513) |
| 0.5 | 8.1 | GO:0001867 | complement activation, lectin pathway(GO:0001867) |
| 0.5 | 4.0 | GO:0040038 | meiotic cytokinesis(GO:0033206) polar body extrusion after meiotic divisions(GO:0040038) |
| 0.5 | 3.5 | GO:0006556 | S-adenosylmethionine biosynthetic process(GO:0006556) |
| 0.5 | 2.3 | GO:0043029 | T cell homeostasis(GO:0043029) |
| 0.4 | 1.8 | GO:0030718 | germ-line stem cell population maintenance(GO:0030718) |
| 0.4 | 2.2 | GO:0035461 | vitamin transmembrane transport(GO:0035461) vitamin A transport(GO:0071938) vitamin A import(GO:0071939) |
| 0.4 | 2.6 | GO:0034389 | lipid particle organization(GO:0034389) |
| 0.4 | 2.1 | GO:0055071 | manganese ion homeostasis(GO:0055071) |
| 0.4 | 1.7 | GO:0060092 | regulation of synaptic transmission, glycinergic(GO:0060092) |
| 0.4 | 2.1 | GO:0000738 | DNA catabolic process, exonucleolytic(GO:0000738) |
| 0.4 | 6.7 | GO:0097324 | melanocyte migration(GO:0097324) |
| 0.4 | 1.5 | GO:0051563 | smooth endoplasmic reticulum calcium ion homeostasis(GO:0051563) |
| 0.4 | 10.2 | GO:0070936 | protein K48-linked ubiquitination(GO:0070936) |
| 0.4 | 6.8 | GO:0019471 | peptidyl-proline hydroxylation to 4-hydroxy-L-proline(GO:0018401) 4-hydroxyproline metabolic process(GO:0019471) |
| 0.3 | 6.5 | GO:0043433 | negative regulation of sequence-specific DNA binding transcription factor activity(GO:0043433) |
| 0.3 | 5.4 | GO:0010738 | regulation of protein kinase A signaling(GO:0010738) |
| 0.3 | 9.8 | GO:0016925 | protein sumoylation(GO:0016925) |
| 0.3 | 1.1 | GO:0060829 | negative regulation of canonical Wnt signaling pathway involved in neural plate anterior/posterior pattern formation(GO:0060829) |
| 0.3 | 1.9 | GO:1901673 | regulation of mitotic spindle assembly(GO:1901673) |
| 0.3 | 1.5 | GO:1900029 | positive regulation of ruffle assembly(GO:1900029) |
| 0.3 | 3.8 | GO:0048096 | chromatin-mediated maintenance of transcription(GO:0048096) |
| 0.2 | 3.2 | GO:0003207 | cardiac chamber formation(GO:0003207) |
| 0.2 | 0.7 | GO:0034134 | toll-like receptor 2 signaling pathway(GO:0034134) |
| 0.2 | 4.4 | GO:0000463 | maturation of LSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000463) |
| 0.2 | 15.5 | GO:0006414 | translational elongation(GO:0006414) |
| 0.2 | 2.1 | GO:0070475 | rRNA base methylation(GO:0070475) |
| 0.2 | 1.6 | GO:0048660 | regulation of smooth muscle cell proliferation(GO:0048660) negative regulation of smooth muscle cell proliferation(GO:0048662) |
| 0.2 | 2.3 | GO:0033211 | adiponectin-activated signaling pathway(GO:0033211) |
| 0.2 | 10.5 | GO:0019226 | transmission of nerve impulse(GO:0019226) |
| 0.2 | 9.2 | GO:0051016 | barbed-end actin filament capping(GO:0051016) |
| 0.2 | 3.3 | GO:0045974 | miRNA mediated inhibition of translation(GO:0035278) negative regulation of translation, ncRNA-mediated(GO:0040033) regulation of translation, ncRNA-mediated(GO:0045974) |
| 0.2 | 1.4 | GO:0006627 | protein processing involved in protein targeting to mitochondrion(GO:0006627) |
| 0.2 | 3.4 | GO:0016584 | nucleosome positioning(GO:0016584) |
| 0.2 | 2.4 | GO:0032264 | IMP salvage(GO:0032264) |
| 0.2 | 3.3 | GO:1902287 | semaphorin-plexin signaling pathway involved in neuron projection guidance(GO:1902285) semaphorin-plexin signaling pathway involved in axon guidance(GO:1902287) |
| 0.2 | 23.8 | GO:0042742 | defense response to bacterium(GO:0042742) |
| 0.1 | 2.5 | GO:0098962 | regulation of postsynaptic neurotransmitter receptor activity(GO:0098962) |
| 0.1 | 3.1 | GO:0042541 | hemoglobin biosynthetic process(GO:0042541) |
| 0.1 | 4.2 | GO:0016185 | synaptic vesicle budding from presynaptic endocytic zone membrane(GO:0016185) |
| 0.1 | 6.6 | GO:0039021 | pronephric glomerulus development(GO:0039021) |
| 0.1 | 1.1 | GO:1902262 | apoptotic process involved in patterning of blood vessels(GO:1902262) |
| 0.1 | 5.3 | GO:0032007 | negative regulation of TOR signaling(GO:0032007) |
| 0.1 | 1.3 | GO:0050936 | xanthophore differentiation(GO:0050936) |
| 0.1 | 16.2 | GO:0007626 | locomotory behavior(GO:0007626) |
| 0.1 | 2.2 | GO:0002430 | complement receptor mediated signaling pathway(GO:0002430) |
| 0.1 | 0.6 | GO:0031174 | lifelong otolith mineralization(GO:0031174) |
| 0.1 | 4.9 | GO:0002224 | toll-like receptor signaling pathway(GO:0002224) |
| 0.1 | 2.0 | GO:0018095 | protein polyglutamylation(GO:0018095) |
| 0.1 | 1.4 | GO:0036158 | outer dynein arm assembly(GO:0036158) |
| 0.1 | 9.3 | GO:0045732 | positive regulation of protein catabolic process(GO:0045732) |
| 0.1 | 0.3 | GO:0097250 | mitochondrial respiratory chain supercomplex assembly(GO:0097250) |
| 0.1 | 0.9 | GO:0044805 | late nucleophagy(GO:0044805) |
| 0.1 | 7.5 | GO:0007605 | sensory perception of sound(GO:0007605) |
| 0.1 | 29.4 | GO:0046777 | protein autophosphorylation(GO:0046777) |
| 0.1 | 0.3 | GO:0009162 | deoxyribonucleoside monophosphate metabolic process(GO:0009162) |
| 0.1 | 9.1 | GO:0006979 | response to oxidative stress(GO:0006979) |
| 0.1 | 0.4 | GO:0039528 | cytoplasmic pattern recognition receptor signaling pathway in response to virus(GO:0039528) cellular response to virus(GO:0098586) |
| 0.1 | 3.9 | GO:0030318 | melanocyte differentiation(GO:0030318) |
| 0.1 | 6.1 | GO:0035249 | synaptic transmission, glutamatergic(GO:0035249) |
| 0.1 | 5.7 | GO:0033339 | pectoral fin development(GO:0033339) |
| 0.1 | 5.9 | GO:1902284 | axon extension involved in axon guidance(GO:0048846) neuron projection extension involved in neuron projection guidance(GO:1902284) |
| 0.1 | 3.9 | GO:0006119 | oxidative phosphorylation(GO:0006119) |
| 0.1 | 3.7 | GO:0015914 | phospholipid transport(GO:0015914) |
| 0.1 | 0.7 | GO:0035914 | skeletal muscle cell differentiation(GO:0035914) |
| 0.1 | 2.8 | GO:0018107 | peptidyl-threonine phosphorylation(GO:0018107) |
| 0.0 | 4.7 | GO:0006637 | acyl-CoA metabolic process(GO:0006637) thioester metabolic process(GO:0035383) |
| 0.0 | 1.2 | GO:0035845 | photoreceptor cell outer segment organization(GO:0035845) |
| 0.0 | 0.6 | GO:0021772 | olfactory bulb development(GO:0021772) olfactory lobe development(GO:0021988) |
| 0.0 | 0.4 | GO:0030324 | respiratory tube development(GO:0030323) lung development(GO:0030324) |
| 0.0 | 1.0 | GO:0097120 | receptor localization to synapse(GO:0097120) |
| 0.0 | 6.7 | GO:0048704 | embryonic skeletal system morphogenesis(GO:0048704) |
| 0.0 | 0.2 | GO:0003365 | establishment of cell polarity involved in ameboidal cell migration(GO:0003365) |
| 0.0 | 3.1 | GO:0044782 | cilium organization(GO:0044782) |
| 0.0 | 4.8 | GO:0007601 | visual perception(GO:0007601) |
| 0.0 | 0.5 | GO:0002098 | tRNA wobble uridine modification(GO:0002098) |
| 0.0 | 0.5 | GO:0043252 | sodium-independent organic anion transport(GO:0043252) |
| 0.0 | 2.4 | GO:0006888 | ER to Golgi vesicle-mediated transport(GO:0006888) |
| 0.0 | 1.0 | GO:0006937 | regulation of muscle contraction(GO:0006937) |
| 0.0 | 2.1 | GO:0051260 | protein homooligomerization(GO:0051260) |
| 0.0 | 0.3 | GO:0097320 | membrane tubulation(GO:0097320) |
| 0.0 | 4.8 | GO:0006412 | translation(GO:0006412) |
| 0.0 | 6.3 | GO:0016567 | protein ubiquitination(GO:0016567) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 6.5 | 26.1 | GO:0000275 | mitochondrial proton-transporting ATP synthase complex, catalytic core F(1)(GO:0000275) |
| 5.1 | 25.4 | GO:0034359 | mature chylomicron(GO:0034359) |
| 2.2 | 15.5 | GO:0005853 | eukaryotic translation elongation factor 1 complex(GO:0005853) |
| 1.5 | 7.3 | GO:0030314 | junctional membrane complex(GO:0030314) |
| 1.0 | 3.1 | GO:0097541 | axonemal basal plate(GO:0097541) |
| 0.9 | 15.3 | GO:0031680 | G-protein beta/gamma-subunit complex(GO:0031680) |
| 0.6 | 2.5 | GO:0031429 | box H/ACA snoRNP complex(GO:0031429) box H/ACA RNP complex(GO:0072588) |
| 0.6 | 4.0 | GO:0035253 | ciliary rootlet(GO:0035253) |
| 0.4 | 20.5 | GO:0031941 | filamentous actin(GO:0031941) |
| 0.4 | 6.4 | GO:0000276 | mitochondrial proton-transporting ATP synthase complex, coupling factor F(o)(GO:0000276) |
| 0.4 | 4.2 | GO:0098888 | presynaptic endocytic zone(GO:0098833) presynaptic endocytic zone membrane(GO:0098835) extrinsic component of presynaptic membrane(GO:0098888) extrinsic component of presynaptic endocytic zone(GO:0098894) |
| 0.4 | 1.8 | GO:1990923 | PET complex(GO:1990923) |
| 0.3 | 8.7 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 0.3 | 1.4 | GO:0042720 | mitochondrial inner membrane peptidase complex(GO:0042720) |
| 0.2 | 27.2 | GO:0016324 | apical plasma membrane(GO:0016324) |
| 0.2 | 5.2 | GO:0005791 | rough endoplasmic reticulum(GO:0005791) |
| 0.2 | 3.3 | GO:0002116 | semaphorin receptor complex(GO:0002116) |
| 0.1 | 10.4 | GO:0030018 | Z disc(GO:0030018) |
| 0.1 | 1.4 | GO:0036157 | outer dynein arm(GO:0036157) |
| 0.1 | 5.7 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.1 | 13.9 | GO:0016459 | myosin complex(GO:0016459) |
| 0.1 | 5.9 | GO:0005793 | endoplasmic reticulum-Golgi intermediate compartment(GO:0005793) |
| 0.1 | 9.6 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.1 | 3.7 | GO:0005758 | mitochondrial intermembrane space(GO:0005758) organelle envelope lumen(GO:0031970) |
| 0.1 | 4.3 | GO:0099634 | postsynaptic density membrane(GO:0098839) postsynaptic specialization membrane(GO:0099634) |
| 0.1 | 164.8 | GO:0005576 | extracellular region(GO:0005576) |
| 0.1 | 4.4 | GO:0022625 | cytosolic large ribosomal subunit(GO:0022625) |
| 0.1 | 5.1 | GO:0034705 | voltage-gated potassium channel complex(GO:0008076) potassium channel complex(GO:0034705) |
| 0.1 | 2.5 | GO:0098978 | glutamatergic synapse(GO:0098978) |
| 0.1 | 2.1 | GO:0032040 | small-subunit processome(GO:0032040) |
| 0.1 | 0.5 | GO:0033588 | Elongator holoenzyme complex(GO:0033588) |
| 0.0 | 2.0 | GO:0005778 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) |
| 0.0 | 4.1 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.0 | 1.0 | GO:0005763 | organellar small ribosomal subunit(GO:0000314) mitochondrial small ribosomal subunit(GO:0005763) |
| 0.0 | 3.8 | GO:0030027 | lamellipodium(GO:0030027) |
| 0.0 | 3.1 | GO:0042579 | peroxisome(GO:0005777) microbody(GO:0042579) |
| 0.0 | 1.5 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.0 | 3.3 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.0 | 6.7 | GO:0048471 | perinuclear region of cytoplasm(GO:0048471) |
| 0.0 | 1.9 | GO:0000313 | organellar ribosome(GO:0000313) mitochondrial ribosome(GO:0005761) |
| 0.0 | 4.6 | GO:0005765 | lysosomal membrane(GO:0005765) lytic vacuole membrane(GO:0098852) |
| 0.0 | 0.9 | GO:0000421 | autophagosome membrane(GO:0000421) |
| 0.0 | 3.4 | GO:0000786 | nucleosome(GO:0000786) |
| 0.0 | 3.0 | GO:0005759 | mitochondrial matrix(GO:0005759) |
| 0.0 | 2.2 | GO:0099501 | synaptic vesicle membrane(GO:0030672) exocytic vesicle membrane(GO:0099501) |
| 0.0 | 5.8 | GO:0005911 | cell-cell junction(GO:0005911) |
| 0.0 | 4.9 | GO:0000785 | chromatin(GO:0000785) |
| 0.0 | 1.0 | GO:0005861 | troponin complex(GO:0005861) |
| 0.0 | 0.3 | GO:0005964 | phosphorylase kinase complex(GO:0005964) |
| 0.0 | 2.0 | GO:0036064 | ciliary basal body(GO:0036064) |
| 0.0 | 0.6 | GO:0005844 | polysome(GO:0005844) |
| 0.0 | 21.4 | GO:0005829 | cytosol(GO:0005829) |
| 0.0 | 0.2 | GO:1990589 | ATF4-CREB1 transcription factor complex(GO:1990589) |
| 0.0 | 8.5 | GO:0005789 | endoplasmic reticulum membrane(GO:0005789) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 9.6 | 28.8 | GO:0016019 | peptidoglycan receptor activity(GO:0016019) |
| 7.5 | 29.8 | GO:0043891 | glyceraldehyde-3-phosphate dehydrogenase (NAD+) (phosphorylating) activity(GO:0004365) glyceraldehyde-3-phosphate dehydrogenase (NAD(P)+) (phosphorylating) activity(GO:0043891) |
| 6.2 | 31.2 | GO:0051903 | alcohol dehydrogenase activity, zinc-dependent(GO:0004024) S-(hydroxymethyl)glutathione dehydrogenase activity(GO:0051903) |
| 5.5 | 27.4 | GO:0004032 | alditol:NADP+ 1-oxidoreductase activity(GO:0004032) |
| 5.3 | 21.3 | GO:0070324 | thyroid hormone binding(GO:0070324) |
| 3.4 | 23.5 | GO:0004459 | L-lactate dehydrogenase activity(GO:0004459) |
| 3.3 | 29.5 | GO:0016714 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced pteridine as one donor, and incorporation of one atom of oxygen(GO:0016714) |
| 2.9 | 14.3 | GO:0017113 | uracil binding(GO:0002058) pyrimidine nucleobase binding(GO:0002061) dihydropyrimidine dehydrogenase (NADP+) activity(GO:0017113) |
| 2.8 | 25.4 | GO:0050750 | low-density lipoprotein particle receptor binding(GO:0050750) lipoprotein particle receptor binding(GO:0070325) |
| 2.6 | 10.5 | GO:0003989 | acetyl-CoA carboxylase activity(GO:0003989) |
| 2.4 | 26.1 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 2.0 | 8.1 | GO:0008887 | glycerate kinase activity(GO:0008887) |
| 1.9 | 9.7 | GO:0047066 | phospholipid-hydroperoxide glutathione peroxidase activity(GO:0047066) |
| 1.8 | 22.9 | GO:0016405 | CoA-ligase activity(GO:0016405) |
| 1.6 | 6.4 | GO:0008523 | sodium-dependent multivitamin transmembrane transporter activity(GO:0008523) |
| 1.3 | 5.2 | GO:0033745 | L-methionine-(R)-S-oxide reductase activity(GO:0033745) |
| 1.2 | 13.2 | GO:0035804 | structural constituent of egg coat(GO:0035804) |
| 1.0 | 20.5 | GO:0051371 | muscle alpha-actinin binding(GO:0051371) alpha-actinin binding(GO:0051393) |
| 1.0 | 7.0 | GO:0004126 | cytidine deaminase activity(GO:0004126) |
| 0.9 | 3.5 | GO:0004478 | methionine adenosyltransferase activity(GO:0004478) |
| 0.8 | 22.6 | GO:0004550 | nucleoside diphosphate kinase activity(GO:0004550) |
| 0.8 | 8.3 | GO:0031995 | insulin-like growth factor I binding(GO:0031994) insulin-like growth factor II binding(GO:0031995) |
| 0.8 | 9.8 | GO:0061665 | SUMO ligase activity(GO:0061665) |
| 0.7 | 15.3 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.7 | 18.1 | GO:0005112 | Notch binding(GO:0005112) |
| 0.7 | 2.8 | GO:0004464 | leukotriene-C4 synthase activity(GO:0004464) |
| 0.7 | 45.4 | GO:0016712 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced flavin or flavoprotein as one donor, and incorporation of one atom of oxygen(GO:0016712) |
| 0.7 | 3.5 | GO:0070004 | cysteine-type exopeptidase activity(GO:0070004) |
| 0.5 | 3.7 | GO:1990050 | phosphatidic acid transporter activity(GO:1990050) |
| 0.5 | 6.8 | GO:0004656 | procollagen-proline 4-dioxygenase activity(GO:0004656) |
| 0.5 | 4.1 | GO:0004300 | enoyl-CoA hydratase activity(GO:0004300) |
| 0.5 | 3.4 | GO:0010859 | calcium-dependent cysteine-type endopeptidase inhibitor activity(GO:0010859) |
| 0.5 | 4.9 | GO:0051373 | telethonin binding(GO:0031433) FATZ binding(GO:0051373) |
| 0.5 | 54.8 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.5 | 3.2 | GO:0099530 | G-protein coupled receptor activity involved in regulation of postsynaptic membrane potential(GO:0099530) |
| 0.5 | 1.8 | GO:0004968 | gonadotropin-releasing hormone receptor activity(GO:0004968) |
| 0.4 | 4.8 | GO:0015129 | lactate transmembrane transporter activity(GO:0015129) |
| 0.4 | 10.3 | GO:0016289 | CoA hydrolase activity(GO:0016289) |
| 0.4 | 2.1 | GO:0005384 | manganese ion transmembrane transporter activity(GO:0005384) |
| 0.3 | 15.5 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.3 | 2.2 | GO:0034632 | retinol transporter activity(GO:0034632) |
| 0.3 | 21.3 | GO:0003727 | single-stranded RNA binding(GO:0003727) |
| 0.3 | 4.2 | GO:0032050 | clathrin heavy chain binding(GO:0032050) |
| 0.3 | 5.4 | GO:0034237 | protein kinase A regulatory subunit binding(GO:0034237) |
| 0.3 | 4.7 | GO:0046975 | histone methyltransferase activity (H3-K36 specific)(GO:0046975) |
| 0.3 | 6.4 | GO:0016888 | endodeoxyribonuclease activity, producing 5'-phosphomonoesters(GO:0016888) |
| 0.2 | 1.8 | GO:0034584 | piRNA binding(GO:0034584) |
| 0.2 | 9.2 | GO:0005546 | phosphatidylinositol-4,5-bisphosphate binding(GO:0005546) |
| 0.2 | 2.4 | GO:0047623 | AMP deaminase activity(GO:0003876) adenosine-phosphate deaminase activity(GO:0047623) |
| 0.2 | 3.3 | GO:0001608 | G-protein coupled nucleotide receptor activity(GO:0001608) G-protein coupled purinergic nucleotide receptor activity(GO:0045028) |
| 0.2 | 11.6 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.2 | 3.8 | GO:0019211 | phosphatase activator activity(GO:0019211) protein phosphatase activator activity(GO:0072542) |
| 0.2 | 3.1 | GO:0015269 | calcium-activated potassium channel activity(GO:0015269) |
| 0.2 | 6.1 | GO:1904315 | neurotransmitter receptor activity involved in regulation of postsynaptic membrane potential(GO:0099529) transmitter-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1904315) |
| 0.2 | 7.4 | GO:0016620 | oxidoreductase activity, acting on the aldehyde or oxo group of donors, NAD or NADP as acceptor(GO:0016620) |
| 0.2 | 1.5 | GO:0042500 | aspartic endopeptidase activity, intramembrane cleaving(GO:0042500) |
| 0.2 | 3.4 | GO:0031492 | nucleosomal DNA binding(GO:0031492) |
| 0.1 | 2.0 | GO:0070739 | protein-glutamic acid ligase activity(GO:0070739) tubulin-glutamic acid ligase activity(GO:0070740) |
| 0.1 | 17.3 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.1 | 3.9 | GO:0015002 | cytochrome-c oxidase activity(GO:0004129) heme-copper terminal oxidase activity(GO:0015002) oxidoreductase activity, acting on a heme group of donors(GO:0016675) oxidoreductase activity, acting on a heme group of donors, oxygen as acceptor(GO:0016676) |
| 0.1 | 5.7 | GO:0005548 | phospholipid transporter activity(GO:0005548) |
| 0.1 | 2.5 | GO:0009982 | pseudouridine synthase activity(GO:0009982) |
| 0.1 | 2.0 | GO:0005324 | long-chain fatty acid transporter activity(GO:0005324) |
| 0.1 | 8.0 | GO:0008188 | neuropeptide receptor activity(GO:0008188) |
| 0.1 | 2.1 | GO:0008649 | rRNA methyltransferase activity(GO:0008649) |
| 0.1 | 1.6 | GO:0004955 | prostaglandin receptor activity(GO:0004955) |
| 0.1 | 1.7 | GO:0015187 | glycine transmembrane transporter activity(GO:0015187) |
| 0.1 | 8.1 | GO:0003823 | antigen binding(GO:0003823) |
| 0.1 | 1.4 | GO:0045503 | dynein light chain binding(GO:0045503) |
| 0.1 | 3.3 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.1 | 2.2 | GO:0004875 | complement receptor activity(GO:0004875) |
| 0.1 | 3.0 | GO:0016790 | thiolester hydrolase activity(GO:0016790) |
| 0.1 | 1.0 | GO:0031013 | troponin C binding(GO:0030172) troponin I binding(GO:0031013) |
| 0.1 | 6.8 | GO:0003730 | mRNA 3'-UTR binding(GO:0003730) |
| 0.1 | 6.4 | GO:0015078 | hydrogen ion transmembrane transporter activity(GO:0015078) |
| 0.1 | 2.3 | GO:0008331 | high voltage-gated calcium channel activity(GO:0008331) |
| 0.1 | 7.3 | GO:0051082 | unfolded protein binding(GO:0051082) |
| 0.1 | 18.0 | GO:0003774 | motor activity(GO:0003774) |
| 0.1 | 2.6 | GO:0016702 | oxidoreductase activity, acting on single donors with incorporation of molecular oxygen, incorporation of two atoms of oxygen(GO:0016702) |
| 0.1 | 2.8 | GO:0004712 | protein serine/threonine/tyrosine kinase activity(GO:0004712) |
| 0.1 | 12.5 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.1 | 1.1 | GO:0005520 | insulin-like growth factor binding(GO:0005520) |
| 0.1 | 0.4 | GO:0016918 | retinal binding(GO:0016918) |
| 0.1 | 9.9 | GO:0015293 | symporter activity(GO:0015293) |
| 0.0 | 6.5 | GO:0003714 | transcription corepressor activity(GO:0003714) |
| 0.0 | 26.3 | GO:0004842 | ubiquitin-protein transferase activity(GO:0004842) |
| 0.0 | 0.2 | GO:0004999 | vasoactive intestinal polypeptide receptor activity(GO:0004999) |
| 0.0 | 5.8 | GO:0016853 | isomerase activity(GO:0016853) |
| 0.0 | 2.1 | GO:0008408 | 3'-5' exonuclease activity(GO:0008408) |
| 0.0 | 0.4 | GO:0004526 | ribonuclease P activity(GO:0004526) |
| 0.0 | 27.6 | GO:0004674 | protein serine/threonine kinase activity(GO:0004674) |
| 0.0 | 2.0 | GO:0003724 | RNA helicase activity(GO:0003724) |
| 0.0 | 1.2 | GO:0015485 | cholesterol binding(GO:0015485) |
| 0.0 | 0.5 | GO:0015347 | sodium-independent organic anion transmembrane transporter activity(GO:0015347) |
| 0.0 | 3.9 | GO:0008236 | serine-type peptidase activity(GO:0008236) serine hydrolase activity(GO:0017171) |
| 0.0 | 2.1 | GO:0001228 | transcriptional activator activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001228) |
| 0.0 | 0.3 | GO:0019239 | deaminase activity(GO:0019239) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 15.3 | PID RHODOPSIN PATHWAY | Visual signal transduction: Rods |
| 0.5 | 31.8 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.5 | 21.1 | PID HNF3B PATHWAY | FOXA2 and FOXA3 transcription factor networks |
| 0.4 | 61.7 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.2 | 7.3 | PID IL6 7 PATHWAY | IL6-mediated signaling events |
| 0.2 | 3.4 | PID NEPHRIN NEPH1 PATHWAY | Nephrin/Neph1 signaling in the kidney podocyte |
| 0.2 | 5.9 | NABA COLLAGENS | Genes encoding collagen proteins |
| 0.1 | 3.0 | PID HNF3A PATHWAY | FOXA1 transcription factor network |
| 0.1 | 2.1 | PID IL4 2PATHWAY | IL4-mediated signaling events |
| 0.1 | 14.9 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
| 0.1 | 1.5 | PID SYNDECAN 3 PATHWAY | Syndecan-3-mediated signaling events |
| 0.1 | 5.6 | PID P73PATHWAY | p73 transcription factor network |
| 0.1 | 1.0 | PID TRAIL PATHWAY | TRAIL signaling pathway |
| 0.1 | 0.7 | PID TOLL ENDOGENOUS PATHWAY | Endogenous TLR signaling |
| 0.1 | 2.5 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.0 | 0.6 | PID TXA2PATHWAY | Thromboxane A2 receptor signaling |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.8 | 15.3 | REACTOME OLFACTORY SIGNALING PATHWAY | Genes involved in Olfactory Signaling Pathway |
| 1.1 | 22.6 | REACTOME SYNTHESIS AND INTERCONVERSION OF NUCLEOTIDE DI AND TRIPHOSPHATES | Genes involved in Synthesis and interconversion of nucleotide di- and triphosphates |
| 0.9 | 9.1 | REACTOME ALPHA LINOLENIC ACID ALA METABOLISM | Genes involved in alpha-linolenic acid (ALA) metabolism |
| 0.9 | 21.3 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.6 | 9.4 | REACTOME ACTIVATED AMPK STIMULATES FATTY ACID OXIDATION IN MUSCLE | Genes involved in Activated AMPK stimulates fatty-acid oxidation in muscle |
| 0.6 | 7.3 | REACTOME TRAFFICKING AND PROCESSING OF ENDOSOMAL TLR | Genes involved in Trafficking and processing of endosomal TLR |
| 0.5 | 11.5 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.3 | 1.6 | REACTOME PROSTANOID LIGAND RECEPTORS | Genes involved in Prostanoid ligand receptors |
| 0.3 | 4.1 | REACTOME MITOCHONDRIAL FATTY ACID BETA OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.3 | 2.8 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.3 | 3.3 | REACTOME P2Y RECEPTORS | Genes involved in P2Y receptors |
| 0.2 | 39.6 | REACTOME METABOLISM OF AMINO ACIDS AND DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |
| 0.2 | 2.5 | REACTOME FATTY ACYL COA BIOSYNTHESIS | Genes involved in Fatty Acyl-CoA Biosynthesis |
| 0.2 | 6.4 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |
| 0.2 | 5.6 | REACTOME STRIATED MUSCLE CONTRACTION | Genes involved in Striated Muscle Contraction |
| 0.2 | 3.1 | REACTOME CGMP EFFECTS | Genes involved in cGMP effects |
| 0.2 | 4.6 | REACTOME SIGNALING BY FGFR1 FUSION MUTANTS | Genes involved in Signaling by FGFR1 fusion mutants |
| 0.1 | 1.5 | REACTOME SIGNALING BY NOTCH4 | Genes involved in Signaling by NOTCH4 |
| 0.1 | 0.7 | REACTOME BETA DEFENSINS | Genes involved in Beta defensins |
| 0.1 | 5.9 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.1 | 1.7 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.1 | 3.8 | REACTOME INTEGRATION OF ENERGY METABOLISM | Genes involved in Integration of energy metabolism |
| 0.1 | 9.6 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 0.1 | 2.5 | REACTOME EXTENSION OF TELOMERES | Genes involved in Extension of Telomeres |
| 0.1 | 12.6 | REACTOME MRNA PROCESSING | Genes involved in mRNA Processing |
| 0.1 | 3.9 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.0 | 2.5 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.0 | 1.9 | REACTOME LATE PHASE OF HIV LIFE CYCLE | Genes involved in Late Phase of HIV Life Cycle |
| 0.0 | 0.7 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.0 | 3.0 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.0 | 0.4 | REACTOME TRAF3 DEPENDENT IRF ACTIVATION PATHWAY | Genes involved in TRAF3-dependent IRF activation pathway |
| 0.0 | 0.3 | REACTOME GLYCOGEN BREAKDOWN GLYCOGENOLYSIS | Genes involved in Glycogen breakdown (glycogenolysis) |
| 0.0 | 0.3 | REACTOME PYRIMIDINE METABOLISM | Genes involved in Pyrimidine metabolism |
| 0.0 | 0.2 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |