Project
PRJDB7713: Age associated transcriptome analysis in 5 tissues of zebrafish
Navigation
Downloads
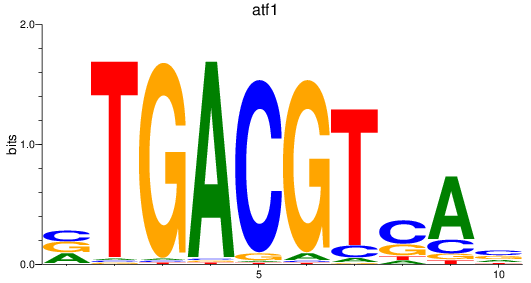
Results for atf1
Z-value: 2.02
Transcription factors associated with atf1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
atf1
|
ENSDARG00000044301 | activating transcription factor 1 |
|
atf1
|
ENSDARG00000109865 | activating transcription factor 1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| atf1 | dr11_v1_chr6_-_39521832_39521832 | 0.61 | 6.4e-11 | Click! |
Activity profile of atf1 motif
Sorted Z-values of atf1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr17_-_12385308 | 38.69 |
ENSDART00000080927
|
snap25b
|
synaptosomal-associated protein, 25b |
| chr6_+_27667359 | 30.65 |
ENSDART00000159624
ENSDART00000049177 |
rab6ba
|
RAB6B, member RAS oncogene family a |
| chr2_-_11512819 | 30.23 |
ENSDART00000142013
|
penka
|
proenkephalin a |
| chr20_+_34913069 | 24.62 |
ENSDART00000007584
|
snap25a
|
synaptosomal-associated protein, 25a |
| chr16_-_29437373 | 22.12 |
ENSDART00000148405
|
si:ch211-113g11.6
|
si:ch211-113g11.6 |
| chr5_+_3501859 | 20.90 |
ENSDART00000080486
|
ywhag1
|
3-monooxygenase/tryptophan 5-monooxygenase activation protein, gamma polypeptide 1 |
| chr23_-_4915118 | 20.88 |
ENSDART00000060714
|
atp6ap1a
|
ATPase H+ transporting accessory protein 1a |
| chr24_-_7632187 | 19.60 |
ENSDART00000041714
|
atp6v0a1b
|
ATPase H+ transporting V0 subunit a1b |
| chr24_-_6158933 | 19.57 |
ENSDART00000021609
|
gad2
|
glutamate decarboxylase 2 |
| chr23_-_29667716 | 19.38 |
ENSDART00000158302
ENSDART00000133902 |
clstn1
|
calsyntenin 1 |
| chr9_-_296169 | 18.70 |
ENSDART00000165228
|
kif5aa
|
kinesin family member 5A, a |
| chr23_+_28731379 | 18.34 |
ENSDART00000047378
|
cort
|
cortistatin |
| chr2_-_42415902 | 17.84 |
ENSDART00000142489
|
slco5a1b
|
solute carrier organic anion transporter family member 5A1b |
| chr12_-_19103490 | 17.67 |
ENSDART00000060561
|
csdc2a
|
cold shock domain containing C2, RNA binding a |
| chr7_+_29133321 | 17.48 |
ENSDART00000052346
|
gnao1b
|
guanine nucleotide binding protein (G protein), alpha activating activity polypeptide O, b |
| chr4_+_9669717 | 17.43 |
ENSDART00000004604
|
si:dkey-153k10.9
|
si:dkey-153k10.9 |
| chr7_+_39166460 | 17.29 |
ENSDART00000052318
ENSDART00000146635 ENSDART00000173877 ENSDART00000173767 ENSDART00000173600 |
mdka
|
midkine a |
| chr10_-_22845485 | 17.13 |
ENSDART00000079454
|
vamp2
|
vesicle-associated membrane protein 2 |
| chr7_+_26224211 | 16.86 |
ENSDART00000173999
|
vgf
|
VGF nerve growth factor inducible |
| chr3_+_15907297 | 16.86 |
ENSDART00000139206
|
mapk8ip3
|
mitogen-activated protein kinase 8 interacting protein 3 |
| chr25_-_37084032 | 16.58 |
ENSDART00000025494
|
hprt1l
|
hypoxanthine phosphoribosyltransferase 1, like |
| chr19_+_24882845 | 16.48 |
ENSDART00000010580
|
si:ch211-195b13.1
|
si:ch211-195b13.1 |
| chr20_+_20637866 | 15.99 |
ENSDART00000060203
ENSDART00000079079 |
rtn1b
|
reticulon 1b |
| chr7_+_40228422 | 15.65 |
ENSDART00000052222
|
ptprn2
|
protein tyrosine phosphatase, receptor type, N polypeptide 2 |
| chr19_-_43552252 | 15.52 |
ENSDART00000138308
|
gpr186
|
G protein-coupled receptor 186 |
| chr15_-_163586 | 15.44 |
ENSDART00000163597
|
SEPT4
|
septin-4 |
| chr4_-_12388535 | 14.94 |
ENSDART00000017180
|
rergla
|
RERG/RAS-like a |
| chr10_-_24371312 | 14.83 |
ENSDART00000149362
|
pitpnab
|
phosphatidylinositol transfer protein, alpha b |
| chr20_+_29743904 | 14.78 |
ENSDART00000146366
ENSDART00000153154 |
kidins220b
|
kinase D-interacting substrate 220b |
| chr1_-_56223913 | 14.76 |
ENSDART00000019573
|
zgc:65894
|
zgc:65894 |
| chr20_+_20638034 | 14.23 |
ENSDART00000189759
|
rtn1b
|
reticulon 1b |
| chr6_-_31348999 | 14.20 |
ENSDART00000153734
|
dnajc6
|
DnaJ (Hsp40) homolog, subfamily C, member 6 |
| chr17_-_20979077 | 14.09 |
ENSDART00000006676
|
phyhipla
|
phytanoyl-CoA 2-hydroxylase interacting protein-like a |
| chr16_+_34523515 | 14.01 |
ENSDART00000041007
|
stmn1b
|
stathmin 1b |
| chr15_-_27710513 | 13.95 |
ENSDART00000005641
ENSDART00000134373 |
lhx1a
|
LIM homeobox 1a |
| chr5_+_37056818 | 13.83 |
ENSDART00000036760
|
tppp2
|
tubulin polymerization-promoting protein family member 2 |
| chr5_-_45877387 | 13.76 |
ENSDART00000183714
ENSDART00000041503 |
slc4a4a
|
solute carrier family 4 (sodium bicarbonate cotransporter), member 4a |
| chr3_-_49566364 | 13.47 |
ENSDART00000161507
|
zgc:153426
|
zgc:153426 |
| chr16_-_9869056 | 13.17 |
ENSDART00000149312
|
ncalda
|
neurocalcin delta a |
| chr19_+_10396042 | 13.14 |
ENSDART00000028048
ENSDART00000151735 |
necap1
|
NECAP endocytosis associated 1 |
| chr1_-_38756870 | 12.89 |
ENSDART00000130324
ENSDART00000148404 |
gpm6ab
|
glycoprotein M6Ab |
| chr14_-_2270973 | 12.67 |
ENSDART00000180729
|
pcdh2ab9
|
protocadherin 2 alpha b 9 |
| chr17_-_26911852 | 12.59 |
ENSDART00000045842
|
rcan3
|
regulator of calcineurin 3 |
| chr12_-_30443562 | 12.47 |
ENSDART00000020769
|
adrb1
|
adrenoceptor beta 1 |
| chr15_+_28685892 | 12.46 |
ENSDART00000155815
ENSDART00000060244 |
nova2
|
neuro-oncological ventral antigen 2 |
| chr16_+_36906693 | 12.37 |
ENSDART00000160645
|
si:ch73-215d9.1
|
si:ch73-215d9.1 |
| chr20_+_41549200 | 12.26 |
ENSDART00000135715
|
fam184a
|
family with sequence similarity 184, member A |
| chr17_+_15535501 | 12.23 |
ENSDART00000002932
|
marcksb
|
myristoylated alanine-rich protein kinase C substrate b |
| chr20_+_38724575 | 12.18 |
ENSDART00000015095
ENSDART00000152972 |
uts1
|
urotensin 1 |
| chr14_-_7888748 | 12.08 |
ENSDART00000166293
|
ppp3cb
|
protein phosphatase 3, catalytic subunit, beta isozyme |
| chr5_-_45876908 | 11.78 |
ENSDART00000188229
|
slc4a4a
|
solute carrier family 4 (sodium bicarbonate cotransporter), member 4a |
| chr22_-_29922872 | 11.53 |
ENSDART00000020249
|
dusp5
|
dual specificity phosphatase 5 |
| chr6_+_24817852 | 11.47 |
ENSDART00000165609
|
barhl2
|
BarH-like homeobox 2 |
| chr25_-_31863374 | 11.45 |
ENSDART00000028338
|
scamp5a
|
secretory carrier membrane protein 5a |
| chr21_-_43606502 | 11.27 |
ENSDART00000151030
|
si:ch73-362m14.4
|
si:ch73-362m14.4 |
| chr18_-_14743659 | 11.05 |
ENSDART00000125979
|
tshz3a
|
teashirt zinc finger homeobox 3a |
| chr23_-_29667544 | 10.96 |
ENSDART00000059339
|
clstn1
|
calsyntenin 1 |
| chr14_-_48939560 | 10.94 |
ENSDART00000021736
|
scocb
|
short coiled-coil protein b |
| chr24_-_21172122 | 10.90 |
ENSDART00000154259
|
atp6v1ab
|
ATPase H+ transporting V1 subunit Ab |
| chr22_+_1796057 | 10.87 |
ENSDART00000170834
|
znf1179
|
zinc finger protein 1179 |
| chr8_+_21588067 | 10.84 |
ENSDART00000172190
|
ajap1
|
adherens junctions associated protein 1 |
| chr18_+_40381102 | 10.80 |
ENSDART00000136588
|
sema6dl
|
sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6D, like |
| chr21_-_23308286 | 10.78 |
ENSDART00000184419
|
zbtb16a
|
zinc finger and BTB domain containing 16a |
| chr11_-_13341483 | 10.73 |
ENSDART00000164978
|
mast3b
|
microtubule associated serine/threonine kinase 3b |
| chr18_-_22094102 | 10.71 |
ENSDART00000100904
|
pard6a
|
par-6 family cell polarity regulator alpha |
| chr18_+_21408794 | 10.71 |
ENSDART00000140161
|
necab2
|
N-terminal EF-hand calcium binding protein 2 |
| chr19_+_46222428 | 10.52 |
ENSDART00000183984
|
vps28
|
vacuolar protein sorting 28 (yeast) |
| chr19_+_16222618 | 10.46 |
ENSDART00000137189
ENSDART00000169246 ENSDART00000190583 ENSDART00000189521 |
ptprua
|
protein tyrosine phosphatase, receptor type, U, a |
| chr20_-_9462433 | 10.22 |
ENSDART00000152674
ENSDART00000040557 |
zgc:101840
|
zgc:101840 |
| chr3_+_32933663 | 10.22 |
ENSDART00000112742
|
nbr1b
|
neighbor of brca1 gene 1b |
| chr5_-_20814576 | 10.21 |
ENSDART00000098682
ENSDART00000147639 |
si:ch211-225b11.1
|
si:ch211-225b11.1 |
| chr2_+_20332044 | 10.11 |
ENSDART00000112131
|
plppr4a
|
phospholipid phosphatase related 4a |
| chr3_-_1204341 | 10.11 |
ENSDART00000089646
|
fam234b
|
family with sequence similarity 234, member B |
| chr10_-_36591511 | 10.06 |
ENSDART00000063347
|
slc1a3b
|
solute carrier family 1 (glial high affinity glutamate transporter), member 3b |
| chr3_-_13147310 | 10.06 |
ENSDART00000160840
|
prkar1b
|
protein kinase, cAMP-dependent, regulatory, type I, beta |
| chr16_-_45069882 | 9.98 |
ENSDART00000058384
|
gapdhs
|
glyceraldehyde-3-phosphate dehydrogenase, spermatogenic |
| chr1_-_22861348 | 9.94 |
ENSDART00000139412
|
SMIM18
|
si:dkey-92j12.6 |
| chr20_+_3108597 | 9.89 |
ENSDART00000133435
|
CEP170B (1 of many)
|
si:ch73-212j7.1 |
| chr10_+_23022263 | 9.88 |
ENSDART00000138955
|
si:dkey-175g6.2
|
si:dkey-175g6.2 |
| chr8_-_17067364 | 9.85 |
ENSDART00000132687
|
rab3c
|
RAB3C, member RAS oncogene family |
| chr7_-_12065668 | 9.74 |
ENSDART00000101537
|
mex3b
|
mex-3 RNA binding family member B |
| chr8_-_54223316 | 9.67 |
ENSDART00000018054
|
trh
|
thyrotropin-releasing hormone |
| chr1_+_45323400 | 9.65 |
ENSDART00000148906
ENSDART00000132366 |
emp1
|
epithelial membrane protein 1 |
| chr11_+_4026229 | 9.64 |
ENSDART00000041417
|
camk1b
|
calcium/calmodulin-dependent protein kinase Ib |
| chr1_+_9290103 | 9.60 |
ENSDART00000055009
|
uncx4.1
|
Unc4.1 homeobox (C. elegans) |
| chr2_+_35603637 | 9.57 |
ENSDART00000147278
|
plk3
|
polo-like kinase 3 (Drosophila) |
| chr11_+_40649412 | 9.57 |
ENSDART00000043016
ENSDART00000134560 |
slc45a1
|
solute carrier family 45, member 1 |
| chr4_-_69189894 | 9.49 |
ENSDART00000169596
|
si:ch211-209j12.1
|
si:ch211-209j12.1 |
| chr10_+_37145007 | 9.36 |
ENSDART00000131777
|
cuedc1a
|
CUE domain containing 1a |
| chr17_-_15382704 | 9.34 |
ENSDART00000005313
|
zgc:85722
|
zgc:85722 |
| chr25_-_31763897 | 9.33 |
ENSDART00000041740
|
ubl7a
|
ubiquitin-like 7a (bone marrow stromal cell-derived) |
| chr6_+_23809163 | 9.30 |
ENSDART00000170402
|
glulb
|
glutamate-ammonia ligase (glutamine synthase) b |
| chr1_+_10305611 | 9.27 |
ENSDART00000043881
|
zgc:77880
|
zgc:77880 |
| chr4_-_5764255 | 9.20 |
ENSDART00000113864
|
faxca
|
failed axon connections homolog a |
| chr2_+_20331445 | 9.17 |
ENSDART00000186880
|
plppr4a
|
phospholipid phosphatase related 4a |
| chr9_+_22632126 | 9.07 |
ENSDART00000139434
|
etv5a
|
ets variant 5a |
| chr9_-_54840124 | 8.95 |
ENSDART00000137214
ENSDART00000085693 |
gpm6bb
|
glycoprotein M6Bb |
| chr25_+_35683956 | 8.72 |
ENSDART00000149768
|
kif21a
|
kinesin family member 21A |
| chr9_+_22631672 | 8.64 |
ENSDART00000101770
ENSDART00000126015 ENSDART00000145005 |
etv5a
|
ets variant 5a |
| chr19_-_32888758 | 8.62 |
ENSDART00000052080
|
laptm4b
|
lysosomal protein transmembrane 4 beta |
| chr6_-_25952848 | 8.61 |
ENSDART00000076997
ENSDART00000148748 |
lmo4b
|
LIM domain only 4b |
| chr17_+_8184649 | 8.61 |
ENSDART00000091818
|
tulp4b
|
tubby like protein 4b |
| chr25_-_23052707 | 8.58 |
ENSDART00000024633
|
dusp8a
|
dual specificity phosphatase 8a |
| chr17_-_11329959 | 8.49 |
ENSDART00000015418
|
irf2bpl
|
interferon regulatory factor 2 binding protein-like |
| chr16_+_25137483 | 8.46 |
ENSDART00000155666
|
znf576.1
|
zinc finger protein 576, tandem duplicate 1 |
| chr12_+_27243059 | 8.42 |
ENSDART00000066269
|
arl4d
|
ADP-ribosylation factor-like 4D |
| chr13_-_40499296 | 8.41 |
ENSDART00000158338
|
CNNM1
|
Danio rerio cyclin and CBS domain divalent metal cation transport mediator 1 (cnnm1), mRNA. |
| chr12_-_41618844 | 8.34 |
ENSDART00000160054
|
dpysl4
|
dihydropyrimidinase-like 4 |
| chr4_+_73973242 | 8.32 |
ENSDART00000182529
|
LO018181.1
|
|
| chr8_-_39984593 | 8.26 |
ENSDART00000140127
|
asphd2
|
aspartate beta-hydroxylase domain containing 2 |
| chr22_-_38621438 | 8.15 |
ENSDART00000098330
|
nppc
|
natriuretic peptide C |
| chr12_+_16953415 | 8.09 |
ENSDART00000020824
|
pank1b
|
pantothenate kinase 1b |
| chr5_-_24000211 | 8.09 |
ENSDART00000188865
|
map7d2a
|
MAP7 domain containing 2a |
| chr16_-_17162843 | 8.04 |
ENSDART00000089386
|
iffo1b
|
intermediate filament family orphan 1b |
| chr20_-_1191910 | 7.93 |
ENSDART00000043218
|
ube2j1
|
ubiquitin-conjugating enzyme E2, J1 |
| chr20_+_45741566 | 7.92 |
ENSDART00000113454
|
chgb
|
chromogranin B |
| chr8_-_17771755 | 7.92 |
ENSDART00000063592
|
prkcz
|
protein kinase C, zeta |
| chr13_-_4333181 | 7.85 |
ENSDART00000122406
|
znf318
|
zinc finger protein 318 |
| chr5_-_9540641 | 7.84 |
ENSDART00000124384
ENSDART00000160079 |
gak
|
cyclin G associated kinase |
| chr17_+_28005763 | 7.82 |
ENSDART00000155838
|
luzp1
|
leucine zipper protein 1 |
| chr5_+_62374092 | 7.78 |
ENSDART00000082965
|
CU928117.1
|
|
| chr16_-_54942532 | 7.77 |
ENSDART00000078887
ENSDART00000101402 |
tmem222a
|
transmembrane protein 222a |
| chr6_+_23809501 | 7.73 |
ENSDART00000168701
|
glulb
|
glutamate-ammonia ligase (glutamine synthase) b |
| chr24_-_6375774 | 7.70 |
ENSDART00000013294
|
anxa13
|
annexin A13 |
| chr7_+_13382852 | 7.70 |
ENSDART00000166318
|
dagla
|
diacylglycerol lipase, alpha |
| chr21_-_14878220 | 7.68 |
ENSDART00000131237
|
ulk1b
|
unc-51 like autophagy activating kinase 1 |
| chr9_-_48296469 | 7.61 |
ENSDART00000058255
|
bbs5
|
Bardet-Biedl syndrome 5 |
| chr21_+_11684830 | 7.60 |
ENSDART00000147473
|
pcsk1
|
proprotein convertase subtilisin/kexin type 1 |
| chr23_+_17354154 | 7.55 |
ENSDART00000155808
|
SRMS
|
zgc:194282 |
| chr19_-_12193622 | 7.48 |
ENSDART00000041960
|
ywhaz
|
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, zeta polypeptide |
| chr7_-_48805181 | 7.44 |
ENSDART00000015884
|
mfge8a
|
milk fat globule-EGF factor 8 protein a |
| chr10_+_42374957 | 7.41 |
ENSDART00000147926
|
COX5B
|
zgc:86599 |
| chr4_-_16124417 | 7.41 |
ENSDART00000128079
ENSDART00000077664 |
atp2b1a
|
ATPase plasma membrane Ca2+ transporting 1a |
| chr4_-_25515513 | 7.40 |
ENSDART00000142276
ENSDART00000044043 |
rbm17
|
RNA binding motif protein 17 |
| chr25_-_24074500 | 7.36 |
ENSDART00000040410
|
th
|
tyrosine hydroxylase |
| chr20_-_44576949 | 7.36 |
ENSDART00000148639
|
ubxn2a
|
UBX domain protein 2A |
| chr13_+_28705143 | 7.33 |
ENSDART00000183338
|
ldb1a
|
LIM domain binding 1a |
| chr15_-_762319 | 7.26 |
ENSDART00000154306
ENSDART00000157492 |
si:dkey-7i4.16
znf1011
|
si:dkey-7i4.16 zinc finger protein 1011 |
| chr2_+_22694382 | 7.23 |
ENSDART00000139196
|
kif1ab
|
kinesin family member 1Ab |
| chr24_+_12989727 | 7.16 |
ENSDART00000126842
ENSDART00000129309 |
flj11011l
|
hypothetical protein FLJ11011-like (H. sapiens) |
| chr4_+_77060861 | 7.13 |
ENSDART00000174271
ENSDART00000174393 ENSDART00000150450 |
si:dkey-240n22.8
|
si:dkey-240n22.8 |
| chr15_-_1687204 | 7.13 |
ENSDART00000155579
ENSDART00000181382 |
sptssb
|
serine palmitoyltransferase, small subunit B |
| chr17_+_32500387 | 7.09 |
ENSDART00000018423
|
ywhaqb
|
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, theta polypeptide b |
| chr24_-_8261786 | 7.06 |
ENSDART00000106388
|
eef1e1
|
eukaryotic translation elongation factor 1 epsilon 1 |
| chr23_+_19590006 | 7.05 |
ENSDART00000021231
|
slmapb
|
sarcolemma associated protein b |
| chr23_-_30764319 | 6.97 |
ENSDART00000075918
|
pcmtd2
|
protein-L-isoaspartate (D-aspartate) O-methyltransferase domain containing 2 |
| chr3_-_21137362 | 6.94 |
ENSDART00000104051
|
cdipt
|
CDP-diacylglycerol--inositol 3-phosphatidyltransferase (phosphatidylinositol synthase) |
| chr2_+_41524238 | 6.90 |
ENSDART00000122860
ENSDART00000017977 |
acvr1l
|
activin A receptor, type 1 like |
| chr22_-_36829006 | 6.90 |
ENSDART00000170256
ENSDART00000086064 |
map1sa
|
microtubule-associated protein 1Sa |
| chr14_+_31618982 | 6.89 |
ENSDART00000026195
|
slc9a6a
|
solute carrier family 9, subfamily A (NHE6, cation proton antiporter 6), member 6a |
| chr5_+_57743815 | 6.88 |
ENSDART00000005090
|
alg9
|
ALG9, alpha-1,2-mannosyltransferase |
| chr2_+_12255568 | 6.88 |
ENSDART00000184164
ENSDART00000013454 |
prtfdc1
|
phosphoribosyl transferase domain containing 1 |
| chr11_-_6970107 | 6.87 |
ENSDART00000171255
|
COMP
|
si:ch211-43f4.1 |
| chr3_-_11624694 | 6.80 |
ENSDART00000127157
|
hlfa
|
hepatic leukemia factor a |
| chr9_-_18743012 | 6.80 |
ENSDART00000131626
|
tsc22d1
|
TSC22 domain family, member 1 |
| chr3_-_21037840 | 6.79 |
ENSDART00000002393
|
rundc3aa
|
RUN domain containing 3Aa |
| chr19_+_45970692 | 6.79 |
ENSDART00000158781
|
si:ch211-153f2.7
|
si:ch211-153f2.7 |
| chr6_-_46403475 | 6.78 |
ENSDART00000154148
|
camk1a
|
calcium/calmodulin-dependent protein kinase Ia |
| chr22_+_2709467 | 6.77 |
ENSDART00000141416
|
znf1171
|
zinc finger protein 1171 |
| chr7_+_15872357 | 6.76 |
ENSDART00000165757
|
pax6b
|
paired box 6b |
| chr14_+_23970818 | 6.75 |
ENSDART00000123338
ENSDART00000124944 |
kif3a
|
kinesin family member 3A |
| chr3_-_45308394 | 6.74 |
ENSDART00000155324
|
pdpk1a
|
3-phosphoinositide dependent protein kinase 1a |
| chr9_-_18742704 | 6.70 |
ENSDART00000145401
|
tsc22d1
|
TSC22 domain family, member 1 |
| chr2_-_58201173 | 6.67 |
ENSDART00000166282
|
pnp5b
|
purine nucleoside phosphorylase 5b |
| chr9_+_6997861 | 6.62 |
ENSDART00000190491
|
inpp4aa
|
inositol polyphosphate-4-phosphatase type I Aa |
| chr19_-_10395683 | 6.62 |
ENSDART00000109488
|
zgc:194578
|
zgc:194578 |
| chr11_-_18800299 | 6.61 |
ENSDART00000156276
|
pfkfb2b
|
6-phosphofructo-2-kinase/fructose-2,6-biphosphatase 2b |
| chr24_+_4977862 | 6.59 |
ENSDART00000114537
|
zic4
|
zic family member 4 |
| chr9_+_6998082 | 6.54 |
ENSDART00000092480
ENSDART00000135576 ENSDART00000188884 |
inpp4aa
|
inositol polyphosphate-4-phosphatase type I Aa |
| chr17_+_29345606 | 6.53 |
ENSDART00000086164
|
kctd3
|
potassium channel tetramerization domain containing 3 |
| chr6_-_58828398 | 6.53 |
ENSDART00000090634
|
kif5ab
|
kinesin family member 5A, b |
| chr9_+_42546504 | 6.50 |
ENSDART00000192224
|
gulp1a
|
GULP, engulfment adaptor PTB domain containing 1a |
| chr15_+_1004680 | 6.50 |
ENSDART00000157310
|
si:dkey-77f5.8
|
si:dkey-77f5.8 |
| chr24_+_32472155 | 6.48 |
ENSDART00000098859
|
neurod6a
|
neuronal differentiation 6a |
| chr15_-_739229 | 6.46 |
ENSDART00000153874
|
si:dkey-7i4.19
|
si:dkey-7i4.19 |
| chr5_-_23999777 | 6.46 |
ENSDART00000085969
|
map7d2a
|
MAP7 domain containing 2a |
| chr9_-_31524907 | 6.45 |
ENSDART00000142904
ENSDART00000127214 ENSDART00000133427 ENSDART00000146268 ENSDART00000182541 ENSDART00000184736 |
tmtc4
|
transmembrane and tetratricopeptide repeat containing 4 |
| chr18_+_17493859 | 6.44 |
ENSDART00000090754
|
si:dkey-102f14.5
|
si:dkey-102f14.5 |
| chr18_-_21859019 | 6.43 |
ENSDART00000100885
|
nrn1la
|
neuritin 1-like a |
| chr14_-_33454595 | 6.41 |
ENSDART00000109615
ENSDART00000173267 ENSDART00000185737 ENSDART00000190989 |
tmem255a
|
transmembrane protein 255A |
| chr12_-_10421955 | 6.41 |
ENSDART00000052004
|
zgc:153595
|
zgc:153595 |
| chr14_+_36226243 | 6.37 |
ENSDART00000052569
|
pitx2
|
paired-like homeodomain 2 |
| chr7_+_67486807 | 6.36 |
ENSDART00000159989
|
cpne7
|
copine VII |
| chr9_-_11655031 | 6.30 |
ENSDART00000044314
|
itgav
|
integrin, alpha V |
| chr16_+_24681177 | 6.24 |
ENSDART00000058956
ENSDART00000189335 |
ywhabl
|
tyrosine 3-monooxygenase/tryptophan 5-monooxygenase activation protein, beta polypeptide like |
| chr3_-_62393449 | 6.24 |
ENSDART00000101870
ENSDART00000140782 ENSDART00000181704 |
proza
|
protein Z, vitamin K-dependent plasma glycoprotein a |
| chr12_+_32159272 | 6.19 |
ENSDART00000153167
|
hlfb
|
hepatic leukemia factor b |
| chr12_-_35505610 | 6.14 |
ENSDART00000105518
|
camk2g1
|
calcium/calmodulin-dependent protein kinase (CaM kinase) II gamma 1 |
| chr1_+_41666611 | 6.14 |
ENSDART00000145789
|
fbxo41
|
F-box protein 41 |
| chr1_-_19215336 | 6.10 |
ENSDART00000162949
ENSDART00000170680 |
ptprdb
|
protein tyrosine phosphatase, receptor type, D, b |
| chr21_+_11685009 | 6.10 |
ENSDART00000014668
|
pcsk1
|
proprotein convertase subtilisin/kexin type 1 |
| chr12_-_41619257 | 6.08 |
ENSDART00000162967
|
dpysl4
|
dihydropyrimidinase-like 4 |
| chr4_-_2525916 | 6.07 |
ENSDART00000134123
ENSDART00000132581 ENSDART00000019508 |
csrp2
|
cysteine and glycine-rich protein 2 |
| chr15_+_5116179 | 6.06 |
ENSDART00000101937
|
pgm2l1
|
phosphoglucomutase 2-like 1 |
| chr6_-_6673813 | 5.99 |
ENSDART00000150995
|
si:dkey-261j15.2
|
si:dkey-261j15.2 |
| chr20_-_21994901 | 5.96 |
ENSDART00000004984
|
daam1b
|
dishevelled associated activator of morphogenesis 1b |
| chr8_+_44714336 | 5.95 |
ENSDART00000145801
|
elmod3
|
ELMO/CED-12 domain containing 3 |
| chr13_+_23843712 | 5.95 |
ENSDART00000057611
|
oprm1
|
opioid receptor, mu 1 |
| chr15_-_28200049 | 5.92 |
ENSDART00000004200
|
sarm1
|
sterile alpha and TIR motif containing 1 |
| chr22_-_718615 | 5.87 |
ENSDART00000149320
|
arl8a
|
ADP-ribosylation factor-like 8A |
Network of associatons between targets according to the STRING database.
First level regulatory network of atf1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 7.6 | 30.3 | GO:0030951 | establishment or maintenance of microtubule cytoskeleton polarity(GO:0030951) |
| 6.5 | 19.6 | GO:0009449 | gamma-aminobutyric acid biosynthetic process(GO:0009449) |
| 6.2 | 18.6 | GO:1903792 | negative regulation of anion transport(GO:1903792) |
| 5.5 | 16.6 | GO:0032263 | guanine salvage(GO:0006178) GMP salvage(GO:0032263) guanine biosynthetic process(GO:0046099) |
| 5.2 | 47.0 | GO:0099640 | axo-dendritic protein transport(GO:0099640) |
| 5.0 | 15.1 | GO:0061217 | regulation of morphogenesis of a branching structure(GO:0060688) positive regulation of mesonephros development(GO:0061213) regulation of mesonephros development(GO:0061217) regulation of kidney development(GO:0090183) positive regulation of kidney development(GO:0090184) regulation of branching involved in ureteric bud morphogenesis(GO:0090189) positive regulation of branching involved in ureteric bud morphogenesis(GO:0090190) |
| 3.7 | 22.0 | GO:0072318 | clathrin coat disassembly(GO:0072318) vesicle uncoating(GO:0072319) |
| 2.8 | 38.7 | GO:0045162 | clustering of voltage-gated sodium channels(GO:0045162) |
| 2.5 | 12.5 | GO:0002025 | vasodilation by norepinephrine-epinephrine involved in regulation of systemic arterial blood pressure(GO:0002025) |
| 2.5 | 7.4 | GO:0042416 | dopamine biosynthetic process from tyrosine(GO:0006585) dopamine biosynthetic process(GO:0042416) |
| 2.4 | 17.0 | GO:0006542 | glutamine biosynthetic process(GO:0006542) |
| 2.4 | 9.6 | GO:0048313 | organelle inheritance(GO:0048308) Golgi inheritance(GO:0048313) |
| 2.0 | 5.9 | GO:0019364 | pyridine nucleotide catabolic process(GO:0019364) NAD catabolic process(GO:0019677) pyridine-containing compound catabolic process(GO:0072526) |
| 1.9 | 5.8 | GO:1901223 | negative regulation of NIK/NF-kappaB signaling(GO:1901223) |
| 1.9 | 11.7 | GO:0021534 | cell proliferation in hindbrain(GO:0021534) |
| 1.9 | 7.7 | GO:0098920 | endocannabinoid signaling pathway(GO:0071926) retrograde trans-synaptic signaling by lipid(GO:0098920) retrograde trans-synaptic signaling by endocannabinoid(GO:0098921) |
| 1.9 | 7.6 | GO:0072116 | kidney rudiment formation(GO:0072003) pronephros formation(GO:0072116) |
| 1.9 | 5.6 | GO:0031630 | regulation of synaptic vesicle fusion to presynaptic membrane(GO:0031630) |
| 1.8 | 20.0 | GO:0060217 | hemangioblast cell differentiation(GO:0060217) |
| 1.8 | 7.3 | GO:0021742 | abducens nucleus development(GO:0021742) |
| 1.8 | 16.2 | GO:0043328 | protein targeting to vacuole involved in ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway(GO:0043328) |
| 1.7 | 6.9 | GO:0071788 | endoplasmic reticulum tubular network maintenance(GO:0071788) |
| 1.6 | 10.9 | GO:0061635 | regulation of protein complex stability(GO:0061635) |
| 1.5 | 10.7 | GO:0042983 | amyloid precursor protein biosynthetic process(GO:0042983) regulation of amyloid precursor protein biosynthetic process(GO:0042984) |
| 1.5 | 5.9 | GO:0038003 | opioid receptor signaling pathway(GO:0038003) |
| 1.5 | 7.4 | GO:0034505 | tooth mineralization(GO:0034505) |
| 1.4 | 4.1 | GO:0015760 | hexose phosphate transport(GO:0015712) glucose-6-phosphate transport(GO:0015760) |
| 1.3 | 18.9 | GO:0035970 | peptidyl-threonine dephosphorylation(GO:0035970) |
| 1.3 | 13.1 | GO:0061303 | cornea development in camera-type eye(GO:0061303) |
| 1.2 | 3.7 | GO:0044878 | abscission(GO:0009838) mitotic cytokinesis checkpoint(GO:0044878) |
| 1.2 | 8.3 | GO:1901798 | positive regulation of DNA damage response, signal transduction by p53 class mediator(GO:0043517) positive regulation of signal transduction by p53 class mediator(GO:1901798) |
| 1.1 | 17.1 | GO:0035493 | SNARE complex assembly(GO:0035493) |
| 1.1 | 4.5 | GO:0035124 | embryonic caudal fin morphogenesis(GO:0035124) |
| 1.1 | 10.0 | GO:0008216 | spermidine metabolic process(GO:0008216) |
| 1.1 | 12.2 | GO:0050714 | positive regulation of protein secretion(GO:0050714) |
| 1.1 | 25.5 | GO:0015701 | bicarbonate transport(GO:0015701) |
| 1.1 | 4.4 | GO:0071632 | optomotor response(GO:0071632) |
| 1.1 | 14.4 | GO:0032481 | positive regulation of type I interferon production(GO:0032481) |
| 1.1 | 3.3 | GO:1903003 | positive regulation of protein deubiquitination(GO:1903003) |
| 1.1 | 3.3 | GO:0048205 | COPI-coated vesicle budding(GO:0035964) Golgi transport vesicle coating(GO:0048200) COPI coating of Golgi vesicle(GO:0048205) |
| 1.1 | 36.1 | GO:0006891 | intra-Golgi vesicle-mediated transport(GO:0006891) |
| 1.1 | 3.2 | GO:0097067 | cellular response to thyroid hormone stimulus(GO:0097067) |
| 1.1 | 6.4 | GO:0010752 | regulation of cGMP-mediated signaling(GO:0010752) |
| 1.0 | 6.3 | GO:0090245 | axis elongation involved in somitogenesis(GO:0090245) |
| 1.0 | 16.6 | GO:0006003 | fructose 2,6-bisphosphate metabolic process(GO:0006003) |
| 1.0 | 17.3 | GO:0070654 | sensory epithelium regeneration(GO:0070654) epithelium regeneration(GO:1990399) |
| 1.0 | 5.0 | GO:1903146 | regulation of mitophagy(GO:1903146) |
| 1.0 | 5.0 | GO:0070973 | protein localization to endoplasmic reticulum exit site(GO:0070973) |
| 1.0 | 28.0 | GO:0016082 | synaptic vesicle priming(GO:0016082) |
| 1.0 | 13.7 | GO:0045851 | vacuolar acidification(GO:0007035) pH reduction(GO:0045851) intracellular pH reduction(GO:0051452) |
| 1.0 | 3.9 | GO:0006272 | leading strand elongation(GO:0006272) |
| 0.9 | 3.7 | GO:0035520 | monoubiquitinated protein deubiquitination(GO:0035520) monoubiquitinated histone deubiquitination(GO:0035521) monoubiquitinated histone H2A deubiquitination(GO:0035522) |
| 0.9 | 13.7 | GO:0016486 | peptide hormone processing(GO:0016486) |
| 0.9 | 4.5 | GO:1990564 | protein polyufmylation(GO:1990564) protein K69-linked ufmylation(GO:1990592) |
| 0.9 | 27.8 | GO:0030641 | regulation of cellular pH(GO:0030641) |
| 0.9 | 4.4 | GO:0042772 | DNA damage response, signal transduction by p53 class mediator resulting in transcription of p21 class mediator(GO:0006978) DNA damage response, signal transduction resulting in transcription(GO:0042772) |
| 0.9 | 4.3 | GO:0071543 | diphosphoinositol polyphosphate metabolic process(GO:0071543) diadenosine pentaphosphate metabolic process(GO:1901906) diadenosine pentaphosphate catabolic process(GO:1901907) diadenosine hexaphosphate metabolic process(GO:1901908) diadenosine hexaphosphate catabolic process(GO:1901909) adenosine 5'-(hexahydrogen pentaphosphate) metabolic process(GO:1901910) adenosine 5'-(hexahydrogen pentaphosphate) catabolic process(GO:1901911) |
| 0.9 | 5.1 | GO:0048260 | positive regulation of receptor-mediated endocytosis(GO:0048260) |
| 0.8 | 4.2 | GO:0007221 | positive regulation of transcription of Notch receptor target(GO:0007221) |
| 0.8 | 3.2 | GO:0048210 | Golgi vesicle fusion to target membrane(GO:0048210) |
| 0.8 | 14.4 | GO:0006208 | pyrimidine nucleobase catabolic process(GO:0006208) |
| 0.8 | 3.2 | GO:0046391 | 5-phosphoribose 1-diphosphate biosynthetic process(GO:0006015) 5-phosphoribose 1-diphosphate metabolic process(GO:0046391) |
| 0.8 | 12.6 | GO:0010738 | regulation of protein kinase A signaling(GO:0010738) |
| 0.8 | 3.1 | GO:0019884 | antigen processing and presentation of exogenous peptide antigen(GO:0002478) antigen processing and presentation of exogenous antigen(GO:0019884) antigen processing and presentation of exogenous peptide antigen via MHC class I(GO:0042590) |
| 0.8 | 2.3 | GO:0000103 | sulfate assimilation(GO:0000103) purine ribonucleoside bisphosphate metabolic process(GO:0034035) purine ribonucleoside bisphosphate biosynthetic process(GO:0034036) 3'-phosphoadenosine 5'-phosphosulfate metabolic process(GO:0050427) 3'-phosphoadenosine 5'-phosphosulfate biosynthetic process(GO:0050428) |
| 0.8 | 3.1 | GO:0010737 | protein kinase A signaling(GO:0010737) |
| 0.8 | 11.3 | GO:2000601 | positive regulation of actin nucleation(GO:0051127) positive regulation of Arp2/3 complex-mediated actin nucleation(GO:2000601) |
| 0.7 | 3.0 | GO:0070987 | error-free translesion synthesis(GO:0070987) |
| 0.7 | 2.1 | GO:0048569 | post-embryonic organ development(GO:0048569) |
| 0.7 | 2.1 | GO:1901407 | regulation of phosphorylation of RNA polymerase II C-terminal domain(GO:1901407) |
| 0.7 | 3.5 | GO:0045737 | positive regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0045737) positive regulation of cyclin-dependent protein kinase activity(GO:1904031) |
| 0.7 | 4.0 | GO:0033169 | histone H3-K9 demethylation(GO:0033169) |
| 0.7 | 5.3 | GO:1902624 | positive regulation of neutrophil migration(GO:1902624) |
| 0.7 | 2.0 | GO:0097676 | histone H3-K36 dimethylation(GO:0097676) |
| 0.6 | 5.1 | GO:0033137 | negative regulation of peptidyl-serine phosphorylation(GO:0033137) |
| 0.6 | 10.7 | GO:0008089 | anterograde axonal transport(GO:0008089) |
| 0.6 | 6.3 | GO:0006054 | N-acetylneuraminate metabolic process(GO:0006054) |
| 0.6 | 12.9 | GO:0051489 | regulation of filopodium assembly(GO:0051489) |
| 0.6 | 6.1 | GO:0042982 | amyloid precursor protein metabolic process(GO:0042982) |
| 0.6 | 2.4 | GO:0071387 | cellular response to cortisol stimulus(GO:0071387) |
| 0.6 | 5.5 | GO:0016237 | lysosomal microautophagy(GO:0016237) piecemeal microautophagy of nucleus(GO:0034727) single-organism membrane invagination(GO:1902534) |
| 0.6 | 3.6 | GO:0045899 | regulation of RNA polymerase II transcriptional preinitiation complex assembly(GO:0045898) positive regulation of RNA polymerase II transcriptional preinitiation complex assembly(GO:0045899) |
| 0.6 | 2.9 | GO:0000012 | single strand break repair(GO:0000012) |
| 0.6 | 4.5 | GO:0014004 | microglia differentiation(GO:0014004) |
| 0.6 | 7.9 | GO:0090049 | regulation of cell migration involved in sprouting angiogenesis(GO:0090049) |
| 0.6 | 5.6 | GO:0070262 | peptidyl-serine dephosphorylation(GO:0070262) |
| 0.6 | 2.2 | GO:0030213 | hyaluronan biosynthetic process(GO:0030213) |
| 0.5 | 2.7 | GO:0046485 | ether lipid metabolic process(GO:0046485) |
| 0.5 | 4.8 | GO:0015015 | heparan sulfate proteoglycan biosynthetic process, enzymatic modification(GO:0015015) |
| 0.5 | 3.2 | GO:0080009 | mRNA methylation(GO:0080009) |
| 0.5 | 4.7 | GO:1904825 | protein localization to microtubule(GO:0035372) protein localization to microtubule plus-end(GO:1904825) |
| 0.5 | 2.6 | GO:0030219 | megakaryocyte differentiation(GO:0030219) |
| 0.5 | 2.0 | GO:0061709 | reticulophagy(GO:0061709) |
| 0.5 | 4.4 | GO:0043162 | ubiquitin-dependent protein catabolic process via the multivesicular body sorting pathway(GO:0043162) |
| 0.5 | 3.3 | GO:2001271 | negative regulation of execution phase of apoptosis(GO:1900118) regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:2001270) negative regulation of cysteine-type endopeptidase activity involved in execution phase of apoptosis(GO:2001271) |
| 0.5 | 6.9 | GO:0043001 | Golgi to plasma membrane protein transport(GO:0043001) |
| 0.5 | 30.9 | GO:0048841 | regulation of axon extension involved in axon guidance(GO:0048841) |
| 0.4 | 1.8 | GO:0070129 | regulation of mitochondrial translation(GO:0070129) positive regulation of mitochondrial translation(GO:0070131) |
| 0.4 | 6.7 | GO:0033173 | calcineurin-NFAT signaling cascade(GO:0033173) |
| 0.4 | 20.9 | GO:0007019 | microtubule depolymerization(GO:0007019) |
| 0.4 | 8.6 | GO:0071108 | protein K48-linked deubiquitination(GO:0071108) |
| 0.4 | 7.1 | GO:0043506 | activation of JUN kinase activity(GO:0007257) regulation of JUN kinase activity(GO:0043506) positive regulation of JUN kinase activity(GO:0043507) |
| 0.4 | 5.7 | GO:0034030 | coenzyme A biosynthetic process(GO:0015937) nucleoside bisphosphate biosynthetic process(GO:0033866) ribonucleoside bisphosphate biosynthetic process(GO:0034030) purine nucleoside bisphosphate biosynthetic process(GO:0034033) |
| 0.4 | 25.8 | GO:0015914 | phospholipid transport(GO:0015914) |
| 0.4 | 4.9 | GO:0048696 | collateral sprouting in absence of injury(GO:0048669) regulation of collateral sprouting in absence of injury(GO:0048696) |
| 0.4 | 4.0 | GO:0000305 | response to superoxide(GO:0000303) response to oxygen radical(GO:0000305) removal of superoxide radicals(GO:0019430) cellular response to oxygen radical(GO:0071450) cellular response to superoxide(GO:0071451) cellular oxidant detoxification(GO:0098869) cellular detoxification(GO:1990748) |
| 0.4 | 2.0 | GO:0023041 | neuronal signal transduction(GO:0023041) |
| 0.4 | 14.4 | GO:0018230 | peptidyl-L-cysteine S-palmitoylation(GO:0018230) peptidyl-S-diacylglycerol-L-cysteine biosynthetic process from peptidyl-cysteine(GO:0018231) |
| 0.4 | 4.4 | GO:0007023 | post-chaperonin tubulin folding pathway(GO:0007023) |
| 0.4 | 4.4 | GO:0034312 | diol biosynthetic process(GO:0034312) sphingosine biosynthetic process(GO:0046512) |
| 0.4 | 5.1 | GO:0042572 | retinol metabolic process(GO:0042572) |
| 0.4 | 12.1 | GO:0048263 | determination of dorsal identity(GO:0048263) |
| 0.4 | 16.3 | GO:0007212 | dopamine receptor signaling pathway(GO:0007212) |
| 0.4 | 1.5 | GO:0010259 | multicellular organism aging(GO:0010259) |
| 0.4 | 4.6 | GO:0051893 | cell-substrate adherens junction assembly(GO:0007045) focal adhesion assembly(GO:0048041) regulation of focal adhesion assembly(GO:0051893) regulation of cell-substrate junction assembly(GO:0090109) |
| 0.4 | 3.8 | GO:0035479 | angioblast cell migration from lateral mesoderm to midline(GO:0035479) |
| 0.4 | 5.3 | GO:0043562 | cellular response to nitrogen starvation(GO:0006995) cellular response to nitrogen levels(GO:0043562) |
| 0.4 | 2.9 | GO:0006513 | protein monoubiquitination(GO:0006513) |
| 0.4 | 1.8 | GO:0019348 | dolichol metabolic process(GO:0019348) |
| 0.4 | 1.8 | GO:0072575 | hepatocyte proliferation(GO:0072574) epithelial cell proliferation involved in liver morphogenesis(GO:0072575) |
| 0.3 | 1.4 | GO:0010586 | miRNA metabolic process(GO:0010586) |
| 0.3 | 4.8 | GO:1905150 | regulation of voltage-gated sodium channel activity(GO:1905150) |
| 0.3 | 1.7 | GO:0051591 | negative regulation of neurotransmitter secretion(GO:0046929) negative regulation of neurotransmitter transport(GO:0051589) response to cAMP(GO:0051591) cellular response to cAMP(GO:0071320) negative regulation of synaptic vesicle transport(GO:1902804) negative regulation of synaptic vesicle exocytosis(GO:2000301) |
| 0.3 | 3.1 | GO:0000056 | ribosomal small subunit export from nucleus(GO:0000056) |
| 0.3 | 1.7 | GO:0097033 | respiratory chain complex III assembly(GO:0017062) mitochondrial respiratory chain complex III assembly(GO:0034551) mitochondrial respiratory chain complex III biogenesis(GO:0097033) |
| 0.3 | 7.3 | GO:0060541 | respiratory system development(GO:0060541) |
| 0.3 | 3.3 | GO:2000780 | negative regulation of DNA repair(GO:0045738) negative regulation of double-strand break repair(GO:2000780) |
| 0.3 | 33.8 | GO:0048675 | axon extension(GO:0048675) |
| 0.3 | 3.8 | GO:2000766 | negative regulation of cytoplasmic translation(GO:2000766) |
| 0.3 | 5.1 | GO:0016926 | protein desumoylation(GO:0016926) |
| 0.3 | 0.9 | GO:0046048 | ribonucleoside diphosphate catabolic process(GO:0009191) pyrimidine ribonucleoside diphosphate metabolic process(GO:0009193) UDP metabolic process(GO:0046048) |
| 0.3 | 1.5 | GO:0010447 | response to acidic pH(GO:0010447) |
| 0.3 | 2.1 | GO:0010172 | embryonic body morphogenesis(GO:0010172) |
| 0.3 | 1.5 | GO:1901841 | regulation of high voltage-gated calcium channel activity(GO:1901841) negative regulation of high voltage-gated calcium channel activity(GO:1901842) |
| 0.3 | 8.7 | GO:0006904 | vesicle docking involved in exocytosis(GO:0006904) |
| 0.3 | 30.2 | GO:0007218 | neuropeptide signaling pathway(GO:0007218) |
| 0.3 | 0.8 | GO:0008612 | peptidyl-lysine modification to peptidyl-hypusine(GO:0008612) |
| 0.3 | 1.3 | GO:0071881 | adenylate cyclase-inhibiting adrenergic receptor signaling pathway(GO:0071881) |
| 0.3 | 1.5 | GO:0036372 | opsin transport(GO:0036372) |
| 0.3 | 11.9 | GO:0001841 | neural tube formation(GO:0001841) |
| 0.2 | 7.4 | GO:0000413 | protein peptidyl-prolyl isomerization(GO:0000413) |
| 0.2 | 3.9 | GO:0031468 | nuclear envelope reassembly(GO:0031468) |
| 0.2 | 1.9 | GO:0021550 | medulla oblongata development(GO:0021550) dorsal motor nucleus of vagus nerve development(GO:0021744) |
| 0.2 | 24.5 | GO:0000380 | alternative mRNA splicing, via spliceosome(GO:0000380) |
| 0.2 | 3.1 | GO:0006346 | methylation-dependent chromatin silencing(GO:0006346) positive regulation of chromatin silencing(GO:0031937) regulation of methylation-dependent chromatin silencing(GO:0090308) positive regulation of methylation-dependent chromatin silencing(GO:0090309) |
| 0.2 | 3.6 | GO:0035269 | protein O-linked mannosylation(GO:0035269) |
| 0.2 | 4.8 | GO:0034626 | fatty acid elongation, saturated fatty acid(GO:0019367) fatty acid elongation, unsaturated fatty acid(GO:0019368) fatty acid elongation, monounsaturated fatty acid(GO:0034625) fatty acid elongation, polyunsaturated fatty acid(GO:0034626) |
| 0.2 | 1.6 | GO:1903729 | regulation of plasma membrane organization(GO:1903729) |
| 0.2 | 2.2 | GO:0042989 | sequestering of actin monomers(GO:0042989) |
| 0.2 | 17.2 | GO:0019722 | calcium-mediated signaling(GO:0019722) |
| 0.2 | 0.7 | GO:0065001 | specification of axis polarity(GO:0065001) positive regulation of feeding behavior(GO:2000253) |
| 0.2 | 3.7 | GO:1900151 | regulation of nuclear-transcribed mRNA poly(A) tail shortening(GO:0060211) positive regulation of nuclear-transcribed mRNA poly(A) tail shortening(GO:0060213) positive regulation of mRNA catabolic process(GO:0061014) regulation of nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay(GO:1900151) positive regulation of nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay(GO:1900153) |
| 0.2 | 1.7 | GO:0051597 | response to methylmercury(GO:0051597) |
| 0.2 | 2.1 | GO:0008655 | pyrimidine-containing compound salvage(GO:0008655) pyrimidine ribonucleotide salvage(GO:0010138) pyrimidine nucleotide salvage(GO:0032262) pyrimidine nucleoside salvage(GO:0043097) UMP salvage(GO:0044206) |
| 0.2 | 2.3 | GO:0030719 | P granule organization(GO:0030719) |
| 0.2 | 1.6 | GO:0042438 | melanin biosynthetic process(GO:0042438) |
| 0.2 | 1.2 | GO:0032488 | Cdc42 protein signal transduction(GO:0032488) |
| 0.2 | 2.2 | GO:0006893 | Golgi to plasma membrane transport(GO:0006893) |
| 0.2 | 2.6 | GO:0002084 | protein depalmitoylation(GO:0002084) macromolecule depalmitoylation(GO:0098734) |
| 0.2 | 3.5 | GO:0048844 | artery morphogenesis(GO:0048844) |
| 0.2 | 1.9 | GO:0050684 | regulation of mRNA processing(GO:0050684) |
| 0.2 | 11.8 | GO:0006096 | glycolytic process(GO:0006096) |
| 0.2 | 6.1 | GO:0072114 | pronephros morphogenesis(GO:0072114) |
| 0.2 | 0.6 | GO:0032239 | regulation of nucleobase-containing compound transport(GO:0032239) positive regulation of nucleobase-containing compound transport(GO:0032241) positive regulation of nucleocytoplasmic transport(GO:0046824) regulation of RNA export from nucleus(GO:0046831) positive regulation of RNA export from nucleus(GO:0046833) |
| 0.2 | 10.7 | GO:0043488 | regulation of mRNA stability(GO:0043488) |
| 0.2 | 2.6 | GO:0040019 | positive regulation of embryonic development(GO:0040019) |
| 0.2 | 1.7 | GO:0060575 | intestinal epithelial cell differentiation(GO:0060575) |
| 0.2 | 3.1 | GO:0051865 | protein autoubiquitination(GO:0051865) |
| 0.2 | 13.6 | GO:0046839 | phospholipid dephosphorylation(GO:0046839) |
| 0.2 | 10.2 | GO:0001819 | positive regulation of cytokine production(GO:0001819) |
| 0.2 | 2.5 | GO:0042407 | cristae formation(GO:0042407) |
| 0.2 | 4.0 | GO:0007168 | receptor guanylyl cyclase signaling pathway(GO:0007168) |
| 0.2 | 2.6 | GO:0032483 | regulation of Rab protein signal transduction(GO:0032483) |
| 0.2 | 1.9 | GO:0071479 | cellular response to ionizing radiation(GO:0071479) |
| 0.2 | 5.2 | GO:0098840 | protein transport along microtubule(GO:0098840) |
| 0.2 | 2.1 | GO:0061026 | cardiac muscle tissue regeneration(GO:0061026) |
| 0.2 | 0.6 | GO:0051881 | regulation of mitochondrial membrane potential(GO:0051881) |
| 0.2 | 1.7 | GO:0010960 | magnesium ion homeostasis(GO:0010960) |
| 0.2 | 0.8 | GO:1990481 | mRNA pseudouridine synthesis(GO:1990481) |
| 0.2 | 1.2 | GO:0010507 | negative regulation of autophagy(GO:0010507) |
| 0.2 | 1.8 | GO:0045842 | positive regulation of mitotic metaphase/anaphase transition(GO:0045842) positive regulation of mitotic sister chromatid separation(GO:1901970) positive regulation of metaphase/anaphase transition of cell cycle(GO:1902101) |
| 0.1 | 41.1 | GO:0006470 | protein dephosphorylation(GO:0006470) |
| 0.1 | 0.6 | GO:0051661 | Golgi localization(GO:0051645) maintenance of centrosome location(GO:0051661) |
| 0.1 | 1.8 | GO:0006896 | Golgi to vacuole transport(GO:0006896) |
| 0.1 | 1.0 | GO:0045736 | negative regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0045736) negative regulation of cyclin-dependent protein kinase activity(GO:1904030) |
| 0.1 | 1.7 | GO:0009313 | oligosaccharide catabolic process(GO:0009313) |
| 0.1 | 10.7 | GO:0030433 | ER-associated ubiquitin-dependent protein catabolic process(GO:0030433) |
| 0.1 | 0.6 | GO:0006531 | aspartate metabolic process(GO:0006531) |
| 0.1 | 1.0 | GO:0001780 | neutrophil homeostasis(GO:0001780) |
| 0.1 | 1.4 | GO:0006685 | sphingomyelin catabolic process(GO:0006685) |
| 0.1 | 0.5 | GO:0046324 | regulation of glucose import(GO:0046324) |
| 0.1 | 2.4 | GO:0016584 | nucleosome positioning(GO:0016584) |
| 0.1 | 6.3 | GO:0042147 | retrograde transport, endosome to Golgi(GO:0042147) |
| 0.1 | 0.6 | GO:0009268 | response to pH(GO:0009268) |
| 0.1 | 1.1 | GO:0031954 | positive regulation of protein autophosphorylation(GO:0031954) |
| 0.1 | 1.9 | GO:0035459 | cargo loading into vesicle(GO:0035459) |
| 0.1 | 4.5 | GO:0042752 | regulation of circadian rhythm(GO:0042752) |
| 0.1 | 4.5 | GO:0007029 | endoplasmic reticulum organization(GO:0007029) |
| 0.1 | 3.6 | GO:0048278 | vesicle docking(GO:0048278) |
| 0.1 | 6.0 | GO:0030514 | negative regulation of BMP signaling pathway(GO:0030514) |
| 0.1 | 3.4 | GO:0060972 | left/right pattern formation(GO:0060972) |
| 0.1 | 2.1 | GO:0010043 | response to zinc ion(GO:0010043) |
| 0.1 | 3.3 | GO:0008214 | protein demethylation(GO:0006482) protein dealkylation(GO:0008214) |
| 0.1 | 2.1 | GO:0030183 | B cell differentiation(GO:0030183) |
| 0.1 | 6.4 | GO:0071277 | cellular response to calcium ion(GO:0071277) |
| 0.1 | 2.5 | GO:0051085 | chaperone mediated protein folding requiring cofactor(GO:0051085) |
| 0.1 | 10.3 | GO:0016236 | macroautophagy(GO:0016236) |
| 0.1 | 0.7 | GO:0072028 | nephron morphogenesis(GO:0072028) |
| 0.1 | 2.7 | GO:0032922 | circadian regulation of gene expression(GO:0032922) |
| 0.1 | 3.8 | GO:0016311 | dephosphorylation(GO:0016311) |
| 0.1 | 6.1 | GO:0045214 | sarcomere organization(GO:0045214) |
| 0.1 | 2.9 | GO:0048268 | clathrin coat assembly(GO:0048268) |
| 0.1 | 2.2 | GO:0006378 | mRNA polyadenylation(GO:0006378) RNA polyadenylation(GO:0043631) |
| 0.1 | 18.6 | GO:0018105 | peptidyl-serine phosphorylation(GO:0018105) |
| 0.1 | 3.6 | GO:0030901 | midbrain development(GO:0030901) |
| 0.1 | 1.6 | GO:0006654 | phosphatidic acid biosynthetic process(GO:0006654) phosphatidic acid metabolic process(GO:0046473) |
| 0.1 | 1.5 | GO:0060841 | venous blood vessel development(GO:0060841) |
| 0.1 | 1.2 | GO:0046549 | retinal cone cell development(GO:0046549) |
| 0.1 | 0.7 | GO:0033314 | mitotic DNA replication checkpoint(GO:0033314) |
| 0.1 | 3.7 | GO:0032012 | ARF protein signal transduction(GO:0032011) regulation of ARF protein signal transduction(GO:0032012) |
| 0.1 | 3.5 | GO:0007254 | JNK cascade(GO:0007254) |
| 0.1 | 2.1 | GO:0032786 | positive regulation of DNA-templated transcription, elongation(GO:0032786) |
| 0.1 | 0.7 | GO:0045947 | negative regulation of translational initiation(GO:0045947) |
| 0.1 | 2.7 | GO:0008286 | insulin receptor signaling pathway(GO:0008286) |
| 0.1 | 1.8 | GO:0051932 | synaptic transmission, GABAergic(GO:0051932) |
| 0.1 | 13.3 | GO:0007189 | adenylate cyclase-activating G-protein coupled receptor signaling pathway(GO:0007189) |
| 0.1 | 1.2 | GO:0071025 | RNA surveillance(GO:0071025) |
| 0.1 | 3.0 | GO:0010950 | positive regulation of endopeptidase activity(GO:0010950) positive regulation of peptidase activity(GO:0010952) |
| 0.1 | 6.0 | GO:0006487 | protein N-linked glycosylation(GO:0006487) |
| 0.1 | 6.9 | GO:0016579 | protein deubiquitination(GO:0016579) |
| 0.1 | 15.2 | GO:0007156 | homophilic cell adhesion via plasma membrane adhesion molecules(GO:0007156) |
| 0.1 | 3.1 | GO:0007338 | single fertilization(GO:0007338) |
| 0.1 | 0.2 | GO:0006369 | termination of RNA polymerase II transcription(GO:0006369) |
| 0.1 | 2.6 | GO:0018298 | protein-chromophore linkage(GO:0018298) |
| 0.1 | 0.5 | GO:0000491 | small nucleolar ribonucleoprotein complex assembly(GO:0000491) |
| 0.1 | 1.7 | GO:0030968 | endoplasmic reticulum unfolded protein response(GO:0030968) |
| 0.1 | 0.2 | GO:0010939 | regulation of necrotic cell death(GO:0010939) regulation of necroptotic process(GO:0060544) negative regulation of necroptotic process(GO:0060546) negative regulation of necrotic cell death(GO:0060547) |
| 0.1 | 6.0 | GO:0046488 | phosphatidylinositol metabolic process(GO:0046488) |
| 0.1 | 8.6 | GO:0030334 | regulation of cell migration(GO:0030334) |
| 0.1 | 3.2 | GO:0008217 | regulation of blood pressure(GO:0008217) |
| 0.1 | 1.3 | GO:0060028 | convergent extension involved in axis elongation(GO:0060028) |
| 0.1 | 1.5 | GO:1903322 | positive regulation of protein ubiquitination(GO:0031398) positive regulation of protein modification by small protein conjugation or removal(GO:1903322) |
| 0.1 | 1.9 | GO:0006367 | transcription initiation from RNA polymerase II promoter(GO:0006367) |
| 0.0 | 0.2 | GO:0015919 | peroxisomal membrane transport(GO:0015919) protein import into peroxisome membrane(GO:0045046) |
| 0.0 | 0.6 | GO:0006596 | polyamine biosynthetic process(GO:0006596) |
| 0.0 | 14.5 | GO:0008380 | RNA splicing(GO:0008380) |
| 0.0 | 2.4 | GO:0042738 | drug metabolic process(GO:0017144) drug catabolic process(GO:0042737) exogenous drug catabolic process(GO:0042738) |
| 0.0 | 4.7 | GO:0006821 | chloride transport(GO:0006821) |
| 0.0 | 5.2 | GO:0006936 | muscle contraction(GO:0006936) |
| 0.0 | 4.8 | GO:0035725 | sodium ion transmembrane transport(GO:0035725) |
| 0.0 | 1.0 | GO:0016574 | histone ubiquitination(GO:0016574) |
| 0.0 | 1.2 | GO:0051028 | mRNA transport(GO:0051028) |
| 0.0 | 0.8 | GO:0070286 | axonemal dynein complex assembly(GO:0070286) |
| 0.0 | 0.5 | GO:0045746 | negative regulation of Notch signaling pathway(GO:0045746) |
| 0.0 | 3.6 | GO:0007626 | locomotory behavior(GO:0007626) |
| 0.0 | 0.3 | GO:0006611 | protein export from nucleus(GO:0006611) |
| 0.0 | 2.6 | GO:0050769 | positive regulation of neurogenesis(GO:0050769) |
| 0.0 | 0.2 | GO:0090311 | regulation of histone deacetylation(GO:0031063) regulation of protein deacetylation(GO:0090311) |
| 0.0 | 0.2 | GO:1905066 | regulation of canonical Wnt signaling pathway involved in heart development(GO:1905066) |
| 0.0 | 1.1 | GO:0006405 | RNA export from nucleus(GO:0006405) |
| 0.0 | 0.4 | GO:0036342 | post-anal tail morphogenesis(GO:0036342) |
| 0.0 | 4.8 | GO:0007018 | microtubule-based movement(GO:0007018) |
| 0.0 | 0.6 | GO:0006165 | nucleoside diphosphate phosphorylation(GO:0006165) |
| 0.0 | 0.6 | GO:0001757 | somite specification(GO:0001757) segment specification(GO:0007379) |
| 0.0 | 0.5 | GO:0032402 | melanosome transport(GO:0032402) pigment granule transport(GO:0051904) |
| 0.0 | 0.7 | GO:0016079 | synaptic vesicle exocytosis(GO:0016079) |
| 0.0 | 1.3 | GO:0007062 | sister chromatid cohesion(GO:0007062) |
| 0.0 | 0.2 | GO:0042403 | thyroid hormone metabolic process(GO:0042403) |
| 0.0 | 1.2 | GO:0006888 | ER to Golgi vesicle-mediated transport(GO:0006888) |
| 0.0 | 0.4 | GO:0006198 | cAMP catabolic process(GO:0006198) |
| 0.0 | 1.6 | GO:0044782 | cilium organization(GO:0044782) |
| 0.0 | 1.7 | GO:0016573 | histone acetylation(GO:0016573) |
| 0.0 | 2.6 | GO:0006338 | chromatin remodeling(GO:0006338) |
| 0.0 | 1.1 | GO:0007498 | mesoderm development(GO:0007498) |
| 0.0 | 5.8 | GO:0000226 | microtubule cytoskeleton organization(GO:0000226) |
| 0.0 | 3.2 | GO:0006869 | lipid transport(GO:0006869) |
| 0.0 | 1.7 | GO:0051260 | protein homooligomerization(GO:0051260) |
| 0.0 | 0.3 | GO:2001243 | negative regulation of intrinsic apoptotic signaling pathway(GO:2001243) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 10.6 | 63.3 | GO:0070032 | synaptobrevin 2-SNAP-25-syntaxin-1a-complexin I complex(GO:0070032) |
| 3.5 | 20.9 | GO:0033181 | plasma membrane proton-transporting V-type ATPase complex(GO:0033181) |
| 2.7 | 10.8 | GO:0044214 | spanning component of plasma membrane(GO:0044214) spanning component of membrane(GO:0089717) |
| 2.0 | 12.1 | GO:0005955 | calcineurin complex(GO:0005955) |
| 1.4 | 5.8 | GO:0032281 | AMPA glutamate receptor complex(GO:0032281) |
| 1.4 | 12.6 | GO:0005952 | cAMP-dependent protein kinase complex(GO:0005952) |
| 1.4 | 6.9 | GO:0070724 | BMP receptor complex(GO:0070724) |
| 1.4 | 13.7 | GO:0000220 | vacuolar proton-transporting V-type ATPase, V0 domain(GO:0000220) |
| 1.3 | 5.2 | GO:0008622 | epsilon DNA polymerase complex(GO:0008622) |
| 1.2 | 4.6 | GO:0016460 | myosin II complex(GO:0016460) |
| 1.1 | 8.6 | GO:0034464 | BBSome(GO:0034464) |
| 1.1 | 12.9 | GO:0044295 | axonal growth cone(GO:0044295) |
| 1.0 | 17.8 | GO:0000813 | ESCRT I complex(GO:0000813) |
| 1.0 | 9.8 | GO:0017059 | serine C-palmitoyltransferase complex(GO:0017059) endoplasmic reticulum palmitoyltransferase complex(GO:0031211) |
| 0.9 | 4.7 | GO:0033503 | HULC complex(GO:0033503) |
| 0.9 | 10.9 | GO:0033180 | proton-transporting V-type ATPase, V1 domain(GO:0033180) |
| 0.9 | 3.5 | GO:1990316 | ATG1/ULK1 kinase complex(GO:1990316) |
| 0.8 | 3.2 | GO:0002189 | ribose phosphate diphosphokinase complex(GO:0002189) |
| 0.8 | 30.8 | GO:0043679 | axon terminus(GO:0043679) |
| 0.8 | 3.0 | GO:0098574 | cytoplasmic side of lysosomal membrane(GO:0098574) |
| 0.8 | 3.0 | GO:0036396 | MIS complex(GO:0036396) mRNA editing complex(GO:0045293) |
| 0.7 | 3.0 | GO:0008537 | proteasome activator complex(GO:0008537) |
| 0.7 | 11.4 | GO:0032588 | trans-Golgi network membrane(GO:0032588) |
| 0.7 | 11.3 | GO:0031209 | SCAR complex(GO:0031209) |
| 0.7 | 4.8 | GO:0030131 | clathrin adaptor complex(GO:0030131) |
| 0.7 | 4.1 | GO:0000938 | GARP complex(GO:0000938) |
| 0.6 | 26.3 | GO:0030125 | clathrin vesicle coat(GO:0030125) |
| 0.6 | 5.6 | GO:0089701 | U2AF(GO:0089701) |
| 0.6 | 50.8 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.6 | 3.5 | GO:0070062 | extracellular exosome(GO:0070062) |
| 0.5 | 3.7 | GO:0035517 | PR-DUB complex(GO:0035517) |
| 0.5 | 17.1 | GO:0032154 | cleavage furrow(GO:0032154) cell surface furrow(GO:0097610) |
| 0.5 | 12.2 | GO:0000407 | pre-autophagosomal structure(GO:0000407) |
| 0.5 | 1.9 | GO:0030896 | checkpoint clamp complex(GO:0030896) |
| 0.4 | 3.6 | GO:0031597 | cytosolic proteasome complex(GO:0031597) |
| 0.4 | 3.7 | GO:0008024 | cyclin/CDK positive transcription elongation factor complex(GO:0008024) nuclear cyclin-dependent protein kinase holoenzyme complex(GO:0019908) |
| 0.4 | 26.2 | GO:0031902 | late endosome membrane(GO:0031902) |
| 0.4 | 3.2 | GO:0030140 | trans-Golgi network transport vesicle(GO:0030140) |
| 0.4 | 2.4 | GO:0016589 | NURF complex(GO:0016589) |
| 0.4 | 1.9 | GO:0070971 | endoplasmic reticulum exit site(GO:0070971) |
| 0.4 | 2.6 | GO:0071818 | BAT3 complex(GO:0071818) |
| 0.4 | 15.1 | GO:0043204 | perikaryon(GO:0043204) |
| 0.4 | 31.9 | GO:0030141 | secretory granule(GO:0030141) |
| 0.4 | 23.8 | GO:0005834 | heterotrimeric G-protein complex(GO:0005834) |
| 0.4 | 4.3 | GO:0030126 | COPI vesicle coat(GO:0030126) |
| 0.3 | 3.4 | GO:0005744 | mitochondrial inner membrane presequence translocase complex(GO:0005744) |
| 0.3 | 5.2 | GO:0030992 | intraciliary transport particle B(GO:0030992) |
| 0.3 | 3.7 | GO:0030015 | CCR4-NOT core complex(GO:0030015) |
| 0.3 | 2.1 | GO:0005682 | U5 snRNP(GO:0005682) |
| 0.2 | 23.5 | GO:0016323 | basolateral plasma membrane(GO:0016323) |
| 0.2 | 26.6 | GO:0005802 | trans-Golgi network(GO:0005802) |
| 0.2 | 3.0 | GO:0071007 | U2-type catalytic step 2 spliceosome(GO:0071007) |
| 0.2 | 2.4 | GO:0061617 | MICOS complex(GO:0061617) |
| 0.2 | 4.6 | GO:0071339 | MLL1/2 complex(GO:0044665) MLL1 complex(GO:0071339) |
| 0.2 | 11.1 | GO:0005776 | autophagosome(GO:0005776) |
| 0.2 | 4.3 | GO:0005686 | U2 snRNP(GO:0005686) |
| 0.2 | 6.5 | GO:0000159 | protein phosphatase type 2A complex(GO:0000159) |
| 0.2 | 0.8 | GO:0070765 | gamma-secretase complex(GO:0070765) |
| 0.2 | 0.6 | GO:0034518 | RNA cap binding complex(GO:0034518) |
| 0.2 | 2.3 | GO:0071014 | post-mRNA release spliceosomal complex(GO:0071014) |
| 0.2 | 6.0 | GO:0031463 | Cul3-RING ubiquitin ligase complex(GO:0031463) |
| 0.2 | 7.6 | GO:0030426 | growth cone(GO:0030426) |
| 0.2 | 1.5 | GO:0031314 | extrinsic component of mitochondrial inner membrane(GO:0031314) |
| 0.2 | 3.9 | GO:0008278 | cohesin complex(GO:0008278) |
| 0.2 | 4.7 | GO:0005881 | cytoplasmic microtubule(GO:0005881) |
| 0.2 | 1.4 | GO:0035145 | exon-exon junction complex(GO:0035145) |
| 0.2 | 39.9 | GO:0045211 | postsynaptic membrane(GO:0045211) |
| 0.2 | 26.3 | GO:0048471 | perinuclear region of cytoplasm(GO:0048471) |
| 0.1 | 5.0 | GO:0030658 | transport vesicle membrane(GO:0030658) |
| 0.1 | 2.3 | GO:0005869 | dynactin complex(GO:0005869) |
| 0.1 | 4.0 | GO:0005876 | spindle microtubule(GO:0005876) |
| 0.1 | 3.2 | GO:0005849 | mRNA cleavage factor complex(GO:0005849) |
| 0.1 | 2.1 | GO:0015030 | Cajal body(GO:0015030) |
| 0.1 | 21.3 | GO:0008021 | synaptic vesicle(GO:0008021) |
| 0.1 | 2.7 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.1 | 2.9 | GO:0042734 | presynaptic membrane(GO:0042734) |
| 0.1 | 6.9 | GO:0043025 | neuronal cell body(GO:0043025) |
| 0.1 | 3.7 | GO:0031901 | early endosome membrane(GO:0031901) |
| 0.1 | 9.4 | GO:0030133 | transport vesicle(GO:0030133) |
| 0.1 | 4.5 | GO:0031201 | SNARE complex(GO:0031201) |
| 0.1 | 1.9 | GO:0034993 | microtubule organizing center attachment site(GO:0034992) LINC complex(GO:0034993) |
| 0.1 | 0.6 | GO:0031616 | spindle pole centrosome(GO:0031616) |
| 0.1 | 1.1 | GO:0017119 | Golgi transport complex(GO:0017119) |
| 0.1 | 10.3 | GO:0016324 | apical plasma membrane(GO:0016324) |
| 0.1 | 4.9 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.1 | 4.6 | GO:0000922 | spindle pole(GO:0000922) |
| 0.1 | 1.9 | GO:0005669 | transcription factor TFIID complex(GO:0005669) |
| 0.1 | 19.6 | GO:0005874 | microtubule(GO:0005874) |
| 0.1 | 2.0 | GO:0030867 | rough endoplasmic reticulum membrane(GO:0030867) |
| 0.1 | 1.4 | GO:0000178 | exosome (RNase complex)(GO:0000178) |
| 0.1 | 4.5 | GO:0016592 | mediator complex(GO:0016592) |
| 0.1 | 21.2 | GO:0043005 | neuron projection(GO:0043005) |
| 0.1 | 3.4 | GO:0008305 | integrin complex(GO:0008305) |
| 0.1 | 5.7 | GO:0045111 | intermediate filament(GO:0005882) intermediate filament cytoskeleton(GO:0045111) |
| 0.1 | 1.2 | GO:0030496 | midbody(GO:0030496) |
| 0.1 | 4.0 | GO:0030018 | Z disc(GO:0030018) |
| 0.1 | 0.4 | GO:0016593 | Cdc73/Paf1 complex(GO:0016593) |
| 0.1 | 0.2 | GO:1990604 | IRE1-TRAF2-ASK1 complex(GO:1990604) |
| 0.1 | 77.1 | GO:0031226 | intrinsic component of plasma membrane(GO:0031226) |
| 0.0 | 5.3 | GO:0030176 | integral component of endoplasmic reticulum membrane(GO:0030176) |
| 0.0 | 0.5 | GO:0005956 | protein kinase CK2 complex(GO:0005956) |
| 0.0 | 1.2 | GO:0035861 | site of double-strand break(GO:0035861) |
| 0.0 | 3.3 | GO:0016607 | nuclear speck(GO:0016607) |
| 0.0 | 20.3 | GO:0005794 | Golgi apparatus(GO:0005794) |
| 0.0 | 0.3 | GO:0019773 | proteasome core complex, alpha-subunit complex(GO:0019773) |
| 0.0 | 0.3 | GO:0016580 | Sin3 complex(GO:0016580) |
| 0.0 | 1.5 | GO:0032432 | actin filament bundle(GO:0032432) |
| 0.0 | 1.7 | GO:0000123 | histone acetyltransferase complex(GO:0000123) |
| 0.0 | 0.4 | GO:0080008 | Cul4-RING E3 ubiquitin ligase complex(GO:0080008) |
| 0.0 | 0.3 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.0 | 0.2 | GO:1990907 | beta-catenin-TCF complex(GO:1990907) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 6.5 | 19.6 | GO:0004351 | glutamate decarboxylase activity(GO:0004351) |
| 5.5 | 16.6 | GO:0004422 | hypoxanthine phosphoribosyltransferase activity(GO:0004422) |
| 3.9 | 19.5 | GO:0008432 | JUN kinase binding(GO:0008432) |
| 3.6 | 46.6 | GO:0031628 | opioid receptor binding(GO:0031628) |
| 2.5 | 15.2 | GO:0008597 | calcium-dependent protein serine/threonine phosphatase regulator activity(GO:0008597) |
| 2.5 | 10.0 | GO:0043891 | glyceraldehyde-3-phosphate dehydrogenase (NAD+) (phosphorylating) activity(GO:0004365) glyceraldehyde-3-phosphate dehydrogenase (NAD(P)+) (phosphorylating) activity(GO:0043891) |
| 2.5 | 12.5 | GO:0004939 | beta-adrenergic receptor activity(GO:0004939) |
| 2.4 | 17.0 | GO:0004356 | glutamate-ammonia ligase activity(GO:0004356) ammonia ligase activity(GO:0016211) acid-ammonia (or amide) ligase activity(GO:0016880) |
| 2.1 | 14.8 | GO:0008526 | phosphatidylinositol transporter activity(GO:0008526) |
| 2.0 | 12.1 | GO:0033192 | calcium-dependent protein serine/threonine phosphatase activity(GO:0004723) calmodulin-dependent protein phosphatase activity(GO:0033192) |
| 2.0 | 80.4 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 2.0 | 5.9 | GO:0038046 | enkephalin receptor activity(GO:0038046) |
| 1.9 | 13.2 | GO:0016316 | phosphatidylinositol-3,4-bisphosphate 4-phosphatase activity(GO:0016316) |
| 1.8 | 25.5 | GO:0008510 | sodium:bicarbonate symporter activity(GO:0008510) |
| 1.8 | 12.6 | GO:0008603 | cAMP-dependent protein kinase regulator activity(GO:0008603) |
| 1.7 | 5.1 | GO:0047173 | phosphatidylcholine-retinol O-acyltransferase activity(GO:0047173) |
| 1.6 | 11.5 | GO:0016454 | serine C-palmitoyltransferase activity(GO:0004758) C-palmitoyltransferase activity(GO:0016454) |
| 1.6 | 4.8 | GO:0004394 | heparan sulfate 2-O-sulfotransferase activity(GO:0004394) |
| 1.5 | 4.5 | GO:0071568 | UFM1 transferase activity(GO:0071568) |
| 1.4 | 6.9 | GO:0000026 | alpha-1,2-mannosyltransferase activity(GO:0000026) |
| 1.4 | 4.1 | GO:0015526 | hexose phosphate transmembrane transporter activity(GO:0015119) organophosphate:inorganic phosphate antiporter activity(GO:0015315) hexose-phosphate:inorganic phosphate antiporter activity(GO:0015526) glucose 6-phosphate:inorganic phosphate antiporter activity(GO:0061513) |
| 1.3 | 18.9 | GO:0008330 | protein tyrosine/threonine phosphatase activity(GO:0008330) |
| 1.3 | 11.3 | GO:0071933 | Arp2/3 complex binding(GO:0071933) |
| 1.2 | 14.4 | GO:0004157 | dihydropyrimidinase activity(GO:0004157) |
| 1.2 | 3.5 | GO:0001605 | adrenomedullin receptor activity(GO:0001605) calcitonin gene-related peptide receptor activity(GO:0001635) |
| 1.2 | 4.6 | GO:0032028 | myosin head/neck binding(GO:0032028) myosin II head/neck binding(GO:0032034) myosin II heavy chain binding(GO:0032038) |
| 1.1 | 3.4 | GO:0015562 | inositol hexakisphosphate binding(GO:0000822) efflux transmembrane transporter activity(GO:0015562) |
| 1.1 | 37.3 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 1.1 | 10.1 | GO:0016714 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced pteridine as one donor, and incorporation of one atom of oxygen(GO:0016714) |
| 1.1 | 5.5 | GO:0070004 | cysteine-type exopeptidase activity(GO:0070004) |
| 1.0 | 7.3 | GO:0030274 | LIM domain binding(GO:0030274) |
| 1.0 | 16.6 | GO:0004331 | fructose-2,6-bisphosphate 2-phosphatase activity(GO:0004331) |
| 1.0 | 4.0 | GO:0035575 | histone demethylase activity (H4-K20 specific)(GO:0035575) |
| 1.0 | 3.0 | GO:0001734 | mRNA (N6-adenosine)-methyltransferase activity(GO:0001734) |
| 1.0 | 2.9 | GO:0030623 | U5 snRNA binding(GO:0030623) |
| 1.0 | 5.7 | GO:0004594 | pantothenate kinase activity(GO:0004594) |
| 0.9 | 13.7 | GO:0051117 | ATPase binding(GO:0051117) |
| 0.9 | 7.0 | GO:0005402 | sugar:proton symporter activity(GO:0005351) cation:sugar symporter activity(GO:0005402) sucrose:proton symporter activity(GO:0008506) sucrose transmembrane transporter activity(GO:0008515) disaccharide transmembrane transporter activity(GO:0015154) oligosaccharide transmembrane transporter activity(GO:0015157) |
| 0.9 | 4.3 | GO:0008486 | endopolyphosphatase activity(GO:0000298) diphosphoinositol-polyphosphate diphosphatase activity(GO:0008486) bis(5'-adenosyl)-hexaphosphatase activity(GO:0034431) bis(5'-adenosyl)-pentaphosphatase activity(GO:0034432) |
| 0.8 | 11.9 | GO:0004169 | dolichyl-phosphate-mannose-protein mannosyltransferase activity(GO:0004169) |
| 0.8 | 3.2 | GO:0004749 | ribose phosphate diphosphokinase activity(GO:0004749) |
| 0.8 | 7.8 | GO:0016909 | JUN kinase activity(GO:0004705) SAP kinase activity(GO:0016909) |
| 0.8 | 4.7 | GO:0032052 | bile acid binding(GO:0032052) |
| 0.8 | 2.3 | GO:0004779 | adenylylsulfate kinase activity(GO:0004020) sulfate adenylyltransferase activity(GO:0004779) sulfate adenylyltransferase (ATP) activity(GO:0004781) |
| 0.7 | 8.2 | GO:0046933 | proton-transporting ATP synthase activity, rotational mechanism(GO:0046933) |
| 0.7 | 6.7 | GO:0004731 | purine-nucleoside phosphorylase activity(GO:0004731) |
| 0.7 | 22.2 | GO:0030165 | PDZ domain binding(GO:0030165) |
| 0.7 | 2.2 | GO:0055105 | ubiquitin-protein transferase inhibitor activity(GO:0055105) |
| 0.7 | 7.7 | GO:0004465 | lipoprotein lipase activity(GO:0004465) |
| 0.7 | 4.1 | GO:1990247 | N6-methyladenosine-containing RNA binding(GO:1990247) |
| 0.7 | 3.4 | GO:0015450 | P-P-bond-hydrolysis-driven protein transmembrane transporter activity(GO:0015450) |
| 0.7 | 2.0 | GO:0051500 | D-aminoacyl-tRNA deacylase activity(GO:0051499) D-tyrosyl-tRNA(Tyr) deacylase activity(GO:0051500) |
| 0.7 | 26.7 | GO:0005184 | neuropeptide hormone activity(GO:0005184) |
| 0.6 | 3.2 | GO:0042809 | vitamin D receptor binding(GO:0042809) |
| 0.6 | 5.1 | GO:0034485 | phosphatidylinositol-3,4,5-trisphosphate 5-phosphatase activity(GO:0034485) |
| 0.6 | 6.9 | GO:0005451 | monovalent cation:proton antiporter activity(GO:0005451) sodium:proton antiporter activity(GO:0015385) potassium:proton antiporter activity(GO:0015386) |
| 0.6 | 3.1 | GO:0004679 | AMP-activated protein kinase activity(GO:0004679) |
| 0.6 | 4.2 | GO:0034648 | histone demethylase activity (H3-dimethyl-K4 specific)(GO:0034648) |
| 0.6 | 10.7 | GO:0005080 | protein kinase C binding(GO:0005080) |
| 0.6 | 4.7 | GO:0017169 | CDP-alcohol phosphatidyltransferase activity(GO:0017169) |
| 0.6 | 6.9 | GO:0005025 | transforming growth factor beta receptor activity, type I(GO:0005025) |
| 0.6 | 9.1 | GO:0004724 | magnesium-dependent protein serine/threonine phosphatase activity(GO:0004724) |
| 0.6 | 2.2 | GO:0003958 | NADPH-hemoprotein reductase activity(GO:0003958) |
| 0.6 | 5.6 | GO:0030628 | pre-mRNA 3'-splice site binding(GO:0030628) |
| 0.5 | 4.8 | GO:0016922 | ligand-dependent nuclear receptor binding(GO:0016922) |
| 0.5 | 13.4 | GO:0042577 | lipid phosphatase activity(GO:0042577) |
| 0.5 | 6.1 | GO:0016868 | intramolecular transferase activity, phosphotransferases(GO:0016868) |
| 0.5 | 4.0 | GO:0004784 | superoxide dismutase activity(GO:0004784) oxidoreductase activity, acting on superoxide radicals as acceptor(GO:0016721) |
| 0.5 | 3.0 | GO:0061133 | endopeptidase activator activity(GO:0061133) |
| 0.5 | 4.5 | GO:0004706 | JUN kinase kinase kinase activity(GO:0004706) |
| 0.5 | 22.7 | GO:0004683 | calmodulin-dependent protein kinase activity(GO:0004683) |
| 0.5 | 7.4 | GO:0016018 | cyclosporin A binding(GO:0016018) |
| 0.4 | 2.2 | GO:0050699 | WW domain binding(GO:0050699) |
| 0.4 | 2.7 | GO:0001159 | core promoter proximal region sequence-specific DNA binding(GO:0000987) core promoter proximal region DNA binding(GO:0001159) |
| 0.4 | 1.3 | GO:0008517 | folic acid transporter activity(GO:0008517) |
| 0.4 | 4.8 | GO:0070567 | cytidylyltransferase activity(GO:0070567) |
| 0.4 | 6.1 | GO:0008307 | structural constituent of muscle(GO:0008307) |
| 0.4 | 26.3 | GO:0030215 | semaphorin receptor binding(GO:0030215) |
| 0.4 | 5.1 | GO:0004711 | ribosomal protein S6 kinase activity(GO:0004711) |
| 0.4 | 4.2 | GO:0005391 | sodium:potassium-exchanging ATPase activity(GO:0005391) |
| 0.4 | 1.6 | GO:0070404 | 6,7-dihydropteridine reductase activity(GO:0004155) NADH binding(GO:0070404) |
| 0.4 | 2.0 | GO:0004385 | guanylate kinase activity(GO:0004385) |
| 0.4 | 4.7 | GO:0015299 | solute:proton antiporter activity(GO:0015299) |
| 0.4 | 5.1 | GO:0047372 | acylglycerol lipase activity(GO:0047372) |
| 0.4 | 5.1 | GO:0042826 | histone deacetylase binding(GO:0042826) |
| 0.4 | 15.1 | GO:0015459 | potassium channel regulator activity(GO:0015459) |
| 0.4 | 4.4 | GO:0043014 | alpha-tubulin binding(GO:0043014) |
| 0.4 | 1.5 | GO:0005021 | vascular endothelial growth factor-activated receptor activity(GO:0005021) |
| 0.4 | 5.1 | GO:0070840 | dynein complex binding(GO:0070840) |
| 0.4 | 5.1 | GO:0016929 | SUMO-specific protease activity(GO:0016929) |
| 0.4 | 2.2 | GO:0036402 | proteasome-activating ATPase activity(GO:0036402) |
| 0.4 | 5.3 | GO:0031682 | G-protein gamma-subunit binding(GO:0031682) |
| 0.4 | 4.9 | GO:0097199 | cysteine-type endopeptidase activity involved in apoptotic signaling pathway(GO:0097199) |
| 0.3 | 3.1 | GO:0046974 | histone methyltransferase activity (H3-K9 specific)(GO:0046974) |
| 0.3 | 1.4 | GO:0070513 | death domain binding(GO:0070513) |
| 0.3 | 4.5 | GO:0070577 | lysine-acetylated histone binding(GO:0070577) |
| 0.3 | 38.6 | GO:0005179 | hormone activity(GO:0005179) |
| 0.3 | 30.2 | GO:0043130 | ubiquitin binding(GO:0043130) |
| 0.3 | 3.7 | GO:0061575 | cyclin-dependent protein serine/threonine kinase activator activity(GO:0061575) |
| 0.3 | 9.1 | GO:0004697 | protein kinase C activity(GO:0004697) calcium-dependent protein kinase C activity(GO:0004698) |
| 0.3 | 15.9 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.3 | 3.1 | GO:0005049 | nuclear export signal receptor activity(GO:0005049) |
| 0.3 | 1.8 | GO:0004743 | pyruvate kinase activity(GO:0004743) |
| 0.3 | 8.6 | GO:0032266 | phosphatidylinositol-3-phosphate binding(GO:0032266) |
| 0.3 | 5.9 | GO:0035591 | signaling adaptor activity(GO:0035591) |
| 0.3 | 1.5 | GO:0008430 | selenium binding(GO:0008430) |
| 0.3 | 1.6 | GO:0016744 | transferase activity, transferring aldehyde or ketonic groups(GO:0016744) |
| 0.3 | 3.1 | GO:0005158 | insulin receptor binding(GO:0005158) |
| 0.3 | 26.8 | GO:0003730 | mRNA 3'-UTR binding(GO:0003730) |
| 0.3 | 6.8 | GO:0015248 | sterol transporter activity(GO:0015248) |
| 0.3 | 14.8 | GO:0005200 | structural constituent of cytoskeleton(GO:0005200) |
| 0.2 | 3.7 | GO:0030544 | Hsp70 protein binding(GO:0030544) |
| 0.2 | 3.7 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.2 | 1.2 | GO:0030116 | glial cell-derived neurotrophic factor receptor binding(GO:0030116) |
| 0.2 | 4.8 | GO:0102338 | fatty acid elongase activity(GO:0009922) 3-oxo-arachidoyl-CoA synthase activity(GO:0102336) 3-oxo-cerotoyl-CoA synthase activity(GO:0102337) 3-oxo-lignoceronyl-CoA synthase activity(GO:0102338) |
| 0.2 | 10.7 | GO:0019706 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.2 | 14.2 | GO:0030276 | clathrin binding(GO:0030276) |
| 0.2 | 2.4 | GO:1990239 | steroid hormone binding(GO:1990239) |
| 0.2 | 1.8 | GO:0003993 | acid phosphatase activity(GO:0003993) |
| 0.2 | 2.8 | GO:0008195 | phosphatidate phosphatase activity(GO:0008195) |
| 0.2 | 3.7 | GO:0005005 | transmembrane-ephrin receptor activity(GO:0005005) |
| 0.2 | 1.3 | GO:0051379 | alpha2-adrenergic receptor activity(GO:0004938) epinephrine binding(GO:0051379) |
| 0.2 | 11.1 | GO:0008013 | beta-catenin binding(GO:0008013) |
| 0.2 | 2.9 | GO:0032051 | clathrin light chain binding(GO:0032051) |
| 0.2 | 7.6 | GO:0016307 | phosphatidylinositol phosphate kinase activity(GO:0016307) |
| 0.2 | 2.1 | GO:0015924 | mannosyl-oligosaccharide 1,2-alpha-mannosidase activity(GO:0004571) mannosyl-oligosaccharide mannosidase activity(GO:0015924) |
| 0.2 | 3.1 | GO:0061578 | Lys63-specific deubiquitinase activity(GO:0061578) |
| 0.2 | 13.5 | GO:0003777 | microtubule motor activity(GO:0003777) |
| 0.2 | 1.9 | GO:0043495 | protein anchor(GO:0043495) |
| 0.2 | 15.3 | GO:0008201 | heparin binding(GO:0008201) |
| 0.2 | 1.7 | GO:0052796 | exo-alpha-(2->3)-sialidase activity(GO:0052794) exo-alpha-(2->6)-sialidase activity(GO:0052795) exo-alpha-(2->8)-sialidase activity(GO:0052796) |
| 0.2 | 0.6 | GO:0008073 | ornithine decarboxylase inhibitor activity(GO:0008073) |
| 0.2 | 15.8 | GO:0005544 | calcium-dependent phospholipid binding(GO:0005544) |
| 0.2 | 28.4 | GO:0004725 | protein tyrosine phosphatase activity(GO:0004725) |
| 0.1 | 1.1 | GO:0043035 | chromatin insulator sequence binding(GO:0043035) |
| 0.1 | 0.6 | GO:0004069 | L-aspartate:2-oxoglutarate aminotransferase activity(GO:0004069) |
| 0.1 | 1.6 | GO:0004622 | lysophospholipase activity(GO:0004622) |
| 0.1 | 4.1 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
| 0.1 | 1.8 | GO:0008503 | benzodiazepine receptor activity(GO:0008503) GABA-gated chloride ion channel activity(GO:0022851) |
| 0.1 | 3.0 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.1 | 0.9 | GO:0004128 | cytochrome-b5 reductase activity, acting on NAD(P)H(GO:0004128) |
| 0.1 | 0.6 | GO:0004127 | cytidylate kinase activity(GO:0004127) |
| 0.1 | 0.6 | GO:0044736 | acid-sensing ion channel activity(GO:0044736) |
| 0.1 | 2.0 | GO:0046975 | histone methyltransferase activity (H3-K36 specific)(GO:0046975) |
| 0.1 | 1.8 | GO:0015271 | outward rectifier potassium channel activity(GO:0015271) |
| 0.1 | 1.1 | GO:0005161 | platelet-derived growth factor receptor binding(GO:0005161) |
| 0.1 | 2.4 | GO:0015095 | magnesium ion transmembrane transporter activity(GO:0015095) |
| 0.1 | 32.9 | GO:0015631 | tubulin binding(GO:0015631) |
| 0.1 | 0.8 | GO:0004126 | cytidine deaminase activity(GO:0004126) |
| 0.1 | 3.1 | GO:0005520 | insulin-like growth factor binding(GO:0005520) |
| 0.1 | 2.2 | GO:0003785 | actin monomer binding(GO:0003785) |
| 0.1 | 0.8 | GO:0019809 | spermidine binding(GO:0019809) |
| 0.1 | 41.4 | GO:0003924 | GTPase activity(GO:0003924) |
| 0.1 | 2.1 | GO:0031681 | G-protein beta-subunit binding(GO:0031681) |
| 0.1 | 12.3 | GO:0005516 | calmodulin binding(GO:0005516) |
| 0.1 | 0.7 | GO:0008190 | eukaryotic initiation factor 4E binding(GO:0008190) |
| 0.1 | 10.6 | GO:0004843 | thiol-dependent ubiquitin-specific protease activity(GO:0004843) thiol-dependent ubiquitinyl hydrolase activity(GO:0036459) ubiquitinyl hydrolase activity(GO:0101005) |
| 0.1 | 6.1 | GO:0031490 | chromatin DNA binding(GO:0031490) |
| 0.1 | 10.0 | GO:0019888 | protein phosphatase regulator activity(GO:0019888) |
| 0.1 | 1.4 | GO:0004767 | sphingomyelin phosphodiesterase activity(GO:0004767) |
| 0.1 | 3.1 | GO:0003755 | peptidyl-prolyl cis-trans isomerase activity(GO:0003755) |
| 0.1 | 0.6 | GO:0034237 | protein kinase A regulatory subunit binding(GO:0034237) |
| 0.1 | 13.2 | GO:0000287 | magnesium ion binding(GO:0000287) |
| 0.1 | 4.0 | GO:0019905 | syntaxin binding(GO:0019905) |
| 0.1 | 1.0 | GO:0035259 | glucocorticoid receptor binding(GO:0035259) |
| 0.1 | 3.5 | GO:0030295 | kinase activator activity(GO:0019209) protein kinase activator activity(GO:0030295) |
| 0.1 | 1.3 | GO:0019894 | kinesin binding(GO:0019894) |
| 0.1 | 2.8 | GO:0051087 | chaperone binding(GO:0051087) |
| 0.1 | 0.4 | GO:0045309 | protein phosphorylated amino acid binding(GO:0045309) phosphoprotein binding(GO:0051219) |
| 0.1 | 0.2 | GO:0001160 | transcription termination site sequence-specific DNA binding(GO:0001147) transcription termination site DNA binding(GO:0001160) |
| 0.1 | 11.5 | GO:0003714 | transcription corepressor activity(GO:0003714) |
| 0.1 | 3.8 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.1 | 15.6 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.1 | 2.9 | GO:0008408 | 3'-5' exonuclease activity(GO:0008408) |
| 0.1 | 1.1 | GO:0031729 | CCR1 chemokine receptor binding(GO:0031726) CCR4 chemokine receptor binding(GO:0031729) |
| 0.1 | 0.7 | GO:0003689 | DNA clamp loader activity(GO:0003689) protein-DNA loading ATPase activity(GO:0033170) |
| 0.1 | 0.7 | GO:0030971 | receptor tyrosine kinase binding(GO:0030971) |
| 0.1 | 0.9 | GO:0004190 | aspartic-type endopeptidase activity(GO:0004190) aspartic-type peptidase activity(GO:0070001) |
| 0.1 | 3.6 | GO:0000149 | SNARE binding(GO:0000149) |
| 0.1 | 0.9 | GO:0045134 | guanosine-diphosphatase activity(GO:0004382) uridine-diphosphatase activity(GO:0045134) |
| 0.1 | 13.7 | GO:0003682 | chromatin binding(GO:0003682) |
| 0.1 | 0.9 | GO:0050811 | GABA receptor binding(GO:0050811) |
| 0.0 | 2.0 | GO:0019957 | C-C chemokine binding(GO:0019957) |
| 0.0 | 0.1 | GO:0003868 | 4-hydroxyphenylpyruvate dioxygenase activity(GO:0003868) |
| 0.0 | 42.2 | GO:0005509 | calcium ion binding(GO:0005509) |
| 0.0 | 1.5 | GO:0003684 | damaged DNA binding(GO:0003684) |
| 0.0 | 1.1 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.0 | 6.8 | GO:0004497 | monooxygenase activity(GO:0004497) |
| 0.0 | 1.0 | GO:0005104 | fibroblast growth factor receptor binding(GO:0005104) |
| 0.0 | 0.6 | GO:0000340 | RNA 7-methylguanosine cap binding(GO:0000340) |
| 0.0 | 2.1 | GO:0016763 | transferase activity, transferring pentosyl groups(GO:0016763) |
| 0.0 | 7.1 | GO:0000989 | transcription factor activity, transcription factor binding(GO:0000989) transcription cofactor activity(GO:0003712) |
| 0.0 | 0.1 | GO:0008174 | mRNA methyltransferase activity(GO:0008174) |
| 0.0 | 0.3 | GO:0016833 | oxo-acid-lyase activity(GO:0016833) |
| 0.0 | 0.3 | GO:0032182 | ubiquitin-like protein binding(GO:0032182) |
| 0.0 | 0.7 | GO:0004016 | adenylate cyclase activity(GO:0004016) |
| 0.0 | 0.6 | GO:0008378 | galactosyltransferase activity(GO:0008378) |
| 0.0 | 1.0 | GO:0005548 | phospholipid transporter activity(GO:0005548) |
| 0.0 | 0.2 | GO:0004438 | phosphatidylinositol-3-phosphatase activity(GO:0004438) |
| 0.0 | 0.2 | GO:0043027 | cysteine-type endopeptidase inhibitor activity involved in apoptotic process(GO:0043027) cysteine-type endopeptidase regulator activity involved in apoptotic process(GO:0043028) |
| 0.0 | 2.4 | GO:0008081 | phosphoric diester hydrolase activity(GO:0008081) |
| 0.0 | 1.4 | GO:0016765 | transferase activity, transferring alkyl or aryl (other than methyl) groups(GO:0016765) |
| 0.0 | 0.2 | GO:0000268 | peroxisome targeting sequence binding(GO:0000268) |
| 0.0 | 0.1 | GO:0050508 | glucuronosyl-N-acetylglucosaminyl-proteoglycan 4-alpha-N-acetylglucosaminyltransferase activity(GO:0050508) |
| 0.0 | 4.7 | GO:0005085 | guanyl-nucleotide exchange factor activity(GO:0005085) |
| 0.0 | 38.4 | GO:0003700 | nucleic acid binding transcription factor activity(GO:0001071) transcription factor activity, sequence-specific DNA binding(GO:0003700) |
| 0.0 | 4.0 | GO:0005096 | GTPase activator activity(GO:0005096) |
| 0.0 | 1.3 | GO:0016791 | phosphatase activity(GO:0016791) |
| 0.0 | 7.4 | GO:0004930 | G-protein coupled receptor activity(GO:0004930) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.8 | 38.7 | PID INSULIN GLUCOSE PATHWAY | Insulin-mediated glucose transport |
| 1.2 | 18.8 | ST MYOCYTE AD PATHWAY | Myocyte Adrenergic Pathway is a specific case of the generalized Adrenergic Pathway. |
| 1.1 | 14.7 | PID TCR CALCIUM PATHWAY | Calcium signaling in the CD4+ TCR pathway |
| 1.0 | 21.6 | ST GRANULE CELL SURVIVAL PATHWAY | Granule Cell Survival Pathway is a specific case of more general PAC1 Receptor Pathway. |
| 0.7 | 5.3 | PID VEGFR1 PATHWAY | VEGFR1 specific signals |
| 0.6 | 7.4 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.6 | 4.9 | SA PROGRAMMED CELL DEATH | Programmed cell death, or apoptosis, eliminates damaged or unneeded cells. |
| 0.6 | 10.1 | PID INTEGRIN A4B1 PATHWAY | Alpha4 beta1 integrin signaling events |
| 0.6 | 7.8 | PID S1P S1P3 PATHWAY | S1P3 pathway |
| 0.5 | 10.7 | PID NEPHRIN NEPH1 PATHWAY | Nephrin/Neph1 signaling in the kidney podocyte |
| 0.4 | 8.0 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.3 | 12.1 | PID ATF2 PATHWAY | ATF-2 transcription factor network |
| 0.3 | 7.7 | PID NCADHERIN PATHWAY | N-cadherin signaling events |
| 0.2 | 1.7 | PID NECTIN PATHWAY | Nectin adhesion pathway |
| 0.2 | 5.3 | PID IL8 CXCR2 PATHWAY | IL8- and CXCR2-mediated signaling events |
| 0.2 | 6.8 | PID HEDGEHOG GLI PATHWAY | Hedgehog signaling events mediated by Gli proteins |
| 0.2 | 1.1 | PID SMAD2 3PATHWAY | Regulation of cytoplasmic and nuclear SMAD2/3 signaling |
| 0.2 | 8.0 | ST JNK MAPK PATHWAY | JNK MAPK Pathway |
| 0.1 | 8.7 | PID AR PATHWAY | Coregulation of Androgen receptor activity |
| 0.1 | 2.4 | PID ALPHA SYNUCLEIN PATHWAY | Alpha-synuclein signaling |
| 0.1 | 0.6 | PID IL1 PATHWAY | IL1-mediated signaling events |
| 0.1 | 4.0 | PID IL4 2PATHWAY | IL4-mediated signaling events |
| 0.1 | 1.7 | PID RHODOPSIN PATHWAY | Visual signal transduction: Rods |
| 0.1 | 5.1 | NABA ECM AFFILIATED | Genes encoding proteins affiliated structurally or functionally to extracellular matrix proteins |
| 0.1 | 1.0 | PID GLYPICAN 1PATHWAY | Glypican 1 network |
| 0.1 | 0.8 | PID SYNDECAN 3 PATHWAY | Syndecan-3-mediated signaling events |
| 0.1 | 0.2 | PID ERBB NETWORK PATHWAY | ErbB receptor signaling network |
| 0.0 | 2.8 | PID P73PATHWAY | p73 transcription factor network |
| 0.0 | 1.2 | PID DELTA NP63 PATHWAY | Validated transcriptional targets of deltaNp63 isoforms |
| 0.0 | 0.2 | PID CD40 PATHWAY | CD40/CD40L signaling |
| 0.0 | 0.4 | SIG INSULIN RECEPTOR PATHWAY IN CARDIAC MYOCYTES | Genes related to the insulin receptor pathway |
| 0.0 | 0.5 | PID HIF1 TFPATHWAY | HIF-1-alpha transcription factor network |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 5.7 | 17.1 | REACTOME ACETYLCHOLINE NEUROTRANSMITTER RELEASE CYCLE | Genes involved in Acetylcholine Neurotransmitter Release Cycle |
| 3.7 | 18.6 | REACTOME TGF BETA RECEPTOR SIGNALING IN EMT EPITHELIAL TO MESENCHYMAL TRANSITION | Genes involved in TGF-beta receptor signaling in EMT (epithelial to mesenchymal transition) |
| 1.9 | 18.8 | REACTOME MEMBRANE BINDING AND TARGETTING OF GAG PROTEINS | Genes involved in Membrane binding and targetting of GAG proteins |
| 1.8 | 19.6 | REACTOME GABA SYNTHESIS RELEASE REUPTAKE AND DEGRADATION | Genes involved in GABA synthesis, release, reuptake and degradation |
| 1.6 | 27.5 | REACTOME DARPP 32 EVENTS | Genes involved in DARPP-32 events |
| 1.4 | 14.4 | REACTOME CRMPS IN SEMA3A SIGNALING | Genes involved in CRMPs in Sema3A signaling |
| 1.4 | 13.7 | REACTOME PEPTIDE HORMONE BIOSYNTHESIS | Genes involved in Peptide hormone biosynthesis |
| 1.0 | 9.2 | REACTOME RAP1 SIGNALLING | Genes involved in Rap1 signalling |
| 1.0 | 17.1 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 1.0 | 12.5 | REACTOME AMINE LIGAND BINDING RECEPTORS | Genes involved in Amine ligand-binding receptors |
| 0.8 | 7.5 | REACTOME AMINE DERIVED HORMONES | Genes involved in Amine-derived hormones |
| 0.7 | 6.3 | REACTOME HORMONE SENSITIVE LIPASE HSL MEDIATED TRIACYLGLYCEROL HYDROLYSIS | Genes involved in Hormone-sensitive lipase (HSL)-mediated triacylglycerol hydrolysis |
| 0.7 | 6.8 | REACTOME ROLE OF DCC IN REGULATING APOPTOSIS | Genes involved in Role of DCC in regulating apoptosis |
| 0.7 | 9.8 | REACTOME METABOLISM OF POLYAMINES | Genes involved in Metabolism of polyamines |
| 0.6 | 28.5 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.6 | 6.3 | REACTOME PECAM1 INTERACTIONS | Genes involved in PECAM1 interactions |
| 0.5 | 4.2 | REACTOME TIGHT JUNCTION INTERACTIONS | Genes involved in Tight junction interactions |
| 0.5 | 4.6 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.5 | 4.8 | REACTOME ACTIVATION OF THE AP1 FAMILY OF TRANSCRIPTION FACTORS | Genes involved in Activation of the AP-1 family of transcription factors |
| 0.4 | 3.8 | REACTOME COPI MEDIATED TRANSPORT | Genes involved in COPI Mediated Transport |
| 0.4 | 8.0 | REACTOME YAP1 AND WWTR1 TAZ STIMULATED GENE EXPRESSION | Genes involved in YAP1- and WWTR1 (TAZ)-stimulated gene expression |
| 0.4 | 4.2 | REACTOME NOTCH HLH TRANSCRIPTION PATHWAY | Genes involved in Notch-HLH transcription pathway |
| 0.3 | 7.7 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |
| 0.3 | 4.4 | REACTOME PROTEOLYTIC CLEAVAGE OF SNARE COMPLEX PROTEINS | Genes involved in Proteolytic cleavage of SNARE complex proteins |
| 0.3 | 2.1 | REACTOME PYRIMIDINE CATABOLISM | Genes involved in Pyrimidine catabolism |
| 0.3 | 4.3 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.2 | 5.9 | REACTOME G PROTEIN ACTIVATION | Genes involved in G-protein activation |
| 0.2 | 29.3 | REACTOME MRNA SPLICING | Genes involved in mRNA Splicing |
| 0.2 | 5.5 | REACTOME SIGNALING BY FGFR1 FUSION MUTANTS | Genes involved in Signaling by FGFR1 fusion mutants |
| 0.2 | 1.5 | REACTOME ERKS ARE INACTIVATED | Genes involved in ERKs are inactivated |
| 0.2 | 2.8 | REACTOME SYNTHESIS OF PE | Genes involved in Synthesis of PE |
| 0.2 | 7.2 | REACTOME BIOSYNTHESIS OF THE N GLYCAN PRECURSOR DOLICHOL LIPID LINKED OLIGOSACCHARIDE LLO AND TRANSFER TO A NASCENT PROTEIN | Genes involved in Biosynthesis of the N-glycan precursor (dolichol lipid-linked oligosaccharide, LLO) and transfer to a nascent protein |
| 0.2 | 1.5 | REACTOME VEGF LIGAND RECEPTOR INTERACTIONS | Genes involved in VEGF ligand-receptor interactions |
| 0.2 | 2.9 | REACTOME RESOLUTION OF AP SITES VIA THE SINGLE NUCLEOTIDE REPLACEMENT PATHWAY | Genes involved in Resolution of AP sites via the single-nucleotide replacement pathway |
| 0.2 | 2.5 | REACTOME PRE NOTCH PROCESSING IN GOLGI | Genes involved in Pre-NOTCH Processing in Golgi |
| 0.2 | 1.6 | REACTOME SIGNALING BY NOTCH3 | Genes involved in Signaling by NOTCH3 |
| 0.2 | 1.0 | REACTOME SIGNALING BY ACTIVATED POINT MUTANTS OF FGFR1 | Genes involved in Signaling by activated point mutants of FGFR1 |
| 0.1 | 5.2 | REACTOME TRANSPORT OF MATURE MRNA DERIVED FROM AN INTRONLESS TRANSCRIPT | Genes involved in Transport of Mature mRNA Derived from an Intronless Transcript |
| 0.1 | 2.1 | REACTOME CALNEXIN CALRETICULIN CYCLE | Genes involved in Calnexin/calreticulin cycle |
| 0.1 | 1.9 | REACTOME DEADENYLATION OF MRNA | Genes involved in Deadenylation of mRNA |
| 0.1 | 2.1 | REACTOME DOWNREGULATION OF SMAD2 3 SMAD4 TRANSCRIPTIONAL ACTIVITY | Genes involved in Downregulation of SMAD2/3:SMAD4 transcriptional activity |
| 0.1 | 8.5 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |
| 0.1 | 5.0 | REACTOME ION TRANSPORT BY P TYPE ATPASES | Genes involved in Ion transport by P-type ATPases |
| 0.1 | 0.5 | REACTOME CTLA4 INHIBITORY SIGNALING | Genes involved in CTLA4 inhibitory signaling |
| 0.1 | 6.9 | REACTOME GLYCEROPHOSPHOLIPID BIOSYNTHESIS | Genes involved in Glycerophospholipid biosynthesis |
| 0.1 | 2.4 | REACTOME DEPOSITION OF NEW CENPA CONTAINING NUCLEOSOMES AT THE CENTROMERE | Genes involved in Deposition of New CENPA-containing Nucleosomes at the Centromere |
| 0.1 | 1.8 | REACTOME CONVERSION FROM APC C CDC20 TO APC C CDH1 IN LATE ANAPHASE | Genes involved in Conversion from APC/C:Cdc20 to APC/C:Cdh1 in late anaphase |
| 0.1 | 13.0 | REACTOME ANTIGEN PROCESSING UBIQUITINATION PROTEASOME DEGRADATION | Genes involved in Antigen processing: Ubiquitination & Proteasome degradation |
| 0.1 | 3.0 | REACTOME G ALPHA1213 SIGNALLING EVENTS | Genes involved in G alpha (12/13) signalling events |
| 0.1 | 0.7 | REACTOME SIGNAL ATTENUATION | Genes involved in Signal attenuation |
| 0.1 | 1.9 | REACTOME RNA POL II TRANSCRIPTION PRE INITIATION AND PROMOTER OPENING | Genes involved in RNA Polymerase II Transcription Pre-Initiation And Promoter Opening |
| 0.1 | 2.2 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |
| 0.0 | 2.3 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.0 | 0.6 | REACTOME KERATAN SULFATE BIOSYNTHESIS | Genes involved in Keratan sulfate biosynthesis |
| 0.0 | 0.8 | REACTOME PTM GAMMA CARBOXYLATION HYPUSINE FORMATION AND ARYLSULFATASE ACTIVATION | Genes involved in PTM: gamma carboxylation, hypusine formation and arylsulfatase activation |
| 0.0 | 0.2 | REACTOME ABCA TRANSPORTERS IN LIPID HOMEOSTASIS | Genes involved in ABCA transporters in lipid homeostasis |
| 0.0 | 0.9 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.0 | 0.9 | REACTOME GENERIC TRANSCRIPTION PATHWAY | Genes involved in Generic Transcription Pathway |