Project
PRJDB7713: Age associated transcriptome analysis in 5 tissues of zebrafish
Navigation
Downloads
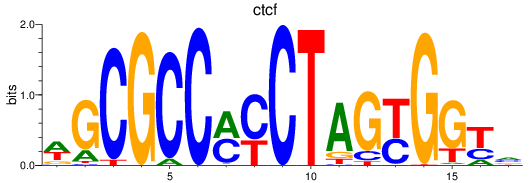
Results for ctcf
Z-value: 1.28
Transcription factors associated with ctcf
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
ctcf
|
ENSDARG00000056621 | CCCTC-binding factor (zinc finger protein) |
|
ctcf
|
ENSDARG00000113285 | CCCTC-binding factor (zinc finger protein) |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| ctcf | dr11_v1_chr18_+_22285992_22286029 | 0.42 | 2.5e-05 | Click! |
Activity profile of ctcf motif
Sorted Z-values of ctcf motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr22_-_2886937 | 10.83 |
ENSDART00000063533
|
aqp12
|
aquaporin 12 |
| chr15_+_47525073 | 10.22 |
ENSDART00000067583
|
sidt2
|
SID1 transmembrane family, member 2 |
| chr4_-_17409533 | 9.24 |
ENSDART00000011943
|
pah
|
phenylalanine hydroxylase |
| chr20_-_30931139 | 8.18 |
ENSDART00000006778
ENSDART00000146376 |
acat2
|
acetyl-CoA acetyltransferase 2 |
| chr16_+_11558868 | 8.00 |
ENSDART00000112497
ENSDART00000180445 |
zgc:198329
|
zgc:198329 |
| chr9_+_21148318 | 7.46 |
ENSDART00000035428
|
hao2
|
hydroxyacid oxidase 2 (long chain) |
| chr6_-_42388608 | 7.06 |
ENSDART00000049425
|
sec61a1l
|
Sec61 translocon alpha 1 subunit, like |
| chr21_-_19918286 | 6.72 |
ENSDART00000180816
|
ppp1r3b
|
protein phosphatase 1, regulatory subunit 3B |
| chr13_+_1369003 | 6.31 |
ENSDART00000146082
|
ifngr1
|
interferon gamma receptor 1 |
| chr8_-_38317914 | 6.31 |
ENSDART00000125920
|
pdlim2
|
PDZ and LIM domain 2 (mystique) |
| chr6_-_27139396 | 6.11 |
ENSDART00000055848
|
zgc:103559
|
zgc:103559 |
| chr24_-_8887825 | 6.09 |
ENSDART00000066781
|
elovl2
|
ELOVL fatty acid elongase 2 |
| chr17_+_30894431 | 5.93 |
ENSDART00000127996
|
degs2
|
delta(4)-desaturase, sphingolipid 2 |
| chr2_-_37883256 | 5.86 |
ENSDART00000035685
|
hbl4
|
hexose-binding lectin 4 |
| chr9_-_48736388 | 5.76 |
ENSDART00000022074
|
dhrs9
|
dehydrogenase/reductase (SDR family) member 9 |
| chr9_-_34269066 | 5.41 |
ENSDART00000059955
|
ildr1b
|
immunoglobulin-like domain containing receptor 1b |
| chr16_+_11585576 | 5.37 |
ENSDART00000172967
|
si:dkey-11o1.6
|
si:dkey-11o1.6 |
| chr3_+_14641962 | 5.37 |
ENSDART00000091070
|
zgc:158403
|
zgc:158403 |
| chr23_-_13875252 | 5.10 |
ENSDART00000104834
ENSDART00000193807 |
g6pd
|
glucose-6-phosphate dehydrogenase |
| chr22_+_5135884 | 5.08 |
ENSDART00000141276
|
mydgf
|
myeloid-derived growth factor |
| chr13_-_42306348 | 5.03 |
ENSDART00000003706
|
kmo
|
kynurenine 3-monooxygenase |
| chr12_+_31783066 | 4.76 |
ENSDART00000105584
|
lrrc59
|
leucine rich repeat containing 59 |
| chr12_-_4220713 | 4.67 |
ENSDART00000129427
|
vkorc1
|
vitamin K epoxide reductase complex, subunit 1 |
| chr19_+_20785724 | 4.29 |
ENSDART00000038942
|
adnp2b
|
ADNP homeobox 2b |
| chr16_+_11573471 | 4.24 |
ENSDART00000145858
|
si:dkey-11o1.2
|
si:dkey-11o1.2 |
| chr21_+_15713097 | 4.08 |
ENSDART00000015841
|
gstt1b
|
glutathione S-transferase theta 1b |
| chr9_-_48700806 | 4.01 |
ENSDART00000026210
|
rdh1
|
retinol dehydrogenase 1 |
| chr2_+_22659787 | 3.90 |
ENSDART00000043956
|
zgc:161973
|
zgc:161973 |
| chr2_+_11028923 | 3.87 |
ENSDART00000076725
|
acot11a
|
acyl-CoA thioesterase 11a |
| chr19_-_33274978 | 3.83 |
ENSDART00000020301
ENSDART00000114714 |
fam92a1
|
family with sequence similarity 92, member A1 |
| chr3_+_25849560 | 3.73 |
ENSDART00000007119
|
mfsd6l
|
major facilitator superfamily domain containing 6-like |
| chr6_-_37749711 | 3.72 |
ENSDART00000078324
|
nipa1
|
non imprinted in Prader-Willi/Angelman syndrome 1 |
| chr2_-_42492445 | 3.71 |
ENSDART00000139929
|
esyt2a
|
extended synaptotagmin-like protein 2a |
| chr9_+_21843915 | 3.66 |
ENSDART00000101977
|
kctd12.1
|
potassium channel tetramerisation domain containing 12.1 |
| chr5_-_31773208 | 3.61 |
ENSDART00000137556
ENSDART00000122066 |
fam102ab
|
family with sequence similarity 102, member Ab |
| chr24_-_17400143 | 3.50 |
ENSDART00000134947
|
cul1b
|
cullin 1b |
| chr21_+_6114709 | 3.43 |
ENSDART00000065858
|
fpgs
|
folylpolyglutamate synthase |
| chr1_-_46663997 | 3.37 |
ENSDART00000134450
|
ebpl
|
emopamil binding protein-like |
| chr6_-_35046735 | 3.34 |
ENSDART00000143649
|
uap1
|
UDP-N-acetylglucosamine pyrophosphorylase 1 |
| chr12_-_7639120 | 3.34 |
ENSDART00000126712
ENSDART00000126219 |
ccdc6b
|
coiled-coil domain containing 6b |
| chr22_-_20403194 | 3.29 |
ENSDART00000010048
|
map2k2a
|
mitogen-activated protein kinase kinase 2a |
| chr9_-_29643628 | 3.28 |
ENSDART00000101177
|
spryd7b
|
SPRY domain containing 7b |
| chr2_+_3823813 | 3.28 |
ENSDART00000103596
ENSDART00000161880 ENSDART00000185408 |
npc1
|
Niemann-Pick disease, type C1 |
| chr13_+_35765317 | 3.23 |
ENSDART00000100156
ENSDART00000167650 |
agpat4
|
1-acylglycerol-3-phosphate O-acyltransferase 4 (lysophosphatidic acid acyltransferase, delta) |
| chr21_+_6114305 | 3.19 |
ENSDART00000141607
|
fpgs
|
folylpolyglutamate synthase |
| chr9_-_11263228 | 3.18 |
ENSDART00000113847
|
chpfa
|
chondroitin polymerizing factor a |
| chr8_+_47099033 | 3.14 |
ENSDART00000142979
|
arhgef16
|
Rho guanine nucleotide exchange factor (GEF) 16 |
| chr19_+_18903533 | 3.11 |
ENSDART00000157523
ENSDART00000166562 |
slc39a7
|
solute carrier family 39 (zinc transporter), member 7 |
| chr7_+_52154215 | 3.10 |
ENSDART00000098712
|
TMEM208
|
zgc:77041 |
| chr13_-_46429220 | 3.07 |
ENSDART00000149125
ENSDART00000098269 ENSDART00000150061 ENSDART00000080916 |
fgfr2
|
fibroblast growth factor receptor 2 |
| chr1_+_19849168 | 3.06 |
ENSDART00000111454
|
si:dkey-82j4.2
|
si:dkey-82j4.2 |
| chr19_-_3886991 | 3.01 |
ENSDART00000162085
|
thrap3b
|
thyroid hormone receptor associated protein 3b |
| chr3_-_45777226 | 3.00 |
ENSDART00000192849
|
h3f3b.1
|
H3 histone, family 3B.1 |
| chr12_-_17492852 | 3.00 |
ENSDART00000012421
ENSDART00000138766 ENSDART00000130735 |
minpp1b
|
multiple inositol-polyphosphate phosphatase 1b |
| chr21_+_43702016 | 2.93 |
ENSDART00000017176
|
dkc1
|
dyskeratosis congenita 1, dyskerin |
| chr9_-_41507712 | 2.87 |
ENSDART00000135821
|
mfsd6b
|
major facilitator superfamily domain containing 6b |
| chr10_+_2842923 | 2.86 |
ENSDART00000181895
|
ykt6
|
YKT6 v-SNARE homolog (S. cerevisiae) |
| chr11_-_41853874 | 2.84 |
ENSDART00000002556
|
mrto4
|
MRT4 homolog, ribosome maturation factor |
| chr6_+_45934682 | 2.84 |
ENSDART00000103489
|
cenps
|
centromere protein S |
| chr7_+_34236238 | 2.84 |
ENSDART00000052474
|
tipin
|
timeless interacting protein |
| chr2_+_23701613 | 2.80 |
ENSDART00000047073
|
oxsr1a
|
oxidative stress responsive 1a |
| chr24_-_13349802 | 2.79 |
ENSDART00000164729
|
terf1
|
telomeric repeat binding factor (NIMA-interacting) 1 |
| chr1_-_24255149 | 2.77 |
ENSDART00000146960
|
lrba
|
LPS-responsive vesicle trafficking, beach and anchor containing |
| chr13_-_35765028 | 2.77 |
ENSDART00000157391
|
erlec1
|
endoplasmic reticulum lectin 1 |
| chr16_-_31686602 | 2.76 |
ENSDART00000170357
|
c1s
|
complement component 1, s subcomponent |
| chr17_+_33158350 | 2.72 |
ENSDART00000104476
|
snx9a
|
sorting nexin 9a |
| chr12_+_19976400 | 2.72 |
ENSDART00000153177
|
mkl2a
|
MKL/myocardin-like 2a |
| chr13_-_33822550 | 2.69 |
ENSDART00000143703
|
flrt3
|
fibronectin leucine rich transmembrane 3 |
| chr6_+_60125033 | 2.67 |
ENSDART00000148557
ENSDART00000008224 |
aurka
|
aurora kinase A |
| chr1_+_40613297 | 2.66 |
ENSDART00000040798
ENSDART00000168067 ENSDART00000130490 |
naa15b
|
N(alpha)-acetyltransferase 15, NatA auxiliary subunit b |
| chr19_-_1947403 | 2.64 |
ENSDART00000113951
ENSDART00000151293 ENSDART00000134074 |
znrf2a
|
zinc and ring finger 2a |
| chr12_-_31726748 | 2.63 |
ENSDART00000153174
|
srsf2a
|
serine/arginine-rich splicing factor 2a |
| chr21_-_23331619 | 2.60 |
ENSDART00000007806
|
zbtb16a
|
zinc finger and BTB domain containing 16a |
| chr1_+_55752593 | 2.59 |
ENSDART00000108838
ENSDART00000134770 |
tecrb
|
trans-2,3-enoyl-CoA reductase b |
| chr24_-_38644937 | 2.58 |
ENSDART00000170194
|
slc6a16b
|
solute carrier family 6, member 16b |
| chr24_-_13349464 | 2.57 |
ENSDART00000134482
ENSDART00000139212 |
terf1
|
telomeric repeat binding factor (NIMA-interacting) 1 |
| chr19_+_24068223 | 2.54 |
ENSDART00000141351
ENSDART00000100420 |
pex11b
|
peroxisomal biogenesis factor 11 beta |
| chr8_-_34427364 | 2.42 |
ENSDART00000112854
ENSDART00000161282 ENSDART00000113230 |
gapvd1
|
GTPase activating protein and VPS9 domains 1 |
| chr5_+_58687541 | 2.41 |
ENSDART00000083015
ENSDART00000181902 |
ccdc84
|
coiled-coil domain containing 84 |
| chr13_+_9819194 | 2.41 |
ENSDART00000091595
|
fam45a
|
family with sequence similarity 45, member A |
| chr11_+_22623284 | 2.40 |
ENSDART00000111850
|
si:ch211-86h15.1
|
si:ch211-86h15.1 |
| chr14_+_31473866 | 2.40 |
ENSDART00000173088
|
ccdc160
|
coiled-coil domain containing 160 |
| chr21_-_32462856 | 2.37 |
ENSDART00000147318
|
zgc:123105
|
zgc:123105 |
| chr19_-_3896514 | 2.36 |
ENSDART00000161346
|
si:ch73-281i18.3
|
si:ch73-281i18.3 |
| chr5_-_1869982 | 2.32 |
ENSDART00000055878
|
rcl1
|
RNA terminal phosphate cyclase-like 1 |
| chr9_+_29985010 | 2.32 |
ENSDART00000020743
|
cmss1
|
cms1 ribosomal small subunit homolog (yeast) |
| chr2_-_20666920 | 2.29 |
ENSDART00000143437
ENSDART00000114546 ENSDART00000136113 ENSDART00000179247 |
dusp12
|
dual specificity phosphatase 12 |
| chr9_+_22929675 | 2.29 |
ENSDART00000061299
|
tsn
|
translin |
| chr7_+_13995792 | 2.28 |
ENSDART00000091470
|
furina
|
furin (paired basic amino acid cleaving enzyme) a |
| chr10_-_40826657 | 2.26 |
ENSDART00000076304
|
pcna
|
proliferating cell nuclear antigen |
| chr5_+_61941610 | 2.23 |
ENSDART00000168808
|
si:dkeyp-117b8.4
|
si:dkeyp-117b8.4 |
| chr5_-_26764298 | 2.21 |
ENSDART00000189373
|
rnf181
|
ring finger protein 181 |
| chr18_+_35229115 | 2.18 |
ENSDART00000129624
ENSDART00000184596 |
tbrg1
|
transforming growth factor beta regulator 1 |
| chr3_+_31093455 | 2.15 |
ENSDART00000153074
|
si:dkey-66i24.9
|
si:dkey-66i24.9 |
| chr16_+_11623956 | 2.14 |
ENSDART00000137788
|
cxcr3.1
|
chemokine (C-X-C motif) receptor 3, tandem duplicate 1 |
| chr19_-_3925801 | 2.12 |
ENSDART00000129570
ENSDART00000163138 |
si:ch73-281i18.6
|
si:ch73-281i18.6 |
| chr8_+_26007988 | 2.10 |
ENSDART00000193948
ENSDART00000058100 |
xpc
|
xeroderma pigmentosum, complementation group C |
| chr2_+_57149002 | 2.07 |
ENSDART00000168497
|
eef2b
|
eukaryotic translation elongation factor 2b |
| chr7_-_71384391 | 2.06 |
ENSDART00000112841
|
ccdc149a
|
coiled-coil domain containing 149a |
| chr24_-_17400472 | 2.06 |
ENSDART00000024691
|
cul1b
|
cullin 1b |
| chr16_-_32672883 | 2.03 |
ENSDART00000124515
ENSDART00000190920 ENSDART00000188776 |
pnisr
|
PNN-interacting serine/arginine-rich protein |
| chr5_+_41496490 | 2.02 |
ENSDART00000039369
|
fancg
|
Fanconi anemia, complementation group G |
| chr5_-_58832332 | 2.01 |
ENSDART00000161230
|
arhgef12b
|
Rho guanine nucleotide exchange factor (GEF) 12b |
| chr6_+_47846366 | 2.01 |
ENSDART00000064842
|
padi2
|
peptidyl arginine deiminase, type II |
| chr5_+_20319519 | 1.93 |
ENSDART00000004217
|
coro1ca
|
coronin, actin binding protein, 1Ca |
| chr16_-_14587332 | 1.91 |
ENSDART00000012479
|
dscc1
|
DNA replication and sister chromatid cohesion 1 |
| chr4_-_75158035 | 1.88 |
ENSDART00000174353
|
CABZ01066312.1
|
|
| chr3_-_33417826 | 1.87 |
ENSDART00000084284
|
abi3a
|
ABI family, member 3a |
| chr3_+_36671585 | 1.87 |
ENSDART00000159033
|
nde1
|
nudE neurodevelopment protein 1 |
| chr1_-_56948694 | 1.86 |
ENSDART00000152597
|
si:ch211-1f22.1
|
si:ch211-1f22.1 |
| chr5_+_57320113 | 1.85 |
ENSDART00000036331
|
atp6v1g1
|
ATPase H+ transporting V1 subunit G1 |
| chr10_-_35186310 | 1.81 |
ENSDART00000127805
|
pom121
|
POM121 transmembrane nucleoporin |
| chr19_+_1688727 | 1.80 |
ENSDART00000115136
ENSDART00000166744 |
dennd3a
|
DENN/MADD domain containing 3a |
| chr9_-_11655031 | 1.75 |
ENSDART00000044314
|
itgav
|
integrin, alpha V |
| chr12_-_18898413 | 1.74 |
ENSDART00000181281
ENSDART00000121866 |
desi1b
|
desumoylating isopeptidase 1b |
| chr2_+_54482603 | 1.74 |
ENSDART00000130977
ENSDART00000183090 |
mtcl1
|
microtubule crosslinking factor 1 |
| chr8_+_42917515 | 1.74 |
ENSDART00000021715
|
slc23a2
|
solute carrier family 23 (ascorbic acid transporter), member 2 |
| chr5_-_9948497 | 1.74 |
ENSDART00000183196
|
tor4ab
|
torsin family 4, member Ab |
| chr8_-_11202378 | 1.72 |
ENSDART00000147817
ENSDART00000174039 |
fam208b
|
family with sequence similarity 208, member B |
| chr15_-_43978141 | 1.71 |
ENSDART00000041249
|
chordc1a
|
cysteine and histidine-rich domain (CHORD) containing 1a |
| chr16_+_40340222 | 1.71 |
ENSDART00000190631
|
mettl6
|
methyltransferase like 6 |
| chr8_+_19621511 | 1.69 |
ENSDART00000017128
|
slc35a3a
|
solute carrier family 35 (UDP-N-acetylglucosamine (UDP-GlcNAc) transporter), member A3a |
| chr25_+_8921425 | 1.69 |
ENSDART00000128591
|
accs
|
1-aminocyclopropane-1-carboxylate synthase homolog (Arabidopsis)(non-functional) |
| chr8_-_17184482 | 1.66 |
ENSDART00000025803
|
pola2
|
polymerase (DNA directed), alpha 2 |
| chr1_+_45958904 | 1.65 |
ENSDART00000108528
|
arhgef7b
|
Rho guanine nucleotide exchange factor (GEF) 7b |
| chr9_-_12594759 | 1.64 |
ENSDART00000021266
|
tra2b
|
transformer 2 beta homolog |
| chr9_+_35832294 | 1.62 |
ENSDART00000130549
ENSDART00000122169 |
si:dkey-67c22.2
|
si:dkey-67c22.2 |
| chr2_+_43204919 | 1.61 |
ENSDART00000160077
ENSDART00000018729 ENSDART00000129134 ENSDART00000056402 |
pard3ab
|
par-3 family cell polarity regulator alpha, b |
| chr22_+_20135443 | 1.60 |
ENSDART00000143641
|
eef2a.1
|
eukaryotic translation elongation factor 2a, tandem duplicate 1 |
| chr14_-_31715373 | 1.60 |
ENSDART00000127303
ENSDART00000173274 ENSDART00000173435 ENSDART00000172876 ENSDART00000173036 |
map7d3
|
MAP7 domain containing 3 |
| chr1_-_45577980 | 1.59 |
ENSDART00000160961
|
atf7ip
|
activating transcription factor 7 interacting protein |
| chr22_-_718615 | 1.56 |
ENSDART00000149320
|
arl8a
|
ADP-ribosylation factor-like 8A |
| chr12_-_999762 | 1.55 |
ENSDART00000127003
ENSDART00000084076 ENSDART00000152425 |
mettl9
|
methyltransferase like 9 |
| chr17_-_25303486 | 1.53 |
ENSDART00000162235
|
ppie
|
peptidylprolyl isomerase E (cyclophilin E) |
| chr16_+_40340523 | 1.53 |
ENSDART00000102571
|
mettl6
|
methyltransferase like 6 |
| chr3_-_37148594 | 1.51 |
ENSDART00000140855
|
mlx
|
MLX, MAX dimerization protein |
| chr15_+_17441734 | 1.51 |
ENSDART00000153729
|
snx19b
|
sorting nexin 19b |
| chr15_+_41815703 | 1.51 |
ENSDART00000059508
|
pxylp1
|
2-phosphoxylose phosphatase 1 |
| chr2_-_40135942 | 1.50 |
ENSDART00000176951
ENSDART00000098632 ENSDART00000148563 ENSDART00000149895 |
epha4a
|
eph receptor A4a |
| chr14_-_45595711 | 1.50 |
ENSDART00000074038
|
scyl1
|
SCY1-like, kinase-like 1 |
| chr25_-_35963158 | 1.49 |
ENSDART00000153612
|
snx20
|
sorting nexin 20 |
| chr3_+_22057109 | 1.48 |
ENSDART00000153709
|
kansl1b
|
KAT8 regulatory NSL complex subunit 1b |
| chr15_-_26887028 | 1.48 |
ENSDART00000156292
|
si:dkey-243i1.1
|
si:dkey-243i1.1 |
| chr16_+_23799622 | 1.47 |
ENSDART00000046922
|
rab13
|
RAB13, member RAS oncogene family |
| chr15_+_6786757 | 1.46 |
ENSDART00000013278
|
sirt2
|
sirtuin 2 (silent mating type information regulation 2, homolog) 2 (S. cerevisiae) |
| chr8_+_36983559 | 1.44 |
ENSDART00000127053
|
kdm5c
|
lysine (K)-specific demethylase 5C |
| chr16_+_11834516 | 1.44 |
ENSDART00000146611
|
cxcr3.3
|
chemokine (C-X-C motif) receptor 3, tandem duplicate 3 |
| chr5_+_31036911 | 1.44 |
ENSDART00000141463
ENSDART00000162121 |
zzef1
|
zinc finger, ZZ-type with EF hand domain 1 |
| chr7_+_29080684 | 1.43 |
ENSDART00000173709
ENSDART00000173576 |
acd
|
ACD, shelterin complex subunit and telomerase recruitment factor |
| chr3_+_37790351 | 1.42 |
ENSDART00000151506
|
si:dkey-260c8.8
|
si:dkey-260c8.8 |
| chr13_+_13033424 | 1.41 |
ENSDART00000159441
|
letm1
|
leucine zipper-EF-hand containing transmembrane protein 1 |
| chr5_+_54497475 | 1.40 |
ENSDART00000158149
ENSDART00000163968 ENSDART00000160446 |
tmem203
|
transmembrane protein 203 |
| chr21_+_17005737 | 1.39 |
ENSDART00000101246
|
vps29
|
vacuolar protein sorting 29 homolog (S. cerevisiae) |
| chr8_+_19621731 | 1.38 |
ENSDART00000144667
|
slc35a3a
|
solute carrier family 35 (UDP-N-acetylglucosamine (UDP-GlcNAc) transporter), member A3a |
| chr18_-_14677936 | 1.38 |
ENSDART00000111995
|
si:dkey-238o13.4
|
si:dkey-238o13.4 |
| chr10_+_11355841 | 1.37 |
ENSDART00000193067
ENSDART00000064215 |
cops4
|
COP9 constitutive photomorphogenic homolog subunit 4 (Arabidopsis) |
| chr16_+_28881235 | 1.35 |
ENSDART00000146525
|
chtopb
|
chromatin target of PRMT1b |
| chr19_-_3906369 | 1.35 |
ENSDART00000160299
ENSDART00000169205 |
si:ch73-281i18.4
|
si:ch73-281i18.4 |
| chr2_-_26596794 | 1.33 |
ENSDART00000134685
ENSDART00000056787 |
zgc:113691
|
zgc:113691 |
| chr5_+_55984270 | 1.29 |
ENSDART00000047358
ENSDART00000138191 |
fkbp6
|
FK506 binding protein 6 |
| chr8_+_23703680 | 1.28 |
ENSDART00000141099
ENSDART00000135394 |
ppardb
|
peroxisome proliferator-activated receptor delta b |
| chr10_+_17681074 | 1.27 |
ENSDART00000057500
|
drg1
|
developmentally regulated GTP binding protein 1 |
| chr4_+_12931763 | 1.26 |
ENSDART00000016382
|
wif1
|
wnt inhibitory factor 1 |
| chr7_+_66634167 | 1.26 |
ENSDART00000027616
|
eif4g2a
|
eukaryotic translation initiation factor 4, gamma 2a |
| chr6_+_48041759 | 1.26 |
ENSDART00000140086
|
si:dkey-92f12.2
|
si:dkey-92f12.2 |
| chr3_-_30248277 | 1.25 |
ENSDART00000133516
ENSDART00000077029 |
emc10
|
ER membrane protein complex subunit 10 |
| chr13_+_13033837 | 1.21 |
ENSDART00000079558
|
letm1
|
leucine zipper-EF-hand containing transmembrane protein 1 |
| chr9_+_38043337 | 1.21 |
ENSDART00000022574
|
stam2
|
signal transducing adaptor molecule (SH3 domain and ITAM motif) 2 |
| chr24_+_26329018 | 1.20 |
ENSDART00000145752
|
mynn
|
myoneurin |
| chr24_-_10897511 | 1.20 |
ENSDART00000145593
ENSDART00000102484 ENSDART00000066784 |
fam49bb
|
family with sequence similarity 49, member Bb |
| chr15_+_31911989 | 1.20 |
ENSDART00000111472
|
brca2
|
breast cancer 2, early onset |
| chr1_+_12049229 | 1.18 |
ENSDART00000103403
ENSDART00000137697 |
saraf
|
store-operated calcium entry-associated regulatory factor |
| chr21_+_26539157 | 1.18 |
ENSDART00000021121
|
stx5al
|
syntaxin 5A, like |
| chr16_-_8132742 | 1.16 |
ENSDART00000104323
|
snrka
|
SNF related kinase a |
| chr13_-_14908999 | 1.15 |
ENSDART00000112771
|
ap5s1
|
adaptor-related protein complex 5, sigma 1 subunit |
| chr25_+_5972690 | 1.14 |
ENSDART00000067517
|
si:ch211-11i22.4
|
si:ch211-11i22.4 |
| chr12_-_5418340 | 1.14 |
ENSDART00000028043
|
noc3l
|
NOC3-like DNA replication regulator |
| chr8_+_25900049 | 1.13 |
ENSDART00000124300
ENSDART00000127618 ENSDART00000024009 |
rhoab
|
ras homolog gene family, member Ab |
| chr13_-_36761379 | 1.13 |
ENSDART00000131534
ENSDART00000029824 |
map4k5
|
mitogen-activated protein kinase kinase kinase kinase 5 |
| chr19_+_20274944 | 1.11 |
ENSDART00000151237
|
oxnad1
|
oxidoreductase NAD-binding domain containing 1 |
| chr15_-_4967302 | 1.11 |
ENSDART00000101992
|
lipt2
|
lipoyl(octanoyl) transferase 2 |
| chr1_+_17527342 | 1.06 |
ENSDART00000139702
ENSDART00000140076 ENSDART00000005593 |
casp3a
|
caspase 3, apoptosis-related cysteine peptidase a |
| chr5_+_35955209 | 1.05 |
ENSDART00000074630
|
zgc:103697
|
zgc:103697 |
| chr8_+_37749263 | 1.04 |
ENSDART00000108556
ENSDART00000147942 |
npm2a
|
nucleophosmin/nucleoplasmin, 2a |
| chr21_+_25120546 | 1.04 |
ENSDART00000149507
|
ddx10
|
DEAD (Asp-Glu-Ala-Asp) box polypeptide 10 |
| chr24_+_26328787 | 1.03 |
ENSDART00000003884
|
mynn
|
myoneurin |
| chr12_+_46967789 | 1.02 |
ENSDART00000114866
|
oat
|
ornithine aminotransferase |
| chr5_+_32076109 | 1.02 |
ENSDART00000051357
ENSDART00000144510 |
zmat5
|
zinc finger, matrin-type 5 |
| chr19_-_3916075 | 1.01 |
ENSDART00000161799
|
si:ch73-281i18.7
|
si:ch73-281i18.7 |
| chr12_+_8569685 | 0.98 |
ENSDART00000031676
|
nrbf2b
|
nuclear receptor binding factor 2b |
| chr1_+_26676758 | 0.97 |
ENSDART00000152299
|
si:dkey-25o16.4
|
si:dkey-25o16.4 |
| chr18_+_41542542 | 0.96 |
ENSDART00000087445
|
tsen34
|
TSEN34 tRNA splicing endonuclease subunit |
| chr2_+_42724404 | 0.96 |
ENSDART00000075392
|
basp1
|
brain abundant, membrane attached signal protein 1 |
| chr19_+_14113886 | 0.96 |
ENSDART00000169343
|
kdf1b
|
keratinocyte differentiation factor 1b |
| chr4_-_18512550 | 0.95 |
ENSDART00000045639
|
rint1
|
RAD50 interactor 1 |
| chr18_-_40753583 | 0.94 |
ENSDART00000026767
|
akt2
|
v-akt murine thymoma viral oncogene homolog 2 |
| chr25_+_24250247 | 0.93 |
ENSDART00000064646
|
tmem86a
|
transmembrane protein 86A |
| chr5_+_55934129 | 0.93 |
ENSDART00000050969
|
tmem150ab
|
transmembrane protein 150Ab |
| chr5_-_32396929 | 0.93 |
ENSDART00000023977
|
fbxw2
|
F-box and WD repeat domain containing 2 |
| chr21_+_25533908 | 0.92 |
ENSDART00000185225
|
nlrc3l1
|
NLR family, CARD domain containing 3-like 1 |
Network of associatons between targets according to the STRING database.
First level regulatory network of ctcf
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.2 | 6.7 | GO:0005981 | regulation of glycogen catabolic process(GO:0005981) |
| 2.0 | 6.1 | GO:0006145 | allantoin catabolic process(GO:0000256) purine nucleobase catabolic process(GO:0006145) |
| 1.7 | 6.6 | GO:0046900 | tetrahydrofolylpolyglutamate metabolic process(GO:0046900) |
| 1.6 | 4.7 | GO:0042373 | vitamin K metabolic process(GO:0042373) |
| 1.0 | 7.1 | GO:0031204 | posttranslational protein targeting to membrane, translocation(GO:0031204) |
| 1.0 | 5.0 | GO:0043420 | anthranilate metabolic process(GO:0043420) |
| 1.0 | 2.9 | GO:0031120 | snRNA pseudouridine synthesis(GO:0031120) |
| 0.9 | 2.7 | GO:0000212 | meiotic spindle organization(GO:0000212) |
| 0.9 | 2.6 | GO:0034214 | protein hexamerization(GO:0034214) |
| 0.8 | 9.2 | GO:1902221 | L-phenylalanine metabolic process(GO:0006558) L-phenylalanine catabolic process(GO:0006559) erythrose 4-phosphate/phosphoenolpyruvate family amino acid metabolic process(GO:1902221) erythrose 4-phosphate/phosphoenolpyruvate family amino acid catabolic process(GO:1902222) |
| 0.8 | 9.8 | GO:0042572 | retinol metabolic process(GO:0042572) |
| 0.7 | 2.9 | GO:0019884 | antigen processing and presentation of exogenous peptide antigen(GO:0002478) antigen processing and presentation of exogenous antigen(GO:0019884) antigen processing and presentation of exogenous peptide antigen via MHC class I(GO:0042590) |
| 0.7 | 3.3 | GO:0030219 | megakaryocyte differentiation(GO:0030219) |
| 0.6 | 1.9 | GO:0034088 | maintenance of sister chromatid cohesion(GO:0034086) maintenance of mitotic sister chromatid cohesion(GO:0034088) |
| 0.6 | 1.7 | GO:0015882 | L-ascorbic acid transport(GO:0015882) L-ascorbic acid metabolic process(GO:0019852) |
| 0.6 | 2.8 | GO:0043111 | replication fork arrest(GO:0043111) |
| 0.6 | 2.3 | GO:0006272 | leading strand elongation(GO:0006272) |
| 0.6 | 1.7 | GO:0071896 | protein localization to adherens junction(GO:0071896) |
| 0.5 | 1.5 | GO:0043388 | negative regulation of gliogenesis(GO:0014014) positive regulation of DNA binding(GO:0043388) negative regulation of reactive oxygen species metabolic process(GO:2000378) |
| 0.5 | 1.4 | GO:0051973 | positive regulation of telomerase activity(GO:0051973) |
| 0.5 | 1.4 | GO:2000374 | oxygen metabolic process(GO:0072592) regulation of oxygen metabolic process(GO:2000374) positive regulation of oxygen metabolic process(GO:2000376) |
| 0.4 | 3.1 | GO:0033278 | cell proliferation in midbrain(GO:0033278) |
| 0.4 | 5.1 | GO:0006098 | pentose-phosphate shunt(GO:0006098) |
| 0.4 | 5.9 | GO:0034311 | diol metabolic process(GO:0034311) |
| 0.4 | 2.5 | GO:0016559 | peroxisome fission(GO:0016559) |
| 0.4 | 1.6 | GO:1902414 | establishment of centrosome localization(GO:0051660) protein localization to cell junction(GO:1902414) |
| 0.4 | 2.0 | GO:0019240 | citrulline biosynthetic process(GO:0019240) |
| 0.4 | 2.7 | GO:0017196 | N-terminal peptidyl-methionine acetylation(GO:0017196) |
| 0.4 | 1.5 | GO:0048210 | Golgi vesicle fusion to target membrane(GO:0048210) |
| 0.4 | 1.1 | GO:0071236 | cellular response to antibiotic(GO:0071236) cellular response to toxic substance(GO:0097237) |
| 0.3 | 1.0 | GO:0051988 | regulation of attachment of spindle microtubules to kinetochore(GO:0051988) |
| 0.3 | 1.0 | GO:0000379 | tRNA-type intron splice site recognition and cleavage(GO:0000379) |
| 0.3 | 1.9 | GO:0007100 | mitotic centrosome separation(GO:0007100) centrosome separation(GO:0051299) |
| 0.3 | 5.5 | GO:0048922 | posterior lateral line neuromast deposition(GO:0048922) |
| 0.3 | 1.5 | GO:0042762 | regulation of sulfur metabolic process(GO:0042762) |
| 0.3 | 6.3 | GO:0098508 | endothelial to hematopoietic transition(GO:0098508) |
| 0.3 | 5.4 | GO:0008156 | negative regulation of DNA replication(GO:0008156) |
| 0.3 | 1.8 | GO:0090245 | axis elongation involved in somitogenesis(GO:0090245) |
| 0.3 | 6.1 | GO:0019368 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.3 | 3.3 | GO:0006048 | UDP-N-acetylglucosamine biosynthetic process(GO:0006048) |
| 0.2 | 5.1 | GO:0001938 | positive regulation of endothelial cell proliferation(GO:0001938) |
| 0.2 | 0.9 | GO:0046324 | regulation of glucose import(GO:0046324) |
| 0.2 | 1.4 | GO:0045721 | negative regulation of gluconeogenesis(GO:0045721) |
| 0.2 | 0.7 | GO:0000350 | generation of catalytic spliceosome for second transesterification step(GO:0000350) |
| 0.2 | 3.7 | GO:0015693 | magnesium ion transport(GO:0015693) |
| 0.2 | 10.8 | GO:0071391 | cellular response to estrogen stimulus(GO:0071391) |
| 0.2 | 2.6 | GO:0042761 | very long-chain fatty acid biosynthetic process(GO:0042761) |
| 0.2 | 1.5 | GO:0003404 | optic vesicle morphogenesis(GO:0003404) |
| 0.2 | 5.6 | GO:0038061 | NIK/NF-kappaB signaling(GO:0038061) |
| 0.2 | 3.1 | GO:0015781 | pyrimidine nucleotide-sugar transport(GO:0015781) pyrimidine nucleotide-sugar transmembrane transport(GO:0090481) |
| 0.2 | 0.6 | GO:0071514 | genetic imprinting(GO:0071514) |
| 0.2 | 8.2 | GO:0006635 | fatty acid beta-oxidation(GO:0006635) |
| 0.2 | 2.6 | GO:0032481 | positive regulation of type I interferon production(GO:0032481) |
| 0.2 | 3.2 | GO:0030206 | chondroitin sulfate biosynthetic process(GO:0030206) |
| 0.2 | 1.0 | GO:0010482 | epidermal cell division(GO:0010481) regulation of epidermal cell division(GO:0010482) |
| 0.2 | 1.3 | GO:0045050 | protein insertion into ER membrane by stop-transfer membrane-anchor sequence(GO:0045050) |
| 0.2 | 2.8 | GO:0051382 | kinetochore assembly(GO:0051382) |
| 0.2 | 0.9 | GO:2000389 | neutrophil extravasation(GO:0072672) regulation of neutrophil extravasation(GO:2000389) positive regulation of neutrophil extravasation(GO:2000391) |
| 0.2 | 1.2 | GO:2001256 | regulation of store-operated calcium entry(GO:2001256) |
| 0.2 | 2.3 | GO:0000478 | endonucleolytic cleavage involved in rRNA processing(GO:0000478) endonucleolytic cleavage of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000479) |
| 0.2 | 2.3 | GO:0060971 | embryonic heart tube left/right pattern formation(GO:0060971) |
| 0.2 | 1.1 | GO:1902766 | skeletal muscle satellite cell migration(GO:1902766) |
| 0.2 | 1.0 | GO:0060628 | regulation of ER to Golgi vesicle-mediated transport(GO:0060628) |
| 0.2 | 4.3 | GO:0043249 | erythrocyte maturation(GO:0043249) |
| 0.1 | 0.6 | GO:0042796 | snRNA transcription from RNA polymerase III promoter(GO:0042796) |
| 0.1 | 3.1 | GO:0071577 | zinc II ion transmembrane transport(GO:0071577) |
| 0.1 | 1.3 | GO:0010887 | negative regulation of cholesterol storage(GO:0010887) |
| 0.1 | 3.7 | GO:0021986 | habenula development(GO:0021986) |
| 0.1 | 2.3 | GO:0048026 | positive regulation of mRNA splicing, via spliceosome(GO:0048026) |
| 0.1 | 1.6 | GO:0090309 | methylation-dependent chromatin silencing(GO:0006346) positive regulation of chromatin silencing(GO:0031937) regulation of methylation-dependent chromatin silencing(GO:0090308) positive regulation of methylation-dependent chromatin silencing(GO:0090309) |
| 0.1 | 1.8 | GO:0032483 | regulation of Rab protein signal transduction(GO:0032483) |
| 0.1 | 2.8 | GO:0000027 | ribosomal large subunit assembly(GO:0000027) |
| 0.1 | 0.7 | GO:0045898 | regulation of RNA polymerase II transcriptional preinitiation complex assembly(GO:0045898) positive regulation of RNA polymerase II transcriptional preinitiation complex assembly(GO:0045899) |
| 0.1 | 12.5 | GO:0050658 | nucleic acid transport(GO:0050657) RNA transport(GO:0050658) establishment of RNA localization(GO:0051236) |
| 0.1 | 1.0 | GO:0006561 | proline biosynthetic process(GO:0006561) L-proline biosynthetic process(GO:0055129) |
| 0.1 | 4.8 | GO:0006749 | glutathione metabolic process(GO:0006749) |
| 0.1 | 1.2 | GO:0007530 | sex determination(GO:0007530) |
| 0.1 | 4.8 | GO:0051057 | positive regulation of Ras protein signal transduction(GO:0046579) positive regulation of small GTPase mediated signal transduction(GO:0051057) |
| 0.1 | 0.9 | GO:0097345 | mitochondrial outer membrane permeabilization(GO:0097345) |
| 0.1 | 0.7 | GO:0090179 | establishment of planar polarity of embryonic epithelium(GO:0042249) establishment of planar polarity involved in neural tube closure(GO:0090177) regulation of establishment of planar polarity involved in neural tube closure(GO:0090178) planar cell polarity pathway involved in neural tube closure(GO:0090179) |
| 0.1 | 2.1 | GO:0006298 | mismatch repair(GO:0006298) |
| 0.1 | 2.8 | GO:0006270 | DNA replication initiation(GO:0006270) |
| 0.1 | 0.9 | GO:0070262 | peptidyl-serine dephosphorylation(GO:0070262) |
| 0.1 | 0.4 | GO:0045048 | protein insertion into ER membrane(GO:0045048) |
| 0.1 | 0.9 | GO:0016926 | protein desumoylation(GO:0016926) |
| 0.1 | 1.9 | GO:0090148 | membrane fission(GO:0090148) |
| 0.1 | 3.2 | GO:0051298 | centrosome duplication(GO:0051298) |
| 0.1 | 1.1 | GO:0035493 | SNARE complex assembly(GO:0035493) |
| 0.1 | 1.3 | GO:0034587 | piRNA metabolic process(GO:0034587) |
| 0.1 | 0.8 | GO:0006614 | SRP-dependent cotranslational protein targeting to membrane(GO:0006614) |
| 0.1 | 0.8 | GO:0090503 | RNA phosphodiester bond hydrolysis, exonucleolytic(GO:0090503) |
| 0.1 | 2.0 | GO:0036297 | interstrand cross-link repair(GO:0036297) |
| 0.1 | 0.6 | GO:0015886 | heme transport(GO:0015886) iron coordination entity transport(GO:1901678) |
| 0.1 | 0.7 | GO:0021694 | cerebellar Purkinje cell layer formation(GO:0021694) cerebellar Purkinje cell differentiation(GO:0021702) |
| 0.1 | 3.2 | GO:0000281 | mitotic cytokinesis(GO:0000281) |
| 0.1 | 3.4 | GO:0006414 | translational elongation(GO:0006414) |
| 0.1 | 1.4 | GO:1990126 | retrograde transport, endosome to plasma membrane(GO:1990126) |
| 0.1 | 0.7 | GO:0050688 | regulation of defense response to virus(GO:0050688) |
| 0.1 | 1.2 | GO:0043153 | entrainment of circadian clock by photoperiod(GO:0043153) |
| 0.1 | 1.6 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.1 | 1.1 | GO:0007064 | mitotic sister chromatid cohesion(GO:0007064) |
| 0.0 | 2.8 | GO:0031638 | zymogen activation(GO:0031638) |
| 0.0 | 4.6 | GO:0048024 | regulation of mRNA splicing, via spliceosome(GO:0048024) |
| 0.0 | 0.9 | GO:0048920 | posterior lateral line neuromast primordium migration(GO:0048920) |
| 0.0 | 0.8 | GO:2001235 | positive regulation of apoptotic signaling pathway(GO:2001235) |
| 0.0 | 1.3 | GO:0048794 | swim bladder development(GO:0048794) |
| 0.0 | 0.4 | GO:0035283 | fourth ventricle development(GO:0021592) central nervous system segmentation(GO:0035283) brain segmentation(GO:0035284) |
| 0.0 | 1.3 | GO:0006998 | nuclear envelope organization(GO:0006998) |
| 0.0 | 1.2 | GO:0034112 | positive regulation of homotypic cell-cell adhesion(GO:0034112) positive regulation of T cell activation(GO:0050870) positive regulation of leukocyte cell-cell adhesion(GO:1903039) |
| 0.0 | 2.1 | GO:0002685 | regulation of leukocyte migration(GO:0002685) |
| 0.0 | 2.9 | GO:0035383 | acyl-CoA metabolic process(GO:0006637) thioester metabolic process(GO:0035383) |
| 0.0 | 1.4 | GO:0043524 | negative regulation of neuron apoptotic process(GO:0043524) |
| 0.0 | 3.4 | GO:0016125 | sterol metabolic process(GO:0016125) |
| 0.0 | 0.8 | GO:0007520 | myoblast fusion(GO:0007520) |
| 0.0 | 0.9 | GO:0016180 | snRNA processing(GO:0016180) |
| 0.0 | 1.9 | GO:0030865 | cortical cytoskeleton organization(GO:0030865) |
| 0.0 | 0.4 | GO:0006278 | RNA-dependent DNA biosynthetic process(GO:0006278) telomere maintenance via telomerase(GO:0007004) |
| 0.0 | 0.8 | GO:0003094 | glomerular filtration(GO:0003094) |
| 0.0 | 3.1 | GO:0090630 | activation of GTPase activity(GO:0090630) |
| 0.0 | 0.4 | GO:0010821 | regulation of mitochondrion organization(GO:0010821) |
| 0.0 | 0.5 | GO:0001522 | pseudouridine synthesis(GO:0001522) |
| 0.0 | 0.5 | GO:0006884 | cell volume homeostasis(GO:0006884) |
| 0.0 | 1.9 | GO:1902600 | hydrogen ion transmembrane transport(GO:1902600) |
| 0.0 | 0.6 | GO:0009062 | fatty acid catabolic process(GO:0009062) |
| 0.0 | 1.0 | GO:0000187 | activation of MAPK activity(GO:0000187) |
| 0.0 | 0.7 | GO:0034314 | Arp2/3 complex-mediated actin nucleation(GO:0034314) |
| 0.0 | 4.9 | GO:0008380 | RNA splicing(GO:0008380) |
| 0.0 | 1.3 | GO:0002181 | cytoplasmic translation(GO:0002181) |
| 0.0 | 1.4 | GO:0019722 | calcium-mediated signaling(GO:0019722) |
| 0.0 | 0.0 | GO:0015074 | DNA integration(GO:0015074) |
| 0.0 | 0.9 | GO:0034968 | histone lysine methylation(GO:0034968) |
| 0.0 | 0.1 | GO:0042276 | error-prone translesion synthesis(GO:0042276) |
| 0.0 | 3.0 | GO:0032259 | methylation(GO:0032259) |
| 0.0 | 1.4 | GO:0006888 | ER to Golgi vesicle-mediated transport(GO:0006888) |
| 0.0 | 1.7 | GO:0070646 | protein modification by small protein removal(GO:0070646) |
| 0.0 | 0.8 | GO:0006413 | translational initiation(GO:0006413) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 7.1 | GO:0005784 | Sec61 translocon complex(GO:0005784) |
| 1.0 | 3.1 | GO:0031390 | Ctf18 RFC-like complex(GO:0031390) |
| 0.7 | 2.9 | GO:0031429 | box H/ACA snoRNP complex(GO:0031429) box H/ACA RNP complex(GO:0072588) |
| 0.6 | 2.8 | GO:0031298 | replication fork protection complex(GO:0031298) |
| 0.5 | 2.7 | GO:0031415 | NatA complex(GO:0031415) |
| 0.5 | 1.5 | GO:0043220 | compact myelin(GO:0043218) Schmidt-Lanterman incisure(GO:0043220) |
| 0.4 | 2.7 | GO:0032133 | chromosome passenger complex(GO:0032133) |
| 0.4 | 4.9 | GO:0043240 | Fanconi anaemia nuclear complex(GO:0043240) |
| 0.4 | 6.7 | GO:0000164 | protein phosphatase type 1 complex(GO:0000164) |
| 0.3 | 1.7 | GO:0005658 | alpha DNA polymerase:primase complex(GO:0005658) |
| 0.3 | 2.5 | GO:0005779 | integral component of peroxisomal membrane(GO:0005779) intrinsic component of peroxisomal membrane(GO:0031231) |
| 0.3 | 6.8 | GO:0000783 | telomere cap complex(GO:0000782) nuclear telomere cap complex(GO:0000783) nuclear chromosome, telomeric region(GO:0000784) |
| 0.2 | 1.0 | GO:0000214 | tRNA-intron endonuclease complex(GO:0000214) |
| 0.2 | 0.6 | GO:0019185 | snRNA-activating protein complex(GO:0019185) |
| 0.2 | 0.5 | GO:0034709 | methylosome(GO:0034709) |
| 0.2 | 2.1 | GO:0000109 | nucleotide-excision repair complex(GO:0000109) |
| 0.2 | 3.8 | GO:0005771 | multivesicular body(GO:0005771) |
| 0.1 | 1.5 | GO:0032593 | insulin-responsive compartment(GO:0032593) |
| 0.1 | 6.3 | GO:0031941 | filamentous actin(GO:0031941) |
| 0.1 | 1.4 | GO:0030904 | retromer complex(GO:0030904) |
| 0.1 | 1.4 | GO:0008180 | COP9 signalosome(GO:0008180) |
| 0.1 | 1.0 | GO:0035032 | phosphatidylinositol 3-kinase complex, class III(GO:0035032) |
| 0.1 | 0.6 | GO:0097433 | dense body(GO:0097433) |
| 0.1 | 0.8 | GO:0005786 | signal recognition particle, endoplasmic reticulum targeting(GO:0005786) signal recognition particle(GO:0048500) |
| 0.1 | 1.3 | GO:0072546 | ER membrane protein complex(GO:0072546) |
| 0.1 | 2.8 | GO:0030687 | preribosome, large subunit precursor(GO:0030687) |
| 0.1 | 0.7 | GO:0031464 | Cul4A-RING E3 ubiquitin ligase complex(GO:0031464) |
| 0.1 | 1.5 | GO:0044545 | NSL complex(GO:0044545) |
| 0.1 | 5.5 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.1 | 5.8 | GO:0031228 | integral component of Golgi membrane(GO:0030173) intrinsic component of Golgi membrane(GO:0031228) |
| 0.1 | 1.9 | GO:0016471 | vacuolar proton-transporting V-type ATPase complex(GO:0016471) |
| 0.1 | 0.4 | GO:0070209 | ASTRA complex(GO:0070209) |
| 0.1 | 2.0 | GO:0005689 | U12-type spliceosomal complex(GO:0005689) |
| 0.1 | 0.7 | GO:0031597 | cytosolic proteasome complex(GO:0031597) |
| 0.1 | 9.2 | GO:0005741 | mitochondrial outer membrane(GO:0005741) |
| 0.1 | 11.0 | GO:0031227 | intrinsic component of endoplasmic reticulum membrane(GO:0031227) |
| 0.1 | 4.0 | GO:0016592 | mediator complex(GO:0016592) |
| 0.1 | 0.8 | GO:0000177 | cytoplasmic exosome (RNase complex)(GO:0000177) |
| 0.1 | 5.9 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.1 | 0.7 | GO:0005682 | U5 snRNP(GO:0005682) |
| 0.1 | 1.3 | GO:0034992 | microtubule organizing center attachment site(GO:0034992) LINC complex(GO:0034993) |
| 0.1 | 0.3 | GO:0005673 | transcription factor TFIIE complex(GO:0005673) |
| 0.1 | 0.4 | GO:0071818 | BAT3 complex(GO:0071818) |
| 0.1 | 1.3 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.1 | 0.5 | GO:0031080 | nuclear pore outer ring(GO:0031080) |
| 0.1 | 2.7 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.0 | 1.9 | GO:0097610 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.0 | 0.8 | GO:0071564 | npBAF complex(GO:0071564) |
| 0.0 | 3.7 | GO:0042734 | presynaptic membrane(GO:0042734) |
| 0.0 | 13.0 | GO:0000323 | lytic vacuole(GO:0000323) |
| 0.0 | 0.8 | GO:0070776 | H3 histone acetyltransferase complex(GO:0070775) MOZ/MORF histone acetyltransferase complex(GO:0070776) |
| 0.0 | 2.0 | GO:0048786 | presynaptic active zone(GO:0048786) |
| 0.0 | 2.5 | GO:0032580 | Golgi cisterna membrane(GO:0032580) |
| 0.0 | 1.9 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.0 | 0.9 | GO:0032039 | integrator complex(GO:0032039) |
| 0.0 | 0.7 | GO:0005885 | Arp2/3 protein complex(GO:0005885) |
| 0.0 | 1.8 | GO:0005643 | nuclear pore(GO:0005643) |
| 0.0 | 22.6 | GO:0005789 | endoplasmic reticulum membrane(GO:0005789) |
| 0.0 | 1.9 | GO:0030175 | filopodium(GO:0030175) |
| 0.0 | 0.2 | GO:0030914 | STAGA complex(GO:0030914) |
| 0.0 | 6.2 | GO:0030659 | cytoplasmic vesicle membrane(GO:0030659) |
| 0.0 | 0.2 | GO:0070876 | SOSS complex(GO:0070876) |
| 0.0 | 2.9 | GO:0000786 | nucleosome(GO:0000786) |
| 0.0 | 0.9 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.0 | 0.2 | GO:0089701 | U2AF(GO:0089701) |
| 0.0 | 1.0 | GO:0008305 | integrin complex(GO:0008305) |
| 0.0 | 1.9 | GO:0000776 | kinetochore(GO:0000776) |
| 0.0 | 1.2 | GO:0031201 | SNARE complex(GO:0031201) |
| 0.0 | 2.3 | GO:0005681 | spliceosomal complex(GO:0005681) |
| 0.0 | 3.7 | GO:0005938 | cell cortex(GO:0005938) |
| 0.0 | 4.4 | GO:0005730 | nucleolus(GO:0005730) |
| 0.0 | 5.2 | GO:0005783 | endoplasmic reticulum(GO:0005783) |
| 0.0 | 0.8 | GO:0008021 | synaptic vesicle(GO:0008021) |
| 0.0 | 0.8 | GO:0005814 | centriole(GO:0005814) |
| 0.0 | 1.1 | GO:0044297 | cell body(GO:0044297) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 2.7 | 8.2 | GO:0003985 | acetyl-CoA C-acetyltransferase activity(GO:0003985) |
| 1.8 | 5.4 | GO:0071532 | ankyrin repeat binding(GO:0071532) |
| 1.7 | 5.1 | GO:0004345 | glucose-6-phosphate dehydrogenase activity(GO:0004345) |
| 1.5 | 6.1 | GO:0050897 | cobalt ion binding(GO:0050897) |
| 1.2 | 9.8 | GO:0004745 | retinol dehydrogenase activity(GO:0004745) |
| 1.1 | 17.3 | GO:0022884 | macromolecule transmembrane transporter activity(GO:0022884) |
| 1.0 | 9.2 | GO:0016714 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced pteridine as one donor, and incorporation of one atom of oxygen(GO:0016714) |
| 0.7 | 3.0 | GO:0034416 | bisphosphoglycerate phosphatase activity(GO:0034416) |
| 0.7 | 3.3 | GO:0003977 | UDP-N-acetylglucosamine diphosphorylase activity(GO:0003977) |
| 0.7 | 2.6 | GO:0015369 | calcium:proton antiporter activity(GO:0015369) metal ion:proton antiporter activity(GO:0051139) |
| 0.5 | 3.8 | GO:0036042 | long-chain fatty acyl-CoA binding(GO:0036042) |
| 0.5 | 2.1 | GO:0016494 | C-X-C chemokine receptor activity(GO:0016494) |
| 0.5 | 5.0 | GO:0050664 | oxidoreductase activity, acting on NAD(P)H, oxygen as acceptor(GO:0050664) |
| 0.5 | 3.4 | GO:0016863 | intramolecular oxidoreductase activity, transposing C=C bonds(GO:0016863) |
| 0.5 | 1.4 | GO:0033819 | lipoyl(octanoyl) transferase activity(GO:0033819) |
| 0.4 | 6.7 | GO:2001069 | glycogen binding(GO:2001069) |
| 0.4 | 3.1 | GO:0004839 | ubiquitin activating enzyme activity(GO:0004839) |
| 0.4 | 2.7 | GO:0035173 | histone kinase activity(GO:0035173) |
| 0.4 | 1.5 | GO:0042903 | tubulin deacetylase activity(GO:0042903) |
| 0.4 | 3.2 | GO:0047238 | glucuronosyl-N-acetylgalactosaminyl-proteoglycan 4-beta-N-acetylgalactosaminyltransferase activity(GO:0047238) |
| 0.3 | 7.5 | GO:0010181 | FMN binding(GO:0010181) |
| 0.3 | 3.7 | GO:0008429 | phosphatidylethanolamine binding(GO:0008429) |
| 0.3 | 6.3 | GO:0051393 | muscle alpha-actinin binding(GO:0051371) alpha-actinin binding(GO:0051393) |
| 0.3 | 3.1 | GO:0005459 | UDP-galactose transmembrane transporter activity(GO:0005459) |
| 0.3 | 6.1 | GO:0102338 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.2 | 1.0 | GO:0030626 | U12 snRNA binding(GO:0030626) |
| 0.2 | 3.1 | GO:0005007 | fibroblast growth factor-activated receptor activity(GO:0005007) |
| 0.2 | 2.0 | GO:0016813 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in linear amidines(GO:0016813) |
| 0.2 | 4.7 | GO:0048038 | quinone binding(GO:0048038) |
| 0.2 | 6.6 | GO:0016881 | acid-amino acid ligase activity(GO:0016881) |
| 0.2 | 1.5 | GO:0035035 | histone acetyltransferase binding(GO:0035035) |
| 0.2 | 2.7 | GO:0004596 | peptide alpha-N-acetyltransferase activity(GO:0004596) |
| 0.2 | 2.8 | GO:0003841 | 1-acylglycerol-3-phosphate O-acyltransferase activity(GO:0003841) |
| 0.2 | 1.0 | GO:0016892 | endoribonuclease activity, producing 3'-phosphomonoesters(GO:0016892) |
| 0.2 | 1.1 | GO:0008349 | MAP kinase kinase kinase kinase activity(GO:0008349) |
| 0.2 | 3.7 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.2 | 3.4 | GO:0009982 | pseudouridine synthase activity(GO:0009982) |
| 0.2 | 1.3 | GO:0032977 | membrane insertase activity(GO:0032977) |
| 0.2 | 1.9 | GO:0033170 | DNA clamp loader activity(GO:0003689) protein-DNA loading ATPase activity(GO:0033170) |
| 0.2 | 0.8 | GO:0030942 | endoplasmic reticulum signal peptide binding(GO:0030942) |
| 0.1 | 1.4 | GO:0034647 | histone demethylase activity (H3-trimethyl-K4 specific)(GO:0034647) |
| 0.1 | 3.0 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.1 | 0.6 | GO:0098973 | structural constituent of postsynaptic actin cytoskeleton(GO:0098973) |
| 0.1 | 4.1 | GO:0004364 | glutathione transferase activity(GO:0004364) |
| 0.1 | 0.9 | GO:0016803 | ether hydrolase activity(GO:0016803) |
| 0.1 | 0.9 | GO:0070139 | ubiquitin-like protein-specific endopeptidase activity(GO:0070137) SUMO-specific endopeptidase activity(GO:0070139) |
| 0.1 | 0.4 | GO:0033829 | O-fucosylpeptide 3-beta-N-acetylglucosaminyltransferase activity(GO:0033829) |
| 0.1 | 3.1 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.1 | 1.3 | GO:0043495 | protein anchor(GO:0043495) |
| 0.1 | 1.6 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 0.1 | 1.8 | GO:0008139 | nuclear localization sequence binding(GO:0008139) |
| 0.1 | 0.6 | GO:0015232 | heme transporter activity(GO:0015232) |
| 0.1 | 4.6 | GO:0004521 | endoribonuclease activity(GO:0004521) |
| 0.1 | 3.7 | GO:0043022 | ribosome binding(GO:0043022) |
| 0.1 | 5.6 | GO:0031625 | ubiquitin protein ligase binding(GO:0031625) |
| 0.1 | 6.1 | GO:0001227 | transcriptional repressor activity, RNA polymerase II transcription regulatory region sequence-specific binding(GO:0001227) |
| 0.1 | 7.7 | GO:0004896 | cytokine receptor activity(GO:0004896) |
| 0.1 | 2.2 | GO:0003684 | damaged DNA binding(GO:0003684) |
| 0.1 | 1.0 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.1 | 1.8 | GO:0042162 | telomeric DNA binding(GO:0042162) |
| 0.1 | 1.1 | GO:0097153 | cysteine-type endopeptidase activity involved in apoptotic process(GO:0097153) |
| 0.0 | 0.9 | GO:0002039 | p53 binding(GO:0002039) |
| 0.0 | 2.7 | GO:0030170 | pyridoxal phosphate binding(GO:0030170) |
| 0.0 | 0.7 | GO:0016812 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in cyclic amides(GO:0016812) |
| 0.0 | 2.6 | GO:0016627 | oxidoreductase activity, acting on the CH-CH group of donors(GO:0016627) |
| 0.0 | 1.3 | GO:0003755 | peptidyl-prolyl cis-trans isomerase activity(GO:0003755) |
| 0.0 | 0.8 | GO:0001540 | beta-amyloid binding(GO:0001540) |
| 0.0 | 6.6 | GO:0019901 | protein kinase binding(GO:0019901) |
| 0.0 | 0.1 | GO:0004743 | pyruvate kinase activity(GO:0004743) |
| 0.0 | 3.3 | GO:0005319 | lipid transporter activity(GO:0005319) |
| 0.0 | 0.2 | GO:0030628 | pre-mRNA 3'-splice site binding(GO:0030628) |
| 0.0 | 0.8 | GO:0000175 | 3'-5'-exoribonuclease activity(GO:0000175) |
| 0.0 | 0.5 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.0 | 0.8 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.0 | 7.5 | GO:0005085 | guanyl-nucleotide exchange factor activity(GO:0005085) |
| 0.0 | 0.2 | GO:0051864 | histone demethylase activity (H3-K36 specific)(GO:0051864) |
| 0.0 | 4.0 | GO:0016705 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen(GO:0016705) |
| 0.0 | 1.2 | GO:0005484 | SNAP receptor activity(GO:0005484) |
| 0.0 | 0.7 | GO:0005109 | frizzled binding(GO:0005109) |
| 0.0 | 3.4 | GO:0035091 | phosphatidylinositol binding(GO:0035091) |
| 0.0 | 0.8 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.0 | 5.0 | GO:0003682 | chromatin binding(GO:0003682) |
| 0.0 | 1.9 | GO:0016820 | P-P-bond-hydrolysis-driven transmembrane transporter activity(GO:0015405) hydrolase activity, acting on acid anhydrides, catalyzing transmembrane movement of substances(GO:0016820) ATPase activity, coupled to transmembrane movement of substances(GO:0042626) ATPase activity, coupled to movement of substances(GO:0043492) |
| 0.0 | 1.0 | GO:0003724 | RNA helicase activity(GO:0003724) |
| 0.0 | 3.0 | GO:0008168 | methyltransferase activity(GO:0008168) |
| 0.0 | 0.9 | GO:0019888 | protein phosphatase regulator activity(GO:0019888) |
| 0.0 | 4.6 | GO:0003712 | transcription factor activity, transcription factor binding(GO:0000989) transcription cofactor activity(GO:0003712) |
| 0.0 | 0.8 | GO:0003743 | translation initiation factor activity(GO:0003743) |
| 0.0 | 0.7 | GO:0019902 | phosphatase binding(GO:0019902) |
| 0.0 | 0.4 | GO:0031624 | ubiquitin conjugating enzyme binding(GO:0031624) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.3 | 6.3 | ST INTERFERON GAMMA PATHWAY | Interferon gamma pathway. |
| 0.2 | 4.3 | PID BARD1 PATHWAY | BARD1 signaling events |
| 0.2 | 1.8 | PID INTEGRIN5 PATHWAY | Beta5 beta6 beta7 and beta8 integrin cell surface interactions |
| 0.1 | 9.6 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.1 | 2.7 | PID AURORA A PATHWAY | Aurora A signaling |
| 0.1 | 4.0 | PID ATR PATHWAY | ATR signaling pathway |
| 0.1 | 1.5 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.1 | 0.8 | PID CIRCADIAN PATHWAY | Circadian rhythm pathway |
| 0.1 | 2.2 | ST WNT BETA CATENIN PATHWAY | Wnt/beta-catenin Pathway |
| 0.1 | 1.2 | PID IL2 1PATHWAY | IL2-mediated signaling events |
| 0.1 | 3.1 | PID FGF PATHWAY | FGF signaling pathway |
| 0.0 | 0.9 | PID IL8 CXCR2 PATHWAY | IL8- and CXCR2-mediated signaling events |
| 0.0 | 0.8 | PID VEGFR1 2 PATHWAY | Signaling events mediated by VEGFR1 and VEGFR2 |
| 0.0 | 0.7 | PID RAC1 PATHWAY | RAC1 signaling pathway |
| 0.0 | 0.8 | PID RHOA REG PATHWAY | Regulation of RhoA activity |
| 0.0 | 0.8 | PID TNF PATHWAY | TNF receptor signaling pathway |
| 0.0 | 1.0 | WNT SIGNALING | Genes related to Wnt-mediated signal transduction |
| 0.0 | 0.8 | PID REG GR PATHWAY | Glucocorticoid receptor regulatory network |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.9 | 6.3 | REACTOME REGULATION OF IFNG SIGNALING | Genes involved in Regulation of IFNG signaling |
| 0.8 | 6.1 | REACTOME SYNTHESIS OF VERY LONG CHAIN FATTY ACYL COAS | Genes involved in Synthesis of very long-chain fatty acyl-CoAs |
| 0.5 | 4.7 | REACTOME GAMMA CARBOXYLATION TRANSPORT AND AMINO TERMINAL CLEAVAGE OF PROTEINS | Genes involved in Gamma-carboxylation, transport, and amino-terminal cleavage of proteins |
| 0.5 | 6.8 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |
| 0.4 | 5.0 | REACTOME TRYPTOPHAN CATABOLISM | Genes involved in Tryptophan catabolism |
| 0.4 | 3.1 | REACTOME FGFR2C LIGAND BINDING AND ACTIVATION | Genes involved in FGFR2c ligand binding and activation |
| 0.2 | 3.9 | REACTOME POL SWITCHING | Genes involved in Polymerase switching |
| 0.2 | 3.1 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.2 | 2.4 | REACTOME CREATION OF C4 AND C2 ACTIVATORS | Genes involved in Creation of C4 and C2 activators |
| 0.2 | 5.9 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.2 | 1.8 | REACTOME PECAM1 INTERACTIONS | Genes involved in PECAM1 interactions |
| 0.2 | 1.9 | REACTOME INSULIN RECEPTOR RECYCLING | Genes involved in Insulin receptor recycling |
| 0.2 | 2.9 | REACTOME TELOMERE MAINTENANCE | Genes involved in Telomere Maintenance |
| 0.2 | 3.2 | REACTOME FANCONI ANEMIA PATHWAY | Genes involved in Fanconi Anemia pathway |
| 0.2 | 0.9 | REACTOME NEGATIVE REGULATION OF THE PI3K AKT NETWORK | Genes involved in Negative regulation of the PI3K/AKT network |
| 0.1 | 3.2 | REACTOME SYNTHESIS OF PA | Genes involved in Synthesis of PA |
| 0.1 | 8.4 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.1 | 1.0 | REACTOME AMINO ACID SYNTHESIS AND INTERCONVERSION TRANSAMINATION | Genes involved in Amino acid synthesis and interconversion (transamination) |
| 0.1 | 2.7 | REACTOME APC C CDH1 MEDIATED DEGRADATION OF CDC20 AND OTHER APC C CDH1 TARGETED PROTEINS IN LATE MITOSIS EARLY G1 | Genes involved in APC/C:Cdh1 mediated degradation of Cdc20 and other APC/C:Cdh1 targeted proteins in late mitosis/early G1 |
| 0.1 | 0.7 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.1 | 2.1 | REACTOME FORMATION OF INCISION COMPLEX IN GG NER | Genes involved in Formation of incision complex in GG-NER |
| 0.1 | 1.3 | REACTOME MEIOTIC SYNAPSIS | Genes involved in Meiotic Synapsis |
| 0.1 | 0.9 | REACTOME SEMA4D INDUCED CELL MIGRATION AND GROWTH CONE COLLAPSE | Genes involved in Sema4D induced cell migration and growth-cone collapse |
| 0.1 | 10.1 | REACTOME METABOLISM OF AMINO ACIDS AND DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |
| 0.1 | 3.2 | REACTOME NRAGE SIGNALS DEATH THROUGH JNK | Genes involved in NRAGE signals death through JNK |
| 0.1 | 0.7 | REACTOME ACTIVATION OF IRF3 IRF7 MEDIATED BY TBK1 IKK EPSILON | Genes involved in Activation of IRF3/IRF7 mediated by TBK1/IKK epsilon |
| 0.1 | 0.7 | REACTOME THE ACTIVATION OF ARYLSULFATASES | Genes involved in The activation of arylsulfatases |
| 0.1 | 2.3 | REACTOME NEP NS2 INTERACTS WITH THE CELLULAR EXPORT MACHINERY | Genes involved in NEP/NS2 Interacts with the Cellular Export Machinery |
| 0.1 | 1.2 | REACTOME EGFR DOWNREGULATION | Genes involved in EGFR downregulation |
| 0.0 | 0.4 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.0 | 4.4 | REACTOME METABOLISM OF CARBOHYDRATES | Genes involved in Metabolism of carbohydrates |
| 0.0 | 0.8 | REACTOME BMAL1 CLOCK NPAS2 ACTIVATES CIRCADIAN EXPRESSION | Genes involved in BMAL1:CLOCK/NPAS2 Activates Circadian Expression |
| 0.0 | 0.5 | REACTOME METABOLISM OF NON CODING RNA | Genes involved in Metabolism of non-coding RNA |
| 0.0 | 0.7 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.0 | 0.4 | REACTOME RNA POL II TRANSCRIPTION PRE INITIATION AND PROMOTER OPENING | Genes involved in RNA Polymerase II Transcription Pre-Initiation And Promoter Opening |