Project
PRJDB7713: Age associated transcriptome analysis in 5 tissues of zebrafish
Navigation
Downloads
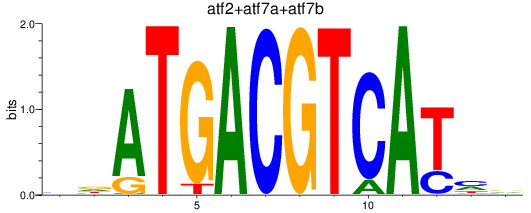
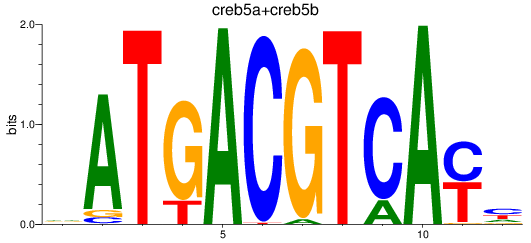
Results for atf2+atf7a+atf7b_creb5a+creb5b
Z-value: 2.53
Transcription factors associated with atf2+atf7a+atf7b_creb5a+creb5b
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
atf7a
|
ENSDARG00000011298 | activating transcription factor 7a |
|
atf2
|
ENSDARG00000023903 | activating transcription factor 2 |
|
atf7b
|
ENSDARG00000055481 | activating transcription factor 7b |
|
atf7b
|
ENSDARG00000114492 | activating transcription factor 7b |
|
atf7b
|
ENSDARG00000115171 | activating transcription factor 7b |
|
creb5b
|
ENSDARG00000070536 | cAMP responsive element binding protein 5b |
|
creb5a
|
ENSDARG00000099002 | cAMP responsive element binding protein 5a |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| atf7b | dr11_v1_chr6_-_39631164_39631181 | 0.75 | 1.5e-18 | Click! |
| atf2 | dr11_v1_chr9_+_2343096_2343172 | 0.63 | 1.1e-11 | Click! |
| creb5b | dr11_v1_chr16_-_20707742_20707742 | 0.59 | 4.0e-10 | Click! |
| creb5a | dr11_v1_chr19_-_19505167_19505167 | 0.30 | 2.9e-03 | Click! |
| atf7a | dr11_v1_chr23_-_27479558_27479558 | -0.22 | 3.0e-02 | Click! |
Activity profile of atf2+atf7a+atf7b_creb5a+creb5b motif
Sorted Z-values of atf2+atf7a+atf7b_creb5a+creb5b motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr16_+_34523515 | 65.64 |
ENSDART00000041007
|
stmn1b
|
stathmin 1b |
| chr21_-_43606502 | 57.71 |
ENSDART00000151030
|
si:ch73-362m14.4
|
si:ch73-362m14.4 |
| chr3_-_49566364 | 57.56 |
ENSDART00000161507
|
zgc:153426
|
zgc:153426 |
| chr24_-_7632187 | 49.95 |
ENSDART00000041714
|
atp6v0a1b
|
ATPase H+ transporting V0 subunit a1b |
| chr1_-_22861348 | 48.29 |
ENSDART00000139412
|
SMIM18
|
si:dkey-92j12.6 |
| chr16_-_12173554 | 46.17 |
ENSDART00000110567
ENSDART00000155935 |
clstn3
|
calsyntenin 3 |
| chr9_+_7548533 | 45.03 |
ENSDART00000081543
|
ptprna
|
protein tyrosine phosphatase, receptor type, Na |
| chr7_+_40228422 | 41.20 |
ENSDART00000052222
|
ptprn2
|
protein tyrosine phosphatase, receptor type, N polypeptide 2 |
| chr6_+_27667359 | 40.60 |
ENSDART00000159624
ENSDART00000049177 |
rab6ba
|
RAB6B, member RAS oncogene family a |
| chr6_+_13920479 | 39.10 |
ENSDART00000155480
|
ptprnb
|
protein tyrosine phosphatase, receptor type, Nb |
| chr9_-_296169 | 38.05 |
ENSDART00000165228
|
kif5aa
|
kinesin family member 5A, a |
| chr24_-_24163201 | 37.89 |
ENSDART00000140170
|
map7d2b
|
MAP7 domain containing 2b |
| chr10_-_24371312 | 37.81 |
ENSDART00000149362
|
pitpnab
|
phosphatidylinositol transfer protein, alpha b |
| chr20_+_20637866 | 37.01 |
ENSDART00000060203
ENSDART00000079079 |
rtn1b
|
reticulon 1b |
| chr15_+_28685892 | 36.40 |
ENSDART00000155815
ENSDART00000060244 |
nova2
|
neuro-oncological ventral antigen 2 |
| chr13_+_36146415 | 34.12 |
ENSDART00000140301
|
TTC9
|
si:ch211-259k16.3 |
| chr20_+_20638034 | 32.35 |
ENSDART00000189759
|
rtn1b
|
reticulon 1b |
| chr1_-_38756870 | 31.21 |
ENSDART00000130324
ENSDART00000148404 |
gpm6ab
|
glycoprotein M6Ab |
| chr24_-_24162930 | 30.61 |
ENSDART00000080602
|
map7d2b
|
MAP7 domain containing 2b |
| chr8_+_21588067 | 30.37 |
ENSDART00000172190
|
ajap1
|
adherens junctions associated protein 1 |
| chr25_+_35683956 | 30.03 |
ENSDART00000149768
|
kif21a
|
kinesin family member 21A |
| chr20_+_29743904 | 29.58 |
ENSDART00000146366
ENSDART00000153154 |
kidins220b
|
kinase D-interacting substrate 220b |
| chr2_-_42415902 | 28.81 |
ENSDART00000142489
|
slco5a1b
|
solute carrier organic anion transporter family member 5A1b |
| chr17_-_26911852 | 28.60 |
ENSDART00000045842
|
rcan3
|
regulator of calcineurin 3 |
| chr2_+_20332044 | 28.59 |
ENSDART00000112131
|
plppr4a
|
phospholipid phosphatase related 4a |
| chr17_+_8184649 | 28.36 |
ENSDART00000091818
|
tulp4b
|
tubby like protein 4b |
| chr13_-_40499296 | 27.86 |
ENSDART00000158338
|
CNNM1
|
Danio rerio cyclin and CBS domain divalent metal cation transport mediator 1 (cnnm1), mRNA. |
| chr5_-_23999777 | 25.82 |
ENSDART00000085969
|
map7d2a
|
MAP7 domain containing 2a |
| chr13_+_32446169 | 25.44 |
ENSDART00000143325
|
nt5c1ba
|
5'-nucleotidase, cytosolic IB a |
| chr23_-_29667716 | 25.43 |
ENSDART00000158302
ENSDART00000133902 |
clstn1
|
calsyntenin 1 |
| chr2_+_20331445 | 25.36 |
ENSDART00000186880
|
plppr4a
|
phospholipid phosphatase related 4a |
| chr8_+_44714336 | 24.91 |
ENSDART00000145801
|
elmod3
|
ELMO/CED-12 domain containing 3 |
| chr10_+_37145007 | 24.03 |
ENSDART00000131777
|
cuedc1a
|
CUE domain containing 1a |
| chr8_+_39607466 | 23.23 |
ENSDART00000097427
|
msi1
|
musashi RNA-binding protein 1 |
| chr16_+_25137483 | 22.96 |
ENSDART00000155666
|
znf576.1
|
zinc finger protein 576, tandem duplicate 1 |
| chr7_-_12065668 | 21.81 |
ENSDART00000101537
|
mex3b
|
mex-3 RNA binding family member B |
| chr6_-_48311 | 21.49 |
ENSDART00000131010
|
zgc:114175
|
zgc:114175 |
| chr16_-_9869056 | 21.38 |
ENSDART00000149312
|
ncalda
|
neurocalcin delta a |
| chr2_+_35603637 | 21.25 |
ENSDART00000147278
|
plk3
|
polo-like kinase 3 (Drosophila) |
| chr14_-_48939560 | 21.12 |
ENSDART00000021736
|
scocb
|
short coiled-coil protein b |
| chr1_+_41666611 | 20.63 |
ENSDART00000145789
|
fbxo41
|
F-box protein 41 |
| chr8_+_29267093 | 20.45 |
ENSDART00000077647
|
grid2
|
glutamate receptor, ionotropic, delta 2 |
| chr23_-_4915118 | 20.18 |
ENSDART00000060714
|
atp6ap1a
|
ATPase H+ transporting accessory protein 1a |
| chr15_-_11341635 | 20.10 |
ENSDART00000055220
|
rab30
|
RAB30, member RAS oncogene family |
| chr17_+_29345606 | 19.67 |
ENSDART00000086164
|
kctd3
|
potassium channel tetramerization domain containing 3 |
| chr20_+_25911342 | 18.71 |
ENSDART00000146004
|
ttbk2b
|
tau tubulin kinase 2b |
| chr13_+_23843712 | 18.49 |
ENSDART00000057611
|
oprm1
|
opioid receptor, mu 1 |
| chr15_-_12319065 | 18.47 |
ENSDART00000162973
ENSDART00000170543 |
fxyd6
|
FXYD domain containing ion transport regulator 6 |
| chr11_-_44163164 | 18.40 |
ENSDART00000047126
|
clcn4
|
chloride channel, voltage-sensitive 4 |
| chr2_+_12255568 | 18.23 |
ENSDART00000184164
ENSDART00000013454 |
prtfdc1
|
phosphoribosyl transferase domain containing 1 |
| chr16_-_32649929 | 18.22 |
ENSDART00000136161
|
faxcb
|
failed axon connections homolog b |
| chr22_+_1796057 | 17.91 |
ENSDART00000170834
|
znf1179
|
zinc finger protein 1179 |
| chr20_-_34801181 | 17.83 |
ENSDART00000048375
ENSDART00000132426 |
stmn4
|
stathmin-like 4 |
| chr4_-_69189894 | 17.72 |
ENSDART00000169596
|
si:ch211-209j12.1
|
si:ch211-209j12.1 |
| chr3_+_20156956 | 17.71 |
ENSDART00000125281
|
ngfra
|
nerve growth factor receptor a (TNFR superfamily, member 16) |
| chr17_-_15382704 | 17.67 |
ENSDART00000005313
|
zgc:85722
|
zgc:85722 |
| chr25_-_37084032 | 17.63 |
ENSDART00000025494
|
hprt1l
|
hypoxanthine phosphoribosyltransferase 1, like |
| chr15_+_28685625 | 17.18 |
ENSDART00000188797
ENSDART00000166036 |
nova2
|
neuro-oncological ventral antigen 2 |
| chr13_+_28705143 | 17.06 |
ENSDART00000183338
|
ldb1a
|
LIM domain binding 1a |
| chr15_+_5028608 | 16.86 |
ENSDART00000092809
|
abcg1
|
ATP-binding cassette, sub-family G (WHITE), member 1 |
| chr13_-_40754499 | 16.84 |
ENSDART00000111641
ENSDART00000159255 |
morn4
|
MORN repeat containing 4 |
| chr21_-_23308286 | 16.30 |
ENSDART00000184419
|
zbtb16a
|
zinc finger and BTB domain containing 16a |
| chr24_+_744713 | 16.18 |
ENSDART00000067764
|
stk17a
|
serine/threonine kinase 17a |
| chr3_-_5964557 | 16.18 |
ENSDART00000184738
|
BX284638.1
|
|
| chr15_+_1004680 | 15.98 |
ENSDART00000157310
|
si:dkey-77f5.8
|
si:dkey-77f5.8 |
| chr23_-_29667544 | 15.78 |
ENSDART00000059339
|
clstn1
|
calsyntenin 1 |
| chr18_+_12058403 | 15.31 |
ENSDART00000140854
ENSDART00000193632 ENSDART00000190519 ENSDART00000190685 ENSDART00000112671 |
bicd1a
|
bicaudal D homolog 1a |
| chr25_+_28893615 | 15.29 |
ENSDART00000156994
ENSDART00000075151 |
amn1
|
antagonist of mitotic exit network 1 homolog (S. cerevisiae) |
| chr14_+_11950011 | 15.11 |
ENSDART00000188138
|
frmpd3
|
FERM and PDZ domain containing 3 |
| chr12_-_35505610 | 15.06 |
ENSDART00000105518
|
camk2g1
|
calcium/calmodulin-dependent protein kinase (CaM kinase) II gamma 1 |
| chr2_-_56635744 | 15.00 |
ENSDART00000167790
ENSDART00000168160 |
pip5k1cb
|
phosphatidylinositol-4-phosphate 5-kinase, type I, gamma b |
| chr7_-_27686021 | 14.38 |
ENSDART00000079112
ENSDART00000100989 |
calca
|
calcitonin/calcitonin-related polypeptide, alpha |
| chr1_+_10305611 | 14.07 |
ENSDART00000043881
|
zgc:77880
|
zgc:77880 |
| chr2_+_38147761 | 14.05 |
ENSDART00000135307
|
sall2
|
spalt-like transcription factor 2 |
| chr16_+_5184402 | 13.79 |
ENSDART00000156685
|
soga3a
|
SOGA family member 3a |
| chr7_+_30493684 | 13.77 |
ENSDART00000027466
|
mindy2
|
MINDY lysine 48 deubiquitinase 2 |
| chr5_-_41494831 | 13.71 |
ENSDART00000051081
|
eef2l2
|
eukaryotic translation elongation factor 2, like 2 |
| chr14_+_597532 | 13.59 |
ENSDART00000159805
|
LO018568.2
|
|
| chr8_-_24113575 | 13.52 |
ENSDART00000099692
ENSDART00000186211 |
dclre1b
|
DNA cross-link repair 1B |
| chr10_-_17170086 | 13.49 |
ENSDART00000020122
|
ywhah
|
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, eta polypeptide |
| chr12_-_32066469 | 13.49 |
ENSDART00000140685
ENSDART00000062185 |
rab40b
|
RAB40B, member RAS oncogene family |
| chr15_-_20412286 | 13.48 |
ENSDART00000008589
|
chp2
|
calcineurin-like EF-hand protein 2 |
| chr19_-_1961024 | 13.37 |
ENSDART00000108784
|
mturn
|
maturin, neural progenitor differentiation regulator homolog (Xenopus) |
| chr21_-_3770636 | 13.35 |
ENSDART00000053596
|
scamp1
|
secretory carrier membrane protein 1 |
| chr8_-_25846188 | 13.05 |
ENSDART00000128829
|
efhd2
|
EF-hand domain family, member D2 |
| chr8_-_9118958 | 12.99 |
ENSDART00000037922
|
slc6a8
|
solute carrier family 6 (neurotransmitter transporter), member 8 |
| chr21_-_25395223 | 12.90 |
ENSDART00000016219
|
ppme1
|
protein phosphatase methylesterase 1 |
| chr16_+_46111849 | 12.81 |
ENSDART00000172232
|
sv2a
|
synaptic vesicle glycoprotein 2A |
| chr20_-_9462433 | 12.49 |
ENSDART00000152674
ENSDART00000040557 |
zgc:101840
|
zgc:101840 |
| chr25_-_7686201 | 12.38 |
ENSDART00000157267
ENSDART00000155094 |
si:ch211-286c4.6
|
si:ch211-286c4.6 |
| chr5_+_57743815 | 12.38 |
ENSDART00000005090
|
alg9
|
ALG9, alpha-1,2-mannosyltransferase |
| chr5_-_24000211 | 12.27 |
ENSDART00000188865
|
map7d2a
|
MAP7 domain containing 2a |
| chr1_-_17693273 | 12.23 |
ENSDART00000146258
|
cfap97
|
cilia and flagella associated protein 97 |
| chr15_-_27710513 | 12.13 |
ENSDART00000005641
ENSDART00000134373 |
lhx1a
|
LIM homeobox 1a |
| chr20_-_32446406 | 12.10 |
ENSDART00000026635
|
nr2e1
|
nuclear receptor subfamily 2, group E, member 1 |
| chr15_-_739229 | 11.98 |
ENSDART00000153874
|
si:dkey-7i4.19
|
si:dkey-7i4.19 |
| chr2_+_22694382 | 11.94 |
ENSDART00000139196
|
kif1ab
|
kinesin family member 1Ab |
| chr4_+_77060861 | 11.70 |
ENSDART00000174271
ENSDART00000174393 ENSDART00000150450 |
si:dkey-240n22.8
|
si:dkey-240n22.8 |
| chr11_-_22372072 | 11.55 |
ENSDART00000065996
|
tmem183a
|
transmembrane protein 183A |
| chr20_-_3319642 | 11.55 |
ENSDART00000186743
ENSDART00000123096 |
marcksa
|
myristoylated alanine-rich protein kinase C substrate a |
| chr17_-_20979077 | 11.53 |
ENSDART00000006676
|
phyhipla
|
phytanoyl-CoA 2-hydroxylase interacting protein-like a |
| chr7_-_33868903 | 11.51 |
ENSDART00000173500
ENSDART00000178746 |
uacab
|
uveal autoantigen with coiled-coil domains and ankyrin repeats b |
| chr9_+_219124 | 11.48 |
ENSDART00000161484
|
map3k12
|
mitogen-activated protein kinase kinase kinase 12 |
| chr10_+_6496185 | 11.46 |
ENSDART00000164770
|
reep5
|
receptor accessory protein 5 |
| chr10_+_44903676 | 11.45 |
ENSDART00000158553
|
zgc:114173
|
zgc:114173 |
| chr20_-_24122881 | 11.20 |
ENSDART00000131857
|
bach2b
|
BTB and CNC homology 1, basic leucine zipper transcription factor 2b |
| chr12_-_30443562 | 10.85 |
ENSDART00000020769
|
adrb1
|
adrenoceptor beta 1 |
| chr21_+_45366229 | 10.82 |
ENSDART00000029946
|
ube2b
|
ubiquitin-conjugating enzyme E2B (RAD6 homolog) |
| chr23_+_23183449 | 10.67 |
ENSDART00000132296
|
klhl17
|
kelch-like family member 17 |
| chr18_-_19103929 | 10.66 |
ENSDART00000188370
ENSDART00000177621 |
dennd4a
|
DENN/MADD domain containing 4A |
| chr7_+_23907692 | 10.63 |
ENSDART00000045479
|
syt4
|
synaptotagmin IV |
| chr8_-_410199 | 10.63 |
ENSDART00000091177
ENSDART00000122979 ENSDART00000151331 ENSDART00000151155 |
trim36
|
tripartite motif containing 36 |
| chr22_+_2254972 | 10.50 |
ENSDART00000144906
|
znf1157
|
zinc finger protein 1157 |
| chr21_-_32374656 | 10.41 |
ENSDART00000112550
|
mapk9
|
mitogen-activated protein kinase 9 |
| chr18_+_17663898 | 10.29 |
ENSDART00000021213
|
cpne2
|
copine II |
| chr17_+_15674052 | 10.14 |
ENSDART00000156726
|
bach2a
|
BTB and CNC homology 1, basic leucine zipper transcription factor 2a |
| chr5_-_24231139 | 10.12 |
ENSDART00000143492
|
senp3a
|
SUMO1/sentrin/SMT3 specific peptidase 3a |
| chr1_+_9290103 | 9.96 |
ENSDART00000055009
|
uncx4.1
|
Unc4.1 homeobox (C. elegans) |
| chr20_+_9211237 | 9.94 |
ENSDART00000139527
|
si:ch211-59d15.4
|
si:ch211-59d15.4 |
| chr7_-_39203799 | 9.91 |
ENSDART00000173727
|
chrm4a
|
cholinergic receptor, muscarinic 4a |
| chr6_-_25952848 | 9.82 |
ENSDART00000076997
ENSDART00000148748 |
lmo4b
|
LIM domain only 4b |
| chr12_+_7491690 | 9.76 |
ENSDART00000152564
|
phyhiplb
|
phytanoyl-CoA 2-hydroxylase interacting protein-like b |
| chr15_-_917274 | 9.65 |
ENSDART00000156624
|
si:dkey-77f5.15
|
si:dkey-77f5.15 |
| chr2_+_42200390 | 9.53 |
ENSDART00000012919
|
ift57
|
intraflagellar transport 57 homolog (Chlamydomonas) |
| chr24_+_4373355 | 9.52 |
ENSDART00000179062
ENSDART00000093256 ENSDART00000138943 |
ccny
|
cyclin Y |
| chr20_-_18915376 | 9.48 |
ENSDART00000063725
|
xkr6b
|
XK, Kell blood group complex subunit-related family, member 6b |
| chr13_-_45475289 | 9.45 |
ENSDART00000043345
|
rsrp1
|
arginine/serine-rich protein 1 |
| chr15_-_1844048 | 9.13 |
ENSDART00000102410
|
taf15
|
TAF15 RNA polymerase II, TATA box binding protein (TBP)-associated factor |
| chr19_+_42227400 | 9.10 |
ENSDART00000131574
ENSDART00000135436 |
jtb
|
jumping translocation breakpoint |
| chr7_-_12909352 | 9.08 |
ENSDART00000172901
|
sh3gl3a
|
SH3-domain GRB2-like 3a |
| chr17_+_32500387 | 9.05 |
ENSDART00000018423
|
ywhaqb
|
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, theta polypeptide b |
| chr16_-_33097398 | 9.01 |
ENSDART00000166617
|
dopey1
|
dopey family member 1 |
| chr22_+_1940595 | 9.01 |
ENSDART00000163506
|
znf1167
|
zinc finger protein 1167 |
| chr6_-_6673813 | 9.00 |
ENSDART00000150995
|
si:dkey-261j15.2
|
si:dkey-261j15.2 |
| chr15_-_591308 | 9.00 |
ENSDART00000153884
|
si:ch73-144d13.5
|
si:ch73-144d13.5 |
| chr4_-_73739119 | 8.85 |
ENSDART00000108669
|
zgc:171551
|
zgc:171551 |
| chr17_+_12698532 | 8.66 |
ENSDART00000064509
ENSDART00000136830 |
stmn4l
|
stathmin-like 4, like |
| chr22_+_2239254 | 8.60 |
ENSDART00000131396
ENSDART00000135320 |
znf1144
|
zinc finger protein 1144 |
| chr9_-_6502491 | 8.56 |
ENSDART00000102672
|
nck2a
|
NCK adaptor protein 2a |
| chr5_-_32882162 | 8.52 |
ENSDART00000085769
|
lrsam1
|
leucine rich repeat and sterile alpha motif containing 1 |
| chr20_+_41549200 | 8.39 |
ENSDART00000135715
|
fam184a
|
family with sequence similarity 184, member A |
| chr12_-_5455936 | 8.37 |
ENSDART00000109305
|
tbc1d12b
|
TBC1 domain family, member 12b |
| chr21_-_5007109 | 8.27 |
ENSDART00000187042
ENSDART00000097796 ENSDART00000146766 |
rnf165a
|
ring finger protein 165a |
| chr15_-_28200049 | 8.26 |
ENSDART00000004200
|
sarm1
|
sterile alpha and TIR motif containing 1 |
| chr4_-_73825089 | 8.25 |
ENSDART00000174207
|
si:dkey-262g12.12
|
si:dkey-262g12.12 |
| chr7_+_22616212 | 8.10 |
ENSDART00000052844
|
cldn7a
|
claudin 7a |
| chr22_-_4760187 | 8.09 |
ENSDART00000081969
|
rad23ab
|
RAD23 homolog A, nucleotide excision repair protein b |
| chr25_-_31763897 | 8.02 |
ENSDART00000041740
|
ubl7a
|
ubiquitin-like 7a (bone marrow stromal cell-derived) |
| chr6_+_21227621 | 8.01 |
ENSDART00000193583
|
prkca
|
protein kinase C, alpha |
| chr4_+_40953320 | 8.01 |
ENSDART00000151912
|
znf1136
|
zinc finger protein 1136 |
| chr13_-_31470439 | 8.01 |
ENSDART00000076574
|
rtn1a
|
reticulon 1a |
| chr4_+_2655358 | 7.98 |
ENSDART00000007638
|
bcap29
|
B cell receptor associated protein 29 |
| chr12_+_19305390 | 7.86 |
ENSDART00000183987
ENSDART00000066391 |
csnk1e
|
casein kinase 1, epsilon |
| chr14_+_16813816 | 7.82 |
ENSDART00000161201
|
limch1b
|
LIM and calponin homology domains 1b |
| chr7_-_11605185 | 7.74 |
ENSDART00000169291
ENSDART00000113904 |
stard5
|
StAR-related lipid transfer (START) domain containing 5 |
| chr8_+_21406769 | 7.71 |
ENSDART00000135766
|
si:dkey-163f12.6
|
si:dkey-163f12.6 |
| chr22_+_2315996 | 7.64 |
ENSDART00000132489
|
znf1175
|
zinc finger protein 1175 |
| chr15_-_762319 | 7.63 |
ENSDART00000154306
ENSDART00000157492 |
si:dkey-7i4.16
znf1011
|
si:dkey-7i4.16 zinc finger protein 1011 |
| chr6_+_58832155 | 7.56 |
ENSDART00000144842
|
dctn2
|
dynactin 2 (p50) |
| chr4_-_30362840 | 7.55 |
ENSDART00000165929
|
znf1083
|
zinc finger protein 1083 |
| chr15_+_618376 | 7.55 |
ENSDART00000156007
|
si:ch73-144d13.8
|
si:ch73-144d13.8 |
| chr4_-_64482414 | 7.53 |
ENSDART00000159000
|
znf1105
|
zinc finger protein 1105 |
| chr7_-_26532089 | 7.37 |
ENSDART00000121698
|
senp3b
|
SUMO1/sentrin/SMT3 specific peptidase 3b |
| chr20_+_39223235 | 7.35 |
ENSDART00000132132
|
reps1
|
RALBP1 associated Eps domain containing 1 |
| chr6_-_48473395 | 7.34 |
ENSDART00000185096
|
ppm1j
|
protein phosphatase, Mg2+/Mn2+ dependent, 1J |
| chr25_+_7321675 | 7.31 |
ENSDART00000104712
ENSDART00000142934 |
hmg20a
|
high mobility group 20A |
| chr22_+_2819613 | 7.31 |
ENSDART00000131234
|
si:dkey-20i20.3
|
si:dkey-20i20.3 |
| chr3_+_32403758 | 7.30 |
ENSDART00000156982
|
si:ch211-195b15.8
|
si:ch211-195b15.8 |
| chr19_+_9174166 | 7.17 |
ENSDART00000104637
ENSDART00000150968 |
si:ch211-81a5.8
|
si:ch211-81a5.8 |
| chr13_+_15816573 | 7.08 |
ENSDART00000137061
|
klc1a
|
kinesin light chain 1a |
| chr17_+_15535501 | 7.06 |
ENSDART00000002932
|
marcksb
|
myristoylated alanine-rich protein kinase C substrate b |
| chr1_-_44037843 | 7.04 |
ENSDART00000160276
|
si:ch73-109d9.3
|
si:ch73-109d9.3 |
| chr25_-_12906872 | 6.93 |
ENSDART00000165156
ENSDART00000167449 |
sept15
|
septin 15 |
| chr15_+_872085 | 6.93 |
ENSDART00000193867
ENSDART00000157483 ENSDART00000156324 |
si:dkey-7i4.9
si:dkey-77f5.13
si:dkey-7i4.11
|
si:dkey-7i4.9 si:dkey-77f5.13 si:dkey-7i4.11 |
| chr23_-_17509656 | 6.82 |
ENSDART00000148423
|
dnajc5ab
|
DnaJ (Hsp40) homolog, subfamily C, member 5ab |
| chr19_+_2279051 | 6.75 |
ENSDART00000182103
|
itgb8
|
integrin, beta 8 |
| chr15_-_1687204 | 6.66 |
ENSDART00000155579
ENSDART00000181382 |
sptssb
|
serine palmitoyltransferase, small subunit B |
| chr13_+_17702853 | 6.62 |
ENSDART00000145438
|
si:dkey-27m7.4
|
si:dkey-27m7.4 |
| chr21_+_9576176 | 6.59 |
ENSDART00000161289
ENSDART00000159899 ENSDART00000162834 |
mapk10
|
mitogen-activated protein kinase 10 |
| chr15_+_903743 | 6.59 |
ENSDART00000155783
|
si:dkey-77f5.13
|
si:dkey-77f5.13 |
| chr15_-_9421481 | 6.58 |
ENSDART00000189045
ENSDART00000177158 |
sacs
|
sacsin molecular chaperone |
| chr18_+_8917766 | 6.57 |
ENSDART00000145226
|
si:ch211-233h19.2
|
si:ch211-233h19.2 |
| chr5_-_20814576 | 6.53 |
ENSDART00000098682
ENSDART00000147639 |
si:ch211-225b11.1
|
si:ch211-225b11.1 |
| chr6_-_39893501 | 6.51 |
ENSDART00000141611
ENSDART00000135631 ENSDART00000077662 ENSDART00000130613 |
myl6
|
myosin, light chain 6, alkali, smooth muscle and non-muscle |
| chr2_+_301898 | 6.49 |
ENSDART00000157246
|
znf1008
|
zinc finger protein 1008 |
| chr15_-_923981 | 6.44 |
ENSDART00000155285
|
si:dkey-77f5.14
|
si:dkey-77f5.14 |
| chr3_-_1204341 | 6.42 |
ENSDART00000089646
|
fam234b
|
family with sequence similarity 234, member B |
| chr13_-_50672341 | 6.36 |
ENSDART00000190498
|
FP102122.1
|
|
| chr6_+_6802582 | 6.28 |
ENSDART00000189422
|
dtd1
|
D-tyrosyl-tRNA deacylase 1 |
| chr22_+_30330574 | 6.27 |
ENSDART00000104751
|
mxi1
|
max interactor 1, dimerization protein |
| chr9_+_48219111 | 6.27 |
ENSDART00000111225
ENSDART00000145972 |
ccdc173
|
coiled-coil domain containing 173 |
| chr24_-_18877118 | 6.23 |
ENSDART00000092783
|
arfgef1
|
ADP-ribosylation factor guanine nucleotide-exchange factor 1 (brefeldin A-inhibited) |
| chr4_-_30422325 | 6.21 |
ENSDART00000158444
|
znf1114
|
zinc finger protein 1114 |
| chr15_-_727479 | 6.18 |
ENSDART00000157279
|
si:dkey-7i4.21
|
si:dkey-7i4.21 |
| chr17_-_8673567 | 6.16 |
ENSDART00000192714
ENSDART00000012546 |
ctbp2a
|
C-terminal binding protein 2a |
| chr25_+_16945348 | 6.12 |
ENSDART00000016591
|
fgf6a
|
fibroblast growth factor 6a |
| chr4_+_41528590 | 6.08 |
ENSDART00000164004
|
znf1094
|
zinc finger protein 1094 |
| chr25_-_32751982 | 6.04 |
ENSDART00000012862
|
isl2a
|
ISL LIM homeobox 2a |
| chr16_-_41131578 | 5.99 |
ENSDART00000102649
ENSDART00000145956 |
ptpn23a
|
protein tyrosine phosphatase, non-receptor type 23, a |
| chr20_-_44576949 | 5.94 |
ENSDART00000148639
|
ubxn2a
|
UBX domain protein 2A |
Network of associatons between targets according to the STRING database.
First level regulatory network of atf2+atf7a+atf7b_creb5a+creb5b
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 10.3 | 41.2 | GO:0030951 | establishment or maintenance of microtubule cytoskeleton polarity(GO:0030951) |
| 9.5 | 38.0 | GO:1990535 | neuron projection maintenance(GO:1990535) |
| 7.7 | 46.2 | GO:0051965 | positive regulation of synapse assembly(GO:0051965) |
| 5.9 | 17.6 | GO:0046099 | guanine salvage(GO:0006178) GMP salvage(GO:0032263) guanine biosynthetic process(GO:0046099) |
| 5.3 | 21.3 | GO:0048308 | organelle inheritance(GO:0048308) Golgi inheritance(GO:0048313) |
| 4.6 | 18.5 | GO:0038003 | opioid receptor signaling pathway(GO:0038003) |
| 4.2 | 16.9 | GO:0010874 | regulation of cholesterol efflux(GO:0010874) |
| 4.0 | 12.1 | GO:0060688 | regulation of morphogenesis of a branching structure(GO:0060688) positive regulation of mesonephros development(GO:0061213) regulation of mesonephros development(GO:0061217) regulation of kidney development(GO:0090183) positive regulation of kidney development(GO:0090184) regulation of branching involved in ureteric bud morphogenesis(GO:0090189) positive regulation of branching involved in ureteric bud morphogenesis(GO:0090190) interneuron axon guidance(GO:0097376) spinal cord interneuron axon guidance(GO:0097377) dorsal spinal cord interneuron axon guidance(GO:0097378) |
| 3.8 | 57.7 | GO:0051127 | positive regulation of actin nucleation(GO:0051127) positive regulation of Arp2/3 complex-mediated actin nucleation(GO:2000601) |
| 3.8 | 11.5 | GO:1901223 | negative regulation of NIK/NF-kappaB signaling(GO:1901223) |
| 3.0 | 21.1 | GO:0061635 | regulation of protein complex stability(GO:0061635) |
| 2.8 | 8.3 | GO:0072526 | pyridine nucleotide catabolic process(GO:0019364) NAD catabolic process(GO:0019677) pyridine-containing compound catabolic process(GO:0072526) |
| 2.7 | 13.5 | GO:0031627 | telomeric loop formation(GO:0031627) |
| 2.7 | 13.4 | GO:0030219 | megakaryocyte differentiation(GO:0030219) |
| 2.6 | 12.8 | GO:1904031 | positive regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0045737) positive regulation of cyclin-dependent protein kinase activity(GO:1904031) |
| 2.6 | 15.3 | GO:0048260 | positive regulation of receptor-mediated endocytosis(GO:0048260) |
| 2.5 | 12.3 | GO:0002025 | vasodilation by norepinephrine-epinephrine involved in regulation of systemic arterial blood pressure(GO:0002025) |
| 2.3 | 31.8 | GO:0051452 | vacuolar acidification(GO:0007035) pH reduction(GO:0045851) intracellular pH reduction(GO:0051452) |
| 2.0 | 25.4 | GO:0046085 | adenosine metabolic process(GO:0046085) |
| 1.9 | 92.1 | GO:0007019 | microtubule depolymerization(GO:0007019) |
| 1.6 | 8.0 | GO:0070973 | protein localization to endoplasmic reticulum exit site(GO:0070973) |
| 1.4 | 18.5 | GO:0032481 | positive regulation of type I interferon production(GO:0032481) |
| 1.4 | 31.2 | GO:0051489 | regulation of filopodium assembly(GO:0051489) |
| 1.2 | 40.6 | GO:0006891 | intra-Golgi vesicle-mediated transport(GO:0006891) |
| 1.2 | 5.9 | GO:0071881 | adenylate cyclase-inhibiting adrenergic receptor signaling pathway(GO:0071881) |
| 1.2 | 3.5 | GO:1902410 | mitotic cytokinetic process(GO:1902410) mitotic cleavage furrow formation(GO:1903673) |
| 1.1 | 20.4 | GO:0035235 | ionotropic glutamate receptor signaling pathway(GO:0035235) |
| 1.1 | 17.2 | GO:0016926 | protein desumoylation(GO:0016926) |
| 1.1 | 6.4 | GO:0080009 | mRNA methylation(GO:0080009) |
| 1.1 | 5.3 | GO:0034505 | tooth mineralization(GO:0034505) |
| 1.0 | 18.6 | GO:0035914 | skeletal muscle cell differentiation(GO:0035914) |
| 0.9 | 9.9 | GO:0007197 | adenylate cyclase-inhibiting G-protein coupled acetylcholine receptor signaling pathway(GO:0007197) |
| 0.9 | 27.9 | GO:0032482 | Rab protein signal transduction(GO:0032482) |
| 0.9 | 9.5 | GO:0070782 | phosphatidylserine exposure on apoptotic cell surface(GO:0070782) |
| 0.8 | 3.8 | GO:0060784 | regulation of cell proliferation involved in tissue homeostasis(GO:0060784) |
| 0.8 | 16.7 | GO:0071108 | protein K48-linked deubiquitination(GO:0071108) |
| 0.7 | 4.3 | GO:0098937 | dendritic transport(GO:0098935) anterograde dendritic transport(GO:0098937) anterograde dendritic transport of neurotransmitter receptor complex(GO:0098971) |
| 0.7 | 7.8 | GO:0048041 | cell-substrate adherens junction assembly(GO:0007045) focal adhesion assembly(GO:0048041) regulation of focal adhesion assembly(GO:0051893) regulation of cell-substrate junction assembly(GO:0090109) |
| 0.7 | 2.6 | GO:0030213 | hyaluronan biosynthetic process(GO:0030213) |
| 0.6 | 5.8 | GO:1902038 | positive regulation of hematopoietic stem cell differentiation(GO:1902038) |
| 0.6 | 5.1 | GO:0021744 | medulla oblongata development(GO:0021550) dorsal motor nucleus of vagus nerve development(GO:0021744) |
| 0.6 | 5.0 | GO:0099517 | anterograde synaptic vesicle transport(GO:0048490) synaptic vesicle cytoskeletal transport(GO:0099514) synaptic vesicle transport along microtubule(GO:0099517) |
| 0.6 | 25.8 | GO:0072114 | pronephros morphogenesis(GO:0072114) |
| 0.6 | 37.8 | GO:0015914 | phospholipid transport(GO:0015914) |
| 0.6 | 53.6 | GO:0000381 | regulation of alternative mRNA splicing, via spliceosome(GO:0000381) |
| 0.6 | 3.7 | GO:0010799 | regulation of peptidyl-threonine phosphorylation(GO:0010799) negative regulation of peptidyl-threonine phosphorylation(GO:0010801) |
| 0.6 | 6.0 | GO:0048845 | venous blood vessel morphogenesis(GO:0048845) |
| 0.6 | 8.4 | GO:2000785 | regulation of autophagosome assembly(GO:2000785) |
| 0.6 | 47.9 | GO:0046839 | phospholipid dephosphorylation(GO:0046839) |
| 0.6 | 5.1 | GO:0061709 | reticulophagy(GO:0061709) |
| 0.5 | 5.9 | GO:0031468 | nuclear envelope reassembly(GO:0031468) |
| 0.5 | 11.5 | GO:0007257 | activation of JUN kinase activity(GO:0007257) positive regulation of JUN kinase activity(GO:0043507) |
| 0.5 | 8.8 | GO:1900151 | regulation of nuclear-transcribed mRNA poly(A) tail shortening(GO:0060211) positive regulation of nuclear-transcribed mRNA poly(A) tail shortening(GO:0060213) positive regulation of mRNA catabolic process(GO:0061014) regulation of nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay(GO:1900151) positive regulation of nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay(GO:1900153) |
| 0.5 | 2.0 | GO:0048210 | Golgi vesicle fusion to target membrane(GO:0048210) |
| 0.5 | 3.4 | GO:0070073 | clustering of voltage-gated calcium channels(GO:0070073) |
| 0.5 | 18.1 | GO:0018230 | peptidyl-L-cysteine S-palmitoylation(GO:0018230) peptidyl-S-diacylglycerol-L-cysteine biosynthetic process from peptidyl-cysteine(GO:0018231) |
| 0.5 | 5.8 | GO:0042572 | retinol metabolic process(GO:0042572) |
| 0.5 | 14.4 | GO:0055064 | chloride ion homeostasis(GO:0055064) |
| 0.5 | 145.1 | GO:0006470 | protein dephosphorylation(GO:0006470) |
| 0.5 | 1.4 | GO:0045724 | positive regulation of cilium assembly(GO:0045724) |
| 0.4 | 1.8 | GO:0010259 | multicellular organism aging(GO:0010259) |
| 0.4 | 15.0 | GO:0008214 | protein demethylation(GO:0006482) protein dealkylation(GO:0008214) |
| 0.4 | 2.8 | GO:0034334 | adherens junction maintenance(GO:0034334) |
| 0.4 | 20.2 | GO:0030641 | regulation of cellular pH(GO:0030641) |
| 0.4 | 3.1 | GO:0001743 | optic placode formation(GO:0001743) optic placode formation involved in camera-type eye formation(GO:0046619) |
| 0.4 | 18.5 | GO:2000649 | regulation of sodium ion transmembrane transporter activity(GO:2000649) |
| 0.4 | 20.9 | GO:0071277 | cellular response to calcium ion(GO:0071277) |
| 0.3 | 17.0 | GO:0007254 | JNK cascade(GO:0007254) |
| 0.3 | 1.9 | GO:0072318 | clathrin coat disassembly(GO:0072318) vesicle uncoating(GO:0072319) |
| 0.3 | 5.0 | GO:0071712 | ER-associated misfolded protein catabolic process(GO:0071712) |
| 0.3 | 3.0 | GO:0000303 | response to superoxide(GO:0000303) response to oxygen radical(GO:0000305) removal of superoxide radicals(GO:0019430) cellular response to oxygen radical(GO:0071450) cellular response to superoxide(GO:0071451) cellular oxidant detoxification(GO:0098869) cellular detoxification(GO:1990748) |
| 0.3 | 28.8 | GO:0019722 | calcium-mediated signaling(GO:0019722) |
| 0.3 | 1.5 | GO:0006121 | mitochondrial electron transport, succinate to ubiquinone(GO:0006121) |
| 0.3 | 9.5 | GO:0042073 | intraciliary transport(GO:0042073) protein transport along microtubule(GO:0098840) |
| 0.3 | 4.2 | GO:0007214 | gamma-aminobutyric acid signaling pathway(GO:0007214) |
| 0.3 | 9.1 | GO:0000281 | mitotic cytokinesis(GO:0000281) |
| 0.3 | 2.9 | GO:0030719 | P granule organization(GO:0030719) |
| 0.3 | 1.8 | GO:0097428 | protein maturation by iron-sulfur cluster transfer(GO:0097428) |
| 0.3 | 5.8 | GO:0030032 | lamellipodium assembly(GO:0030032) |
| 0.3 | 5.3 | GO:0008625 | extrinsic apoptotic signaling pathway via death domain receptors(GO:0008625) |
| 0.2 | 11.0 | GO:0032012 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.2 | 4.4 | GO:0006995 | cellular response to nitrogen starvation(GO:0006995) cellular response to nitrogen levels(GO:0043562) |
| 0.2 | 0.7 | GO:0045448 | regulation of mitotic cell cycle, embryonic(GO:0009794) mitotic cell cycle, embryonic(GO:0045448) |
| 0.2 | 14.3 | GO:0090263 | positive regulation of canonical Wnt signaling pathway(GO:0090263) |
| 0.2 | 2.6 | GO:0070307 | lens fiber cell development(GO:0070307) lens fiber cell morphogenesis(GO:0070309) |
| 0.2 | 0.6 | GO:1903069 | regulation of ER-associated ubiquitin-dependent protein catabolic process(GO:1903069) |
| 0.2 | 2.6 | GO:0050908 | detection of light stimulus involved in visual perception(GO:0050908) detection of light stimulus involved in sensory perception(GO:0050962) |
| 0.2 | 1.9 | GO:0071260 | cellular response to mechanical stimulus(GO:0071260) |
| 0.2 | 6.8 | GO:0033627 | cell adhesion mediated by integrin(GO:0033627) |
| 0.2 | 1.5 | GO:0042403 | thyroid hormone metabolic process(GO:0042403) |
| 0.2 | 6.8 | GO:0016574 | histone ubiquitination(GO:0016574) |
| 0.2 | 137.4 | GO:0007017 | microtubule-based process(GO:0007017) |
| 0.2 | 19.5 | GO:0035725 | sodium ion transmembrane transport(GO:0035725) |
| 0.2 | 0.4 | GO:0044878 | abscission(GO:0009838) mitotic cytokinesis checkpoint(GO:0044878) |
| 0.2 | 8.1 | GO:0043297 | apical junction assembly(GO:0043297) bicellular tight junction assembly(GO:0070830) |
| 0.2 | 0.9 | GO:0055071 | manganese ion homeostasis(GO:0055071) |
| 0.2 | 0.6 | GO:0008612 | peptidyl-lysine modification to peptidyl-hypusine(GO:0008612) |
| 0.2 | 8.1 | GO:0006289 | nucleotide-excision repair(GO:0006289) |
| 0.2 | 22.1 | GO:0051260 | protein homooligomerization(GO:0051260) |
| 0.2 | 6.7 | GO:0007029 | endoplasmic reticulum organization(GO:0007029) |
| 0.2 | 0.8 | GO:0032475 | otolith formation(GO:0032475) |
| 0.2 | 4.1 | GO:0007168 | receptor guanylyl cyclase signaling pathway(GO:0007168) |
| 0.2 | 3.6 | GO:0021952 | central nervous system projection neuron axonogenesis(GO:0021952) |
| 0.2 | 1.3 | GO:0014004 | microglia differentiation(GO:0014004) |
| 0.2 | 6.8 | GO:0030433 | ER-associated ubiquitin-dependent protein catabolic process(GO:0030433) |
| 0.2 | 13.8 | GO:0010506 | regulation of autophagy(GO:0010506) |
| 0.2 | 16.6 | GO:0006821 | chloride transport(GO:0006821) |
| 0.1 | 7.2 | GO:0046928 | regulation of neurotransmitter secretion(GO:0046928) |
| 0.1 | 4.8 | GO:0009072 | aromatic amino acid family metabolic process(GO:0009072) |
| 0.1 | 2.3 | GO:0006265 | DNA topological change(GO:0006265) |
| 0.1 | 1.4 | GO:0032262 | pyrimidine-containing compound salvage(GO:0008655) pyrimidine ribonucleotide salvage(GO:0010138) pyrimidine nucleotide salvage(GO:0032262) pyrimidine nucleoside salvage(GO:0043097) UMP salvage(GO:0044206) |
| 0.1 | 12.4 | GO:0006487 | protein N-linked glycosylation(GO:0006487) |
| 0.1 | 25.2 | GO:0018105 | peptidyl-serine phosphorylation(GO:0018105) |
| 0.1 | 0.3 | GO:0051898 | negative regulation of protein kinase B signaling(GO:0051898) |
| 0.1 | 2.9 | GO:0000187 | activation of MAPK activity(GO:0000187) |
| 0.1 | 0.5 | GO:0060546 | negative regulation of necroptotic process(GO:0060546) negative regulation of necrotic cell death(GO:0060547) |
| 0.1 | 1.6 | GO:0015937 | coenzyme A biosynthetic process(GO:0015937) |
| 0.1 | 3.2 | GO:0008286 | insulin receptor signaling pathway(GO:0008286) |
| 0.1 | 1.2 | GO:0070932 | histone H3 deacetylation(GO:0070932) |
| 0.1 | 3.1 | GO:0042752 | regulation of circadian rhythm(GO:0042752) |
| 0.1 | 15.0 | GO:0046488 | phosphatidylinositol metabolic process(GO:0046488) |
| 0.1 | 7.1 | GO:0031146 | SCF-dependent proteasomal ubiquitin-dependent protein catabolic process(GO:0031146) |
| 0.1 | 13.5 | GO:0072659 | protein localization to plasma membrane(GO:0072659) protein localization to cell periphery(GO:1990778) |
| 0.1 | 8.5 | GO:0060218 | hematopoietic stem cell differentiation(GO:0060218) |
| 0.1 | 1.3 | GO:0098508 | endothelial to hematopoietic transition(GO:0098508) |
| 0.1 | 0.5 | GO:0035021 | negative regulation of Rac protein signal transduction(GO:0035021) wound healing, spreading of epidermal cells(GO:0035313) negative regulation of ruffle assembly(GO:1900028) |
| 0.1 | 0.7 | GO:0033314 | mitotic DNA replication checkpoint(GO:0033314) |
| 0.1 | 4.6 | GO:0006096 | glycolytic process(GO:0006096) ATP generation from ADP(GO:0006757) |
| 0.1 | 0.9 | GO:0006896 | Golgi to vacuole transport(GO:0006896) |
| 0.1 | 1.0 | GO:0043162 | ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway(GO:0043162) |
| 0.1 | 1.2 | GO:0001843 | neural tube closure(GO:0001843) primary neural tube formation(GO:0014020) |
| 0.0 | 3.5 | GO:0006836 | neurotransmitter transport(GO:0006836) |
| 0.0 | 2.1 | GO:0016575 | histone deacetylation(GO:0016575) |
| 0.0 | 2.3 | GO:0009583 | detection of light stimulus(GO:0009583) |
| 0.0 | 5.4 | GO:0007266 | Rho protein signal transduction(GO:0007266) |
| 0.0 | 0.7 | GO:0030150 | protein import into mitochondrial matrix(GO:0030150) |
| 0.0 | 0.8 | GO:0048264 | determination of ventral identity(GO:0048264) |
| 0.0 | 4.8 | GO:0006399 | tRNA metabolic process(GO:0006399) |
| 0.0 | 1.2 | GO:0010842 | retina layer formation(GO:0010842) |
| 0.0 | 2.4 | GO:0006898 | receptor-mediated endocytosis(GO:0006898) |
| 0.0 | 2.0 | GO:0010942 | positive regulation of cell death(GO:0010942) positive regulation of apoptotic process(GO:0043065) positive regulation of programmed cell death(GO:0043068) |
| 0.0 | 1.2 | GO:0008543 | fibroblast growth factor receptor signaling pathway(GO:0008543) |
| 0.0 | 4.5 | GO:0018193 | peptidyl-amino acid modification(GO:0018193) |
| 0.0 | 49.6 | GO:0006355 | regulation of transcription, DNA-templated(GO:0006355) |
| 0.0 | 0.1 | GO:0042670 | retinal cone cell differentiation(GO:0042670) retinal cone cell development(GO:0046549) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 7.6 | 30.4 | GO:0089717 | spanning component of plasma membrane(GO:0044214) spanning component of membrane(GO:0089717) |
| 3.6 | 57.7 | GO:0031209 | SCAR complex(GO:0031209) |
| 3.4 | 20.2 | GO:0033181 | plasma membrane proton-transporting V-type ATPase complex(GO:0033181) |
| 3.2 | 31.8 | GO:0000220 | vacuolar proton-transporting V-type ATPase, V0 domain(GO:0000220) |
| 2.6 | 31.2 | GO:0044295 | axonal growth cone(GO:0044295) |
| 2.2 | 10.8 | GO:0033503 | HULC complex(GO:0033503) |
| 2.0 | 7.8 | GO:0016460 | myosin II complex(GO:0016460) |
| 1.3 | 5.0 | GO:0098574 | cytoplasmic side of lysosomal membrane(GO:0098574) |
| 1.2 | 109.4 | GO:0030141 | secretory granule(GO:0030141) |
| 1.1 | 4.2 | GO:0045293 | MIS complex(GO:0036396) mRNA editing complex(GO:0045293) |
| 1.1 | 89.3 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.8 | 17.5 | GO:0044665 | MLL1/2 complex(GO:0044665) MLL1 complex(GO:0071339) |
| 0.7 | 5.1 | GO:0071818 | BAT3 complex(GO:0071818) |
| 0.7 | 8.8 | GO:0030015 | CCR4-NOT core complex(GO:0030015) |
| 0.7 | 13.4 | GO:0032588 | trans-Golgi network membrane(GO:0032588) |
| 0.6 | 5.8 | GO:0032021 | NELF complex(GO:0032021) |
| 0.5 | 9.5 | GO:0030992 | intraciliary transport particle B(GO:0030992) |
| 0.5 | 4.2 | GO:0097648 | G-protein coupled receptor dimeric complex(GO:0038037) G-protein coupled receptor heterodimeric complex(GO:0038039) G-protein coupled receptor complex(GO:0097648) |
| 0.5 | 5.0 | GO:0017059 | serine C-palmitoyltransferase complex(GO:0017059) endoplasmic reticulum palmitoyltransferase complex(GO:0031211) |
| 0.5 | 34.1 | GO:0031902 | late endosome membrane(GO:0031902) |
| 0.4 | 117.7 | GO:0045211 | postsynaptic membrane(GO:0045211) |
| 0.4 | 18.5 | GO:0043204 | perikaryon(GO:0043204) |
| 0.4 | 2.9 | GO:0035517 | PR-DUB complex(GO:0035517) |
| 0.4 | 41.2 | GO:0005802 | trans-Golgi network(GO:0005802) |
| 0.4 | 18.5 | GO:0030496 | midbody(GO:0030496) |
| 0.3 | 13.5 | GO:0000781 | chromosome, telomeric region(GO:0000781) |
| 0.3 | 0.9 | GO:0030121 | AP-1 adaptor complex(GO:0030121) |
| 0.3 | 7.2 | GO:0043195 | terminal bouton(GO:0043195) |
| 0.3 | 10.6 | GO:0035869 | ciliary transition zone(GO:0035869) |
| 0.3 | 0.9 | GO:0030289 | protein phosphatase 4 complex(GO:0030289) |
| 0.3 | 1.5 | GO:0045257 | mitochondrial respiratory chain complex II, succinate dehydrogenase complex (ubiquinone)(GO:0005749) succinate dehydrogenase complex (ubiquinone)(GO:0045257) respiratory chain complex II(GO:0045273) succinate dehydrogenase complex(GO:0045281) fumarate reductase complex(GO:0045283) |
| 0.3 | 1.2 | GO:0070209 | ASTRA complex(GO:0070209) |
| 0.3 | 46.7 | GO:0008021 | synaptic vesicle(GO:0008021) |
| 0.3 | 0.8 | GO:0071141 | SMAD protein complex(GO:0071141) |
| 0.3 | 3.3 | GO:0071007 | U2-type catalytic step 2 spliceosome(GO:0071007) |
| 0.2 | 21.8 | GO:0000922 | spindle pole(GO:0000922) |
| 0.2 | 12.8 | GO:0005776 | autophagosome(GO:0005776) |
| 0.2 | 9.1 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.2 | 130.9 | GO:0043005 | neuron projection(GO:0043005) |
| 0.2 | 1.8 | GO:0031314 | extrinsic component of mitochondrial inner membrane(GO:0031314) |
| 0.2 | 3.8 | GO:1990907 | beta-catenin-TCF complex(GO:1990907) |
| 0.2 | 2.9 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.2 | 2.1 | GO:0016580 | Sin3 complex(GO:0016580) |
| 0.2 | 5.3 | GO:0016605 | PML body(GO:0016605) |
| 0.2 | 95.5 | GO:0015630 | microtubule cytoskeleton(GO:0015630) |
| 0.1 | 6.8 | GO:0008305 | integrin complex(GO:0008305) |
| 0.1 | 16.9 | GO:0048471 | perinuclear region of cytoplasm(GO:0048471) |
| 0.1 | 11.3 | GO:0005882 | intermediate filament(GO:0005882) |
| 0.1 | 0.8 | GO:0030130 | trans-Golgi network transport vesicle membrane(GO:0012510) clathrin coat of trans-Golgi network vesicle(GO:0030130) |
| 0.1 | 3.5 | GO:0016342 | catenin complex(GO:0016342) |
| 0.1 | 0.7 | GO:0005744 | mitochondrial inner membrane presequence translocase complex(GO:0005744) |
| 0.1 | 4.8 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.1 | 1.2 | GO:0080008 | Cul4-RING E3 ubiquitin ligase complex(GO:0080008) |
| 0.1 | 71.0 | GO:0005887 | integral component of plasma membrane(GO:0005887) |
| 0.1 | 18.1 | GO:0005667 | transcription factor complex(GO:0005667) |
| 0.0 | 1.2 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.0 | 1.1 | GO:0031226 | intrinsic component of plasma membrane(GO:0031226) |
| 0.0 | 0.1 | GO:0016589 | NURF complex(GO:0016589) |
| 0.0 | 2.8 | GO:0043235 | receptor complex(GO:0043235) |
| 0.0 | 3.1 | GO:0030176 | integral component of endoplasmic reticulum membrane(GO:0030176) |
| 0.0 | 0.4 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.0 | 3.9 | GO:0009986 | cell surface(GO:0009986) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 6.4 | 57.7 | GO:0071933 | Arp2/3 complex binding(GO:0071933) |
| 6.2 | 18.5 | GO:0038046 | enkephalin receptor activity(GO:0038046) |
| 5.9 | 17.6 | GO:0004422 | hypoxanthine phosphoribosyltransferase activity(GO:0004422) |
| 5.4 | 37.8 | GO:0008526 | phosphatidylinositol transporter activity(GO:0008526) |
| 4.8 | 14.4 | GO:0031716 | calcitonin receptor binding(GO:0031716) |
| 4.8 | 28.6 | GO:0008597 | calcium-dependent protein serine/threonine phosphatase regulator activity(GO:0008597) |
| 2.8 | 13.8 | GO:0070004 | cysteine-type exopeptidase activity(GO:0070004) |
| 2.5 | 17.7 | GO:0005035 | death receptor activity(GO:0005035) |
| 2.5 | 12.4 | GO:0000026 | alpha-1,2-mannosyltransferase activity(GO:0000026) |
| 2.5 | 12.3 | GO:0004939 | beta-adrenergic receptor activity(GO:0004939) |
| 2.4 | 17.1 | GO:0030274 | LIM domain binding(GO:0030274) |
| 2.1 | 31.8 | GO:0051117 | ATPase binding(GO:0051117) |
| 2.1 | 6.3 | GO:0051500 | D-aminoacyl-tRNA deacylase activity(GO:0051499) D-tyrosyl-tRNA(Tyr) deacylase activity(GO:0051500) |
| 2.0 | 7.8 | GO:0032038 | myosin head/neck binding(GO:0032028) myosin II head/neck binding(GO:0032034) myosin II heavy chain binding(GO:0032038) |
| 1.9 | 5.8 | GO:0047173 | phosphatidylcholine-retinol O-acyltransferase activity(GO:0047173) |
| 1.9 | 15.3 | GO:0034452 | dynactin binding(GO:0034452) |
| 1.8 | 47.9 | GO:0042577 | lipid phosphatase activity(GO:0042577) |
| 1.7 | 17.0 | GO:0004705 | JUN kinase activity(GO:0004705) SAP kinase activity(GO:0016909) |
| 1.7 | 13.5 | GO:0035312 | 5'-3' exodeoxyribonuclease activity(GO:0035312) |
| 1.6 | 54.3 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 1.6 | 4.8 | GO:0003868 | 4-hydroxyphenylpyruvate dioxygenase activity(GO:0003868) |
| 1.4 | 4.2 | GO:0001734 | mRNA (N6-adenosine)-methyltransferase activity(GO:0001734) |
| 1.3 | 7.7 | GO:0032052 | bile acid binding(GO:0032052) |
| 1.3 | 11.5 | GO:0004706 | JUN kinase kinase kinase activity(GO:0004706) |
| 1.2 | 25.4 | GO:0008253 | 5'-nucleotidase activity(GO:0008253) |
| 1.2 | 16.9 | GO:0015665 | alcohol transmembrane transporter activity(GO:0015665) |
| 1.2 | 34.9 | GO:0030165 | PDZ domain binding(GO:0030165) |
| 1.1 | 3.3 | GO:0030623 | U5 snRNA binding(GO:0030623) |
| 1.1 | 17.2 | GO:0016929 | SUMO-specific protease activity(GO:0016929) |
| 1.0 | 5.9 | GO:0004938 | alpha2-adrenergic receptor activity(GO:0004938) epinephrine binding(GO:0051379) |
| 0.9 | 4.7 | GO:0004385 | guanylate kinase activity(GO:0004385) |
| 0.9 | 16.6 | GO:0005247 | voltage-gated chloride channel activity(GO:0005247) |
| 0.9 | 9.9 | GO:0016907 | G-protein coupled acetylcholine receptor activity(GO:0016907) |
| 0.8 | 4.6 | GO:0004743 | pyruvate kinase activity(GO:0004743) |
| 0.7 | 10.2 | GO:0070628 | proteasome binding(GO:0070628) |
| 0.7 | 5.0 | GO:0004758 | serine C-palmitoyltransferase activity(GO:0004758) C-palmitoyltransferase activity(GO:0016454) |
| 0.7 | 18.5 | GO:0017080 | sodium channel regulator activity(GO:0017080) |
| 0.7 | 10.8 | GO:0004724 | magnesium-dependent protein serine/threonine phosphatase activity(GO:0004724) |
| 0.7 | 34.2 | GO:0008013 | beta-catenin binding(GO:0008013) |
| 0.6 | 129.6 | GO:0004725 | protein tyrosine phosphatase activity(GO:0004725) |
| 0.6 | 20.4 | GO:0099529 | neurotransmitter receptor activity involved in regulation of postsynaptic membrane potential(GO:0099529) transmitter-gated ion channel activity involved in regulation of postsynaptic membrane potential(GO:1904315) |
| 0.6 | 8.8 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.5 | 4.2 | GO:0004965 | G-protein coupled GABA receptor activity(GO:0004965) |
| 0.4 | 2.2 | GO:0008174 | mRNA methyltransferase activity(GO:0008174) |
| 0.4 | 5.3 | GO:0016868 | intramolecular transferase activity, phosphotransferases(GO:0016868) |
| 0.4 | 6.6 | GO:0030544 | Hsp70 protein binding(GO:0030544) |
| 0.4 | 19.5 | GO:0004683 | calmodulin-dependent protein kinase activity(GO:0004683) |
| 0.4 | 15.0 | GO:0016307 | phosphatidylinositol phosphate kinase activity(GO:0016307) |
| 0.4 | 18.1 | GO:0019707 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.4 | 3.0 | GO:0004784 | superoxide dismutase activity(GO:0004784) oxidoreductase activity, acting on superoxide radicals as acceptor(GO:0016721) |
| 0.4 | 2.6 | GO:0001642 | group III metabotropic glutamate receptor activity(GO:0001642) |
| 0.4 | 1.1 | GO:0015562 | inositol hexakisphosphate binding(GO:0000822) efflux transmembrane transporter activity(GO:0015562) |
| 0.3 | 1.7 | GO:1990756 | protein binding, bridging involved in substrate recognition for ubiquitination(GO:1990756) |
| 0.3 | 8.3 | GO:0035591 | signaling adaptor activity(GO:0035591) |
| 0.3 | 13.7 | GO:0003746 | translation elongation factor activity(GO:0003746) |
| 0.3 | 26.1 | GO:0003777 | microtubule motor activity(GO:0003777) |
| 0.3 | 1.4 | GO:0046592 | polyamine oxidase activity(GO:0046592) |
| 0.3 | 1.6 | GO:0004594 | pantothenate kinase activity(GO:0004594) |
| 0.3 | 91.9 | GO:0015631 | tubulin binding(GO:0015631) |
| 0.3 | 3.2 | GO:0005158 | insulin receptor binding(GO:0005158) |
| 0.2 | 24.9 | GO:0005544 | calcium-dependent phospholipid binding(GO:0005544) |
| 0.2 | 5.2 | GO:0050321 | tau-protein kinase activity(GO:0050321) |
| 0.2 | 5.9 | GO:0004707 | MAP kinase activity(GO:0004707) |
| 0.2 | 10.8 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.2 | 3.6 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.2 | 2.9 | GO:0004169 | dolichyl-phosphate-mannose-protein mannosyltransferase activity(GO:0004169) |
| 0.2 | 2.3 | GO:0003917 | DNA topoisomerase type I activity(GO:0003917) |
| 0.2 | 7.2 | GO:0031593 | polyubiquitin binding(GO:0031593) |
| 0.2 | 0.9 | GO:0005384 | manganese ion transmembrane transporter activity(GO:0005384) |
| 0.2 | 5.9 | GO:0043130 | ubiquitin binding(GO:0043130) |
| 0.1 | 0.7 | GO:0015450 | P-P-bond-hydrolysis-driven protein transmembrane transporter activity(GO:0015450) |
| 0.1 | 48.6 | GO:0003729 | mRNA binding(GO:0003729) poly(A) RNA binding(GO:0044822) |
| 0.1 | 55.0 | GO:0003924 | GTPase activity(GO:0003924) |
| 0.1 | 6.8 | GO:0005178 | integrin binding(GO:0005178) |
| 0.1 | 17.5 | GO:0019901 | protein kinase binding(GO:0019901) |
| 0.1 | 102.3 | GO:0005509 | calcium ion binding(GO:0005509) |
| 0.1 | 19.5 | GO:0015293 | symporter activity(GO:0015293) |
| 0.1 | 4.9 | GO:0004722 | protein serine/threonine phosphatase activity(GO:0004722) |
| 0.1 | 2.4 | GO:0010857 | protein kinase C activity(GO:0004697) calcium-dependent protein kinase C activity(GO:0004698) calcium-dependent protein serine/threonine kinase activity(GO:0009931) calcium-dependent protein kinase activity(GO:0010857) |
| 0.1 | 9.5 | GO:0005516 | calmodulin binding(GO:0005516) |
| 0.1 | 0.3 | GO:0005547 | phosphatidylinositol-3,4,5-trisphosphate binding(GO:0005547) |
| 0.1 | 0.7 | GO:0043035 | chromatin insulator sequence binding(GO:0043035) |
| 0.1 | 1.2 | GO:0070403 | NAD+ binding(GO:0070403) |
| 0.1 | 26.7 | GO:0005096 | GTPase activator activity(GO:0005096) |
| 0.1 | 0.7 | GO:0003689 | DNA clamp loader activity(GO:0003689) protein-DNA loading ATPase activity(GO:0033170) |
| 0.1 | 3.4 | GO:0051287 | NAD binding(GO:0051287) |
| 0.1 | 1.9 | GO:0008381 | mechanically-gated ion channel activity(GO:0008381) mechanically gated channel activity(GO:0022833) |
| 0.1 | 7.7 | GO:0051213 | dioxygenase activity(GO:0051213) |
| 0.1 | 4.1 | GO:0051427 | hormone receptor binding(GO:0051427) |
| 0.1 | 134.3 | GO:0001071 | nucleic acid binding transcription factor activity(GO:0001071) transcription factor activity, sequence-specific DNA binding(GO:0003700) |
| 0.0 | 2.3 | GO:0008020 | G-protein coupled photoreceptor activity(GO:0008020) |
| 0.0 | 1.2 | GO:0005104 | fibroblast growth factor receptor binding(GO:0005104) |
| 0.0 | 0.8 | GO:0004983 | neuropeptide Y receptor activity(GO:0004983) |
| 0.0 | 5.3 | GO:0003714 | transcription corepressor activity(GO:0003714) |
| 0.0 | 0.7 | GO:0045504 | dynein heavy chain binding(GO:0045504) |
| 0.0 | 29.6 | GO:0004674 | protein serine/threonine kinase activity(GO:0004674) |
| 0.0 | 8.9 | GO:0003712 | transcription factor activity, transcription factor binding(GO:0000989) transcription cofactor activity(GO:0003712) |
| 0.0 | 0.4 | GO:0043028 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) cysteine-type endopeptidase regulator activity involved in apoptotic process(GO:0043028) |
| 0.0 | 0.6 | GO:0008137 | NADH dehydrogenase (ubiquinone) activity(GO:0008137) NADH dehydrogenase (quinone) activity(GO:0050136) |
| 0.0 | 1.7 | GO:0019905 | syntaxin binding(GO:0019905) |
| 0.0 | 1.4 | GO:0019888 | protein phosphatase regulator activity(GO:0019888) |
| 0.0 | 0.9 | GO:0008175 | tRNA methyltransferase activity(GO:0008175) |
| 0.0 | 2.9 | GO:0003774 | motor activity(GO:0003774) |
| 0.0 | 1.4 | GO:0016763 | transferase activity, transferring pentosyl groups(GO:0016763) |
| 0.0 | 1.0 | GO:0019902 | phosphatase binding(GO:0019902) |
| 0.0 | 0.4 | GO:0005227 | calcium activated cation channel activity(GO:0005227) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.0 | 13.5 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.8 | 17.0 | ST GRANULE CELL SURVIVAL PATHWAY | Granule Cell Survival Pathway is a specific case of more general PAC1 Receptor Pathway. |
| 0.7 | 5.2 | SA PROGRAMMED CELL DEATH | Programmed cell death, or apoptosis, eliminates damaged or unneeded cells. |
| 0.6 | 10.9 | ST MYOCYTE AD PATHWAY | Myocyte Adrenergic Pathway is a specific case of the generalized Adrenergic Pathway. |
| 0.6 | 10.5 | PID VEGFR1 PATHWAY | VEGFR1 specific signals |
| 0.5 | 7.9 | PID CIRCADIAN PATHWAY | Circadian rhythm pathway |
| 0.4 | 18.5 | PID IL4 2PATHWAY | IL4-mediated signaling events |
| 0.3 | 8.4 | PID P38 ALPHA BETA PATHWAY | Regulation of p38-alpha and p38-beta |
| 0.3 | 6.8 | PID INTEGRIN CS PATHWAY | Integrin family cell surface interactions |
| 0.3 | 28.5 | PID P53 DOWNSTREAM PATHWAY | Direct p53 effectors |
| 0.1 | 4.8 | PID ERBB1 INTERNALIZATION PATHWAY | Internalization of ErbB1 |
| 0.1 | 1.2 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.1 | 2.3 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.1 | 4.0 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.1 | 3.2 | PID CMYB PATHWAY | C-MYB transcription factor network |
| 0.0 | 0.5 | PID CD40 PATHWAY | CD40/CD40L signaling |
| 0.0 | 0.6 | SIG CD40PATHWAYMAP | Genes related to CD40 signaling |
| 0.0 | 0.7 | PID ATR PATHWAY | ATR signaling pathway |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.7 | 17.0 | REACTOME ACTIVATION OF THE AP1 FAMILY OF TRANSCRIPTION FACTORS | Genes involved in Activation of the AP-1 family of transcription factors |
| 1.4 | 16.9 | REACTOME HDL MEDIATED LIPID TRANSPORT | Genes involved in HDL-mediated lipid transport |
| 0.9 | 8.0 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.8 | 10.9 | REACTOME AMINE LIGAND BINDING RECEPTORS | Genes involved in Amine ligand-binding receptors |
| 0.8 | 18.5 | REACTOME G PROTEIN ACTIVATION | Genes involved in G-protein activation |
| 0.6 | 5.8 | REACTOME DCC MEDIATED ATTRACTIVE SIGNALING | Genes involved in DCC mediated attractive signaling |
| 0.6 | 14.4 | REACTOME AMYLOIDS | Genes involved in Amyloids |
| 0.5 | 8.8 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.5 | 4.2 | REACTOME DESTABILIZATION OF MRNA BY AUF1 HNRNP D0 | Genes involved in Destabilization of mRNA by AUF1 (hnRNP D0) |
| 0.4 | 3.4 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 0.4 | 12.4 | REACTOME BIOSYNTHESIS OF THE N GLYCAN PRECURSOR DOLICHOL LIPID LINKED OLIGOSACCHARIDE LLO AND TRANSFER TO A NASCENT PROTEIN | Genes involved in Biosynthesis of the N-glycan precursor (dolichol lipid-linked oligosaccharide, LLO) and transfer to a nascent protein |
| 0.3 | 6.5 | REACTOME SMOOTH MUSCLE CONTRACTION | Genes involved in Smooth Muscle Contraction |
| 0.3 | 12.1 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.2 | 10.8 | REACTOME LOSS OF NLP FROM MITOTIC CENTROSOMES | Genes involved in Loss of Nlp from mitotic centrosomes |
| 0.2 | 2.5 | REACTOME SIGNAL ATTENUATION | Genes involved in Signal attenuation |
| 0.2 | 1.4 | REACTOME PYRIMIDINE CATABOLISM | Genes involved in Pyrimidine catabolism |
| 0.1 | 28.7 | REACTOME ANTIGEN PROCESSING UBIQUITINATION PROTEASOME DEGRADATION | Genes involved in Antigen processing: Ubiquitination & Proteasome degradation |
| 0.1 | 1.5 | REACTOME AMINE DERIVED HORMONES | Genes involved in Amine-derived hormones |
| 0.1 | 6.8 | REACTOME INTEGRIN CELL SURFACE INTERACTIONS | Genes involved in Integrin cell surface interactions |
| 0.1 | 13.0 | REACTOME METABOLISM OF AMINO ACIDS AND DERIVATIVES | Genes involved in Metabolism of amino acids and derivatives |
| 0.1 | 2.7 | REACTOME ENDOSOMAL SORTING COMPLEX REQUIRED FOR TRANSPORT ESCRT | Genes involved in Endosomal Sorting Complex Required For Transport (ESCRT) |
| 0.1 | 1.9 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.1 | 2.1 | REACTOME MRNA SPLICING MINOR PATHWAY | Genes involved in mRNA Splicing - Minor Pathway |
| 0.0 | 0.8 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.0 | 0.6 | REACTOME RIP MEDIATED NFKB ACTIVATION VIA DAI | Genes involved in RIP-mediated NFkB activation via DAI |
| 0.0 | 1.0 | REACTOME NOD1 2 SIGNALING PATHWAY | Genes involved in NOD1/2 Signaling Pathway |
| 0.0 | 0.6 | REACTOME PTM GAMMA CARBOXYLATION HYPUSINE FORMATION AND ARYLSULFATASE ACTIVATION | Genes involved in PTM: gamma carboxylation, hypusine formation and arylsulfatase activation |
| 0.0 | 0.9 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |