Project
PRJDB7713: Age associated transcriptome analysis in 5 tissues of zebrafish
Navigation
Downloads
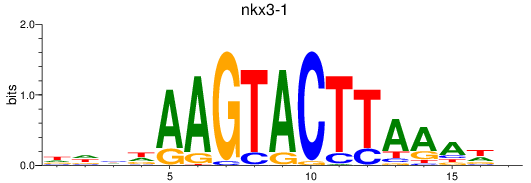
Results for nkx3-1
Z-value: 1.26
Transcription factors associated with nkx3-1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
nkx3-1
|
ENSDARG00000078280 | NK3 homeobox 1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| nkx3-1 | dr11_v1_chr8_-_50259448_50259448 | 0.37 | 2.0e-04 | Click! |
Activity profile of nkx3-1 motif
Sorted Z-values of nkx3-1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr22_-_11623063 | 17.28 |
ENSDART00000145653
|
gcga
|
glucagon a |
| chr11_-_30636163 | 16.19 |
ENSDART00000140516
|
zgc:153665
|
zgc:153665 |
| chr8_-_40464935 | 13.19 |
ENSDART00000040013
|
myl7
|
myosin, light chain 7, regulatory |
| chr19_-_5699703 | 13.08 |
ENSDART00000082050
|
zgc:174904
|
zgc:174904 |
| chr9_+_8396755 | 12.84 |
ENSDART00000043067
|
zgc:171776
|
zgc:171776 |
| chr24_-_26328721 | 11.68 |
ENSDART00000125468
|
apodb
|
apolipoprotein Db |
| chr1_-_5542565 | 11.34 |
ENSDART00000160791
ENSDART00000103754 |
fn1b
|
fibronectin 1b |
| chr7_+_22680560 | 10.63 |
ENSDART00000133761
|
ponzr4
|
plac8 onzin related protein 4 |
| chr11_-_5865744 | 10.48 |
ENSDART00000104360
|
gamt
|
guanidinoacetate N-methyltransferase |
| chr17_+_24615091 | 9.35 |
ENSDART00000064739
|
rpl13a
|
ribosomal protein L13a |
| chr20_+_25552057 | 8.63 |
ENSDART00000102913
|
cyp2v1
|
cytochrome P450, family 2, subfamily V, polypeptide 1 |
| chr8_+_6576940 | 8.46 |
ENSDART00000138135
|
vsig8b
|
V-set and immunoglobulin domain containing 8b |
| chr23_+_31431956 | 8.45 |
ENSDART00000138350
ENSDART00000053542 |
rps12
|
ribosomal protein S12 |
| chr8_-_39739627 | 8.44 |
ENSDART00000135422
ENSDART00000067844 |
si:ch211-170d8.5
|
si:ch211-170d8.5 |
| chr7_+_22657566 | 8.20 |
ENSDART00000141048
|
ponzr5
|
plac8 onzin related protein 5 |
| chr2_+_4208323 | 7.72 |
ENSDART00000167906
|
gata6
|
GATA binding protein 6 |
| chr24_-_30263301 | 7.57 |
ENSDART00000162328
|
snx7
|
sorting nexin 7 |
| chr14_+_52440161 | 7.45 |
ENSDART00000168437
|
b4galt1l
|
DP-Gal:betaGlcNAc beta 1,4- galactosyltransferase, polypeptide 1, like |
| chr2_+_1486822 | 7.40 |
ENSDART00000132500
|
c8a
|
complement component 8, alpha polypeptide |
| chr9_+_44994214 | 7.40 |
ENSDART00000141434
|
retsatl
|
retinol saturase (all-trans-retinol 13,14-reductase) like |
| chr4_-_9891874 | 7.21 |
ENSDART00000067193
|
adm2a
|
adrenomedullin 2a |
| chr2_+_1487118 | 7.20 |
ENSDART00000147283
|
c8a
|
complement component 8, alpha polypeptide |
| chr23_-_10254288 | 7.13 |
ENSDART00000081215
|
krt8
|
keratin 8 |
| chr6_-_11091449 | 7.00 |
ENSDART00000127209
|
pttg1ipa
|
PTTG1 interacting protein a |
| chr6_+_7466223 | 6.98 |
ENSDART00000148908
|
erbb3a
|
erb-b2 receptor tyrosine kinase 3a |
| chr21_-_35419486 | 6.96 |
ENSDART00000138529
|
si:dkeyp-23e4.3
|
si:dkeyp-23e4.3 |
| chr13_-_15700060 | 6.93 |
ENSDART00000170689
ENSDART00000010986 ENSDART00000101741 ENSDART00000139124 |
ckba
|
creatine kinase, brain a |
| chr17_-_1705013 | 6.87 |
ENSDART00000182864
|
CABZ01086293.1
|
|
| chr17_+_30894431 | 6.81 |
ENSDART00000127996
|
degs2
|
delta(4)-desaturase, sphingolipid 2 |
| chr2_+_11031360 | 6.71 |
ENSDART00000180020
ENSDART00000145093 |
acot11a
|
acyl-CoA thioesterase 11a |
| chr3_+_22377312 | 6.50 |
ENSDART00000155597
|
arhgap27l
|
Rho GTPase activating protein 27, like |
| chr3_+_29942338 | 6.38 |
ENSDART00000158220
|
ifi35
|
interferon-induced protein 35 |
| chr6_+_11438972 | 6.34 |
ENSDART00000029314
|
col5a2b
|
collagen, type V, alpha 2b |
| chr8_-_39739056 | 6.22 |
ENSDART00000147992
|
si:ch211-170d8.5
|
si:ch211-170d8.5 |
| chr25_+_31323978 | 6.03 |
ENSDART00000067030
|
lsp1
|
lymphocyte-specific protein 1 |
| chr15_+_29025090 | 6.00 |
ENSDART00000131755
|
si:ch211-137a8.2
|
si:ch211-137a8.2 |
| chr16_+_33142734 | 5.95 |
ENSDART00000138244
|
rhbdl2
|
rhomboid, veinlet-like 2 (Drosophila) |
| chr22_+_12770877 | 5.94 |
ENSDART00000044683
|
ftcd
|
formimidoyltransferase cyclodeaminase |
| chr25_-_13050959 | 5.93 |
ENSDART00000169041
|
ccl35.1
|
chemokine (C-C motif) ligand 35, duplicate 1 |
| chr20_-_22464250 | 5.90 |
ENSDART00000165904
|
pdgfra
|
platelet-derived growth factor receptor, alpha polypeptide |
| chr24_-_27473771 | 5.90 |
ENSDART00000139874
|
cxl34b.11
|
CX chemokine ligand 34b, duplicate 11 |
| chr3_+_3139240 | 5.81 |
ENSDART00000105014
|
si:dkey-30g5.1
|
si:dkey-30g5.1 |
| chr11_-_40128722 | 5.74 |
ENSDART00000165781
|
fam83e
|
family with sequence similarity 83, member E |
| chr12_+_17100021 | 5.74 |
ENSDART00000177923
|
acta2
|
actin, alpha 2, smooth muscle, aorta |
| chr22_+_25236657 | 5.72 |
ENSDART00000138012
|
zgc:172218
|
zgc:172218 |
| chr15_-_39969988 | 5.71 |
ENSDART00000146054
|
rps5
|
ribosomal protein S5 |
| chr5_-_10082244 | 5.57 |
ENSDART00000036421
|
chek2
|
checkpoint kinase 2 |
| chr16_+_31922065 | 5.57 |
ENSDART00000131661
ENSDART00000144194 ENSDART00000145510 |
rps9
|
ribosomal protein S9 |
| chr21_+_25236297 | 5.49 |
ENSDART00000112783
|
tmem45b
|
transmembrane protein 45B |
| chr3_+_16922226 | 5.47 |
ENSDART00000017646
|
atp6v0a1a
|
ATPase H+ transporting V0 subunit a1a |
| chr11_+_24729346 | 5.44 |
ENSDART00000087740
|
zgc:153953
|
zgc:153953 |
| chr18_+_6558338 | 5.41 |
ENSDART00000110892
|
b4galnt3b
|
beta-1,4-N-acetyl-galactosaminyl transferase 3b |
| chr7_-_34192834 | 5.40 |
ENSDART00000125131
|
smad6a
|
SMAD family member 6a |
| chr14_+_3495542 | 5.36 |
ENSDART00000168934
|
gstp2
|
glutathione S-transferase pi 2 |
| chr2_-_7757273 | 5.34 |
ENSDART00000136074
|
b3gnt5b
|
UDP-GlcNAc:betaGal beta-1,3-N-acetylglucosaminyltransferase 5b |
| chr8_-_38317914 | 5.18 |
ENSDART00000125920
|
pdlim2
|
PDZ and LIM domain 2 (mystique) |
| chr23_-_40295610 | 5.17 |
ENSDART00000186719
|
CABZ01067002.1
|
|
| chr4_+_76705830 | 5.15 |
ENSDART00000064312
|
ms4a17a.7
|
membrane-spanning 4-domains, subfamily A, member 17A.7 |
| chr22_+_25249193 | 5.09 |
ENSDART00000171851
|
si:ch211-226h8.11
|
si:ch211-226h8.11 |
| chr14_+_48903640 | 5.03 |
ENSDART00000165323
|
zgc:154054
|
zgc:154054 |
| chr4_-_4570475 | 4.98 |
ENSDART00000184955
|
rassf3
|
Ras association (RalGDS/AF-6) domain family member 3 |
| chr18_+_35861930 | 4.97 |
ENSDART00000185223
|
ppp1r13l
|
protein phosphatase 1, regulatory subunit 13 like |
| chr3_+_25849560 | 4.86 |
ENSDART00000007119
|
mfsd6l
|
major facilitator superfamily domain containing 6-like |
| chr5_-_23795688 | 4.83 |
ENSDART00000099084
|
gbgt1l4
|
globoside alpha-1,3-N-acetylgalactosaminyltransferase 1, like 4 |
| chr5_-_37900350 | 4.81 |
ENSDART00000084839
ENSDART00000084841 ENSDART00000133437 |
tmprss13b
|
transmembrane protease, serine 13b |
| chr9_-_51563575 | 4.74 |
ENSDART00000167034
ENSDART00000148918 |
tank
|
TRAF family member-associated NFKB activator |
| chr10_-_3427589 | 4.73 |
ENSDART00000133452
ENSDART00000037183 |
tmed2
|
transmembrane p24 trafficking protein 2 |
| chr17_-_43031763 | 4.72 |
ENSDART00000132754
ENSDART00000050399 |
npc2
|
Niemann-Pick disease, type C2 |
| chr22_-_10541712 | 4.72 |
ENSDART00000013933
|
si:dkey-42i9.4
|
si:dkey-42i9.4 |
| chr21_+_19547806 | 4.71 |
ENSDART00000159707
ENSDART00000184869 ENSDART00000181321 ENSDART00000058487 ENSDART00000058485 |
rai14
|
retinoic acid induced 14 |
| chr5_-_25037454 | 4.70 |
ENSDART00000027237
|
abo
|
ABO blood group (transferase A, alpha 1-3-N-acetylgalactosaminyltransferase; transferase B, alpha 1-3-galactosyltransferase) |
| chr19_-_42556086 | 4.69 |
ENSDART00000051731
|
si:dkey-267n13.1
|
si:dkey-267n13.1 |
| chr19_-_3821678 | 4.59 |
ENSDART00000169639
|
si:dkey-206d17.12
|
si:dkey-206d17.12 |
| chr18_-_16801033 | 4.54 |
ENSDART00000100100
|
admb
|
adrenomedullin b |
| chr17_+_14886828 | 4.51 |
ENSDART00000010507
ENSDART00000131052 |
ptger2a
|
prostaglandin E receptor 2a (subtype EP2) |
| chr11_+_20899029 | 4.47 |
ENSDART00000163029
|
zgc:162182
|
zgc:162182 |
| chr4_-_4119396 | 4.41 |
ENSDART00000067409
ENSDART00000138221 |
lmod2b
|
leiomodin 2 (cardiac) b |
| chr25_-_13403726 | 4.41 |
ENSDART00000056723
|
gins3
|
GINS complex subunit 3 |
| chr6_-_54826061 | 4.38 |
ENSDART00000149982
|
tnni1b
|
troponin I type 1b (skeletal, slow) |
| chr3_+_29941777 | 4.33 |
ENSDART00000113889
|
ifi35
|
interferon-induced protein 35 |
| chr14_+_14806851 | 4.31 |
ENSDART00000169235
|
fhdc2
|
FH2 domain containing 2 |
| chr7_+_8358547 | 4.27 |
ENSDART00000173016
|
jac7
|
jacalin 7 |
| chr4_+_76671012 | 4.24 |
ENSDART00000005585
|
ms4a17a.2
|
membrane-spanning 4-domains, subfamily A, member 17a.2 |
| chr8_-_38105053 | 4.23 |
ENSDART00000131546
|
adgra2
|
adhesion G protein-coupled receptor A2 |
| chr18_+_40471826 | 4.22 |
ENSDART00000098806
|
ugt5c3
|
UDP glucuronosyltransferase 5 family, polypeptide C3 |
| chr2_+_15100742 | 4.16 |
ENSDART00000027171
|
f3b
|
coagulation factor IIIb |
| chr5_-_67499279 | 4.16 |
ENSDART00000128050
|
si:dkey-251i10.3
|
si:dkey-251i10.3 |
| chr2_-_10877228 | 4.15 |
ENSDART00000138718
ENSDART00000034246 |
cdc7
|
cell division cycle 7 homolog (S. cerevisiae) |
| chr5_+_36666715 | 4.14 |
ENSDART00000097686
|
zgc:153990
|
zgc:153990 |
| chr7_+_17947217 | 4.13 |
ENSDART00000101601
|
cth1
|
cysteine three histidine 1 |
| chr20_-_25902141 | 4.10 |
ENSDART00000142611
ENSDART00000024821 |
elmsan1a
|
ELM2 and Myb/SANT-like domain containing 1a |
| chr9_-_12888082 | 4.07 |
ENSDART00000133135
ENSDART00000134415 |
si:ch211-167j6.3
|
si:ch211-167j6.3 |
| chr24_-_31904924 | 4.04 |
ENSDART00000156060
ENSDART00000129741 ENSDART00000154276 |
si:ch73-78o10.1
|
si:ch73-78o10.1 |
| chr18_+_2228737 | 4.02 |
ENSDART00000165301
|
rab27a
|
RAB27A, member RAS oncogene family |
| chr23_+_25832689 | 4.02 |
ENSDART00000138907
|
hnf4a
|
hepatocyte nuclear factor 4, alpha |
| chr8_+_25616946 | 3.99 |
ENSDART00000133983
|
slc38a5a
|
solute carrier family 38, member 5a |
| chr1_-_58002973 | 3.94 |
ENSDART00000140390
|
si:ch211-114l13.9
|
si:ch211-114l13.9 |
| chr7_+_12950507 | 3.92 |
ENSDART00000067629
ENSDART00000158004 |
saa
|
serum amyloid A |
| chr19_+_4051695 | 3.83 |
ENSDART00000166368
|
btr24
|
bloodthirsty-related gene family, member 24 |
| chr5_+_51594209 | 3.71 |
ENSDART00000164668
ENSDART00000058403 ENSDART00000055857 |
ckmt2b
|
creatine kinase, mitochondrial 2b (sarcomeric) |
| chr9_-_8296723 | 3.57 |
ENSDART00000139867
|
si:ch211-145c1.1
|
si:ch211-145c1.1 |
| chr1_-_354115 | 3.56 |
ENSDART00000141590
ENSDART00000098627 |
pros1
|
protein S |
| chr21_-_30658509 | 3.53 |
ENSDART00000139764
|
si:dkey-22f5.9
|
si:dkey-22f5.9 |
| chr8_+_2319091 | 3.50 |
ENSDART00000140265
|
ikbkb
|
inhibitor of kappa light polypeptide gene enhancer in B-cells, kinase beta |
| chr3_-_20957011 | 3.49 |
ENSDART00000159787
ENSDART00000180531 |
ndufa4l
|
NADH dehydrogenase (ubiquinone) 1 alpha subcomplex 4, like |
| chr22_+_13886821 | 3.47 |
ENSDART00000130585
ENSDART00000105711 |
sh3bp4a
|
SH3-domain binding protein 4a |
| chr19_-_31372896 | 3.46 |
ENSDART00000046609
|
scin
|
scinderin |
| chr6_-_54107269 | 3.42 |
ENSDART00000190017
|
hyal2a
|
hyaluronoglucosaminidase 2a |
| chr1_+_580642 | 3.41 |
ENSDART00000147633
|
mrpl39
|
mitochondrial ribosomal protein L39 |
| chr16_+_54641230 | 3.33 |
ENSDART00000157641
ENSDART00000159540 |
fbxo43
|
F-box protein 43 |
| chr25_-_13703826 | 3.32 |
ENSDART00000163398
|
pla2g15
|
phospholipase A2, group XV |
| chr13_-_37647209 | 3.32 |
ENSDART00000189102
|
si:dkey-188i13.10
|
si:dkey-188i13.10 |
| chr16_-_31686602 | 3.30 |
ENSDART00000170357
|
c1s
|
complement component 1, s subcomponent |
| chr7_+_2259413 | 3.28 |
ENSDART00000173420
|
si:dkey-187j14.7
|
si:dkey-187j14.7 |
| chr24_+_26017094 | 3.28 |
ENSDART00000137851
|
tfr1b
|
transferrin receptor 1b |
| chr4_+_76735113 | 3.25 |
ENSDART00000075602
|
ms4a17a.6
|
membrane-spanning 4-domains, subfamily A, member 17A.6 |
| chr20_+_33904258 | 3.25 |
ENSDART00000170930
|
rxrgb
|
retinoid X receptor, gamma b |
| chr5_+_1933131 | 3.25 |
ENSDART00000061693
|
si:ch73-55i23.1
|
si:ch73-55i23.1 |
| chr15_-_28805493 | 3.23 |
ENSDART00000179617
|
cd3eap
|
CD3e molecule, epsilon associated protein |
| chr13_-_48058779 | 3.21 |
ENSDART00000186623
ENSDART00000045475 |
itga9
|
integrin, alpha 9 |
| chr1_+_32054159 | 3.17 |
ENSDART00000181442
|
sts
|
steroid sulfatase (microsomal), isozyme S |
| chr19_+_43359075 | 3.09 |
ENSDART00000148287
ENSDART00000149856 ENSDART00000188236 ENSDART00000136695 ENSDART00000193859 |
yrk
|
Yes-related kinase |
| chr16_-_51299061 | 3.06 |
ENSDART00000148677
|
serpinb1l4
|
serpin peptidase inhibitor, clade B (ovalbumin), member 1, like 4 |
| chr22_-_23000815 | 2.94 |
ENSDART00000137111
|
ptprc
|
protein tyrosine phosphatase, receptor type, C |
| chr7_+_58179513 | 2.90 |
ENSDART00000123117
|
ggh
|
gamma-glutamyl hydrolase (conjugase, folylpolygammaglutamyl hydrolase) |
| chr9_-_50001606 | 2.80 |
ENSDART00000161648
ENSDART00000168514 |
scn1a
|
sodium channel, voltage-gated, type I, alpha |
| chr3_-_22366562 | 2.78 |
ENSDART00000129447
|
ifnphi1
|
interferon phi 1 |
| chr3_-_13921173 | 2.78 |
ENSDART00000159177
ENSDART00000165174 |
si:dkey-61n16.5
|
si:dkey-61n16.5 |
| chr1_-_2457546 | 2.76 |
ENSDART00000103795
|
ggact.1
|
gamma-glutamylamine cyclotransferase, tandem duplicate 1 |
| chr20_-_33566640 | 2.74 |
ENSDART00000159729
|
si:dkey-65b13.9
|
si:dkey-65b13.9 |
| chr10_+_38512270 | 2.71 |
ENSDART00000109752
|
serpinh1a
|
serpin peptidase inhibitor, clade H (heat shock protein 47), member 1a |
| chr21_+_25198637 | 2.66 |
ENSDART00000164972
|
si:dkey-183i3.6
|
si:dkey-183i3.6 |
| chr5_+_26795773 | 2.58 |
ENSDART00000145631
|
tcn2
|
transcobalamin II |
| chr19_-_10330778 | 2.58 |
ENSDART00000081465
ENSDART00000136653 ENSDART00000171232 |
ccdc106b
|
coiled-coil domain containing 106b |
| chr15_+_37331585 | 2.57 |
ENSDART00000170715
|
zgc:171592
|
zgc:171592 |
| chr7_-_20464468 | 2.54 |
ENSDART00000134700
|
cnpy4
|
canopy4 |
| chr3_-_36419641 | 2.54 |
ENSDART00000173545
|
cog1
|
component of oligomeric golgi complex 1 |
| chr22_-_11626014 | 2.50 |
ENSDART00000063133
ENSDART00000160085 |
gcga
|
glucagon a |
| chr25_+_3318192 | 2.43 |
ENSDART00000146154
|
slc25a3b
|
solute carrier family 25 (mitochondrial carrier; phosphate carrier), member 3b |
| chr13_-_40237121 | 2.41 |
ENSDART00000145635
|
loxl4
|
lysyl oxidase-like 4 |
| chr12_-_13155653 | 2.41 |
ENSDART00000152467
|
cxl34c
|
CX chemokine ligand 34c |
| chr5_-_16475682 | 2.40 |
ENSDART00000090695
|
piwil2
|
piwi-like RNA-mediated gene silencing 2 |
| chr15_+_25452092 | 2.38 |
ENSDART00000009545
|
pak4
|
p21 protein (Cdc42/Rac)-activated kinase 4 |
| chr16_-_41535690 | 2.35 |
ENSDART00000102662
|
rpp25l
|
ribonuclease P/MRP 25 subunit-like |
| chr12_-_31461219 | 2.34 |
ENSDART00000148954
|
acsl5
|
acyl-CoA synthetase long chain family member 5 |
| chr5_-_16475374 | 2.29 |
ENSDART00000134274
ENSDART00000136004 |
piwil2
|
piwi-like RNA-mediated gene silencing 2 |
| chr17_+_6538733 | 2.26 |
ENSDART00000193712
|
slc5a6b
|
solute carrier family 5 (sodium/multivitamin and iodide cotransporter), member 6 |
| chr19_+_7424347 | 2.26 |
ENSDART00000004622
|
sf3b4
|
splicing factor 3b, subunit 4 |
| chr21_-_11286483 | 2.25 |
ENSDART00000074845
|
rtkn2b
|
rhotekin 2b |
| chr21_-_21781158 | 2.25 |
ENSDART00000113734
|
chrdl2
|
chordin-like 2 |
| chr5_+_27429872 | 2.18 |
ENSDART00000087894
|
zgc:165555
|
zgc:165555 |
| chr16_-_4203022 | 2.17 |
ENSDART00000018686
|
rrp15
|
ribosomal RNA processing 15 homolog |
| chr10_+_585719 | 2.16 |
ENSDART00000180167
|
smad4a
|
SMAD family member 4a |
| chr3_-_506241 | 2.14 |
ENSDART00000150141
|
si:dkey-12f6.5
|
si:dkey-12f6.5 |
| chr12_-_28548789 | 2.10 |
ENSDART00000153062
|
si:ch73-180n10.1
|
si:ch73-180n10.1 |
| chr5_-_57204352 | 2.09 |
ENSDART00000171252
ENSDART00000180727 |
man2a1
|
mannosidase, alpha, class 2A, member 1 |
| chr19_+_33139164 | 2.09 |
ENSDART00000043039
|
fam84b
|
family with sequence similarity 84, member B |
| chr13_+_27328098 | 2.07 |
ENSDART00000037585
|
mb21d1
|
Mab-21 domain containing 1 |
| chr20_-_33507458 | 2.06 |
ENSDART00000140287
|
paplnb
|
papilin b, proteoglycan-like sulfated glycoprotein |
| chr16_-_9425451 | 2.05 |
ENSDART00000149163
|
ccr8.1
|
chemokine (C-C motif) receptor 8.1 |
| chr23_+_4362463 | 2.04 |
ENSDART00000039829
|
zgc:112175
|
zgc:112175 |
| chr25_+_10953351 | 2.03 |
ENSDART00000154050
|
mhc1lda
|
major histocompatibility complex class I LDA |
| chr7_-_25697285 | 2.02 |
ENSDART00000082620
|
dysf
|
dysferlin, limb girdle muscular dystrophy 2B (autosomal recessive) |
| chr7_-_26518086 | 2.00 |
ENSDART00000058913
|
eif4a1a
|
eukaryotic translation initiation factor 4A1A |
| chr14_-_498979 | 1.96 |
ENSDART00000171976
|
spry1
|
sprouty homolog 1, antagonist of FGF signaling (Drosophila) |
| chr9_-_52386733 | 1.95 |
ENSDART00000171721
|
dap1b
|
death associated protein 1b |
| chr5_-_24997293 | 1.95 |
ENSDART00000177478
|
emid1
|
EMI domain containing 1 |
| chr23_+_19813677 | 1.93 |
ENSDART00000139192
ENSDART00000142308 |
emd
|
emerin (Emery-Dreifuss muscular dystrophy) |
| chr18_-_1258777 | 1.91 |
ENSDART00000077106
ENSDART00000129065 |
ugt5f1
|
UDP glucuronosyltransferase 5 family, polypeptide F1 |
| chr18_-_6766354 | 1.90 |
ENSDART00000132611
|
adm2b
|
adrenomedullin 2b |
| chr7_+_48667081 | 1.89 |
ENSDART00000083473
|
trpm5
|
transient receptor potential cation channel, subfamily M, member 5 |
| chr12_-_30548244 | 1.86 |
ENSDART00000193616
|
zgc:158404
|
zgc:158404 |
| chr22_-_5252005 | 1.85 |
ENSDART00000132942
ENSDART00000081801 |
ncln
|
nicalin |
| chr2_-_27575803 | 1.81 |
ENSDART00000014568
|
urod
|
uroporphyrinogen decarboxylase |
| chr4_-_67800414 | 1.80 |
ENSDART00000160213
|
si:ch211-66c13.1
|
si:ch211-66c13.1 |
| chr25_-_173165 | 1.79 |
ENSDART00000193594
|
CABZ01114053.1
|
|
| chr25_+_34749187 | 1.74 |
ENSDART00000141473
|
wwp2
|
WW domain containing E3 ubiquitin protein ligase 2 |
| chr9_+_35016201 | 1.74 |
ENSDART00000182404
|
gabpa
|
GA binding protein transcription factor, alpha subunit |
| chr1_-_55248496 | 1.72 |
ENSDART00000098615
|
nanos3
|
nanos homolog 3 |
| chr14_+_14806692 | 1.71 |
ENSDART00000193050
|
fhdc2
|
FH2 domain containing 2 |
| chr23_+_36340520 | 1.70 |
ENSDART00000011201
|
copz1
|
coatomer protein complex, subunit zeta 1 |
| chr4_-_7673165 | 1.69 |
ENSDART00000028171
|
lta4h
|
leukotriene A4 hydrolase |
| chr21_-_40835069 | 1.68 |
ENSDART00000004686
|
limk1b
|
LIM domain kinase 1b |
| chr4_+_7514687 | 1.65 |
ENSDART00000067350
ENSDART00000169998 ENSDART00000162082 |
tnnt2e
|
troponin T2e, cardiac |
| chr19_-_18152942 | 1.63 |
ENSDART00000190182
|
nfe2l3
|
nuclear factor, erythroid 2-like 3 |
| chr4_+_76745015 | 1.61 |
ENSDART00000155883
|
ms4a17a.10
|
membrane-spanning 4-domains, subfamily A, member 17A.10 |
| chr1_+_24365582 | 1.60 |
ENSDART00000169389
|
pimr178
|
Pim proto-oncogene, serine/threonine kinase, related 178 |
| chr14_+_8715527 | 1.58 |
ENSDART00000170754
|
kcnk4a
|
potassium channel, subfamily K, member 4a |
| chr12_+_23850661 | 1.57 |
ENSDART00000152921
|
svila
|
supervillin a |
| chr13_+_7242916 | 1.57 |
ENSDART00000184238
|
aifm2
|
apoptosis-inducing factor, mitochondrion-associated, 2 |
| chr24_+_26328787 | 1.57 |
ENSDART00000003884
|
mynn
|
myoneurin |
| chr1_+_44056129 | 1.56 |
ENSDART00000161114
|
CU915778.1
|
|
| chr20_-_18535502 | 1.55 |
ENSDART00000049437
|
cdc42bpb
|
CDC42 binding protein kinase beta (DMPK-like) |
| chr20_-_13625588 | 1.53 |
ENSDART00000078893
|
sytl3
|
synaptotagmin-like 3 |
| chr13_-_31370184 | 1.51 |
ENSDART00000034829
|
rrp12
|
ribosomal RNA processing 12 homolog |
| chr18_-_49318823 | 1.48 |
ENSDART00000098419
|
sb:cb81
|
sb:cb81 |
| chr13_-_36680531 | 1.47 |
ENSDART00000085298
|
l2hgdh
|
L-2-hydroxyglutarate dehydrogenase |
| chr16_+_11603096 | 1.46 |
ENSDART00000146286
|
si:dkey-11o1.3
|
si:dkey-11o1.3 |
| chr4_-_18434924 | 1.46 |
ENSDART00000190271
|
socs2
|
suppressor of cytokine signaling 2 |
| chr12_-_5418340 | 1.45 |
ENSDART00000028043
|
noc3l
|
NOC3-like DNA replication regulator |
Network of associatons between targets according to the STRING database.
First level regulatory network of nkx3-1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.5 | 10.5 | GO:0006601 | creatine metabolic process(GO:0006600) creatine biosynthetic process(GO:0006601) |
| 2.2 | 13.2 | GO:0055014 | ventricular cardiac myofibril assembly(GO:0055005) atrial cardiac muscle cell development(GO:0055014) |
| 2.2 | 15.3 | GO:0006953 | acute-phase response(GO:0006953) |
| 2.0 | 5.9 | GO:0043606 | formate metabolic process(GO:0015942) histidine catabolic process to glutamate and formamide(GO:0019556) histidine catabolic process to glutamate and formate(GO:0019557) formamide metabolic process(GO:0043606) |
| 1.9 | 5.7 | GO:0090131 | mesenchyme migration(GO:0090131) |
| 1.8 | 9.2 | GO:0043011 | myeloid dendritic cell activation(GO:0001773) myeloid dendritic cell differentiation(GO:0043011) |
| 1.6 | 6.4 | GO:0030718 | germ-line stem cell population maintenance(GO:0030718) |
| 1.4 | 4.2 | GO:0022009 | central nervous system vasculogenesis(GO:0022009) |
| 1.4 | 5.6 | GO:0045903 | positive regulation of translational fidelity(GO:0045903) |
| 1.2 | 3.6 | GO:0042730 | fibrinolysis(GO:0042730) |
| 1.2 | 5.9 | GO:0003261 | cardiac muscle progenitor cell migration to the midline involved in heart field formation(GO:0003261) |
| 1.1 | 4.6 | GO:0061158 | 3'-UTR-mediated mRNA destabilization(GO:0061158) |
| 1.0 | 4.0 | GO:1902024 | L-histidine transmembrane transport(GO:0089709) L-histidine transport(GO:1902024) |
| 0.9 | 4.5 | GO:0071380 | response to prostaglandin E(GO:0034695) cellular response to prostaglandin E stimulus(GO:0071380) |
| 0.9 | 3.5 | GO:0034260 | negative regulation of GTPase activity(GO:0034260) |
| 0.9 | 2.6 | GO:0006824 | cobalt ion transport(GO:0006824) cobalamin transport(GO:0015889) |
| 0.8 | 23.8 | GO:0003323 | type B pancreatic cell development(GO:0003323) |
| 0.8 | 3.3 | GO:0015682 | ferric iron transport(GO:0015682) transferrin transport(GO:0033572) trivalent inorganic cation transport(GO:0072512) |
| 0.8 | 10.6 | GO:0006599 | phosphagen metabolic process(GO:0006599) phosphocreatine metabolic process(GO:0006603) phosphagen biosynthetic process(GO:0042396) phosphocreatine biosynthetic process(GO:0046314) |
| 0.8 | 4.7 | GO:0032367 | intracellular cholesterol transport(GO:0032367) |
| 0.8 | 2.3 | GO:2000193 | regulation of plasma membrane long-chain fatty acid transport(GO:0010746) positive regulation of plasma membrane long-chain fatty acid transport(GO:0010747) plasma membrane long-chain fatty acid transport(GO:0015911) positive regulation of lipid transport(GO:0032370) fatty acid transmembrane transport(GO:1902001) positive regulation of anion transmembrane transport(GO:1903961) regulation of fatty acid transport(GO:2000191) positive regulation of fatty acid transport(GO:2000193) |
| 0.8 | 5.4 | GO:0060394 | negative regulation of pathway-restricted SMAD protein phosphorylation(GO:0060394) |
| 0.7 | 2.9 | GO:0046900 | tetrahydrofolylpolyglutamate metabolic process(GO:0046900) |
| 0.7 | 2.0 | GO:0003156 | regulation of organ formation(GO:0003156) |
| 0.6 | 4.4 | GO:1902315 | cell cycle DNA replication initiation(GO:1902292) nuclear cell cycle DNA replication initiation(GO:1902315) mitotic DNA replication initiation(GO:1902975) |
| 0.6 | 1.2 | GO:0002320 | lymphoid progenitor cell differentiation(GO:0002320) |
| 0.6 | 2.3 | GO:0015887 | biotin transport(GO:0015878) pantothenate transmembrane transport(GO:0015887) coenzyme transport(GO:0051182) |
| 0.6 | 9.5 | GO:0030259 | lipid glycosylation(GO:0030259) |
| 0.5 | 2.0 | GO:0001778 | plasma membrane repair(GO:0001778) monocyte activation(GO:0042117) |
| 0.5 | 4.4 | GO:0051694 | pointed-end actin filament capping(GO:0051694) |
| 0.5 | 6.8 | GO:0034311 | diol metabolic process(GO:0034311) |
| 0.4 | 5.5 | GO:0001845 | phagolysosome assembly(GO:0001845) |
| 0.4 | 12.1 | GO:0019835 | cytolysis(GO:0019835) |
| 0.4 | 3.3 | GO:0045835 | negative regulation of meiotic nuclear division(GO:0045835) |
| 0.4 | 1.2 | GO:0065001 | specification of axis polarity(GO:0065001) |
| 0.4 | 1.9 | GO:1900145 | nodal signaling pathway involved in determination of left/right asymmetry(GO:0038107) regulation of nodal signaling pathway involved in determination of left/right asymmetry(GO:1900145) |
| 0.4 | 11.7 | GO:0007568 | aging(GO:0007568) |
| 0.4 | 1.1 | GO:1902746 | regulation of lens fiber cell differentiation(GO:1902746) |
| 0.4 | 1.1 | GO:0034214 | protein hexamerization(GO:0034214) |
| 0.3 | 2.8 | GO:0046685 | response to arsenic-containing substance(GO:0046685) |
| 0.3 | 1.0 | GO:0071498 | cellular response to fluid shear stress(GO:0071498) |
| 0.3 | 1.7 | GO:0006691 | leukotriene metabolic process(GO:0006691) leukotriene biosynthetic process(GO:0019370) |
| 0.3 | 2.3 | GO:0032185 | septin ring organization(GO:0031106) septin cytoskeleton organization(GO:0032185) |
| 0.3 | 1.9 | GO:0021888 | hypothalamus gonadotrophin-releasing hormone neuron differentiation(GO:0021886) hypothalamus gonadotrophin-releasing hormone neuron development(GO:0021888) |
| 0.3 | 4.1 | GO:0000727 | double-strand break repair via break-induced replication(GO:0000727) |
| 0.3 | 1.4 | GO:0033147 | negative regulation of intracellular estrogen receptor signaling pathway(GO:0033147) |
| 0.3 | 3.2 | GO:0050909 | sensory perception of taste(GO:0050909) |
| 0.3 | 3.4 | GO:0030214 | hyaluronan catabolic process(GO:0030214) |
| 0.3 | 2.3 | GO:0001682 | tRNA 5'-leader removal(GO:0001682) |
| 0.2 | 5.7 | GO:0000028 | ribosomal small subunit assembly(GO:0000028) |
| 0.2 | 1.4 | GO:0017182 | peptidyl-diphthamide metabolic process(GO:0017182) peptidyl-diphthamide biosynthetic process from peptidyl-histidine(GO:0017183) |
| 0.2 | 1.1 | GO:0019482 | uracil catabolic process(GO:0006212) beta-alanine metabolic process(GO:0019482) beta-alanine biosynthetic process(GO:0019483) uracil metabolic process(GO:0019860) 'de novo' UMP biosynthetic process(GO:0044205) |
| 0.2 | 1.5 | GO:1904893 | negative regulation of JAK-STAT cascade(GO:0046426) negative regulation of STAT cascade(GO:1904893) |
| 0.2 | 5.6 | GO:0044773 | mitotic DNA damage checkpoint(GO:0044773) |
| 0.2 | 6.4 | GO:0006891 | intra-Golgi vesicle-mediated transport(GO:0006891) |
| 0.2 | 3.5 | GO:0033209 | tumor necrosis factor-mediated signaling pathway(GO:0033209) |
| 0.2 | 8.6 | GO:0042737 | drug metabolic process(GO:0017144) drug catabolic process(GO:0042737) exogenous drug catabolic process(GO:0042738) |
| 0.2 | 5.0 | GO:0051014 | actin filament severing(GO:0051014) |
| 0.2 | 5.4 | GO:0009636 | response to toxic substance(GO:0009636) |
| 0.1 | 3.4 | GO:0046475 | glycerophospholipid catabolic process(GO:0046475) |
| 0.1 | 2.4 | GO:0018057 | peptidyl-lysine oxidation(GO:0018057) |
| 0.1 | 5.9 | GO:0048247 | lymphocyte chemotaxis(GO:0048247) |
| 0.1 | 1.4 | GO:0043545 | Mo-molybdopterin cofactor biosynthetic process(GO:0006777) Mo-molybdopterin cofactor metabolic process(GO:0019720) molybdopterin cofactor biosynthetic process(GO:0032324) molybdopterin cofactor metabolic process(GO:0043545) prosthetic group metabolic process(GO:0051189) |
| 0.1 | 2.7 | GO:1901888 | regulation of cell junction assembly(GO:1901888) |
| 0.1 | 3.1 | GO:0048246 | macrophage chemotaxis(GO:0048246) |
| 0.1 | 2.3 | GO:0048026 | positive regulation of mRNA splicing, via spliceosome(GO:0048026) |
| 0.1 | 0.7 | GO:0044341 | sodium-dependent phosphate transport(GO:0044341) |
| 0.1 | 2.7 | GO:0030199 | collagen fibril organization(GO:0030199) |
| 0.1 | 8.3 | GO:0006606 | protein import into nucleus(GO:0006606) protein targeting to nucleus(GO:0044744) single-organism nuclear import(GO:1902593) |
| 0.1 | 1.8 | GO:0006782 | protoporphyrinogen IX biosynthetic process(GO:0006782) protoporphyrinogen IX metabolic process(GO:0046501) |
| 0.1 | 5.7 | GO:0007173 | epidermal growth factor receptor signaling pathway(GO:0007173) |
| 0.1 | 3.3 | GO:0032526 | response to retinoic acid(GO:0032526) |
| 0.1 | 0.9 | GO:0018206 | peptidyl-methionine modification(GO:0018206) |
| 0.1 | 10.2 | GO:0006487 | protein N-linked glycosylation(GO:0006487) |
| 0.1 | 4.0 | GO:0051904 | melanosome transport(GO:0032402) pigment granule transport(GO:0051904) |
| 0.1 | 2.4 | GO:0035435 | phosphate ion transmembrane transport(GO:0035435) |
| 0.1 | 2.8 | GO:0042219 | cellular modified amino acid catabolic process(GO:0042219) |
| 0.1 | 0.5 | GO:0071480 | response to gamma radiation(GO:0010332) cellular response to gamma radiation(GO:0071480) |
| 0.1 | 10.0 | GO:0017148 | negative regulation of translation(GO:0017148) |
| 0.1 | 1.1 | GO:0035677 | posterior lateral line neuromast hair cell development(GO:0035677) |
| 0.1 | 0.6 | GO:0046477 | glycosylceramide catabolic process(GO:0046477) |
| 0.1 | 1.3 | GO:0006270 | DNA replication initiation(GO:0006270) |
| 0.1 | 1.9 | GO:0010507 | negative regulation of autophagy(GO:0010507) |
| 0.1 | 3.5 | GO:0017038 | protein import(GO:0017038) |
| 0.1 | 2.2 | GO:0000460 | maturation of 5.8S rRNA(GO:0000460) |
| 0.1 | 4.7 | GO:0007249 | I-kappaB kinase/NF-kappaB signaling(GO:0007249) |
| 0.1 | 0.4 | GO:0090133 | establishment or maintenance of cytoskeleton polarity(GO:0030952) mesendoderm migration(GO:0090133) cell migration involved in mesendoderm migration(GO:0090134) |
| 0.1 | 1.4 | GO:0002098 | tRNA wobble uridine modification(GO:0002098) |
| 0.1 | 0.2 | GO:0070125 | mitochondrial translational elongation(GO:0070125) |
| 0.1 | 1.0 | GO:0060021 | palate development(GO:0060021) |
| 0.1 | 1.6 | GO:0030322 | stabilization of membrane potential(GO:0030322) |
| 0.1 | 4.2 | GO:0007596 | blood coagulation(GO:0007596) |
| 0.1 | 1.4 | GO:0006418 | tRNA aminoacylation for protein translation(GO:0006418) |
| 0.1 | 1.1 | GO:0043162 | ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway(GO:0043162) |
| 0.1 | 1.6 | GO:0008637 | apoptotic mitochondrial changes(GO:0008637) |
| 0.1 | 4.7 | GO:0045930 | negative regulation of mitotic cell cycle(GO:0045930) |
| 0.1 | 1.4 | GO:0007520 | myoblast fusion(GO:0007520) |
| 0.1 | 6.0 | GO:0035023 | regulation of Rho protein signal transduction(GO:0035023) |
| 0.1 | 3.3 | GO:0031638 | zymogen activation(GO:0031638) |
| 0.1 | 2.0 | GO:0002183 | cytoplasmic translational initiation(GO:0002183) |
| 0.1 | 1.4 | GO:0046847 | filopodium assembly(GO:0046847) |
| 0.1 | 0.6 | GO:0042771 | intrinsic apoptotic signaling pathway in response to DNA damage by p53 class mediator(GO:0042771) |
| 0.1 | 7.2 | GO:0006364 | rRNA processing(GO:0006364) |
| 0.1 | 2.0 | GO:0042742 | defense response to bacterium(GO:0042742) |
| 0.1 | 3.2 | GO:0001946 | lymphangiogenesis(GO:0001946) |
| 0.0 | 1.0 | GO:0051156 | glucose 6-phosphate metabolic process(GO:0051156) |
| 0.0 | 4.7 | GO:0006888 | ER to Golgi vesicle-mediated transport(GO:0006888) |
| 0.0 | 1.0 | GO:0035141 | medial fin morphogenesis(GO:0035141) |
| 0.0 | 1.3 | GO:0070536 | protein K63-linked deubiquitination(GO:0070536) |
| 0.0 | 0.6 | GO:0021520 | spinal cord motor neuron cell fate specification(GO:0021520) |
| 0.0 | 0.8 | GO:0060232 | delamination(GO:0060232) |
| 0.0 | 2.9 | GO:0006469 | negative regulation of protein kinase activity(GO:0006469) |
| 0.0 | 3.1 | GO:0010951 | negative regulation of endopeptidase activity(GO:0010951) |
| 0.0 | 2.2 | GO:0060395 | SMAD protein signal transduction(GO:0060395) |
| 0.0 | 1.6 | GO:0018107 | peptidyl-threonine phosphorylation(GO:0018107) |
| 0.0 | 8.7 | GO:0033674 | positive regulation of kinase activity(GO:0033674) |
| 0.0 | 8.4 | GO:0043062 | extracellular matrix organization(GO:0030198) extracellular structure organization(GO:0043062) |
| 0.0 | 1.7 | GO:0045214 | sarcomere organization(GO:0045214) |
| 0.0 | 2.5 | GO:0035383 | acyl-CoA metabolic process(GO:0006637) thioester metabolic process(GO:0035383) |
| 0.0 | 8.5 | GO:0006412 | translation(GO:0006412) |
| 0.0 | 0.1 | GO:0042762 | regulation of sulfur metabolic process(GO:0042762) |
| 0.0 | 9.3 | GO:0043066 | negative regulation of apoptotic process(GO:0043066) |
| 0.0 | 9.8 | GO:0006955 | immune response(GO:0006955) |
| 0.0 | 1.3 | GO:1902476 | chloride transmembrane transport(GO:1902476) |
| 0.0 | 5.8 | GO:0006915 | apoptotic process(GO:0006915) |
| 0.0 | 3.0 | GO:0006486 | protein glycosylation(GO:0006486) macromolecule glycosylation(GO:0043413) |
| 0.0 | 0.3 | GO:0051289 | protein homotetramerization(GO:0051289) |
| 0.0 | 0.2 | GO:0042761 | very long-chain fatty acid biosynthetic process(GO:0042761) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.6 | 6.3 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 1.1 | 4.2 | GO:1990909 | Wnt signalosome(GO:1990909) |
| 0.9 | 4.7 | GO:1990923 | PET complex(GO:1990923) |
| 0.9 | 14.6 | GO:0005579 | membrane attack complex(GO:0005579) |
| 0.9 | 4.4 | GO:0000811 | GINS complex(GO:0000811) |
| 0.6 | 7.1 | GO:0045095 | keratin filament(GO:0045095) |
| 0.6 | 3.5 | GO:0008385 | IkappaB kinase complex(GO:0008385) |
| 0.5 | 5.5 | GO:0000220 | vacuolar proton-transporting V-type ATPase, V0 domain(GO:0000220) |
| 0.4 | 19.7 | GO:0022627 | cytosolic small ribosomal subunit(GO:0022627) |
| 0.4 | 7.6 | GO:0071144 | SMAD2-SMAD3 protein complex(GO:0071144) |
| 0.4 | 5.7 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.3 | 2.3 | GO:0000172 | ribonuclease MRP complex(GO:0000172) |
| 0.3 | 1.0 | GO:0097124 | cyclin A2-CDK2 complex(GO:0097124) |
| 0.3 | 7.0 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.3 | 2.3 | GO:0005826 | actomyosin contractile ring(GO:0005826) |
| 0.3 | 6.0 | GO:0034992 | microtubule organizing center attachment site(GO:0034992) LINC complex(GO:0034993) |
| 0.3 | 3.5 | GO:0017119 | Golgi transport complex(GO:0017119) |
| 0.3 | 3.9 | GO:0034364 | high-density lipoprotein particle(GO:0034364) |
| 0.2 | 3.5 | GO:0002102 | podosome(GO:0002102) |
| 0.2 | 1.4 | GO:0019008 | molybdopterin synthase complex(GO:0019008) |
| 0.2 | 7.2 | GO:0048770 | melanosome(GO:0042470) pigment granule(GO:0048770) |
| 0.2 | 4.7 | GO:0033116 | endoplasmic reticulum-Golgi intermediate compartment membrane(GO:0033116) |
| 0.2 | 10.5 | GO:0032580 | Golgi cisterna membrane(GO:0032580) |
| 0.2 | 10.4 | GO:0036379 | striated muscle thin filament(GO:0005865) myofilament(GO:0036379) |
| 0.2 | 5.6 | GO:0032040 | small-subunit processome(GO:0032040) |
| 0.2 | 1.7 | GO:0030126 | COPI vesicle coat(GO:0030126) |
| 0.1 | 9.3 | GO:0022626 | cytosolic ribosome(GO:0022626) |
| 0.1 | 5.0 | GO:0005793 | endoplasmic reticulum-Golgi intermediate compartment(GO:0005793) |
| 0.1 | 2.9 | GO:0005689 | U12-type spliceosomal complex(GO:0005689) |
| 0.1 | 1.4 | GO:0030015 | CCR4-NOT core complex(GO:0030015) |
| 0.1 | 5.2 | GO:0031941 | filamentous actin(GO:0031941) |
| 0.1 | 0.6 | GO:0005786 | signal recognition particle, endoplasmic reticulum targeting(GO:0005786) signal recognition particle(GO:0048500) |
| 0.1 | 2.2 | GO:0030687 | preribosome, large subunit precursor(GO:0030687) |
| 0.1 | 1.0 | GO:0016328 | lateral plasma membrane(GO:0016328) |
| 0.1 | 3.2 | GO:0008305 | integrin complex(GO:0008305) |
| 0.1 | 1.1 | GO:0000813 | ESCRT I complex(GO:0000813) |
| 0.1 | 1.7 | GO:0043186 | P granule(GO:0043186) |
| 0.1 | 110.6 | GO:0005576 | extracellular region(GO:0005576) |
| 0.1 | 2.2 | GO:0045271 | mitochondrial respiratory chain complex I(GO:0005747) NADH dehydrogenase complex(GO:0030964) respiratory chain complex I(GO:0045271) |
| 0.0 | 2.5 | GO:0031305 | integral component of mitochondrial inner membrane(GO:0031305) |
| 0.0 | 4.6 | GO:0036464 | cytoplasmic ribonucleoprotein granule(GO:0036464) |
| 0.0 | 0.7 | GO:0005903 | brush border(GO:0005903) |
| 0.0 | 0.5 | GO:0016363 | nuclear matrix(GO:0016363) |
| 0.0 | 3.4 | GO:0005840 | ribosome(GO:0005840) |
| 0.0 | 0.9 | GO:0032154 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.0 | 3.3 | GO:0090575 | RNA polymerase II transcription factor complex(GO:0090575) |
| 0.0 | 2.2 | GO:0000786 | nucleosome(GO:0000786) |
| 0.0 | 1.3 | GO:0005643 | nuclear pore(GO:0005643) |
| 0.0 | 1.4 | GO:0030018 | Z disc(GO:0030018) |
| 0.0 | 1.2 | GO:0031228 | integral component of Golgi membrane(GO:0030173) intrinsic component of Golgi membrane(GO:0031228) |
| 0.0 | 0.4 | GO:0031932 | TORC2 complex(GO:0031932) |
| 0.0 | 1.2 | GO:0031514 | motile cilium(GO:0031514) |
| 0.0 | 0.6 | GO:0016605 | PML body(GO:0016605) |
| 0.0 | 1.3 | GO:0005882 | intermediate filament(GO:0005882) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 4.0 | 19.8 | GO:0031769 | glucagon receptor binding(GO:0031769) |
| 2.5 | 7.5 | GO:0003831 | beta-N-acetylglucosaminylglycopeptide beta-1,4-galactosyltransferase activity(GO:0003831) |
| 1.8 | 5.3 | GO:0008457 | beta-galactosyl-N-acetylglucosaminylgalactosylglucosyl-ceramide beta-1,3-acetylglucosaminyltransferase activity(GO:0008457) lactosylceramide 1,3-N-acetyl-beta-D-glucosaminyltransferase activity(GO:0047256) |
| 1.5 | 5.9 | GO:0016841 | ammonia-lyase activity(GO:0016841) |
| 1.5 | 5.9 | GO:0048407 | platelet-derived growth factor-activated receptor activity(GO:0005017) platelet-derived growth factor binding(GO:0048407) |
| 1.4 | 11.3 | GO:0043394 | glycoprotein binding(GO:0001948) proteoglycan binding(GO:0043394) |
| 1.4 | 7.0 | GO:0038131 | neuregulin receptor activity(GO:0038131) neuregulin binding(GO:0038132) |
| 1.1 | 3.3 | GO:0015462 | protein-transmembrane transporting ATPase activity(GO:0015462) |
| 1.1 | 5.4 | GO:0033842 | N-acetyl-beta-glucosaminyl-glycoprotein 4-beta-N-acetylgalactosaminyltransferase activity(GO:0033842) |
| 1.0 | 6.7 | GO:0036042 | long-chain fatty acyl-CoA binding(GO:0036042) |
| 0.9 | 2.8 | GO:0005132 | type I interferon receptor binding(GO:0005132) |
| 0.9 | 3.5 | GO:0008384 | IkappaB kinase activity(GO:0008384) |
| 0.9 | 3.4 | GO:0033906 | hyaluronoglucuronidase activity(GO:0033906) |
| 0.8 | 3.3 | GO:0004998 | transferrin receptor activity(GO:0004998) |
| 0.8 | 10.6 | GO:0004111 | creatine kinase activity(GO:0004111) phosphotransferase activity, nitrogenous group as acceptor(GO:0016775) |
| 0.6 | 4.7 | GO:0034584 | piRNA binding(GO:0034584) |
| 0.6 | 2.3 | GO:0008523 | sodium-dependent multivitamin transmembrane transporter activity(GO:0008523) |
| 0.6 | 1.7 | GO:0004301 | epoxide hydrolase activity(GO:0004301) |
| 0.5 | 1.6 | GO:0004174 | electron-transferring-flavoprotein dehydrogenase activity(GO:0004174) oxidoreductase activity, acting on the CH-NH group of donors, quinone or similar compound as acceptor(GO:0016649) |
| 0.5 | 2.0 | GO:0070513 | death domain binding(GO:0070513) |
| 0.5 | 1.4 | GO:0016300 | tRNA (uracil) methyltransferase activity(GO:0016300) |
| 0.5 | 4.5 | GO:0004957 | prostaglandin E receptor activity(GO:0004957) |
| 0.4 | 7.6 | GO:0070411 | I-SMAD binding(GO:0070411) |
| 0.4 | 5.4 | GO:0051117 | ATPase binding(GO:0051117) |
| 0.4 | 3.3 | GO:0044323 | retinoic acid-responsive element binding(GO:0044323) |
| 0.4 | 7.6 | GO:0097153 | cysteine-type endopeptidase activity involved in apoptotic process(GO:0097153) |
| 0.3 | 5.9 | GO:0031729 | CCR1 chemokine receptor binding(GO:0031726) CCR4 chemokine receptor binding(GO:0031729) |
| 0.3 | 1.4 | GO:0030366 | molybdopterin synthase activity(GO:0030366) |
| 0.3 | 2.7 | GO:0072518 | Rho-dependent protein serine/threonine kinase activity(GO:0072518) |
| 0.3 | 4.0 | GO:0015186 | L-glutamine transmembrane transporter activity(GO:0015186) |
| 0.3 | 2.6 | GO:0031419 | cobalamin binding(GO:0031419) |
| 0.3 | 6.2 | GO:0051393 | muscle alpha-actinin binding(GO:0051371) alpha-actinin binding(GO:0051393) |
| 0.3 | 3.2 | GO:0047498 | calcium-dependent phospholipase A2 activity(GO:0047498) |
| 0.3 | 6.1 | GO:0005523 | tropomyosin binding(GO:0005523) |
| 0.3 | 1.1 | GO:0015369 | calcium:proton antiporter activity(GO:0015369) metal ion:proton antiporter activity(GO:0051139) |
| 0.3 | 1.1 | GO:0016635 | oxidoreductase activity, acting on the CH-CH group of donors, quinone or related compound as acceptor(GO:0016635) |
| 0.3 | 12.7 | GO:0019843 | rRNA binding(GO:0019843) |
| 0.2 | 2.3 | GO:0047676 | arachidonate-CoA ligase activity(GO:0047676) |
| 0.2 | 3.5 | GO:0005092 | GDP-dissociation inhibitor activity(GO:0005092) |
| 0.2 | 1.5 | GO:0005159 | insulin-like growth factor receptor binding(GO:0005159) |
| 0.2 | 8.3 | GO:0008009 | chemokine activity(GO:0008009) |
| 0.2 | 1.1 | GO:0034714 | type III transforming growth factor beta receptor binding(GO:0034714) |
| 0.2 | 2.4 | GO:0005315 | inorganic phosphate transmembrane transporter activity(GO:0005315) |
| 0.2 | 2.9 | GO:0008242 | omega peptidase activity(GO:0008242) |
| 0.1 | 3.2 | GO:0004065 | arylsulfatase activity(GO:0004065) |
| 0.1 | 2.4 | GO:0004720 | protein-lysine 6-oxidase activity(GO:0004720) |
| 0.1 | 8.6 | GO:0016712 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced flavin or flavoprotein as one donor, and incorporation of one atom of oxygen(GO:0016712) |
| 0.1 | 3.3 | GO:0004806 | triglyceride lipase activity(GO:0004806) |
| 0.1 | 0.7 | GO:0005436 | sodium:phosphate symporter activity(GO:0005436) |
| 0.1 | 17.8 | GO:0003735 | structural constituent of ribosome(GO:0003735) |
| 0.1 | 6.1 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.1 | 5.4 | GO:0005546 | phosphatidylinositol-4,5-bisphosphate binding(GO:0005546) |
| 0.1 | 4.8 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.1 | 25.2 | GO:0030246 | carbohydrate binding(GO:0030246) |
| 0.1 | 0.6 | GO:0008312 | 7S RNA binding(GO:0008312) |
| 0.1 | 4.7 | GO:0015485 | cholesterol binding(GO:0015485) |
| 0.1 | 1.3 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.1 | 4.6 | GO:0004521 | endoribonuclease activity(GO:0004521) |
| 0.1 | 0.3 | GO:0072571 | ADP-D-ribose binding(GO:0072570) mono-ADP-D-ribose binding(GO:0072571) |
| 0.1 | 2.7 | GO:0005518 | collagen binding(GO:0005518) |
| 0.1 | 2.4 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.1 | 1.9 | GO:0005227 | calcium activated cation channel activity(GO:0005227) |
| 0.1 | 1.6 | GO:0022840 | leak channel activity(GO:0022840) potassium ion leak channel activity(GO:0022841) narrow pore channel activity(GO:0022842) |
| 0.1 | 1.4 | GO:0016876 | aminoacyl-tRNA ligase activity(GO:0004812) ligase activity, forming carbon-oxygen bonds(GO:0016875) ligase activity, forming aminoacyl-tRNA and related compounds(GO:0016876) |
| 0.1 | 0.6 | GO:0008422 | beta-glucosidase activity(GO:0008422) |
| 0.0 | 6.3 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.0 | 5.1 | GO:0004867 | serine-type endopeptidase inhibitor activity(GO:0004867) |
| 0.0 | 6.6 | GO:0005179 | hormone activity(GO:0005179) |
| 0.0 | 0.5 | GO:0008568 | microtubule-severing ATPase activity(GO:0008568) |
| 0.0 | 5.2 | GO:0004896 | cytokine receptor activity(GO:0004896) |
| 0.0 | 1.4 | GO:0030374 | ligand-dependent nuclear receptor transcription coactivator activity(GO:0030374) |
| 0.0 | 6.8 | GO:0016705 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen(GO:0016705) |
| 0.0 | 1.8 | GO:0016831 | carboxy-lyase activity(GO:0016831) |
| 0.0 | 0.1 | GO:0034246 | core DNA-dependent RNA polymerase binding promoter specificity activity(GO:0000996) mitochondrial RNA polymerase binding promoter specificity activity(GO:0034246) |
| 0.0 | 2.9 | GO:0008375 | acetylglucosaminyltransferase activity(GO:0008375) |
| 0.0 | 3.1 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.0 | 0.3 | GO:0016408 | C-acyltransferase activity(GO:0016408) |
| 0.0 | 0.2 | GO:0017150 | tRNA dihydrouridine synthase activity(GO:0017150) |
| 0.0 | 0.3 | GO:0004861 | cyclin-dependent protein serine/threonine kinase inhibitor activity(GO:0004861) |
| 0.0 | 4.0 | GO:0004879 | RNA polymerase II transcription factor activity, ligand-activated sequence-specific DNA binding(GO:0004879) |
| 0.0 | 0.1 | GO:0015450 | P-P-bond-hydrolysis-driven protein transmembrane transporter activity(GO:0015450) |
| 0.0 | 5.7 | GO:0008757 | S-adenosylmethionine-dependent methyltransferase activity(GO:0008757) |
| 0.0 | 1.3 | GO:0030170 | pyridoxal phosphate binding(GO:0030170) |
| 0.0 | 5.7 | GO:0019901 | protein kinase binding(GO:0019901) |
| 0.0 | 7.1 | GO:0016757 | transferase activity, transferring glycosyl groups(GO:0016757) |
| 0.0 | 1.7 | GO:0003727 | single-stranded RNA binding(GO:0003727) |
| 0.0 | 2.0 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.0 | 0.2 | GO:0015248 | sterol transporter activity(GO:0015248) |
| 0.0 | 1.0 | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity(GO:0016538) |
| 0.0 | 0.6 | GO:0017069 | snRNA binding(GO:0017069) |
| 0.0 | 4.1 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.0 | 6.8 | GO:0005096 | GTPase activator activity(GO:0005096) |
| 0.0 | 0.5 | GO:0042162 | telomeric DNA binding(GO:0042162) |
| 0.0 | 2.9 | GO:0004725 | protein tyrosine phosphatase activity(GO:0004725) |
| 0.0 | 0.3 | GO:1904315 | neurotransmitter receptor activity involved in regulation of postsynaptic membrane potential(GO:0099529) transmitter-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1904315) |
| 0.0 | 1.3 | GO:0005254 | chloride channel activity(GO:0005254) |
| 0.0 | 0.3 | GO:0050136 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.0 | 0.1 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.0 | 0.1 | GO:0043035 | chromatin insulator sequence binding(GO:0043035) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 6.0 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.3 | 3.5 | PID NFKAPPAB ATYPICAL PATHWAY | Atypical NF-kappaB pathway |
| 0.2 | 3.2 | PID INTEGRIN A9B1 PATHWAY | Alpha9 beta1 integrin signaling events |
| 0.2 | 5.9 | PID PDGFRA PATHWAY | PDGFR-alpha signaling pathway |
| 0.2 | 7.1 | PID P38 ALPHA BETA DOWNSTREAM PATHWAY | Signaling mediated by p38-alpha and p38-beta |
| 0.2 | 10.5 | PID P53 REGULATION PATHWAY | p53 pathway |
| 0.2 | 7.7 | PID HES HEY PATHWAY | Notch-mediated HES/HEY network |
| 0.1 | 5.3 | ST T CELL SIGNAL TRANSDUCTION | T Cell Signal Transduction |
| 0.1 | 4.0 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
| 0.1 | 1.0 | SA REG CASCADE OF CYCLIN EXPR | Expression of cyclins regulates progression through the cell cycle by activating cyclin-dependent kinases. |
| 0.1 | 1.5 | PID IL2 1PATHWAY | IL2-mediated signaling events |
| 0.1 | 4.9 | WNT SIGNALING | Genes related to Wnt-mediated signal transduction |
| 0.1 | 0.6 | PID IL3 PATHWAY | IL3-mediated signaling events |
| 0.0 | 0.3 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.0 | 0.5 | PID DNA PK PATHWAY | DNA-PK pathway in nonhomologous end joining |
| 0.0 | 0.8 | PID RB 1PATHWAY | Regulation of retinoblastoma protein |
| 0.0 | 0.4 | PID PI3KCI AKT PATHWAY | Class I PI3K signaling events mediated by Akt |
| 0.0 | 1.6 | PID P53 DOWNSTREAM PATHWAY | Direct p53 effectors |
| 0.0 | 0.8 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.0 | 2.2 | NABA SECRETED FACTORS | Genes encoding secreted soluble factors |
| 0.0 | 1.9 | NABA ECM GLYCOPROTEINS | Genes encoding structural ECM glycoproteins |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 3.6 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 0.8 | 17.9 | REACTOME COMPLEMENT CASCADE | Genes involved in Complement cascade |
| 0.5 | 5.6 | REACTOME G2 M DNA DAMAGE CHECKPOINT | Genes involved in G2/M DNA damage checkpoint |
| 0.4 | 4.7 | REACTOME TRAF6 MEDIATED IRF7 ACTIVATION | Genes involved in TRAF6 mediated IRF7 activation |
| 0.4 | 2.9 | REACTOME PHOSPHORYLATION OF CD3 AND TCR ZETA CHAINS | Genes involved in Phosphorylation of CD3 and TCR zeta chains |
| 0.4 | 3.5 | REACTOME IKK COMPLEX RECRUITMENT MEDIATED BY RIP1 | Genes involved in IKK complex recruitment mediated by RIP1 |
| 0.4 | 19.7 | REACTOME FORMATION OF THE TERNARY COMPLEX AND SUBSEQUENTLY THE 43S COMPLEX | Genes involved in Formation of the ternary complex, and subsequently, the 43S complex |
| 0.4 | 10.7 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 0.3 | 2.3 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.3 | 7.9 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.3 | 3.2 | REACTOME THE ACTIVATION OF ARYLSULFATASES | Genes involved in The activation of arylsulfatases |
| 0.3 | 5.7 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.2 | 4.7 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.2 | 4.0 | REACTOME REGULATION OF GENE EXPRESSION IN BETA CELLS | Genes involved in Regulation of gene expression in beta cells |
| 0.2 | 4.0 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.2 | 1.7 | REACTOME COPI MEDIATED TRANSPORT | Genes involved in COPI Mediated Transport |
| 0.2 | 2.0 | REACTOME EGFR DOWNREGULATION | Genes involved in EGFR downregulation |
| 0.2 | 2.4 | REACTOME ACTIVATION OF RAC | Genes involved in Activation of Rac |
| 0.1 | 9.3 | REACTOME PEPTIDE CHAIN ELONGATION | Genes involved in Peptide chain elongation |
| 0.1 | 1.8 | REACTOME METABOLISM OF PORPHYRINS | Genes involved in Metabolism of porphyrins |
| 0.1 | 0.8 | REACTOME FATTY ACYL COA BIOSYNTHESIS | Genes involved in Fatty Acyl-CoA Biosynthesis |
| 0.1 | 4.1 | REACTOME ACTIVATION OF THE PRE REPLICATIVE COMPLEX | Genes involved in Activation of the pre-replicative complex |
| 0.1 | 0.6 | REACTOME DIGESTION OF DIETARY CARBOHYDRATE | Genes involved in Digestion of dietary carbohydrate |
| 0.1 | 1.5 | REACTOME GROWTH HORMONE RECEPTOR SIGNALING | Genes involved in Growth hormone receptor signaling |
| 0.1 | 1.0 | REACTOME CYCLIN A B1 ASSOCIATED EVENTS DURING G2 M TRANSITION | Genes involved in Cyclin A/B1 associated events during G2/M transition |
| 0.1 | 7.7 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.1 | 0.5 | REACTOME INTEGRATION OF PROVIRUS | Genes involved in Integration of provirus |
| 0.1 | 6.3 | REACTOME DOWNSTREAM SIGNAL TRANSDUCTION | Genes involved in Downstream signal transduction |
| 0.1 | 1.1 | REACTOME PROTEOLYTIC CLEAVAGE OF SNARE COMPLEX PROTEINS | Genes involved in Proteolytic cleavage of SNARE complex proteins |
| 0.1 | 0.3 | REACTOME GAB1 SIGNALOSOME | Genes involved in GAB1 signalosome |
| 0.1 | 3.2 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.0 | 2.2 | REACTOME RESPIRATORY ELECTRON TRANSPORT | Genes involved in Respiratory electron transport |
| 0.0 | 1.5 | REACTOME PYRUVATE METABOLISM AND CITRIC ACID TCA CYCLE | Genes involved in Pyruvate metabolism and Citric Acid (TCA) cycle |
| 0.0 | 1.3 | REACTOME REGULATION OF GLUCOKINASE BY GLUCOKINASE REGULATORY PROTEIN | Genes involved in Regulation of Glucokinase by Glucokinase Regulatory Protein |
| 0.0 | 5.4 | REACTOME METABOLISM OF AMINO ACIDS AND DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |
| 0.0 | 0.6 | REACTOME SRP DEPENDENT COTRANSLATIONAL PROTEIN TARGETING TO MEMBRANE | Genes involved in SRP-dependent cotranslational protein targeting to membrane |
| 0.0 | 0.4 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |