Project
PRJDB7713: Age associated transcriptome analysis in 5 tissues of zebrafish
Navigation
Downloads
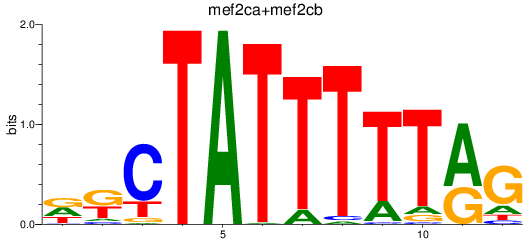
Results for mef2ca+mef2cb
Z-value: 2.80
Transcription factors associated with mef2ca+mef2cb
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
mef2cb
|
ENSDARG00000009418 | myocyte enhancer factor 2cb |
|
mef2ca
|
ENSDARG00000029764 | myocyte enhancer factor 2ca |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| mef2ca | dr11_v1_chr10_-_43568239_43568241 | 0.80 | 9.8e-22 | Click! |
| mef2cb | dr11_v1_chr5_-_48307804_48307804 | 0.55 | 7.9e-09 | Click! |
Activity profile of mef2ca+mef2cb motif
Sorted Z-values of mef2ca+mef2cb motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr11_+_11201096 | 74.60 |
ENSDART00000171916
ENSDART00000171521 ENSDART00000087105 ENSDART00000159603 |
myom2a
|
myomesin 2a |
| chr3_-_61205711 | 57.34 |
ENSDART00000055062
|
pvalb1
|
parvalbumin 1 |
| chr5_+_51597677 | 53.31 |
ENSDART00000048210
ENSDART00000184797 |
ckmt2b
|
creatine kinase, mitochondrial 2b (sarcomeric) |
| chr12_-_26064480 | 51.06 |
ENSDART00000158215
ENSDART00000171206 ENSDART00000171212 ENSDART00000182956 ENSDART00000186779 |
ldb3b
|
LIM domain binding 3b |
| chr25_+_29160102 | 49.82 |
ENSDART00000162854
|
pkmb
|
pyruvate kinase M1/2b |
| chr12_-_26064105 | 49.54 |
ENSDART00000168825
|
ldb3b
|
LIM domain binding 3b |
| chr11_+_11200550 | 48.48 |
ENSDART00000181339
ENSDART00000187116 |
myom2a
|
myomesin 2a |
| chr11_-_18253111 | 47.62 |
ENSDART00000125984
|
mustn1b
|
musculoskeletal, embryonic nuclear protein 1b |
| chr15_-_23645810 | 44.37 |
ENSDART00000168845
|
ckmb
|
creatine kinase, muscle b |
| chr9_-_42873700 | 42.94 |
ENSDART00000125953
|
ttn.1
|
titin, tandem duplicate 1 |
| chr12_+_18524953 | 41.74 |
ENSDART00000090332
|
neurl2
|
neuralized E3 ubiquitin protein ligase 2 |
| chr12_-_17707449 | 41.22 |
ENSDART00000142427
ENSDART00000034914 |
pvalb3
|
parvalbumin 3 |
| chr17_-_5583345 | 38.86 |
ENSDART00000035944
|
clic5a
|
chloride intracellular channel 5a |
| chr13_+_22249636 | 38.57 |
ENSDART00000108472
ENSDART00000173123 |
synpo2la
|
synaptopodin 2-like a |
| chr2_+_30916188 | 37.92 |
ENSDART00000137012
|
myom1a
|
myomesin 1a (skelemin) |
| chr5_-_72125551 | 36.32 |
ENSDART00000149412
|
smyd1a
|
SET and MYND domain containing 1a |
| chr12_+_6002715 | 34.90 |
ENSDART00000114961
|
si:ch211-131k2.3
|
si:ch211-131k2.3 |
| chr6_+_29410986 | 34.38 |
ENSDART00000065293
|
usp13
|
ubiquitin specific peptidase 13 |
| chr16_+_29663809 | 34.38 |
ENSDART00000191336
|
tmod4
|
tropomodulin 4 (muscle) |
| chr16_-_17200120 | 33.97 |
ENSDART00000147739
|
gapdh
|
glyceraldehyde-3-phosphate dehydrogenase |
| chr8_-_26388090 | 33.52 |
ENSDART00000147912
|
si:dkey-20d21.12
|
si:dkey-20d21.12 |
| chr1_+_7546259 | 33.36 |
ENSDART00000015732
|
mylz3
|
myosin, light polypeptide 3, skeletal muscle |
| chr6_-_18199062 | 33.35 |
ENSDART00000167513
|
ppp1r27b
|
protein phosphatase 1, regulatory subunit 27b |
| chr25_-_31396479 | 33.27 |
ENSDART00000156828
|
prr33
|
proline rich 33 |
| chr8_-_18535822 | 31.72 |
ENSDART00000100558
|
nexn
|
nexilin (F actin binding protein) |
| chr12_-_26430507 | 31.33 |
ENSDART00000153214
|
synpo2lb
|
synaptopodin 2-like b |
| chr13_+_22480857 | 31.14 |
ENSDART00000078721
ENSDART00000044719 ENSDART00000130957 ENSDART00000078757 ENSDART00000130424 ENSDART00000078747 |
ldb3a
|
LIM domain binding 3a |
| chr6_+_3680651 | 29.05 |
ENSDART00000013588
|
klhl41b
|
kelch-like family member 41b |
| chr9_-_6927587 | 28.55 |
ENSDART00000059092
|
tmem182a
|
transmembrane protein 182a |
| chr7_-_71758613 | 28.32 |
ENSDART00000166724
|
myom1b
|
myomesin 1b |
| chr16_+_25245857 | 27.87 |
ENSDART00000155220
|
klhl38b
|
kelch-like family member 38b |
| chr4_+_6572364 | 27.81 |
ENSDART00000122574
|
ppp1r3aa
|
protein phosphatase 1, regulatory subunit 3Aa |
| chr21_-_22730832 | 27.61 |
ENSDART00000101797
|
fbxo40.1
|
F-box protein 40, tandem duplicate 1 |
| chr9_-_14108896 | 26.69 |
ENSDART00000135209
|
prkag3b
|
protein kinase, AMP-activated, gamma 3b non-catalytic subunit |
| chr24_+_34089977 | 26.66 |
ENSDART00000157466
|
asb10
|
ankyrin repeat and SOCS box containing 10 |
| chr9_-_43142636 | 26.50 |
ENSDART00000134349
ENSDART00000181835 |
ccdc141
|
coiled-coil domain containing 141 |
| chr24_+_34085940 | 26.05 |
ENSDART00000171189
|
asb10
|
ankyrin repeat and SOCS box containing 10 |
| chr9_-_33107237 | 25.89 |
ENSDART00000013918
|
casq2
|
calsequestrin 2 |
| chr14_-_12307522 | 25.49 |
ENSDART00000163900
|
myot
|
myotilin |
| chr23_-_5685023 | 25.02 |
ENSDART00000148680
ENSDART00000149365 |
tnnt2a
|
troponin T type 2a (cardiac) |
| chr13_-_2189761 | 24.95 |
ENSDART00000166255
|
mlip
|
muscular LMNA-interacting protein |
| chr2_-_23768818 | 24.83 |
ENSDART00000148685
ENSDART00000191167 |
xirp1
|
xin actin binding repeat containing 1 |
| chr25_-_26736088 | 24.79 |
ENSDART00000067114
|
fbxl22
|
F-box and leucine-rich repeat protein 22 |
| chr23_-_21515182 | 24.64 |
ENSDART00000142000
|
rnf207b
|
ring finger protein 207b |
| chr23_+_18722715 | 24.60 |
ENSDART00000137438
|
myh7bb
|
myosin, heavy chain 7B, cardiac muscle, beta b |
| chr7_-_23745984 | 23.98 |
ENSDART00000048050
|
ITGB1BP2
|
zgc:92429 |
| chr4_-_9592402 | 23.10 |
ENSDART00000114060
|
cdnf
|
cerebral dopamine neurotrophic factor |
| chr9_-_34191627 | 22.50 |
ENSDART00000142664
|
dcaf6
|
ddb1 and cul4 associated factor 6 |
| chr24_-_40700596 | 22.44 |
ENSDART00000162635
|
smyhc2
|
slow myosin heavy chain 2 |
| chr5_+_64368770 | 21.98 |
ENSDART00000162246
|
plpp7
|
phospholipid phosphatase 7 (inactive) |
| chr20_-_9980318 | 21.66 |
ENSDART00000080664
|
ACTC1
|
zgc:86709 |
| chr22_+_16308450 | 20.96 |
ENSDART00000105678
|
lrrc39
|
leucine rich repeat containing 39 |
| chr3_+_28953274 | 20.73 |
ENSDART00000133528
ENSDART00000103602 |
lgals2a
|
lectin, galactoside-binding, soluble, 2a |
| chr23_+_24931999 | 20.72 |
ENSDART00000136162
ENSDART00000140335 |
klhl21
|
kelch-like family member 21 |
| chr6_+_14301214 | 20.50 |
ENSDART00000129491
|
tmem182b
|
transmembrane protein 182b |
| chr4_-_4119396 | 20.47 |
ENSDART00000067409
ENSDART00000138221 |
lmod2b
|
leiomodin 2 (cardiac) b |
| chr22_+_16308806 | 20.20 |
ENSDART00000162685
|
lrrc39
|
leucine rich repeat containing 39 |
| chr19_-_9662958 | 20.03 |
ENSDART00000041094
|
clcn1a
|
chloride channel, voltage-sensitive 1a |
| chr24_+_25471196 | 19.74 |
ENSDART00000066625
|
smpx
|
small muscle protein, X-linked |
| chr1_-_38813679 | 19.46 |
ENSDART00000148917
|
asb5b
|
ankyrin repeat and SOCS box containing 5b |
| chr21_-_22737228 | 19.11 |
ENSDART00000151366
|
fbxo40.2
|
F-box protein 40, tandem duplicate 2 |
| chr23_+_18722915 | 18.91 |
ENSDART00000025057
|
myh7bb
|
myosin, heavy chain 7B, cardiac muscle, beta b |
| chr12_+_3078221 | 18.70 |
ENSDART00000148835
ENSDART00000149427 |
sgca
|
sarcoglycan, alpha |
| chr11_-_25213651 | 18.62 |
ENSDART00000097316
ENSDART00000152186 |
myh7ba
|
myosin, heavy chain 7B, cardiac muscle, beta a |
| chr9_-_23894392 | 18.43 |
ENSDART00000133417
|
asb18
|
ankyrin repeat and SOCS box containing 18 |
| chr8_-_17980317 | 18.42 |
ENSDART00000129148
|
tnni3k
|
TNNI3 interacting kinase |
| chr9_-_23891102 | 18.31 |
ENSDART00000186799
|
asb18
|
ankyrin repeat and SOCS box containing 18 |
| chr23_+_17417539 | 18.01 |
ENSDART00000182605
|
BX649300.2
|
|
| chr16_-_43356018 | 17.71 |
ENSDART00000181683
|
FO704821.1
|
|
| chr11_-_43104475 | 17.55 |
ENSDART00000125368
|
acyp2
|
acylphosphatase 2, muscle type |
| chr8_-_11229523 | 17.53 |
ENSDART00000002164
|
unc45b
|
unc-45 myosin chaperone B |
| chr25_+_20089986 | 16.77 |
ENSDART00000143441
ENSDART00000184073 |
tnni4b.2
|
troponin I4b, tandem duplicate 2 |
| chr24_+_9298198 | 16.56 |
ENSDART00000165780
|
otud1
|
OTU deubiquitinase 1 |
| chr22_+_396840 | 15.88 |
ENSDART00000163198
|
capzb
|
capping protein (actin filament) muscle Z-line, beta |
| chr3_+_59851537 | 15.72 |
ENSDART00000180997
|
CU693479.1
|
|
| chr4_-_2350371 | 15.63 |
ENSDART00000166274
|
phlda1
|
pleckstrin homology-like domain, family A, member 1 |
| chr7_-_71758307 | 15.60 |
ENSDART00000161067
ENSDART00000165253 |
myom1b
|
myomesin 1b |
| chr2_-_6182098 | 15.52 |
ENSDART00000156167
|
si:ch73-182a11.2
|
si:ch73-182a11.2 |
| chr14_+_29780113 | 14.86 |
ENSDART00000173195
|
zgc:153146
|
zgc:153146 |
| chr4_+_21717793 | 14.86 |
ENSDART00000040266
|
myf6
|
myogenic factor 6 |
| chr20_+_34455645 | 14.73 |
ENSDART00000135789
|
mettl11b
|
methyltransferase like 11B |
| chr1_+_6171585 | 14.58 |
ENSDART00000024358
|
prkag3a
|
protein kinase, AMP-activated, gamma 3a non-catalytic subunit |
| chr23_+_6077503 | 14.40 |
ENSDART00000081714
ENSDART00000139834 |
mybpha
|
myosin binding protein Ha |
| chr21_-_22951604 | 14.35 |
ENSDART00000083449
ENSDART00000180129 |
dub
|
duboraya |
| chr23_-_9925568 | 14.23 |
ENSDART00000081268
|
si:ch211-220i18.4
|
si:ch211-220i18.4 |
| chr8_-_32497815 | 14.15 |
ENSDART00000122359
|
si:dkey-164f24.2
|
si:dkey-164f24.2 |
| chr17_-_20118145 | 14.13 |
ENSDART00000149737
ENSDART00000165606 |
ryr2b
|
ryanodine receptor 2b (cardiac) |
| chr20_-_36575475 | 13.80 |
ENSDART00000062893
|
enah
|
enabled homolog (Drosophila) |
| chr24_+_20575259 | 13.54 |
ENSDART00000010488
|
klhl40b
|
kelch-like family member 40b |
| chr14_-_30366196 | 13.46 |
ENSDART00000007022
|
pdgfrl
|
platelet-derived growth factor receptor-like |
| chr24_+_26276805 | 13.29 |
ENSDART00000089749
|
adipoqa
|
adiponectin, C1Q and collagen domain containing, a |
| chr7_+_22543963 | 13.13 |
ENSDART00000101528
|
chrnb1
|
cholinergic receptor, nicotinic, beta 1 (muscle) |
| chr5_-_48268049 | 13.11 |
ENSDART00000187454
|
mef2cb
|
myocyte enhancer factor 2cb |
| chr2_-_4496358 | 12.87 |
ENSDART00000158216
|
adipoqb
|
adiponectin, C1Q and collagen domain containing, b |
| chr3_+_26135502 | 12.49 |
ENSDART00000146979
|
atp2a1
|
ATPase sarcoplasmic/endoplasmic reticulum Ca2+ transporting 1 |
| chr12_-_19119176 | 12.45 |
ENSDART00000149180
|
aco2
|
aconitase 2, mitochondrial |
| chr3_-_50046004 | 12.43 |
ENSDART00000109544
|
si:ch1073-100f3.2
|
si:ch1073-100f3.2 |
| chr7_+_22718251 | 12.34 |
ENSDART00000027718
ENSDART00000143341 |
fxr2
|
fragile X mental retardation, autosomal homolog 2 |
| chr18_+_20494413 | 12.10 |
ENSDART00000060295
|
rapsn
|
receptor-associated protein of the synapse, 43kD |
| chr8_-_32497581 | 11.96 |
ENSDART00000176298
ENSDART00000183340 |
si:dkey-164f24.2
|
si:dkey-164f24.2 |
| chr2_+_6181383 | 11.55 |
ENSDART00000153307
|
si:ch73-344o19.1
|
si:ch73-344o19.1 |
| chr25_+_5039050 | 11.49 |
ENSDART00000154700
|
parvb
|
parvin, beta |
| chr16_-_31475904 | 11.39 |
ENSDART00000145691
|
si:ch211-251p5.5
|
si:ch211-251p5.5 |
| chr9_-_43073960 | 11.06 |
ENSDART00000059460
|
ttn.2
|
titin, tandem duplicate 2 |
| chr14_+_30285613 | 10.92 |
ENSDART00000173090
|
mtus1a
|
microtubule associated tumor suppressor 1a |
| chr9_-_48281941 | 10.91 |
ENSDART00000099787
|
klhl41a
|
kelch-like family member 41a |
| chr7_-_48263516 | 10.89 |
ENSDART00000006619
ENSDART00000142370 ENSDART00000148273 ENSDART00000147968 |
rbpms2b
|
RNA binding protein with multiple splicing 2b |
| chr3_-_53114299 | 10.78 |
ENSDART00000109390
|
AL954361.1
|
|
| chr25_+_16945348 | 10.46 |
ENSDART00000016591
|
fgf6a
|
fibroblast growth factor 6a |
| chr6_-_20875111 | 10.11 |
ENSDART00000115118
ENSDART00000159916 |
tns1a
|
tensin 1a |
| chr23_-_1557195 | 10.08 |
ENSDART00000136436
|
epm2a
|
epilepsy, progressive myoclonus type 2A, Lafora disease (laforin) |
| chr14_-_29906209 | 9.84 |
ENSDART00000192952
|
sorbs2b
|
sorbin and SH3 domain containing 2b |
| chr14_-_21063977 | 9.45 |
ENSDART00000164373
|
si:dkey-74k8.3
|
si:dkey-74k8.3 |
| chr15_-_9031996 | 9.34 |
ENSDART00000124998
|
rtn2a
|
reticulon 2a |
| chr19_-_38611814 | 9.05 |
ENSDART00000151958
|
col16a1
|
collagen, type XVI, alpha 1 |
| chr14_-_21064199 | 9.03 |
ENSDART00000172099
|
si:dkey-74k8.3
|
si:dkey-74k8.3 |
| chr16_-_17541890 | 8.90 |
ENSDART00000131328
|
clcn1b
|
chloride channel, voltage-sensitive 1b |
| chr9_+_6578580 | 8.70 |
ENSDART00000061577
|
fhl2a
|
four and a half LIM domains 2a |
| chr19_+_44039849 | 8.31 |
ENSDART00000086040
|
lrrc14b
|
leucine rich repeat containing 14B |
| chr8_+_45294767 | 8.30 |
ENSDART00000191527
|
ubap2b
|
ubiquitin associated protein 2b |
| chr20_+_6756247 | 8.18 |
ENSDART00000167344
|
igfbp3
|
insulin-like growth factor binding protein 3 |
| chr12_+_10952794 | 8.14 |
ENSDART00000167090
|
raraa
|
retinoic acid receptor, alpha a |
| chr2_-_31791633 | 8.03 |
ENSDART00000180662
|
retreg1
|
reticulophagy regulator 1 |
| chr16_-_7228276 | 7.86 |
ENSDART00000149030
|
nt5c3a
|
5'-nucleotidase, cytosolic IIIA |
| chr18_-_15467446 | 7.83 |
ENSDART00000187847
|
endouc
|
endonuclease, polyU-specific C |
| chr13_-_11536951 | 7.77 |
ENSDART00000018155
|
adss
|
adenylosuccinate synthase |
| chr23_+_24926407 | 7.54 |
ENSDART00000137486
|
klhl21
|
kelch-like family member 21 |
| chr18_+_23218980 | 7.46 |
ENSDART00000185014
|
mef2aa
|
myocyte enhancer factor 2aa |
| chr14_+_22129096 | 7.29 |
ENSDART00000132514
|
ccng1
|
cyclin G1 |
| chr9_-_7673856 | 7.20 |
ENSDART00000102715
|
tuba8l3
|
tubulin, alpha 8 like 3 |
| chr16_+_31542645 | 7.03 |
ENSDART00000163724
|
SLA (1 of many)
|
Src like adaptor |
| chr22_-_22164338 | 6.56 |
ENSDART00000183840
|
cdc34a
|
cell division cycle 34 homolog (S. cerevisiae) a |
| chr9_+_24065855 | 6.53 |
ENSDART00000161468
ENSDART00000171577 ENSDART00000172743 ENSDART00000159324 ENSDART00000079689 ENSDART00000023196 ENSDART00000101577 |
lrrfip1a
|
leucine rich repeat (in FLII) interacting protein 1a |
| chr18_+_23193567 | 6.46 |
ENSDART00000190072
ENSDART00000171594 ENSDART00000181762 |
mef2aa
|
myocyte enhancer factor 2aa |
| chr20_+_23173710 | 6.36 |
ENSDART00000074172
|
sgcb
|
sarcoglycan, beta (dystrophin-associated glycoprotein) |
| chr6_+_12462079 | 6.32 |
ENSDART00000192029
ENSDART00000065385 |
nr4a2b
|
nuclear receptor subfamily 4, group A, member 2b |
| chr23_-_35347714 | 6.16 |
ENSDART00000161770
ENSDART00000165615 |
cpne9
|
copine family member IX |
| chr13_+_31757331 | 6.11 |
ENSDART00000044282
|
hif1aa
|
hypoxia inducible factor 1, alpha subunit (basic helix-loop-helix transcription factor) a |
| chr22_-_8509215 | 6.07 |
ENSDART00000140146
|
si:ch73-27e22.3
|
si:ch73-27e22.3 |
| chr21_+_40589770 | 6.07 |
ENSDART00000164650
ENSDART00000161584 ENSDART00000161108 |
pdk3b
|
pyruvate dehydrogenase kinase, isozyme 3b |
| chr18_+_23193820 | 6.06 |
ENSDART00000148106
|
mef2aa
|
myocyte enhancer factor 2aa |
| chr4_-_1720648 | 5.95 |
ENSDART00000103484
|
gas2l3
|
growth arrest-specific 2 like 3 |
| chr3_+_59784632 | 5.94 |
ENSDART00000084729
|
pecam1
|
platelet/endothelial cell adhesion molecule 1 |
| chr9_+_31795343 | 5.90 |
ENSDART00000139584
|
itgbl1
|
integrin, beta-like 1 |
| chr5_-_32505109 | 5.90 |
ENSDART00000188219
|
ntmt1
|
N-terminal Xaa-Pro-Lys N-methyltransferase 1 |
| chr6_+_43450221 | 5.87 |
ENSDART00000075521
|
zgc:113054
|
zgc:113054 |
| chr20_-_39271844 | 5.83 |
ENSDART00000192708
|
clu
|
clusterin |
| chr1_+_18965750 | 5.79 |
ENSDART00000132379
|
limch1a
|
LIM and calponin homology domains 1a |
| chr2_-_23349116 | 5.79 |
ENSDART00000099690
|
fam129ab
|
family with sequence similarity 129, member Ab |
| chr6_+_3373665 | 5.71 |
ENSDART00000134133
|
st3gal3a
|
ST3 beta-galactoside alpha-2,3-sialyltransferase 3a |
| chr2_+_20472150 | 5.63 |
ENSDART00000168537
|
agla
|
amylo-alpha-1, 6-glucosidase, 4-alpha-glucanotransferase a |
| chr3_+_53511936 | 5.60 |
ENSDART00000177083
|
zmp:0000001048
|
zmp:0000001048 |
| chr12_+_23866368 | 5.53 |
ENSDART00000188652
ENSDART00000192478 |
svila
|
supervillin a |
| chr5_+_51227147 | 5.31 |
ENSDART00000083340
|
ubac1
|
UBA domain containing 1 |
| chr13_-_42673978 | 5.28 |
ENSDART00000133848
ENSDART00000099738 ENSDART00000099729 ENSDART00000169083 |
lrrfip2
|
leucine rich repeat (in FLII) interacting protein 2 |
| chr25_-_16554757 | 5.20 |
ENSDART00000154480
|
si:ch211-266k8.6
|
si:ch211-266k8.6 |
| chr20_+_18163821 | 5.17 |
ENSDART00000186507
|
aqp4
|
aquaporin 4 |
| chr7_-_30143092 | 5.02 |
ENSDART00000173636
|
frmd5
|
FERM domain containing 5 |
| chr16_+_5259886 | 4.99 |
ENSDART00000186668
|
plecb
|
plectin b |
| chr19_-_12648122 | 4.99 |
ENSDART00000151184
|
fam210aa
|
family with sequence similarity 210, member Aa |
| chr11_-_11336986 | 4.95 |
ENSDART00000016677
|
zgc:77929
|
zgc:77929 |
| chr4_-_72080351 | 4.92 |
ENSDART00000174925
|
LO017820.1
|
|
| chr6_+_11438972 | 4.85 |
ENSDART00000029314
|
col5a2b
|
collagen, type V, alpha 2b |
| chr22_+_20546612 | 4.84 |
ENSDART00000141852
|
si:dkey-172o19.2
|
si:dkey-172o19.2 |
| chr8_+_4337312 | 4.82 |
ENSDART00000182228
|
myl2b
|
myosin, light chain 2b, regulatory, cardiac, slow |
| chr6_-_25201810 | 4.72 |
ENSDART00000168683
|
lrrc8c
|
leucine rich repeat containing 8 VRAC subunit C |
| chr13_-_28674422 | 4.69 |
ENSDART00000122754
ENSDART00000057574 |
nt5c2a
|
5'-nucleotidase, cytosolic IIa |
| chr22_+_16497670 | 4.61 |
ENSDART00000014330
|
ier5
|
immediate early response 5 |
| chr21_-_25685739 | 4.59 |
ENSDART00000129619
ENSDART00000101205 |
phkg1b
|
phosphorylase kinase, gamma 1b (muscle) |
| chr20_-_45423498 | 4.56 |
ENSDART00000098424
|
trib2
|
tribbles pseudokinase 2 |
| chr6_-_46768040 | 4.41 |
ENSDART00000154071
|
igfn1.2
|
immunoglobulin-like and fibronectin type III domain containing 1, tandem duplicate 2 |
| chr15_-_506010 | 4.32 |
ENSDART00000155472
|
nudt8
|
nudix (nucleoside diphosphate linked moiety X)-type motif 8 |
| chr12_+_17106117 | 4.29 |
ENSDART00000149990
|
acta2
|
actin, alpha 2, smooth muscle, aorta |
| chr16_-_29557338 | 4.28 |
ENSDART00000058888
|
hormad1
|
HORMA domain containing 1 |
| chr1_-_49250490 | 4.21 |
ENSDART00000150386
|
si:ch73-6k14.2
|
si:ch73-6k14.2 |
| chr8_-_13362757 | 4.02 |
ENSDART00000188608
|
ccdc124
|
coiled-coil domain containing 124 |
| chr16_+_20895904 | 3.93 |
ENSDART00000052662
|
hoxa13b
|
homeobox A13b |
| chr5_-_32505276 | 3.90 |
ENSDART00000034705
ENSDART00000187597 |
ntmt1
|
N-terminal Xaa-Pro-Lys N-methyltransferase 1 |
| chr16_+_6021908 | 3.80 |
ENSDART00000163786
|
BX511115.1
|
|
| chr16_-_30570161 | 3.79 |
ENSDART00000184045
ENSDART00000191040 |
lmna
|
lamin A |
| chr15_-_31406093 | 3.79 |
ENSDART00000123444
|
or111-8
|
odorant receptor, family D, subfamily 111, member 8 |
| chr12_+_34119439 | 3.75 |
ENSDART00000032821
|
cyth1b
|
cytohesin 1b |
| chr1_-_43915423 | 3.52 |
ENSDART00000181915
ENSDART00000113673 |
scpp5
|
secretory calcium-binding phosphoprotein 5 |
| chr7_+_17534485 | 3.47 |
ENSDART00000170911
|
nitr1k
|
novel immune-type receptor 1k |
| chr20_-_26066020 | 3.25 |
ENSDART00000078559
|
myct1a
|
myc target 1a |
| chr14_-_15699528 | 3.24 |
ENSDART00000161123
|
neurl1b
|
neuralized E3 ubiquitin protein ligase 1B |
| chr20_+_19066858 | 3.12 |
ENSDART00000192086
|
sox7
|
SRY (sex determining region Y)-box 7 |
| chr16_+_36748538 | 3.10 |
ENSDART00000139069
|
decr1
|
2,4-dienoyl CoA reductase 1, mitochondrial |
| chr5_+_51248784 | 3.05 |
ENSDART00000159571
ENSDART00000189513 ENSDART00000092285 |
msh3
|
mutS homolog 3 (E. coli) |
| chr1_+_13930625 | 3.04 |
ENSDART00000111026
|
noctb
|
nocturnin b |
| chr24_-_27409599 | 2.99 |
ENSDART00000041770
|
ccl34b.3
|
chemokine (C-C motif) ligand 34b, duplicate 3 |
| chr9_-_48184823 | 2.99 |
ENSDART00000180264
|
klhl23
|
kelch-like family member 23 |
| chr24_+_21540842 | 2.92 |
ENSDART00000091529
|
wasf3b
|
WAS protein family, member 3b |
| chr20_+_13175379 | 2.90 |
ENSDART00000025644
|
ppp2r5a
|
protein phosphatase 2, regulatory subunit B', alpha isoform |
| chr11_+_25634041 | 2.88 |
ENSDART00000033657
|
grm6b
|
glutamate receptor, metabotropic 6b |
| chr16_+_41570653 | 2.87 |
ENSDART00000102665
|
aste1a
|
asteroid homolog 1a |
| chr23_-_27506161 | 2.86 |
ENSDART00000145007
|
asb8
|
ankyrin repeat and SOCS box containing 8 |
| chr22_-_13649588 | 2.85 |
ENSDART00000131877
|
si:ch211-279g13.1
|
si:ch211-279g13.1 |
| chr12_-_35393211 | 2.82 |
ENSDART00000137139
|
camk2g1
|
calcium/calmodulin-dependent protein kinase (CaM kinase) II gamma 1 |
| chr4_+_33462238 | 2.80 |
ENSDART00000111083
|
si:dkey-247i3.1
|
si:dkey-247i3.1 |
Network of associatons between targets according to the STRING database.
First level regulatory network of mef2ca+mef2cb
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 9.5 | 47.6 | GO:0035988 | chondrocyte proliferation(GO:0035988) |
| 8.2 | 24.6 | GO:1903779 | regulation of cardiac conduction(GO:1903779) |
| 8.2 | 24.5 | GO:0006480 | N-terminal protein amino acid methylation(GO:0006480) |
| 7.5 | 97.7 | GO:0006599 | phosphagen metabolic process(GO:0006599) phosphocreatine metabolic process(GO:0006603) phosphagen biosynthetic process(GO:0042396) phosphocreatine biosynthetic process(GO:0046314) |
| 6.8 | 13.5 | GO:0098528 | skeletal muscle fiber differentiation(GO:0098528) |
| 6.5 | 25.9 | GO:0010882 | regulation of cardiac muscle contraction by calcium ion signaling(GO:0010882) |
| 6.3 | 25.0 | GO:0003228 | atrial cardiac muscle tissue development(GO:0003228) |
| 6.1 | 54.9 | GO:0051694 | pointed-end actin filament capping(GO:0051694) |
| 4.9 | 54.0 | GO:0048769 | sarcomerogenesis(GO:0048769) |
| 4.8 | 14.4 | GO:0061341 | non-canonical Wnt signaling pathway involved in heart development(GO:0061341) |
| 4.6 | 13.8 | GO:0070358 | actin polymerization-dependent cell motility(GO:0070358) |
| 4.5 | 36.3 | GO:0030240 | skeletal muscle thin filament assembly(GO:0030240) |
| 4.5 | 31.7 | GO:0048739 | cardiac muscle fiber development(GO:0048739) |
| 4.5 | 22.4 | GO:0031444 | slow-twitch skeletal muscle fiber contraction(GO:0031444) |
| 3.7 | 18.5 | GO:0010332 | response to gamma radiation(GO:0010332) cellular response to gamma radiation(GO:0071480) |
| 3.4 | 54.8 | GO:0035914 | skeletal muscle cell differentiation(GO:0035914) |
| 3.2 | 15.9 | GO:0010591 | regulation of lamellipodium assembly(GO:0010591) |
| 3.1 | 12.5 | GO:0014724 | regulation of twitch skeletal muscle contraction(GO:0014724) fast-twitch skeletal muscle fiber contraction(GO:0031443) regulation of fast-twitch skeletal muscle fiber contraction(GO:0031446) positive regulation of fast-twitch skeletal muscle fiber contraction(GO:0031448) negative regulation of striated muscle contraction(GO:0045988) relaxation of skeletal muscle(GO:0090076) |
| 2.8 | 24.8 | GO:0086003 | cardiac muscle cell contraction(GO:0086003) |
| 2.7 | 75.4 | GO:0032233 | positive regulation of actin filament bundle assembly(GO:0032233) |
| 2.5 | 12.4 | GO:0015729 | thiosulfate transport(GO:0015709) oxaloacetate transport(GO:0015729) malate transport(GO:0015743) succinate transport(GO:0015744) succinate transmembrane transport(GO:0071422) malate transmembrane transport(GO:0071423) |
| 2.3 | 34.9 | GO:0051131 | chaperone-mediated protein complex assembly(GO:0051131) |
| 2.2 | 10.9 | GO:0051145 | smooth muscle cell differentiation(GO:0051145) negative regulation of muscle cell differentiation(GO:0051148) |
| 2.0 | 68.4 | GO:0050821 | protein stabilization(GO:0050821) |
| 1.7 | 28.3 | GO:0034629 | cellular protein complex localization(GO:0034629) |
| 1.5 | 20.7 | GO:0090303 | positive regulation of wound healing(GO:0090303) |
| 1.4 | 4.3 | GO:0090131 | mesenchyme migration(GO:0090131) |
| 1.4 | 4.3 | GO:0060631 | regulation of meiosis I(GO:0060631) |
| 1.4 | 41.2 | GO:0055008 | cardiac muscle tissue morphogenesis(GO:0055008) |
| 1.3 | 205.6 | GO:0006936 | muscle contraction(GO:0006936) |
| 1.1 | 41.3 | GO:0042149 | cellular response to glucose starvation(GO:0042149) |
| 1.0 | 3.0 | GO:0051095 | regulation of helicase activity(GO:0051095) positive regulation of helicase activity(GO:0051096) |
| 1.0 | 25.5 | GO:0070593 | dendrite self-avoidance(GO:0070593) |
| 1.0 | 21.5 | GO:0010923 | negative regulation of phosphatase activity(GO:0010923) |
| 1.0 | 1.9 | GO:0001778 | plasma membrane repair(GO:0001778) |
| 0.9 | 20.3 | GO:0005979 | regulation of glycogen biosynthetic process(GO:0005979) regulation of glucan biosynthetic process(GO:0010962) |
| 0.9 | 21.9 | GO:0071542 | dopaminergic neuron differentiation(GO:0071542) |
| 0.8 | 5.9 | GO:0021534 | cell proliferation in hindbrain(GO:0021534) |
| 0.8 | 8.0 | GO:0061709 | reticulophagy(GO:0061709) |
| 0.8 | 49.8 | GO:0006096 | glycolytic process(GO:0006096) |
| 0.8 | 2.3 | GO:0031630 | regulation of synaptic vesicle fusion to presynaptic membrane(GO:0031630) |
| 0.7 | 8.2 | GO:0043567 | regulation of insulin-like growth factor receptor signaling pathway(GO:0043567) |
| 0.7 | 5.6 | GO:0006703 | estrogen biosynthetic process(GO:0006703) |
| 0.7 | 11.5 | GO:0030032 | lamellipodium assembly(GO:0030032) |
| 0.7 | 2.0 | GO:0032369 | negative regulation of lipid transport(GO:0032369) |
| 0.7 | 13.1 | GO:0007271 | synaptic transmission, cholinergic(GO:0007271) |
| 0.6 | 3.2 | GO:0070086 | ubiquitin-dependent endocytosis(GO:0070086) |
| 0.6 | 3.8 | GO:0090342 | regulation of cell aging(GO:0090342) |
| 0.6 | 13.1 | GO:0055013 | cardiac muscle cell development(GO:0055013) |
| 0.6 | 16.6 | GO:0070536 | protein K63-linked deubiquitination(GO:0070536) |
| 0.6 | 26.5 | GO:0019226 | transmission of nerve impulse(GO:0019226) |
| 0.6 | 5.2 | GO:0006833 | water transport(GO:0006833) |
| 0.6 | 14.1 | GO:0051209 | release of sequestered calcium ion into cytosol(GO:0051209) |
| 0.5 | 5.6 | GO:0044247 | polysaccharide catabolic process(GO:0000272) glycogen catabolic process(GO:0005980) glucan catabolic process(GO:0009251) cellular polysaccharide catabolic process(GO:0044247) |
| 0.5 | 44.4 | GO:0030239 | myofibril assembly(GO:0030239) |
| 0.5 | 2.4 | GO:0030242 | pexophagy(GO:0030242) |
| 0.5 | 5.6 | GO:0050962 | detection of light stimulus involved in visual perception(GO:0050908) detection of light stimulus involved in sensory perception(GO:0050962) |
| 0.5 | 1.8 | GO:0006116 | NADH oxidation(GO:0006116) glycerol-3-phosphate catabolic process(GO:0046168) |
| 0.5 | 1.8 | GO:0035610 | protein side chain deglutamylation(GO:0035610) |
| 0.4 | 1.8 | GO:0060231 | mesenchymal to epithelial transition(GO:0060231) |
| 0.4 | 2.6 | GO:1900028 | wound healing, spreading of epidermal cells(GO:0035313) negative regulation of ruffle assembly(GO:1900028) |
| 0.4 | 1.3 | GO:0051574 | positive regulation of histone H3-K9 methylation(GO:0051574) |
| 0.4 | 56.9 | GO:0006821 | chloride transport(GO:0006821) |
| 0.4 | 7.9 | GO:0051489 | regulation of filopodium assembly(GO:0051489) |
| 0.4 | 121.7 | GO:0061061 | muscle structure development(GO:0061061) |
| 0.4 | 13.5 | GO:0006099 | tricarboxylic acid cycle(GO:0006099) |
| 0.3 | 2.2 | GO:0032732 | positive regulation of interleukin-1 production(GO:0032732) |
| 0.3 | 1.6 | GO:0042264 | peptidyl-aspartic acid modification(GO:0018197) peptidyl-aspartic acid hydroxylation(GO:0042264) |
| 0.3 | 2.8 | GO:0044805 | late nucleophagy(GO:0044805) |
| 0.3 | 8.1 | GO:0032526 | response to retinoic acid(GO:0032526) |
| 0.3 | 6.1 | GO:0010906 | regulation of cellular carbohydrate metabolic process(GO:0010675) regulation of glucose metabolic process(GO:0010906) |
| 0.3 | 1.6 | GO:0007603 | phototransduction, visible light(GO:0007603) |
| 0.3 | 14.2 | GO:0000245 | spliceosomal complex assembly(GO:0000245) |
| 0.3 | 0.8 | GO:0002155 | regulation of thyroid hormone mediated signaling pathway(GO:0002155) |
| 0.3 | 6.1 | GO:0036294 | cellular response to decreased oxygen levels(GO:0036294) cellular response to hypoxia(GO:0071456) |
| 0.2 | 2.4 | GO:1901571 | icosanoid transport(GO:0071715) fatty acid derivative transport(GO:1901571) |
| 0.2 | 0.9 | GO:2001244 | regulation of release of cytochrome c from mitochondria(GO:0090199) positive regulation of release of cytochrome c from mitochondria(GO:0090200) positive regulation of intrinsic apoptotic signaling pathway(GO:2001244) |
| 0.2 | 31.3 | GO:0021782 | glial cell development(GO:0021782) |
| 0.2 | 4.6 | GO:0005978 | glycogen biosynthetic process(GO:0005978) glucan biosynthetic process(GO:0009250) |
| 0.2 | 0.8 | GO:0003322 | pancreatic A cell development(GO:0003322) |
| 0.2 | 5.9 | GO:0033627 | cell adhesion mediated by integrin(GO:0033627) |
| 0.2 | 2.9 | GO:0051127 | positive regulation of actin nucleation(GO:0051127) positive regulation of Arp2/3 complex-mediated actin nucleation(GO:2000601) |
| 0.2 | 1.2 | GO:0035881 | amacrine cell differentiation(GO:0035881) |
| 0.2 | 1.7 | GO:2000273 | positive regulation of epidermal growth factor-activated receptor activity(GO:0045741) positive regulation of receptor activity(GO:2000273) |
| 0.2 | 5.5 | GO:0051014 | actin filament severing(GO:0051014) |
| 0.2 | 17.2 | GO:0045732 | positive regulation of protein catabolic process(GO:0045732) |
| 0.2 | 2.9 | GO:0031952 | regulation of protein autophosphorylation(GO:0031952) |
| 0.2 | 108.6 | GO:0016567 | protein ubiquitination(GO:0016567) |
| 0.2 | 2.1 | GO:0099638 | neurotransmitter receptor transport, endosome to postsynaptic membrane(GO:0098887) endosome to plasma membrane protein transport(GO:0099638) neurotransmitter receptor transport, endosome to plasma membrane(GO:0099639) |
| 0.1 | 7.3 | GO:1904029 | regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0000079) regulation of cyclin-dependent protein kinase activity(GO:1904029) |
| 0.1 | 2.2 | GO:0007130 | synaptonemal complex assembly(GO:0007130) synaptonemal complex organization(GO:0070193) |
| 0.1 | 12.1 | GO:0007416 | synapse assembly(GO:0007416) |
| 0.1 | 1.8 | GO:0010996 | response to auditory stimulus(GO:0010996) |
| 0.1 | 15.7 | GO:0043068 | positive regulation of cell death(GO:0010942) positive regulation of apoptotic process(GO:0043065) positive regulation of programmed cell death(GO:0043068) |
| 0.1 | 5.8 | GO:1901214 | regulation of neuron death(GO:1901214) |
| 0.1 | 1.9 | GO:0098962 | regulation of postsynaptic neurotransmitter receptor activity(GO:0098962) |
| 0.1 | 1.9 | GO:0007622 | rhythmic behavior(GO:0007622) circadian behavior(GO:0048512) |
| 0.1 | 5.8 | GO:0071277 | cellular response to calcium ion(GO:0071277) |
| 0.1 | 1.3 | GO:0030214 | hyaluronan catabolic process(GO:0030214) |
| 0.1 | 2.8 | GO:0072114 | pronephros morphogenesis(GO:0072114) |
| 0.1 | 1.9 | GO:0097031 | NADH dehydrogenase complex assembly(GO:0010257) mitochondrial respiratory chain complex I assembly(GO:0032981) mitochondrial respiratory chain complex I biogenesis(GO:0097031) |
| 0.1 | 12.2 | GO:0008284 | positive regulation of cell proliferation(GO:0008284) |
| 0.1 | 3.8 | GO:0032011 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.1 | 3.6 | GO:0048247 | lymphocyte chemotaxis(GO:0048247) |
| 0.1 | 1.0 | GO:0035306 | positive regulation of dephosphorylation(GO:0035306) positive regulation of protein dephosphorylation(GO:0035307) |
| 0.1 | 1.9 | GO:0048268 | clathrin coat assembly(GO:0048268) |
| 0.1 | 5.0 | GO:0031032 | actomyosin structure organization(GO:0031032) |
| 0.1 | 1.1 | GO:0007214 | gamma-aminobutyric acid signaling pathway(GO:0007214) |
| 0.1 | 0.3 | GO:0098535 | de novo centriole assembly(GO:0098535) |
| 0.1 | 3.1 | GO:0001570 | vasculogenesis(GO:0001570) |
| 0.1 | 0.2 | GO:0009256 | 10-formyltetrahydrofolate metabolic process(GO:0009256) 10-formyltetrahydrofolate catabolic process(GO:0009258) folic acid-containing compound catabolic process(GO:0009397) pteridine-containing compound catabolic process(GO:0042560) |
| 0.1 | 1.7 | GO:0001732 | formation of cytoplasmic translation initiation complex(GO:0001732) |
| 0.1 | 2.4 | GO:0007019 | microtubule depolymerization(GO:0007019) |
| 0.1 | 0.4 | GO:0034605 | cellular response to heat(GO:0034605) |
| 0.1 | 0.9 | GO:0006073 | glycogen metabolic process(GO:0005977) cellular glucan metabolic process(GO:0006073) glucan metabolic process(GO:0044042) |
| 0.1 | 0.9 | GO:0018057 | peptidyl-lysine oxidation(GO:0018057) |
| 0.1 | 1.1 | GO:0071577 | zinc II ion transmembrane transport(GO:0071577) |
| 0.0 | 1.2 | GO:0007634 | optokinetic behavior(GO:0007634) |
| 0.0 | 0.3 | GO:0032218 | riboflavin transport(GO:0032218) |
| 0.0 | 1.0 | GO:0043486 | histone exchange(GO:0043486) |
| 0.0 | 1.1 | GO:0017121 | phospholipid scrambling(GO:0017121) |
| 0.0 | 2.3 | GO:1990573 | potassium ion import across plasma membrane(GO:1990573) |
| 0.0 | 0.9 | GO:0070831 | basement membrane assembly(GO:0070831) |
| 0.0 | 3.9 | GO:0033333 | fin development(GO:0033333) |
| 0.0 | 5.9 | GO:0043062 | extracellular matrix organization(GO:0030198) extracellular structure organization(GO:0043062) |
| 0.0 | 0.2 | GO:0071392 | cellular response to estradiol stimulus(GO:0071392) |
| 0.0 | 0.9 | GO:0050772 | positive regulation of axonogenesis(GO:0050772) |
| 0.0 | 1.5 | GO:0007218 | neuropeptide signaling pathway(GO:0007218) |
| 0.0 | 0.1 | GO:0002281 | macrophage activation involved in immune response(GO:0002281) |
| 0.0 | 0.8 | GO:0030433 | ER-associated ubiquitin-dependent protein catabolic process(GO:0030433) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 14.4 | 302.1 | GO:0031430 | M band(GO:0031430) |
| 4.4 | 43.6 | GO:0031672 | A band(GO:0031672) |
| 4.0 | 15.9 | GO:0008290 | F-actin capping protein complex(GO:0008290) |
| 3.7 | 25.9 | GO:0033018 | sarcoplasmic reticulum lumen(GO:0033018) |
| 3.4 | 41.3 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 3.1 | 12.3 | GO:1902737 | dendritic spine neck(GO:0044326) dendritic filopodium(GO:1902737) |
| 3.0 | 139.7 | GO:0031941 | filamentous actin(GO:0031941) |
| 3.0 | 20.7 | GO:0016012 | sarcoglycan complex(GO:0016012) |
| 2.6 | 54.1 | GO:0033017 | sarcoplasmic reticulum membrane(GO:0033017) |
| 1.8 | 121.8 | GO:0030018 | Z disc(GO:0030018) |
| 1.6 | 93.8 | GO:0036379 | striated muscle thin filament(GO:0005865) myofilament(GO:0036379) |
| 1.4 | 5.8 | GO:0016460 | myosin II complex(GO:0016460) |
| 1.2 | 4.9 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 1.2 | 20.3 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 1.1 | 13.1 | GO:0005892 | acetylcholine-gated channel complex(GO:0005892) |
| 0.9 | 28.3 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.8 | 23.4 | GO:0080008 | Cul4-RING E3 ubiquitin ligase complex(GO:0080008) |
| 0.6 | 24.9 | GO:0016605 | PML body(GO:0016605) |
| 0.6 | 76.1 | GO:0016459 | myosin complex(GO:0016459) |
| 0.6 | 5.8 | GO:0042583 | chromaffin granule(GO:0042583) |
| 0.6 | 38.2 | GO:0034707 | chloride channel complex(GO:0034707) |
| 0.5 | 2.4 | GO:0034272 | phosphatidylinositol 3-kinase complex, class III, type I(GO:0034271) phosphatidylinositol 3-kinase complex, class III, type II(GO:0034272) |
| 0.5 | 7.6 | GO:0016010 | dystrophin-associated glycoprotein complex(GO:0016010) glycoprotein complex(GO:0090665) |
| 0.5 | 5.5 | GO:0005964 | phosphorylase kinase complex(GO:0005964) |
| 0.4 | 5.0 | GO:0030056 | hemidesmosome(GO:0030056) |
| 0.3 | 24.8 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.3 | 3.0 | GO:0032300 | mismatch repair complex(GO:0032300) |
| 0.3 | 1.8 | GO:0009331 | glycerol-3-phosphate dehydrogenase complex(GO:0009331) |
| 0.3 | 4.3 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.2 | 4.8 | GO:0005921 | gap junction(GO:0005921) |
| 0.2 | 0.8 | GO:0097519 | DNA recombinase complex(GO:0097519) |
| 0.2 | 2.9 | GO:0031209 | SCAR complex(GO:0031209) |
| 0.2 | 20.3 | GO:0030055 | cell-substrate junction(GO:0030055) |
| 0.2 | 7.3 | GO:0000307 | cyclin-dependent protein kinase holoenzyme complex(GO:0000307) |
| 0.2 | 13.7 | GO:0005581 | collagen trimer(GO:0005581) |
| 0.1 | 11.1 | GO:0005814 | centriole(GO:0005814) |
| 0.1 | 1.1 | GO:0038039 | G-protein coupled receptor dimeric complex(GO:0038037) G-protein coupled receptor heterodimeric complex(GO:0038039) G-protein coupled receptor complex(GO:0097648) |
| 0.1 | 1.9 | GO:0030315 | T-tubule(GO:0030315) |
| 0.1 | 0.8 | GO:0071818 | BAT3 complex(GO:0071818) |
| 0.1 | 2.8 | GO:0000407 | pre-autophagosomal structure(GO:0000407) |
| 0.1 | 1.0 | GO:0000812 | Swr1 complex(GO:0000812) |
| 0.1 | 1.3 | GO:0035098 | ESC/E(Z) complex(GO:0035098) |
| 0.1 | 2.9 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.1 | 0.8 | GO:0070937 | CRD-mediated mRNA stability complex(GO:0070937) |
| 0.1 | 4.0 | GO:0030496 | midbody(GO:0030496) |
| 0.1 | 1.7 | GO:0016282 | eukaryotic 43S preinitiation complex(GO:0016282) |
| 0.1 | 1.8 | GO:0048787 | presynaptic active zone membrane(GO:0048787) |
| 0.0 | 0.2 | GO:0033503 | HULC complex(GO:0033503) |
| 0.0 | 0.7 | GO:0031680 | G-protein beta/gamma-subunit complex(GO:0031680) |
| 0.0 | 3.8 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.0 | 0.6 | GO:0097038 | perinuclear endoplasmic reticulum(GO:0097038) |
| 0.0 | 24.7 | GO:0015630 | microtubule cytoskeleton(GO:0015630) |
| 0.0 | 20.5 | GO:0005739 | mitochondrion(GO:0005739) |
| 0.0 | 1.9 | GO:0099572 | postsynaptic specialization(GO:0099572) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 8.5 | 34.0 | GO:0043891 | glyceraldehyde-3-phosphate dehydrogenase (NAD+) (phosphorylating) activity(GO:0004365) glyceraldehyde-3-phosphate dehydrogenase (NAD(P)+) (phosphorylating) activity(GO:0043891) |
| 8.3 | 49.8 | GO:0004743 | pyruvate kinase activity(GO:0004743) |
| 8.2 | 24.5 | GO:0071885 | N-terminal protein N-methyltransferase activity(GO:0071885) |
| 7.5 | 97.7 | GO:0004111 | creatine kinase activity(GO:0004111) phosphotransferase activity, nitrogenous group as acceptor(GO:0016775) |
| 6.6 | 131.7 | GO:0051393 | muscle alpha-actinin binding(GO:0051371) alpha-actinin binding(GO:0051393) |
| 6.2 | 24.9 | GO:0005521 | lamin binding(GO:0005521) |
| 4.6 | 18.4 | GO:0061629 | RNA polymerase II sequence-specific DNA binding transcription factor binding(GO:0061629) |
| 4.2 | 12.5 | GO:0003994 | aconitate hydratase activity(GO:0003994) |
| 4.0 | 12.1 | GO:0033130 | acetylcholine receptor binding(GO:0033130) |
| 3.6 | 79.9 | GO:0005523 | tropomyosin binding(GO:0005523) |
| 3.5 | 52.4 | GO:0051879 | Hsp90 protein binding(GO:0051879) |
| 2.8 | 14.1 | GO:0005219 | ryanodine-sensitive calcium-release channel activity(GO:0005219) calcium-induced calcium release activity(GO:0048763) |
| 2.5 | 12.4 | GO:0015140 | thiosulfate transmembrane transporter activity(GO:0015117) oxaloacetate transmembrane transporter activity(GO:0015131) malate transmembrane transporter activity(GO:0015140) |
| 2.1 | 20.7 | GO:0016936 | galactoside binding(GO:0016936) |
| 2.1 | 24.6 | GO:0030544 | Hsp70 protein binding(GO:0030544) |
| 2.0 | 10.1 | GO:2001070 | starch binding(GO:2001070) |
| 1.7 | 57.2 | GO:0098632 | protein binding involved in cell-cell adhesion(GO:0098632) |
| 1.5 | 4.6 | GO:0031434 | mitogen-activated protein kinase kinase binding(GO:0031434) |
| 1.5 | 41.3 | GO:0016208 | AMP binding(GO:0016208) |
| 1.5 | 20.3 | GO:2001069 | glycogen binding(GO:2001069) |
| 1.4 | 5.8 | GO:0032028 | myosin head/neck binding(GO:0032028) myosin II head/neck binding(GO:0032034) myosin II heavy chain binding(GO:0032038) |
| 1.4 | 5.6 | GO:0004134 | glycogen debranching enzyme activity(GO:0004133) 4-alpha-glucanotransferase activity(GO:0004134) amylo-alpha-1,6-glucosidase activity(GO:0004135) |
| 1.4 | 13.8 | GO:0005522 | profilin binding(GO:0005522) |
| 1.1 | 19.2 | GO:0005247 | voltage-gated chloride channel activity(GO:0005247) |
| 0.9 | 1.9 | GO:0005545 | 1-phosphatidylinositol binding(GO:0005545) |
| 0.9 | 4.6 | GO:0004689 | phosphorylase kinase activity(GO:0004689) |
| 0.8 | 309.1 | GO:0051015 | actin filament binding(GO:0051015) |
| 0.8 | 5.7 | GO:0008118 | N-acetyllactosaminide alpha-2,3-sialyltransferase activity(GO:0008118) |
| 0.8 | 3.0 | GO:0032135 | DNA insertion or deletion binding(GO:0032135) |
| 0.8 | 6.1 | GO:0004740 | pyruvate dehydrogenase (acetyl-transferring) kinase activity(GO:0004740) |
| 0.7 | 8.2 | GO:0031995 | insulin-like growth factor I binding(GO:0031994) insulin-like growth factor II binding(GO:0031995) |
| 0.7 | 12.5 | GO:0008553 | hydrogen-exporting ATPase activity, phosphorylative mechanism(GO:0008553) |
| 0.7 | 5.6 | GO:0004303 | estradiol 17-beta-dehydrogenase activity(GO:0004303) |
| 0.7 | 2.1 | GO:0004377 | GDP-Man:Man3GlcNAc2-PP-Dol alpha-1,2-mannosyltransferase activity(GO:0004377) |
| 0.7 | 33.1 | GO:0042826 | histone deacetylase binding(GO:0042826) |
| 0.6 | 6.3 | GO:0035259 | glucocorticoid receptor binding(GO:0035259) |
| 0.6 | 5.2 | GO:0015250 | water channel activity(GO:0015250) |
| 0.6 | 16.0 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.5 | 13.1 | GO:0022848 | acetylcholine-gated cation channel activity(GO:0022848) |
| 0.5 | 36.3 | GO:0018024 | histone-lysine N-methyltransferase activity(GO:0018024) |
| 0.5 | 37.7 | GO:0005254 | chloride channel activity(GO:0005254) |
| 0.5 | 1.8 | GO:0004367 | glycerol-3-phosphate dehydrogenase [NAD+] activity(GO:0004367) |
| 0.4 | 9.3 | GO:0008253 | 5'-nucleotidase activity(GO:0008253) |
| 0.4 | 2.9 | GO:0001642 | group III metabotropic glutamate receptor activity(GO:0001642) |
| 0.4 | 1.6 | GO:0004597 | peptide-aspartate beta-dioxygenase activity(GO:0004597) |
| 0.4 | 11.2 | GO:0005104 | fibroblast growth factor receptor binding(GO:0005104) |
| 0.4 | 35.2 | GO:0043130 | ubiquitin binding(GO:0043130) |
| 0.3 | 32.0 | GO:0004722 | protein serine/threonine phosphatase activity(GO:0004722) |
| 0.3 | 21.7 | GO:0019902 | phosphatase binding(GO:0019902) |
| 0.3 | 1.6 | GO:0050254 | rhodopsin kinase activity(GO:0050254) |
| 0.3 | 2.9 | GO:0071933 | Arp2/3 complex binding(GO:0071933) |
| 0.3 | 1.3 | GO:0033906 | hyaluronoglucuronidase activity(GO:0033906) |
| 0.3 | 1.2 | GO:0003839 | gamma-glutamylcyclotransferase activity(GO:0003839) |
| 0.3 | 6.2 | GO:0005544 | calcium-dependent phospholipid binding(GO:0005544) |
| 0.3 | 12.3 | GO:0045182 | translation regulator activity(GO:0045182) |
| 0.3 | 5.9 | GO:0048038 | quinone binding(GO:0048038) |
| 0.3 | 8.9 | GO:0030374 | ligand-dependent nuclear receptor transcription coactivator activity(GO:0030374) |
| 0.2 | 2.9 | GO:0019211 | phosphatase activator activity(GO:0019211) protein phosphatase activator activity(GO:0072542) |
| 0.2 | 1.1 | GO:0030060 | L-malate dehydrogenase activity(GO:0030060) |
| 0.2 | 7.2 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.2 | 0.8 | GO:1990238 | double-stranded DNA endodeoxyribonuclease activity(GO:1990238) |
| 0.2 | 52.8 | GO:0003779 | actin binding(GO:0003779) |
| 0.2 | 4.8 | GO:0022829 | wide pore channel activity(GO:0022829) |
| 0.2 | 80.0 | GO:0061630 | ubiquitin protein ligase activity(GO:0061630) |
| 0.2 | 15.6 | GO:1901981 | phosphatidylinositol phosphate binding(GO:1901981) |
| 0.2 | 2.3 | GO:0015272 | ATP-activated inward rectifier potassium channel activity(GO:0015272) |
| 0.2 | 1.9 | GO:0016868 | intramolecular transferase activity, phosphotransferases(GO:0016868) |
| 0.1 | 1.8 | GO:0035612 | AP-2 adaptor complex binding(GO:0035612) |
| 0.1 | 19.3 | GO:0005201 | extracellular matrix structural constituent(GO:0005201) |
| 0.1 | 1.6 | GO:0090482 | vitamin transmembrane transporter activity(GO:0090482) |
| 0.1 | 1.1 | GO:0016917 | G-protein coupled GABA receptor activity(GO:0004965) GABA receptor activity(GO:0016917) |
| 0.1 | 15.6 | GO:0036459 | thiol-dependent ubiquitin-specific protease activity(GO:0004843) thiol-dependent ubiquitinyl hydrolase activity(GO:0036459) ubiquitinyl hydrolase activity(GO:0101005) |
| 0.1 | 5.9 | GO:0005178 | integrin binding(GO:0005178) |
| 0.1 | 1.3 | GO:0034647 | histone demethylase activity (H3-trimethyl-K4 specific)(GO:0034647) |
| 0.1 | 7.3 | GO:0016538 | cyclin-dependent protein serine/threonine kinase regulator activity(GO:0016538) |
| 0.1 | 2.5 | GO:0070492 | oligosaccharide binding(GO:0070492) |
| 0.1 | 103.8 | GO:0005509 | calcium ion binding(GO:0005509) |
| 0.1 | 14.7 | GO:0008083 | growth factor activity(GO:0008083) |
| 0.1 | 2.1 | GO:0030159 | receptor signaling complex scaffold activity(GO:0030159) |
| 0.1 | 1.1 | GO:0017128 | phospholipid scramblase activity(GO:0017128) |
| 0.1 | 0.5 | GO:0030548 | acetylcholine receptor regulator activity(GO:0030548) neurotransmitter receptor regulator activity(GO:0099602) |
| 0.1 | 1.5 | GO:0004983 | neuropeptide Y receptor activity(GO:0004983) |
| 0.1 | 3.6 | GO:0048020 | CCR chemokine receptor binding(GO:0048020) |
| 0.1 | 1.6 | GO:0050321 | tau-protein kinase activity(GO:0050321) |
| 0.1 | 0.9 | GO:0004720 | protein-lysine 6-oxidase activity(GO:0004720) |
| 0.1 | 2.8 | GO:0004683 | calmodulin-dependent protein kinase activity(GO:0004683) |
| 0.1 | 7.4 | GO:0042803 | protein homodimerization activity(GO:0042803) |
| 0.1 | 0.2 | GO:0016155 | formyltetrahydrofolate dehydrogenase activity(GO:0016155) |
| 0.0 | 0.3 | GO:0032217 | riboflavin transporter activity(GO:0032217) |
| 0.0 | 1.1 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.0 | 15.3 | GO:0046983 | protein dimerization activity(GO:0046983) |
| 0.0 | 2.5 | GO:0004521 | endoribonuclease activity(GO:0004521) |
| 0.0 | 0.6 | GO:0031386 | protein tag(GO:0031386) |
| 0.0 | 0.7 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.0 | 0.9 | GO:0045296 | cadherin binding(GO:0045296) |
| 0.0 | 0.2 | GO:0004994 | somatostatin receptor activity(GO:0004994) |
| 0.0 | 6.9 | GO:0015631 | tubulin binding(GO:0015631) |
| 0.0 | 3.3 | GO:0004842 | ubiquitin-protein transferase activity(GO:0004842) |
| 0.0 | 0.1 | GO:0043023 | ribosomal large subunit binding(GO:0043023) |
| 0.0 | 0.9 | GO:0005516 | calmodulin binding(GO:0005516) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.6 | 35.2 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.4 | 13.8 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.3 | 1.7 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.3 | 9.0 | PID SYNDECAN 1 PATHWAY | Syndecan-1-mediated signaling events |
| 0.3 | 8.6 | PID INTEGRIN3 PATHWAY | Beta3 integrin cell surface interactions |
| 0.2 | 8.4 | PID ILK PATHWAY | Integrin-linked kinase signaling |
| 0.2 | 11.1 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.2 | 7.6 | PID P53 REGULATION PATHWAY | p53 pathway |
| 0.1 | 2.0 | PID ALK2 PATHWAY | ALK2 signaling events |
| 0.1 | 5.6 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
| 0.1 | 7.6 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.1 | 1.9 | PID BMP PATHWAY | BMP receptor signaling |
| 0.1 | 3.8 | PID CASPASE PATHWAY | Caspase cascade in apoptosis |
| 0.0 | 1.2 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.6 | 34.0 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 1.1 | 8.8 | REACTOME PLATELET SENSITIZATION BY LDL | Genes involved in Platelet sensitization by LDL |
| 1.0 | 8.2 | REACTOME REGULATION OF INSULIN LIKE GROWTH FACTOR IGF ACTIVITY BY INSULIN LIKE GROWTH FACTOR BINDING PROTEINS IGFBPS | Genes involved in Regulation of Insulin-like Growth Factor (IGF) Activity by Insulin-like Growth Factor Binding Proteins (IGFBPs) |
| 0.8 | 11.5 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.8 | 9.2 | REACTOME GENERATION OF SECOND MESSENGER MOLECULES | Genes involved in Generation of second messenger molecules |
| 0.7 | 12.5 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.7 | 12.5 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.6 | 7.8 | REACTOME PURINE RIBONUCLEOSIDE MONOPHOSPHATE BIOSYNTHESIS | Genes involved in Purine ribonucleoside monophosphate biosynthesis |
| 0.4 | 14.9 | REACTOME MYOGENESIS | Genes involved in Myogenesis |
| 0.2 | 3.1 | REACTOME MITOCHONDRIAL FATTY ACID BETA OXIDATION | Genes involved in Mitochondrial Fatty Acid Beta-Oxidation |
| 0.2 | 2.4 | REACTOME SYNTHESIS OF PIPS AT THE LATE ENDOSOME MEMBRANE | Genes involved in Synthesis of PIPs at the late endosome membrane |
| 0.2 | 4.3 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.2 | 9.0 | REACTOME COLLAGEN FORMATION | Genes involved in Collagen formation |
| 0.2 | 2.3 | REACTOME INWARDLY RECTIFYING K CHANNELS | Genes involved in Inwardly rectifying K+ channels |
| 0.1 | 3.8 | REACTOME APOPTOTIC CLEAVAGE OF CELLULAR PROTEINS | Genes involved in Apoptotic cleavage of cellular proteins |
| 0.1 | 1.9 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |
| 0.1 | 5.2 | REACTOME REGULATION OF WATER BALANCE BY RENAL AQUAPORINS | Genes involved in Regulation of Water Balance by Renal Aquaporins |
| 0.1 | 2.6 | REACTOME INTERFERON ALPHA BETA SIGNALING | Genes involved in Interferon alpha/beta signaling |
| 0.1 | 1.2 | REACTOME REGULATION OF HYPOXIA INDUCIBLE FACTOR HIF BY OXYGEN | Genes involved in Regulation of Hypoxia-inducible Factor (HIF) by Oxygen |
| 0.1 | 0.9 | REACTOME GLYCOGEN BREAKDOWN GLYCOGENOLYSIS | Genes involved in Glycogen breakdown (glycogenolysis) |
| 0.0 | 2.7 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.0 | 1.7 | REACTOME PI3K EVENTS IN ERBB4 SIGNALING | Genes involved in PI3K events in ERBB4 signaling |
| 0.0 | 0.2 | REACTOME ROLE OF DCC IN REGULATING APOPTOSIS | Genes involved in Role of DCC in regulating apoptosis |
| 0.0 | 0.1 | REACTOME TRANSPORT OF ORGANIC ANIONS | Genes involved in Transport of organic anions |
| 0.0 | 0.8 | REACTOME DIABETES PATHWAYS | Genes involved in Diabetes pathways |
| 0.0 | 0.9 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |