Project
PRJDB7713: Age associated transcriptome analysis in 5 tissues of zebrafish
Navigation
Downloads
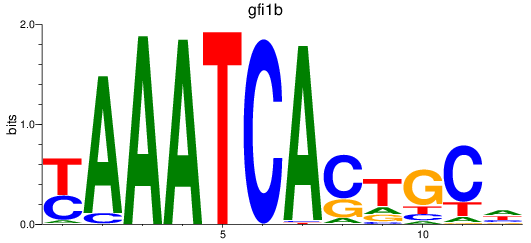
Results for gfi1b
Z-value: 1.48
Transcription factors associated with gfi1b
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
gfi1b
|
ENSDARG00000079947 | growth factor independent 1B transcription repressor |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| gfi1b | dr11_v1_chr21_-_17296789_17296789 | 0.12 | 2.6e-01 | Click! |
Activity profile of gfi1b motif
Sorted Z-values of gfi1b motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr17_-_12385308 | 22.72 |
ENSDART00000080927
|
snap25b
|
synaptosomal-associated protein, 25b |
| chr17_+_15433671 | 20.83 |
ENSDART00000149568
|
fabp7a
|
fatty acid binding protein 7, brain, a |
| chr17_-_12389259 | 20.81 |
ENSDART00000185724
|
snap25b
|
synaptosomal-associated protein, 25b |
| chr17_+_15433518 | 20.81 |
ENSDART00000026180
|
fabp7a
|
fatty acid binding protein 7, brain, a |
| chr25_+_19999623 | 17.86 |
ENSDART00000026401
|
zgc:194665
|
zgc:194665 |
| chr1_+_12763461 | 16.35 |
ENSDART00000159226
ENSDART00000180121 |
pcdh10a
|
protocadherin 10a |
| chr23_-_35694171 | 16.05 |
ENSDART00000077539
|
tuba1c
|
tubulin, alpha 1c |
| chr3_-_36440705 | 14.44 |
ENSDART00000162875
|
rogdi
|
rogdi homolog (Drosophila) |
| chr7_-_26408472 | 14.01 |
ENSDART00000111494
|
gal3st4
|
galactose-3-O-sulfotransferase 4 |
| chr4_-_12007404 | 12.74 |
ENSDART00000092250
|
btbd11a
|
BTB (POZ) domain containing 11a |
| chr24_-_32408404 | 12.22 |
ENSDART00000144157
|
si:ch211-56a11.2
|
si:ch211-56a11.2 |
| chr9_+_34641237 | 11.61 |
ENSDART00000133996
|
shox
|
short stature homeobox |
| chr5_-_40734045 | 10.82 |
ENSDART00000010896
|
isl1
|
ISL LIM homeobox 1 |
| chr25_-_518656 | 10.37 |
ENSDART00000156421
|
myo9ab
|
myosin IXAb |
| chr16_-_45235947 | 10.36 |
ENSDART00000164436
|
si:dkey-33i11.4
|
si:dkey-33i11.4 |
| chr21_+_9628854 | 9.97 |
ENSDART00000161753
ENSDART00000160711 |
mapk10
|
mitogen-activated protein kinase 10 |
| chr7_-_25895189 | 9.92 |
ENSDART00000173599
ENSDART00000079235 ENSDART00000173786 ENSDART00000173602 ENSDART00000079245 ENSDART00000187568 ENSDART00000173505 |
cd99l2
|
CD99 molecule-like 2 |
| chr11_-_6188413 | 9.79 |
ENSDART00000109972
|
ccl44
|
chemokine (C-C motif) ligand 44 |
| chr20_-_29864390 | 9.66 |
ENSDART00000161834
ENSDART00000132278 |
rnf144ab
|
ring finger protein 144ab |
| chr23_-_35694461 | 9.65 |
ENSDART00000185884
|
tuba1c
|
tubulin, alpha 1c |
| chr10_-_34871737 | 9.49 |
ENSDART00000138755
|
dclk1a
|
doublecortin-like kinase 1a |
| chr20_+_50852356 | 9.43 |
ENSDART00000167517
ENSDART00000168396 |
gphnb
|
gephyrin b |
| chr9_-_459910 | 9.37 |
ENSDART00000162551
|
si:dkey-11f4.16
|
si:dkey-11f4.16 |
| chr12_+_21299338 | 9.29 |
ENSDART00000074540
ENSDART00000133188 |
ca10a
|
carbonic anhydrase Xa |
| chr16_+_26612401 | 9.27 |
ENSDART00000145571
|
epb41l4b
|
erythrocyte membrane protein band 4.1 like 4B |
| chr9_+_30108641 | 8.98 |
ENSDART00000060174
|
jagn1a
|
jagunal homolog 1a |
| chr21_-_4032650 | 8.82 |
ENSDART00000151648
|
ntng2b
|
netrin g2b |
| chr23_-_29505463 | 8.75 |
ENSDART00000050915
|
kif1b
|
kinesin family member 1B |
| chr7_-_28148310 | 8.71 |
ENSDART00000044208
|
lmo1
|
LIM domain only 1 |
| chr16_+_43152727 | 8.43 |
ENSDART00000125590
ENSDART00000154493 |
adam22
|
ADAM metallopeptidase domain 22 |
| chr25_-_15049694 | 8.35 |
ENSDART00000162485
ENSDART00000164384 ENSDART00000165632 ENSDART00000159490 |
pax6a
|
paired box 6a |
| chr5_-_23362602 | 7.91 |
ENSDART00000137120
|
gria3a
|
glutamate receptor, ionotropic, AMPA 3a |
| chr16_-_44945224 | 7.83 |
ENSDART00000156921
|
ncam3
|
neural cell adhesion molecule 3 |
| chr14_+_49007501 | 7.70 |
ENSDART00000128508
|
zdhhc5b
|
zinc finger, DHHC-type containing 5b |
| chr2_-_36925561 | 7.68 |
ENSDART00000187690
|
map1sb
|
microtubule-associated protein 1Sb |
| chr2_+_34967022 | 7.56 |
ENSDART00000134926
|
astn1
|
astrotactin 1 |
| chr16_+_39159752 | 7.52 |
ENSDART00000122081
|
sybu
|
syntabulin (syntaxin-interacting) |
| chr7_+_22801465 | 7.35 |
ENSDART00000052862
ENSDART00000173633 |
rbm4.1
|
RNA binding motif protein 4.1 |
| chr18_-_16181952 | 7.20 |
ENSDART00000157824
|
slc6a15
|
solute carrier family 6 (neutral amino acid transporter), member 15 |
| chr20_-_26042070 | 7.17 |
ENSDART00000140255
|
si:dkey-12h9.6
|
si:dkey-12h9.6 |
| chr3_-_42086577 | 7.13 |
ENSDART00000083111
ENSDART00000187312 |
ttyh3a
|
tweety family member 3a |
| chr13_+_15816573 | 7.12 |
ENSDART00000137061
|
klc1a
|
kinesin light chain 1a |
| chr19_-_9503473 | 6.96 |
ENSDART00000091615
|
iffo1a
|
intermediate filament family orphan 1a |
| chr2_+_34967210 | 6.86 |
ENSDART00000141796
|
astn1
|
astrotactin 1 |
| chr2_+_54482603 | 6.83 |
ENSDART00000130977
ENSDART00000183090 |
mtcl1
|
microtubule crosslinking factor 1 |
| chr5_-_62306819 | 6.80 |
ENSDART00000168993
|
spata22
|
spermatogenesis associated 22 |
| chr5_-_40510397 | 6.73 |
ENSDART00000146237
ENSDART00000051065 |
fsta
|
follistatin a |
| chr19_+_29798064 | 6.71 |
ENSDART00000167803
ENSDART00000051804 |
marcksl1b
|
MARCKS-like 1b |
| chr17_+_9308425 | 6.63 |
ENSDART00000188283
ENSDART00000183311 |
NPAS3
|
neuronal PAS domain protein 3 |
| chr16_+_22761846 | 6.50 |
ENSDART00000193028
|
CR854942.1
|
|
| chr7_+_7019911 | 6.48 |
ENSDART00000172421
|
rbm14b
|
RNA binding motif protein 14b |
| chr6_+_11249706 | 6.33 |
ENSDART00000186547
ENSDART00000193287 |
atg9a
|
ATG9 autophagy related 9 homolog A (S. cerevisiae) |
| chr4_-_63923523 | 6.32 |
ENSDART00000144330
|
si:dkey-179k24.1
|
si:dkey-179k24.1 |
| chr17_+_34805897 | 6.25 |
ENSDART00000137090
ENSDART00000077626 |
id2a
|
inhibitor of DNA binding 2a |
| chr21_+_25068215 | 6.02 |
ENSDART00000167523
ENSDART00000189259 |
dixdc1b
|
DIX domain containing 1b |
| chr8_-_52715911 | 6.00 |
ENSDART00000168241
|
tubb2b
|
tubulin, beta 2b |
| chr19_-_12193622 | 5.95 |
ENSDART00000041960
|
ywhaz
|
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, zeta polypeptide |
| chr5_+_57773222 | 5.95 |
ENSDART00000135344
|
ppp2r1ba
|
protein phosphatase 2, regulatory subunit A, beta a |
| chr9_+_44722205 | 5.87 |
ENSDART00000086176
ENSDART00000145271 ENSDART00000132696 |
nckap1
|
NCK-associated protein 1 |
| chr6_+_55277419 | 5.80 |
ENSDART00000083670
|
CABZ01041604.1
|
|
| chr7_+_48319916 | 5.79 |
ENSDART00000052122
|
crabp1b
|
cellular retinoic acid binding protein 1b |
| chr7_+_25920792 | 5.76 |
ENSDART00000026295
|
arrb2b
|
arrestin, beta 2b |
| chr14_-_6943934 | 5.68 |
ENSDART00000126279
|
clk4a
|
CDC-like kinase 4a |
| chr21_-_33995710 | 5.67 |
ENSDART00000100508
ENSDART00000179622 |
ebf1b
|
early B cell factor 1b |
| chr2_+_31838442 | 5.44 |
ENSDART00000066789
|
stard3nl
|
STARD3 N-terminal like |
| chr16_+_10422836 | 5.35 |
ENSDART00000161568
|
ino80e
|
INO80 complex subunit E |
| chr6_-_12851888 | 5.33 |
ENSDART00000056764
|
bmpr2a
|
bone morphogenetic protein receptor, type II a (serine/threonine kinase) |
| chr14_-_47391084 | 5.15 |
ENSDART00000159608
|
fstl5
|
follistatin-like 5 |
| chr5_+_60928576 | 5.12 |
ENSDART00000131041
|
doc2b
|
double C2-like domains, beta |
| chr14_-_47314340 | 5.09 |
ENSDART00000164851
|
fstl5
|
follistatin-like 5 |
| chr3_+_51684963 | 5.06 |
ENSDART00000091180
ENSDART00000183711 ENSDART00000159493 |
baiap2a
|
BAI1-associated protein 2a |
| chr6_+_11250033 | 5.01 |
ENSDART00000065411
ENSDART00000132677 |
atg9a
|
ATG9 autophagy related 9 homolog A (S. cerevisiae) |
| chr6_+_11250316 | 5.00 |
ENSDART00000137122
|
atg9a
|
ATG9 autophagy related 9 homolog A (S. cerevisiae) |
| chr21_-_33995213 | 4.63 |
ENSDART00000140184
|
EBF1 (1 of many)
|
si:ch211-51e8.2 |
| chr25_-_3979583 | 4.62 |
ENSDART00000124749
|
myrf
|
myelin regulatory factor |
| chr5_+_43965078 | 4.60 |
ENSDART00000113502
ENSDART00000187143 |
TMEM8B
|
si:dkey-84j12.1 |
| chr18_+_27511976 | 4.60 |
ENSDART00000132017
ENSDART00000140781 |
tp53i11b
|
tumor protein p53 inducible protein 11b |
| chr19_+_4912817 | 4.59 |
ENSDART00000101658
ENSDART00000165082 |
ppp1r1b
|
protein phosphatase 1, regulatory (inhibitor) subunit 1B |
| chr6_+_29791164 | 4.58 |
ENSDART00000017424
|
ptmaa
|
prothymosin, alpha a |
| chr19_-_28130658 | 4.57 |
ENSDART00000079114
|
irx1b
|
iroquois homeobox 1b |
| chr4_-_5912951 | 4.53 |
ENSDART00000169439
|
arl1
|
ADP-ribosylation factor-like 1 |
| chr11_-_2297832 | 4.46 |
ENSDART00000158266
|
znf740a
|
zinc finger protein 740a |
| chr20_+_28434196 | 4.41 |
ENSDART00000034245
|
dpf3
|
D4, zinc and double PHD fingers, family 3 |
| chr11_+_25044082 | 4.41 |
ENSDART00000123263
|
phf20a
|
PHD finger protein 20, a |
| chr24_+_26402110 | 4.30 |
ENSDART00000133684
|
si:ch211-230g15.5
|
si:ch211-230g15.5 |
| chr13_+_22712406 | 4.21 |
ENSDART00000132847
|
si:ch211-134m17.9
|
si:ch211-134m17.9 |
| chr13_+_15657911 | 4.21 |
ENSDART00000134972
ENSDART00000138991 ENSDART00000133342 |
mark3a
|
MAP/microtubule affinity-regulating kinase 3a |
| chr2_-_15040345 | 4.16 |
ENSDART00000109657
|
si:dkey-10f21.4
|
si:dkey-10f21.4 |
| chr9_-_3653259 | 4.14 |
ENSDART00000140425
ENSDART00000025332 |
gad1a
|
glutamate decarboxylase 1a |
| chr3_+_26223376 | 4.09 |
ENSDART00000128284
|
nudt9
|
nudix (nucleoside diphosphate linked moiety X)-type motif 9 |
| chr2_+_9821757 | 4.04 |
ENSDART00000018408
ENSDART00000141227 ENSDART00000144681 ENSDART00000148227 |
anxa13l
|
annexin A13, like |
| chr4_-_17642168 | 3.99 |
ENSDART00000007030
|
klhl42
|
kelch-like family, member 42 |
| chr3_-_39696066 | 3.95 |
ENSDART00000015393
|
b9d1
|
B9 protein domain 1 |
| chr6_+_35362225 | 3.87 |
ENSDART00000133783
ENSDART00000102483 |
rgs4
|
regulator of G protein signaling 4 |
| chr6_-_10780698 | 3.84 |
ENSDART00000151714
|
gpr155b
|
G protein-coupled receptor 155b |
| chr7_-_33684632 | 3.84 |
ENSDART00000130553
|
tle3b
|
transducin-like enhancer of split 3b |
| chr1_+_18863060 | 3.72 |
ENSDART00000139241
|
rnf38
|
ring finger protein 38 |
| chr8_-_43574935 | 3.69 |
ENSDART00000051139
|
ncor2
|
nuclear receptor corepressor 2 |
| chr3_-_39695856 | 3.65 |
ENSDART00000148247
|
b9d1
|
B9 protein domain 1 |
| chr5_-_36328688 | 3.65 |
ENSDART00000011399
|
efnb1
|
ephrin-B1 |
| chr21_-_2935455 | 3.58 |
ENSDART00000163963
|
CABZ01023255.1
|
|
| chr23_+_7471072 | 3.58 |
ENSDART00000135551
|
si:ch211-200e2.1
|
si:ch211-200e2.1 |
| chr18_-_22735002 | 3.52 |
ENSDART00000023721
|
nudt21
|
nudix hydrolase 21 |
| chr17_+_11675362 | 3.52 |
ENSDART00000157911
|
kif26ba
|
kinesin family member 26Ba |
| chr3_-_54607166 | 3.50 |
ENSDART00000021977
|
dnmt1
|
DNA (cytosine-5-)-methyltransferase 1 |
| chr19_+_21919856 | 3.49 |
ENSDART00000187306
ENSDART00000138544 |
galr1a
|
galanin receptor 1a |
| chr2_+_9822319 | 3.47 |
ENSDART00000144078
ENSDART00000144371 |
anxa13l
|
annexin A13, like |
| chr5_+_20823409 | 3.47 |
ENSDART00000093185
ENSDART00000142894 |
limk2
|
LIM domain kinase 2 |
| chr20_-_43917647 | 3.41 |
ENSDART00000026213
|
mark3b
|
MAP/microtubule affinity-regulating kinase 3b |
| chr21_+_27370671 | 3.39 |
ENSDART00000009234
ENSDART00000142071 |
rbm14a
|
RNA binding motif protein 14a |
| chr23_+_25232411 | 3.34 |
ENSDART00000138974
|
erbb3b
|
erb-b2 receptor tyrosine kinase 3b |
| chr20_-_40487208 | 3.27 |
ENSDART00000075070
ENSDART00000142029 |
hsf2
|
heat shock transcription factor 2 |
| chr8_+_20495889 | 3.26 |
ENSDART00000138794
ENSDART00000021683 |
csnk1g2b
|
casein kinase 1, gamma 2b |
| chr22_+_34784075 | 3.24 |
ENSDART00000167538
|
lcor
|
ligand dependent nuclear receptor corepressor |
| chr5_+_45976130 | 3.23 |
ENSDART00000175670
|
sv2c
|
synaptic vesicle glycoprotein 2C |
| chr20_+_6535176 | 3.13 |
ENSDART00000054652
|
si:ch211-191a24.4
|
si:ch211-191a24.4 |
| chr7_+_35191220 | 3.05 |
ENSDART00000110552
|
zdhhc1
|
zinc finger, DHHC-type containing 1 |
| chr17_+_24687338 | 3.03 |
ENSDART00000135794
|
selenon
|
selenoprotein N |
| chr16_+_10329701 | 3.00 |
ENSDART00000172845
|
mdc1
|
mediator of DNA damage checkpoint 1 |
| chr14_-_21218891 | 2.96 |
ENSDART00000158294
|
ppp2r2cb
|
protein phosphatase 2, regulatory subunit B, gamma b |
| chr4_-_20235904 | 2.95 |
ENSDART00000146621
ENSDART00000193655 |
stk38l
|
serine/threonine kinase 38 like |
| chr25_+_17313568 | 2.94 |
ENSDART00000125459
|
cnot1
|
CCR4-NOT transcription complex, subunit 1 |
| chr23_-_3721444 | 2.94 |
ENSDART00000141682
|
nudt3a
|
nudix (nucleoside diphosphate linked moiety X)-type motif 3a |
| chr2_-_13254821 | 2.86 |
ENSDART00000022621
|
kdsr
|
3-ketodihydrosphingosine reductase |
| chr6_+_50393047 | 2.83 |
ENSDART00000055502
ENSDART00000055511 |
ergic3
|
ERGIC and golgi 3 |
| chr1_+_1805294 | 2.67 |
ENSDART00000103850
|
atp1a1a.3
|
ATPase Na+/K+ transporting subunit alpha 1a, tandem duplicate 3 |
| chr4_+_6833583 | 2.60 |
ENSDART00000165179
ENSDART00000186134 ENSDART00000174507 |
dock4b
|
dedicator of cytokinesis 4b |
| chr22_+_31821815 | 2.59 |
ENSDART00000159825
|
dock3
|
dedicator of cytokinesis 3 |
| chr20_-_16852 | 2.59 |
ENSDART00000129277
ENSDART00000148717 |
zgc:174972
|
zgc:174972 |
| chr12_+_4220353 | 2.57 |
ENSDART00000133675
|
mapk7
|
mitogen-activated protein kinase 7 |
| chr20_-_45772306 | 2.49 |
ENSDART00000062092
|
trmt6
|
tRNA methyltransferase 6 homolog (S. cerevisiae) |
| chr16_-_263658 | 2.42 |
ENSDART00000129303
|
slc6a3
|
solute carrier family 6 (neurotransmitter transporter), member 3 |
| chr6_-_32349153 | 2.38 |
ENSDART00000140004
|
angptl3
|
angiopoietin-like 3 |
| chr23_+_25232711 | 2.31 |
ENSDART00000128510
|
erbb3b
|
erb-b2 receptor tyrosine kinase 3b |
| chr6_-_12912606 | 2.30 |
ENSDART00000164640
|
ical1
|
islet cell autoantigen 1-like |
| chr5_-_58832332 | 2.18 |
ENSDART00000161230
|
arhgef12b
|
Rho guanine nucleotide exchange factor (GEF) 12b |
| chr20_-_28433616 | 2.13 |
ENSDART00000169289
|
wdr21
|
WD repeat domain 21 |
| chr4_-_13567387 | 2.08 |
ENSDART00000132971
ENSDART00000102010 |
mdm1
|
Mdm1 nuclear protein homolog (mouse) |
| chr23_-_29505645 | 2.07 |
ENSDART00000146458
|
kif1b
|
kinesin family member 1B |
| chr7_-_35036770 | 2.05 |
ENSDART00000123174
|
galr1b
|
galanin receptor 1b |
| chr24_-_23716097 | 2.05 |
ENSDART00000084954
ENSDART00000129028 |
pign
|
phosphatidylinositol glycan anchor biosynthesis, class N |
| chr7_+_28960518 | 2.03 |
ENSDART00000182903
ENSDART00000188250 ENSDART00000191830 |
dok4
|
docking protein 4 |
| chr8_-_1266181 | 2.01 |
ENSDART00000148654
ENSDART00000149924 |
cdc14b
|
cell division cycle 14B |
| chr21_+_7605803 | 1.95 |
ENSDART00000121813
|
wdr41
|
WD repeat domain 41 |
| chr1_-_47089818 | 1.94 |
ENSDART00000132378
|
itsn1
|
intersectin 1 (SH3 domain protein) |
| chr5_-_37871526 | 1.91 |
ENSDART00000136450
|
arhgap35b
|
Rho GTPase activating protein 35b |
| chr9_+_48007081 | 1.91 |
ENSDART00000060593
ENSDART00000099835 |
zgc:92380
|
zgc:92380 |
| chr25_-_17918810 | 1.90 |
ENSDART00000023959
|
arntl1a
|
aryl hydrocarbon receptor nuclear translocator-like 1a |
| chr13_+_21779975 | 1.87 |
ENSDART00000021556
|
si:ch211-51a6.2
|
si:ch211-51a6.2 |
| chr4_-_42242844 | 1.87 |
ENSDART00000163476
|
si:ch211-129p6.2
|
si:ch211-129p6.2 |
| chr8_-_16515127 | 1.85 |
ENSDART00000146469
ENSDART00000132681 |
ttc39a
|
tetratricopeptide repeat domain 39A |
| chr7_+_30201611 | 1.85 |
ENSDART00000075588
|
wdr76
|
WD repeat domain 76 |
| chr7_-_72208248 | 1.81 |
ENSDART00000108916
|
zmp:0000001168
|
zmp:0000001168 |
| chr14_-_45512807 | 1.78 |
ENSDART00000173172
|
si:ch211-114c17.1
|
si:ch211-114c17.1 |
| chr12_-_4220713 | 1.77 |
ENSDART00000129427
|
vkorc1
|
vitamin K epoxide reductase complex, subunit 1 |
| chr16_-_41667101 | 1.72 |
ENSDART00000084528
|
atp2c1
|
ATPase secretory pathway Ca2+ transporting 1 |
| chr7_+_13609457 | 1.72 |
ENSDART00000172857
|
ankdd1a
|
ankyrin repeat and death domain containing 1A |
| chr14_+_35023923 | 1.71 |
ENSDART00000172171
|
ebf3a
|
early B cell factor 3a |
| chr6_+_46431848 | 1.69 |
ENSDART00000181056
ENSDART00000144569 ENSDART00000064865 ENSDART00000133992 |
stau1
|
staufen double-stranded RNA binding protein 1 |
| chr7_+_30202104 | 1.69 |
ENSDART00000173525
|
wdr76
|
WD repeat domain 76 |
| chr2_+_37836821 | 1.66 |
ENSDART00000143203
|
parp2
|
poly (ADP-ribose) polymerase 2 |
| chr22_-_4407871 | 1.63 |
ENSDART00000162523
|
kdm4b
|
lysine (K)-specific demethylase 4B |
| chr17_-_33552363 | 1.62 |
ENSDART00000154400
|
tmem121aa
|
transmembrane protein 121Aa |
| chr25_-_17918536 | 1.62 |
ENSDART00000148660
|
arntl1a
|
aryl hydrocarbon receptor nuclear translocator-like 1a |
| chr2_-_13254594 | 1.61 |
ENSDART00000155671
|
kdsr
|
3-ketodihydrosphingosine reductase |
| chr20_+_13783040 | 1.59 |
ENSDART00000115329
ENSDART00000152497 |
lpgat1
|
lysophosphatidylglycerol acyltransferase 1 |
| chr20_-_28433990 | 1.59 |
ENSDART00000182824
ENSDART00000193381 |
wdr21
|
WD repeat domain 21 |
| chr5_+_44064764 | 1.57 |
ENSDART00000143843
|
si:dkey-84j12.1
|
si:dkey-84j12.1 |
| chr19_-_31584444 | 1.57 |
ENSDART00000052183
|
zgc:111986
|
zgc:111986 |
| chr10_-_34867401 | 1.51 |
ENSDART00000145545
|
dclk1a
|
doublecortin-like kinase 1a |
| chr20_+_27464721 | 1.42 |
ENSDART00000189552
|
kif26aa
|
kinesin family member 26Aa |
| chr16_-_6849754 | 1.40 |
ENSDART00000149206
ENSDART00000149778 |
mbpb
|
myelin basic protein b |
| chr16_+_25608778 | 1.39 |
ENSDART00000077484
|
zhx2a
|
zinc fingers and homeoboxes 2a |
| chr7_-_66133786 | 1.36 |
ENSDART00000154961
|
btbd10b
|
BTB (POZ) domain containing 10b |
| chr8_+_23615132 | 1.35 |
ENSDART00000099769
|
ccdc22
|
coiled-coil domain containing 22 |
| chr11_+_31121340 | 1.34 |
ENSDART00000185172
|
stx10
|
syntaxin 10 |
| chr8_+_10862353 | 1.33 |
ENSDART00000140717
|
brpf3b
|
bromodomain and PHD finger containing, 3b |
| chr10_+_8968203 | 1.27 |
ENSDART00000110443
ENSDART00000080772 |
fstb
|
follistatin b |
| chr11_+_18612166 | 1.19 |
ENSDART00000162694
|
ncoa3
|
nuclear receptor coactivator 3 |
| chr23_-_18415872 | 1.11 |
ENSDART00000135430
|
fam120c
|
family with sequence similarity 120C |
| chr11_+_13423776 | 1.06 |
ENSDART00000102553
|
homer3b
|
homer scaffolding protein 3b |
| chr16_-_31824525 | 1.00 |
ENSDART00000058737
|
cdc42l
|
cell division cycle 42, like |
| chr19_-_31707892 | 0.97 |
ENSDART00000088427
|
ripor2
|
RHO family interacting cell polarization regulator 2 |
| chr3_-_12970418 | 0.92 |
ENSDART00000158747
|
pdgfab
|
platelet-derived growth factor alpha polypeptide b |
| chr4_-_27897160 | 0.90 |
ENSDART00000066924
ENSDART00000066925 ENSDART00000193020 |
tbc1d22a
|
TBC1 domain family, member 22a |
| chr4_-_73548389 | 0.89 |
ENSDART00000174327
ENSDART00000150753 ENSDART00000170775 |
si:ch73-266f23.1
|
si:ch73-266f23.1 |
| chr12_+_19199735 | 0.88 |
ENSDART00000066393
|
pdap1a
|
pdgfa associated protein 1a |
| chr21_+_26733529 | 0.88 |
ENSDART00000168379
|
pcxa
|
pyruvate carboxylase a |
| chr17_-_37195163 | 0.87 |
ENSDART00000108514
|
asxl2
|
additional sex combs like transcriptional regulator 2 |
| chr4_-_16706776 | 0.86 |
ENSDART00000079461
|
dennd5b
|
DENN/MADD domain containing 5B |
| chr20_+_34596205 | 0.84 |
ENSDART00000138338
|
si:ch211-242b18.1
|
si:ch211-242b18.1 |
| chr1_+_9860381 | 0.80 |
ENSDART00000054848
|
pmm2
|
phosphomannomutase 2 |
| chr16_+_23282655 | 0.78 |
ENSDART00000015956
|
efna1b
|
ephrin-A1b |
| chr11_+_13424116 | 0.72 |
ENSDART00000125563
|
homer3b
|
homer scaffolding protein 3b |
| chr2_-_10338759 | 0.72 |
ENSDART00000150166
ENSDART00000149584 |
gng12a
|
guanine nucleotide binding protein (G protein), gamma 12a |
| chr6_+_37655078 | 0.70 |
ENSDART00000122199
ENSDART00000065127 |
cyfip1
|
cytoplasmic FMR1 interacting protein 1 |
| chr12_-_26022663 | 0.67 |
ENSDART00000166769
|
bmpr1ab
|
bone morphogenetic protein receptor, type IAb |
| chr8_-_15182232 | 0.67 |
ENSDART00000138855
|
bcar3
|
BCAR3, NSP family adaptor protein |
| chr11_-_42396302 | 0.65 |
ENSDART00000165624
|
slmapa
|
sarcolemma associated protein a |
| chr20_+_35857399 | 0.63 |
ENSDART00000102611
|
cd2ap
|
CD2-associated protein |
Network of associatons between targets according to the STRING database.
First level regulatory network of gfi1b
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.6 | 10.8 | GO:0048917 | posterior lateral line ganglion development(GO:0048917) |
| 3.1 | 9.4 | GO:0072579 | establishment of synaptic specificity at neuromuscular junction(GO:0007529) molybdenum incorporation into molybdenum-molybdopterin complex(GO:0018315) metal incorporation into metallo-molybdopterin complex(GO:0042040) glycine receptor clustering(GO:0072579) |
| 3.1 | 43.5 | GO:0045162 | clustering of voltage-gated sodium channels(GO:0045162) |
| 3.0 | 9.0 | GO:0038158 | granulocyte colony-stimulating factor signaling pathway(GO:0038158) |
| 2.3 | 6.8 | GO:0000711 | meiotic DNA repair synthesis(GO:0000711) |
| 2.1 | 10.4 | GO:0098529 | neuromuscular junction development, skeletal muscle fiber(GO:0098529) |
| 1.9 | 5.7 | GO:0048913 | anterior lateral line nerve glial cell differentiation(GO:0048913) myelination of anterior lateral line nerve axons(GO:0048914) anterior lateral line nerve glial cell development(GO:0048939) anterior lateral line nerve glial cell morphogenesis involved in differentiation(GO:0048940) |
| 1.9 | 7.5 | GO:0060074 | synapse maturation(GO:0060074) |
| 1.8 | 7.2 | GO:0015820 | branched-chain amino acid transport(GO:0015803) leucine transport(GO:0015820) |
| 1.6 | 16.3 | GO:0044805 | late nucleophagy(GO:0044805) |
| 1.4 | 5.8 | GO:1900136 | regulation of cytokine activity(GO:0060300) regulation of receptor binding(GO:1900120) regulation of chemokine activity(GO:1900136) |
| 1.4 | 7.2 | GO:0023041 | neuronal signal transduction(GO:0023041) |
| 1.2 | 8.3 | GO:0060221 | retinal rod cell differentiation(GO:0060221) |
| 1.2 | 3.5 | GO:1990120 | messenger ribonucleoprotein complex assembly(GO:1990120) |
| 1.2 | 5.8 | GO:0016103 | diterpenoid catabolic process(GO:0016103) terpenoid catabolic process(GO:0016115) retinoic acid catabolic process(GO:0034653) |
| 1.1 | 4.5 | GO:0031584 | activation of phospholipase D activity(GO:0031584) |
| 1.1 | 4.5 | GO:0036344 | platelet formation(GO:0030220) platelet morphogenesis(GO:0036344) |
| 1.0 | 14.4 | GO:0007158 | neuron cell-cell adhesion(GO:0007158) |
| 0.9 | 16.4 | GO:0097324 | melanocyte migration(GO:0097324) |
| 0.9 | 6.3 | GO:0046532 | regulation of photoreceptor cell differentiation(GO:0046532) |
| 0.8 | 2.4 | GO:0015874 | norepinephrine transport(GO:0015874) |
| 0.7 | 2.9 | GO:0010606 | positive regulation of cytoplasmic mRNA processing body assembly(GO:0010606) negative regulation of retinoic acid receptor signaling pathway(GO:0048387) |
| 0.7 | 8.0 | GO:0032926 | negative regulation of activin receptor signaling pathway(GO:0032926) |
| 0.7 | 3.5 | GO:2000001 | regulation of DNA damage checkpoint(GO:2000001) |
| 0.7 | 3.5 | GO:0032776 | DNA methylation on cytosine(GO:0032776) |
| 0.6 | 2.4 | GO:0055091 | phospholipid homeostasis(GO:0055091) |
| 0.6 | 1.8 | GO:0042373 | vitamin K metabolic process(GO:0042373) |
| 0.6 | 2.9 | GO:1901910 | diphosphoinositol polyphosphate metabolic process(GO:0071543) diadenosine pentaphosphate metabolic process(GO:1901906) diadenosine pentaphosphate catabolic process(GO:1901907) diadenosine hexaphosphate metabolic process(GO:1901908) diadenosine hexaphosphate catabolic process(GO:1901909) adenosine 5'-(hexahydrogen pentaphosphate) metabolic process(GO:1901910) adenosine 5'-(hexahydrogen pentaphosphate) catabolic process(GO:1901911) |
| 0.6 | 5.1 | GO:2000251 | positive regulation of actin cytoskeleton reorganization(GO:2000251) |
| 0.5 | 3.6 | GO:0003404 | optic vesicle morphogenesis(GO:0003404) |
| 0.5 | 5.1 | GO:0032024 | positive regulation of insulin secretion(GO:0032024) |
| 0.5 | 4.2 | GO:0043217 | myelin maintenance(GO:0043217) |
| 0.4 | 3.0 | GO:0050848 | regulation of calcium-mediated signaling(GO:0050848) |
| 0.4 | 1.6 | GO:0036149 | phosphatidylinositol acyl-chain remodeling(GO:0036149) |
| 0.3 | 1.7 | GO:0055071 | manganese ion homeostasis(GO:0055071) |
| 0.3 | 1.4 | GO:0042787 | protein ubiquitination involved in ubiquitin-dependent protein catabolic process(GO:0042787) regulation of protein ubiquitination involved in ubiquitin-dependent protein catabolic process(GO:2000058) |
| 0.3 | 1.0 | GO:2000402 | response to chemokine(GO:1990868) cellular response to chemokine(GO:1990869) negative regulation of lymphocyte migration(GO:2000402) negative regulation of T cell migration(GO:2000405) |
| 0.3 | 8.8 | GO:0070831 | basement membrane assembly(GO:0070831) |
| 0.3 | 12.2 | GO:0010921 | regulation of phosphatase activity(GO:0010921) |
| 0.3 | 2.1 | GO:0046600 | negative regulation of centriole replication(GO:0046600) |
| 0.3 | 1.8 | GO:0000395 | mRNA 5'-splice site recognition(GO:0000395) |
| 0.3 | 3.0 | GO:0070262 | peptidyl-serine dephosphorylation(GO:0070262) |
| 0.3 | 7.7 | GO:0031114 | regulation of microtubule depolymerization(GO:0031114) |
| 0.3 | 16.1 | GO:0009247 | glycolipid biosynthetic process(GO:0009247) |
| 0.3 | 1.8 | GO:2001256 | regulation of store-operated calcium entry(GO:2001256) |
| 0.2 | 9.8 | GO:0048247 | lymphocyte chemotaxis(GO:0048247) |
| 0.2 | 0.7 | GO:0060912 | cardiac cell fate specification(GO:0060912) |
| 0.2 | 4.2 | GO:0010569 | regulation of double-strand break repair via homologous recombination(GO:0010569) |
| 0.2 | 3.5 | GO:0030818 | negative regulation of adenylate cyclase activity(GO:0007194) negative regulation of cyclic nucleotide metabolic process(GO:0030800) negative regulation of cyclic nucleotide biosynthetic process(GO:0030803) negative regulation of nucleotide biosynthetic process(GO:0030809) negative regulation of cAMP metabolic process(GO:0030815) negative regulation of cAMP biosynthetic process(GO:0030818) negative regulation of cyclase activity(GO:0031280) negative regulation of lyase activity(GO:0051350) negative regulation of purine nucleotide biosynthetic process(GO:1900372) |
| 0.2 | 5.3 | GO:0048570 | notochord morphogenesis(GO:0048570) |
| 0.2 | 11.6 | GO:0030282 | bone mineralization(GO:0030282) |
| 0.2 | 2.0 | GO:0071850 | mitotic cell cycle arrest(GO:0071850) |
| 0.2 | 3.7 | GO:0030225 | macrophage differentiation(GO:0030225) |
| 0.2 | 10.8 | GO:0018345 | protein palmitoylation(GO:0018345) |
| 0.2 | 9.3 | GO:0036269 | swimming behavior(GO:0036269) |
| 0.2 | 9.7 | GO:1901800 | positive regulation of proteasomal ubiquitin-dependent protein catabolic process(GO:0032436) positive regulation of proteasomal protein catabolic process(GO:1901800) positive regulation of proteolysis involved in cellular protein catabolic process(GO:1903052) |
| 0.2 | 0.7 | GO:0099563 | modification of synaptic structure(GO:0099563) |
| 0.2 | 10.0 | GO:0048484 | enteric nervous system development(GO:0048484) |
| 0.2 | 4.4 | GO:0055008 | cardiac muscle tissue morphogenesis(GO:0055008) |
| 0.1 | 2.7 | GO:0071436 | cellular potassium ion homeostasis(GO:0030007) sodium ion export from cell(GO:0036376) sodium ion export(GO:0071436) |
| 0.1 | 1.7 | GO:0070212 | protein poly-ADP-ribosylation(GO:0070212) |
| 0.1 | 0.8 | GO:0009298 | mannose metabolic process(GO:0006013) GDP-mannose biosynthetic process(GO:0009298) |
| 0.1 | 1.0 | GO:0032488 | Cdc42 protein signal transduction(GO:0032488) |
| 0.1 | 0.9 | GO:0046959 | nonassociative learning(GO:0046958) habituation(GO:0046959) |
| 0.1 | 3.5 | GO:0009648 | photoperiodism(GO:0009648) |
| 0.1 | 0.9 | GO:0031954 | positive regulation of protein autophosphorylation(GO:0031954) |
| 0.1 | 1.3 | GO:0044154 | histone H3-K14 acetylation(GO:0044154) |
| 0.1 | 2.6 | GO:0001503 | ossification(GO:0001503) |
| 0.1 | 8.8 | GO:0010506 | regulation of autophagy(GO:0010506) |
| 0.1 | 0.6 | GO:0001672 | regulation of chromatin assembly or disassembly(GO:0001672) |
| 0.1 | 0.5 | GO:1904262 | negative regulation of TORC1 signaling(GO:1904262) |
| 0.1 | 2.8 | GO:0006890 | retrograde vesicle-mediated transport, Golgi to ER(GO:0006890) |
| 0.1 | 6.4 | GO:0007224 | smoothened signaling pathway(GO:0007224) |
| 0.1 | 3.3 | GO:0030866 | cortical actin cytoskeleton organization(GO:0030866) |
| 0.1 | 29.1 | GO:0000226 | microtubule cytoskeleton organization(GO:0000226) |
| 0.1 | 7.0 | GO:0006338 | chromatin remodeling(GO:0006338) |
| 0.1 | 4.4 | GO:0016573 | histone acetylation(GO:0016573) |
| 0.1 | 13.6 | GO:0060070 | canonical Wnt signaling pathway(GO:0060070) |
| 0.1 | 4.1 | GO:0010469 | regulation of receptor activity(GO:0010469) |
| 0.0 | 1.6 | GO:0030488 | tRNA methylation(GO:0030488) |
| 0.0 | 1.8 | GO:0031532 | actin cytoskeleton reorganization(GO:0031532) |
| 0.0 | 8.5 | GO:0031032 | actomyosin structure organization(GO:0031032) |
| 0.0 | 5.4 | GO:0006821 | chloride transport(GO:0006821) |
| 0.0 | 6.3 | GO:0071773 | BMP signaling pathway(GO:0030509) response to BMP(GO:0071772) cellular response to BMP stimulus(GO:0071773) |
| 0.0 | 2.2 | GO:0031638 | zymogen activation(GO:0031638) |
| 0.0 | 0.2 | GO:0006526 | arginine biosynthetic process(GO:0006526) |
| 0.0 | 0.8 | GO:0048013 | ephrin receptor signaling pathway(GO:0048013) |
| 0.0 | 0.7 | GO:0033138 | positive regulation of peptidyl-serine phosphorylation(GO:0033138) |
| 0.0 | 4.6 | GO:0030902 | hindbrain development(GO:0030902) |
| 0.0 | 2.2 | GO:0007626 | locomotory behavior(GO:0007626) |
| 0.0 | 1.3 | GO:0042147 | retrograde transport, endosome to Golgi(GO:0042147) |
| 0.0 | 7.7 | GO:0000377 | RNA splicing, via transesterification reactions(GO:0000375) RNA splicing, via transesterification reactions with bulged adenosine as nucleophile(GO:0000377) mRNA splicing, via spliceosome(GO:0000398) |
| 0.0 | 3.5 | GO:0051216 | cartilage development(GO:0051216) |
| 0.0 | 0.5 | GO:0007205 | protein kinase C-activating G-protein coupled receptor signaling pathway(GO:0007205) |
| 0.0 | 0.6 | GO:0003094 | glomerular filtration(GO:0003094) |
| 0.0 | 0.6 | GO:0006383 | transcription from RNA polymerase III promoter(GO:0006383) |
| 0.0 | 0.5 | GO:0051209 | release of sequestered calcium ion into cytosol(GO:0051209) |
| 0.0 | 1.5 | GO:0007218 | neuropeptide signaling pathway(GO:0007218) |
| 0.0 | 2.4 | GO:0007018 | microtubule-based movement(GO:0007018) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 7.3 | 43.5 | GO:0070032 | synaptobrevin 2-SNAP-25-syntaxin-1a-complexin I complex(GO:0070032) |
| 3.6 | 14.4 | GO:0043291 | RAVE complex(GO:0043291) |
| 1.2 | 3.5 | GO:0042382 | paraspeckles(GO:0042382) |
| 1.0 | 5.9 | GO:0031258 | lamellipodium membrane(GO:0031258) |
| 0.6 | 16.3 | GO:0000407 | pre-autophagosomal structure(GO:0000407) |
| 0.5 | 2.5 | GO:0031515 | tRNA (m1A) methyltransferase complex(GO:0031515) |
| 0.5 | 7.6 | GO:0036038 | MKS complex(GO:0036038) |
| 0.4 | 8.1 | GO:0043256 | basal lamina(GO:0005605) laminin complex(GO:0043256) |
| 0.3 | 4.4 | GO:0044545 | NSL complex(GO:0044545) |
| 0.3 | 5.3 | GO:0033202 | Ino80 complex(GO:0031011) DNA helicase complex(GO:0033202) |
| 0.3 | 7.2 | GO:0030867 | rough endoplasmic reticulum membrane(GO:0030867) |
| 0.3 | 8.9 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.3 | 3.5 | GO:0034751 | aryl hydrocarbon receptor complex(GO:0034751) |
| 0.2 | 1.0 | GO:0060171 | stereocilium membrane(GO:0060171) |
| 0.2 | 20.9 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.2 | 2.9 | GO:0030015 | CCR4-NOT core complex(GO:0030015) |
| 0.2 | 7.5 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.2 | 10.4 | GO:0030426 | growth cone(GO:0030426) |
| 0.2 | 7.9 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 0.2 | 4.3 | GO:0035102 | PRC1 complex(GO:0035102) |
| 0.2 | 4.4 | GO:0071565 | nBAF complex(GO:0071565) |
| 0.1 | 40.3 | GO:0005874 | microtubule(GO:0005874) |
| 0.1 | 4.2 | GO:0035861 | site of double-strand break(GO:0035861) |
| 0.1 | 5.1 | GO:0030175 | filopodium(GO:0030175) |
| 0.1 | 2.4 | GO:0032809 | neuronal cell body membrane(GO:0032809) cell body membrane(GO:0044298) |
| 0.1 | 1.8 | GO:0000243 | commitment complex(GO:0000243) |
| 0.1 | 0.9 | GO:0035517 | PR-DUB complex(GO:0035517) |
| 0.1 | 2.8 | GO:0033116 | endoplasmic reticulum-Golgi intermediate compartment membrane(GO:0033116) |
| 0.1 | 3.7 | GO:0017053 | transcriptional repressor complex(GO:0017053) |
| 0.1 | 1.3 | GO:0030285 | integral component of synaptic vesicle membrane(GO:0030285) intrinsic component of synaptic vesicle membrane(GO:0098563) |
| 0.1 | 1.8 | GO:0009925 | basal plasma membrane(GO:0009925) |
| 0.1 | 1.4 | GO:0043209 | myelin sheath(GO:0043209) |
| 0.1 | 5.4 | GO:0034707 | chloride channel complex(GO:0034707) |
| 0.1 | 5.8 | GO:0043025 | neuronal cell body(GO:0043025) |
| 0.1 | 1.3 | GO:0070776 | H3 histone acetyltransferase complex(GO:0070775) MOZ/MORF histone acetyltransferase complex(GO:0070776) |
| 0.1 | 7.2 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.1 | 15.0 | GO:0036477 | somatodendritic compartment(GO:0036477) |
| 0.1 | 4.0 | GO:0031902 | late endosome membrane(GO:0031902) |
| 0.1 | 0.3 | GO:0071005 | U2-type precatalytic spliceosome(GO:0071005) |
| 0.1 | 2.0 | GO:0072686 | mitotic spindle(GO:0072686) |
| 0.1 | 0.7 | GO:0031209 | SCAR complex(GO:0031209) |
| 0.0 | 6.5 | GO:0005681 | spliceosomal complex(GO:0005681) |
| 0.0 | 4.8 | GO:0016607 | nuclear speck(GO:0016607) |
| 0.0 | 0.7 | GO:0031680 | G-protein beta/gamma-subunit complex(GO:0031680) |
| 0.0 | 17.5 | GO:0005768 | endosome(GO:0005768) |
| 0.0 | 1.9 | GO:0005905 | clathrin-coated pit(GO:0005905) |
| 0.0 | 2.8 | GO:0005802 | trans-Golgi network(GO:0005802) |
| 0.0 | 1.0 | GO:0030496 | midbody(GO:0030496) |
| 0.0 | 0.6 | GO:0005669 | transcription factor TFIID complex(GO:0005669) |
| 0.0 | 7.6 | GO:0000151 | ubiquitin ligase complex(GO:0000151) |
| 0.0 | 0.4 | GO:0045178 | basal part of cell(GO:0045178) |
| 0.0 | 0.6 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.0 | 14.4 | GO:0005789 | endoplasmic reticulum membrane(GO:0005789) |
| 0.0 | 0.4 | GO:1990907 | beta-catenin-TCF complex(GO:1990907) |
| 0.0 | 1.1 | GO:0099512 | supramolecular fiber(GO:0099512) polymeric cytoskeletal fiber(GO:0099513) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.1 | 9.4 | GO:0061598 | molybdopterin adenylyltransferase activity(GO:0061598) molybdopterin molybdotransferase activity(GO:0061599) |
| 2.8 | 41.6 | GO:0005504 | fatty acid binding(GO:0005504) |
| 1.9 | 5.8 | GO:0031701 | angiotensin receptor binding(GO:0031701) |
| 1.6 | 7.8 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 1.6 | 14.0 | GO:0001733 | galactosylceramide sulfotransferase activity(GO:0001733) galactose 3-O-sulfotransferase activity(GO:0050694) |
| 1.4 | 7.2 | GO:0015193 | L-proline transmembrane transporter activity(GO:0015193) |
| 1.4 | 12.2 | GO:0004865 | protein serine/threonine phosphatase inhibitor activity(GO:0004865) |
| 1.3 | 51.0 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 1.3 | 7.6 | GO:0008158 | hedgehog receptor activity(GO:0008158) |
| 1.0 | 10.0 | GO:0004705 | JUN kinase activity(GO:0004705) SAP kinase activity(GO:0016909) |
| 0.9 | 5.5 | GO:0004966 | galanin receptor activity(GO:0004966) |
| 0.8 | 2.4 | GO:0005334 | dopamine transmembrane transporter activity(GO:0005329) dopamine:sodium symporter activity(GO:0005330) norepinephrine transmembrane transporter activity(GO:0005333) norepinephrine:sodium symporter activity(GO:0005334) |
| 0.7 | 5.8 | GO:0001972 | retinoic acid binding(GO:0001972) |
| 0.7 | 4.6 | GO:0016316 | phosphatidylinositol-3,4-bisphosphate 4-phosphatase activity(GO:0016316) |
| 0.6 | 5.4 | GO:0072320 | volume-sensitive chloride channel activity(GO:0072320) |
| 0.6 | 2.9 | GO:0008486 | endopolyphosphatase activity(GO:0000298) diphosphoinositol-polyphosphate diphosphatase activity(GO:0008486) bis(5'-adenosyl)-hexaphosphatase activity(GO:0034431) bis(5'-adenosyl)-pentaphosphatase activity(GO:0034432) |
| 0.5 | 7.9 | GO:0004971 | AMPA glutamate receptor activity(GO:0004971) |
| 0.5 | 2.0 | GO:0051377 | mannose-ethanolamine phosphotransferase activity(GO:0051377) |
| 0.5 | 30.0 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.5 | 2.5 | GO:0016429 | tRNA (adenine-N1-)-methyltransferase activity(GO:0016429) |
| 0.5 | 2.9 | GO:0042974 | estrogen receptor binding(GO:0030331) retinoic acid receptor binding(GO:0042974) |
| 0.4 | 6.0 | GO:0005024 | transforming growth factor beta-activated receptor activity(GO:0005024) |
| 0.4 | 8.0 | GO:0048185 | activin binding(GO:0048185) |
| 0.4 | 1.8 | GO:0038132 | neuregulin receptor activity(GO:0038131) neuregulin binding(GO:0038132) |
| 0.4 | 1.8 | GO:0035256 | G-protein coupled glutamate receptor binding(GO:0035256) |
| 0.4 | 3.5 | GO:0017091 | AU-rich element binding(GO:0017091) mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.3 | 7.6 | GO:0050321 | tau-protein kinase activity(GO:0050321) |
| 0.3 | 5.1 | GO:0008093 | cytoskeletal adaptor activity(GO:0008093) |
| 0.3 | 9.3 | GO:0004089 | carbonate dehydratase activity(GO:0004089) |
| 0.3 | 10.8 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.3 | 0.9 | GO:0004736 | pyruvate carboxylase activity(GO:0004736) |
| 0.3 | 4.1 | GO:0047631 | ADP-ribose diphosphatase activity(GO:0047631) |
| 0.3 | 1.7 | GO:0005384 | manganese ion transmembrane transporter activity(GO:0005384) |
| 0.3 | 3.5 | GO:0003886 | DNA (cytosine-5-)-methyltransferase activity(GO:0003886) |
| 0.3 | 2.7 | GO:0005391 | sodium:potassium-exchanging ATPase activity(GO:0005391) |
| 0.2 | 9.7 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
| 0.2 | 1.0 | GO:0071889 | 14-3-3 protein binding(GO:0071889) |
| 0.2 | 10.8 | GO:0019707 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.2 | 9.8 | GO:0048020 | CCR chemokine receptor binding(GO:0048020) |
| 0.2 | 4.4 | GO:0046875 | ephrin receptor binding(GO:0046875) |
| 0.2 | 0.9 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.1 | 4.9 | GO:0035257 | nuclear hormone receptor binding(GO:0035257) |
| 0.1 | 1.7 | GO:1990404 | protein ADP-ribosylase activity(GO:1990404) |
| 0.1 | 12.6 | GO:0005544 | calcium-dependent phospholipid binding(GO:0005544) |
| 0.1 | 1.4 | GO:0097602 | cullin family protein binding(GO:0097602) |
| 0.1 | 0.9 | GO:0005161 | platelet-derived growth factor receptor binding(GO:0005161) |
| 0.1 | 1.6 | GO:0032454 | histone demethylase activity (H3-K9 specific)(GO:0032454) |
| 0.1 | 2.6 | GO:0004707 | MAP kinase activity(GO:0004707) |
| 0.1 | 1.3 | GO:0043994 | H3 histone acetyltransferase activity(GO:0010484) histone acetyltransferase activity (H3-K23 specific)(GO:0043994) |
| 0.1 | 1.4 | GO:0019911 | structural constituent of myelin sheath(GO:0019911) |
| 0.1 | 3.8 | GO:0070491 | repressing transcription factor binding(GO:0070491) |
| 0.1 | 8.9 | GO:0019888 | protein phosphatase regulator activity(GO:0019888) |
| 0.1 | 1.8 | GO:0048038 | quinone binding(GO:0048038) |
| 0.1 | 0.6 | GO:0000995 | transcription factor activity, core RNA polymerase III binding(GO:0000995) |
| 0.1 | 0.8 | GO:0016868 | intramolecular transferase activity, phosphotransferases(GO:0016868) |
| 0.1 | 0.3 | GO:0030620 | U2 snRNA binding(GO:0030620) |
| 0.1 | 4.1 | GO:0030170 | pyridoxal phosphate binding(GO:0030170) |
| 0.1 | 9.8 | GO:0003774 | motor activity(GO:0003774) |
| 0.1 | 19.2 | GO:0008017 | microtubule binding(GO:0008017) |
| 0.0 | 0.7 | GO:0000340 | RNA 7-methylguanosine cap binding(GO:0000340) |
| 0.0 | 8.7 | GO:0042393 | histone binding(GO:0042393) |
| 0.0 | 6.7 | GO:0005516 | calmodulin binding(GO:0005516) |
| 0.0 | 0.2 | GO:0016842 | amidine-lyase activity(GO:0016842) |
| 0.0 | 0.5 | GO:0005220 | inositol 1,4,5-trisphosphate-sensitive calcium-release channel activity(GO:0005220) |
| 0.0 | 0.7 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.0 | 5.9 | GO:0004497 | monooxygenase activity(GO:0004497) |
| 0.0 | 0.4 | GO:0097199 | cysteine-type endopeptidase activity involved in apoptotic signaling pathway(GO:0097199) |
| 0.0 | 14.1 | GO:0008047 | enzyme activator activity(GO:0008047) |
| 0.0 | 1.9 | GO:0005044 | scavenger receptor activity(GO:0005044) |
| 0.0 | 2.0 | GO:0008138 | protein serine/threonine phosphatase activity(GO:0004722) protein tyrosine/serine/threonine phosphatase activity(GO:0008138) |
| 0.0 | 1.3 | GO:0005484 | SNAP receptor activity(GO:0005484) |
| 0.0 | 0.5 | GO:0004143 | diacylglycerol kinase activity(GO:0004143) |
| 0.0 | 5.6 | GO:0044822 | mRNA binding(GO:0003729) poly(A) RNA binding(GO:0044822) |
| 0.0 | 7.7 | GO:0046983 | protein dimerization activity(GO:0046983) |
| 0.0 | 1.9 | GO:0060090 | binding, bridging(GO:0060090) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 10.0 | ST GRANULE CELL SURVIVAL PATHWAY | Granule Cell Survival Pathway is a specific case of more general PAC1 Receptor Pathway. |
| 0.4 | 5.9 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.4 | 2.0 | PID RET PATHWAY | Signaling events regulated by Ret tyrosine kinase |
| 0.2 | 4.9 | PID RETINOIC ACID PATHWAY | Retinoic acid receptors-mediated signaling |
| 0.2 | 2.6 | ST GA12 PATHWAY | G alpha 12 Pathway |
| 0.2 | 5.6 | PID EPHB FWD PATHWAY | EPHB forward signaling |
| 0.2 | 5.9 | PID ECADHERIN NASCENT AJ PATHWAY | E-cadherin signaling in the nascent adherens junction |
| 0.2 | 3.5 | PID RHOA PATHWAY | RhoA signaling pathway |
| 0.1 | 2.4 | PID ALPHA SYNUCLEIN PATHWAY | Alpha-synuclein signaling |
| 0.1 | 6.5 | PID ERA GENOMIC PATHWAY | Validated nuclear estrogen receptor alpha network |
| 0.1 | 3.0 | PID ATM PATHWAY | ATM pathway |
| 0.1 | 0.6 | PID VEGFR1 PATHWAY | VEGFR1 specific signals |
| 0.1 | 2.0 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.0 | 2.4 | PID AVB3 INTEGRIN PATHWAY | Integrins in angiogenesis |
| 0.0 | 5.5 | NABA ECM REGULATORS | Genes encoding enzymes and their regulators involved in the remodeling of the extracellular matrix |
| 0.0 | 0.5 | ST WNT CA2 CYCLIC GMP PATHWAY | Wnt/Ca2+/cyclic GMP signaling. |
| 0.0 | 1.7 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.0 | 0.7 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.0 | 0.4 | PID HIV NEF PATHWAY | HIV-1 Nef: Negative effector of Fas and TNF-alpha |
| 0.0 | 0.6 | PID MYC REPRESS PATHWAY | Validated targets of C-MYC transcriptional repression |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.0 | 31.7 | REACTOME POST CHAPERONIN TUBULIN FOLDING PATHWAY | Genes involved in Post-chaperonin tubulin folding pathway |
| 1.0 | 10.0 | REACTOME ACTIVATION OF THE AP1 FAMILY OF TRANSCRIPTION FACTORS | Genes involved in Activation of the AP-1 family of transcription factors |
| 0.9 | 10.8 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GIP | Genes involved in Synthesis, Secretion, and Inactivation of Glucose-dependent Insulinotropic Polypeptide (GIP) |
| 0.7 | 5.9 | REACTOME RAP1 SIGNALLING | Genes involved in Rap1 signalling |
| 0.6 | 9.6 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.4 | 2.6 | REACTOME ERKS ARE INACTIVATED | Genes involved in ERKs are inactivated |
| 0.3 | 4.6 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 0.2 | 3.5 | REACTOME SEMA4D INDUCED CELL MIGRATION AND GROWTH CONE COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.2 | 1.6 | REACTOME ACYL CHAIN REMODELLING OF PG | Genes involved in Acyl chain remodelling of PG |
| 0.2 | 1.8 | REACTOME GAMMA CARBOXYLATION TRANSPORT AND AMINO TERMINAL CLEAVAGE OF PROTEINS | Genes involved in Gamma-carboxylation, transport, and amino-terminal cleavage of proteins |
| 0.2 | 3.7 | REACTOME DOWNREGULATION OF SMAD2 3 SMAD4 TRANSCRIPTIONAL ACTIVITY | Genes involved in Downregulation of SMAD2/3:SMAD4 transcriptional activity |
| 0.2 | 3.5 | REACTOME PROCESSING OF INTRONLESS PRE MRNAS | Genes involved in Processing of Intronless Pre-mRNAs |
| 0.2 | 4.5 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.1 | 3.0 | REACTOME HOMOLOGOUS RECOMBINATION REPAIR OF REPLICATION INDEPENDENT DOUBLE STRAND BREAKS | Genes involved in Homologous recombination repair of replication-independent double-strand breaks |
| 0.1 | 2.0 | REACTOME SYNTHESIS OF GLYCOSYLPHOSPHATIDYLINOSITOL GPI | Genes involved in Synthesis of glycosylphosphatidylinositol (GPI) |
| 0.1 | 3.9 | REACTOME G ALPHA Z SIGNALLING EVENTS | Genes involved in G alpha (z) signalling events |
| 0.1 | 1.3 | REACTOME PROTEOLYTIC CLEAVAGE OF SNARE COMPLEX PROTEINS | Genes involved in Proteolytic cleavage of SNARE complex proteins |
| 0.1 | 1.2 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.1 | 0.5 | REACTOME ELEVATION OF CYTOSOLIC CA2 LEVELS | Genes involved in Elevation of cytosolic Ca2+ levels |
| 0.1 | 0.8 | REACTOME SYNTHESIS OF SUBSTRATES IN N GLYCAN BIOSYTHESIS | Genes involved in Synthesis of substrates in N-glycan biosythesis |
| 0.0 | 0.6 | REACTOME NEPHRIN INTERACTIONS | Genes involved in Nephrin interactions |
| 0.0 | 1.9 | REACTOME G ALPHA1213 SIGNALLING EVENTS | Genes involved in G alpha (12/13) signalling events |
| 0.0 | 1.7 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.0 | 2.6 | REACTOME FACTORS INVOLVED IN MEGAKARYOCYTE DEVELOPMENT AND PLATELET PRODUCTION | Genes involved in Factors involved in megakaryocyte development and platelet production |
| 0.0 | 0.6 | REACTOME RNA POL III TRANSCRIPTION INITIATION FROM TYPE 2 PROMOTER | Genes involved in RNA Polymerase III Transcription Initiation From Type 2 Promoter |