Project
PRJDB7713: Age associated transcriptome analysis in 5 tissues of zebrafish
Navigation
Downloads
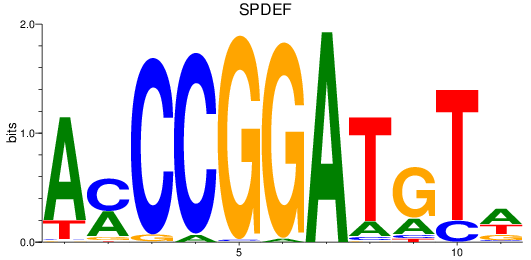
Results for SPDEF
Z-value: 1.38
Transcription factors associated with SPDEF
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
SPDEF
|
ENSDARG00000029930 | SAM pointed domain containing ETS transcription factor |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| SPDEF | dr11_v1_chr6_-_54290227_54290227 | 0.22 | 3.6e-02 | Click! |
Activity profile of SPDEF motif
Sorted Z-values of SPDEF motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr14_+_28438947 | 11.25 |
ENSDART00000006489
|
acsl4a
|
acyl-CoA synthetase long chain family member 4a |
| chr21_+_6751760 | 8.70 |
ENSDART00000135914
|
olfm1b
|
olfactomedin 1b |
| chr21_+_6751405 | 7.96 |
ENSDART00000037265
ENSDART00000146371 |
olfm1b
|
olfactomedin 1b |
| chr3_-_36440705 | 7.65 |
ENSDART00000162875
|
rogdi
|
rogdi homolog (Drosophila) |
| chr8_-_49431939 | 7.48 |
ENSDART00000011453
ENSDART00000088240 ENSDART00000114173 |
sypb
|
synaptophysin b |
| chr1_-_44581937 | 6.95 |
ENSDART00000009858
|
tmx2b
|
thioredoxin-related transmembrane protein 2b |
| chr5_+_64732270 | 6.66 |
ENSDART00000134241
|
olfm1a
|
olfactomedin 1a |
| chr16_-_16212615 | 6.48 |
ENSDART00000059905
|
upp1
|
uridine phosphorylase 1 |
| chr5_+_64732036 | 6.37 |
ENSDART00000073950
|
olfm1a
|
olfactomedin 1a |
| chr12_-_14211293 | 6.32 |
ENSDART00000158399
|
avl9
|
AVL9 homolog (S. cerevisiase) |
| chr13_-_12494575 | 6.08 |
ENSDART00000137761
|
si:dkey-20i10.7
|
si:dkey-20i10.7 |
| chr4_-_12978925 | 5.92 |
ENSDART00000013839
|
tmbim4
|
transmembrane BAX inhibitor motif containing 4 |
| chr16_-_45001842 | 5.91 |
ENSDART00000037797
|
sult2st3
|
sulfotransferase family 2, cytosolic sulfotransferase 3 |
| chr1_+_46598764 | 5.76 |
ENSDART00000053240
|
cab39l
|
calcium binding protein 39-like |
| chr5_+_26795465 | 5.44 |
ENSDART00000053001
|
tcn2
|
transcobalamin II |
| chr25_+_7872135 | 5.44 |
ENSDART00000003042
|
mdkb
|
midkine b |
| chr15_-_31043183 | 5.40 |
ENSDART00000100145
|
lgals9l1
|
lectin, galactoside-binding, soluble, 9 (galectin 9)-like 1 |
| chr20_-_5369105 | 5.18 |
ENSDART00000114316
|
sptlc2b
|
serine palmitoyltransferase, long chain base subunit 2b |
| chr20_+_27087539 | 5.17 |
ENSDART00000062094
|
tmem251
|
transmembrane protein 251 |
| chr2_-_3044947 | 5.06 |
ENSDART00000192642
|
guk1a
|
guanylate kinase 1a |
| chr19_-_1871415 | 5.03 |
ENSDART00000004585
|
clptm1l
|
CLPTM1-like |
| chr13_+_681628 | 4.99 |
ENSDART00000016604
|
abcc2
|
ATP-binding cassette, sub-family C (CFTR/MRP), member 2 |
| chr5_+_26795773 | 4.71 |
ENSDART00000145631
|
tcn2
|
transcobalamin II |
| chr20_-_5291012 | 4.60 |
ENSDART00000122892
|
cyp46a1.3
|
cytochrome P450, family 46, subfamily A, polypeptide 1, tandem duplicate 3 |
| chr18_+_3169579 | 4.58 |
ENSDART00000164724
ENSDART00000186340 ENSDART00000181247 ENSDART00000168056 |
pak1
|
p21 protein (Cdc42/Rac)-activated kinase 1 |
| chr7_+_38762043 | 4.56 |
ENSDART00000036461
|
arhgap1
|
Rho GTPase activating protein 1 |
| chr20_-_20611063 | 4.49 |
ENSDART00000063492
|
ppm1ab
|
protein phosphatase, Mg2+/Mn2+ dependent, 1Ab |
| chr1_+_49435017 | 4.41 |
ENSDART00000124833
|
pdcd11
|
programmed cell death 11 |
| chr12_-_14211067 | 4.36 |
ENSDART00000077903
|
avl9
|
AVL9 homolog (S. cerevisiase) |
| chr23_-_24234371 | 4.36 |
ENSDART00000124539
|
ddost
|
dolichyl-diphosphooligosaccharide--protein glycosyltransferase subunit (non-catalytic) |
| chr11_-_15296805 | 4.35 |
ENSDART00000124968
|
rpn2
|
ribophorin II |
| chr9_-_23824290 | 4.29 |
ENSDART00000059209
|
wdfy2
|
WD repeat and FYVE domain containing 2 |
| chr4_+_16885854 | 4.26 |
ENSDART00000017726
|
etnk1
|
ethanolamine kinase 1 |
| chr21_-_40938382 | 4.21 |
ENSDART00000008593
|
yipf5
|
Yip1 domain family, member 5 |
| chr19_-_27006764 | 4.17 |
ENSDART00000089540
|
sacm1la
|
SAC1 like phosphatidylinositide phosphatase a |
| chr5_+_44346691 | 4.16 |
ENSDART00000034523
|
tars
|
threonyl-tRNA synthetase |
| chr2_+_32846602 | 4.03 |
ENSDART00000056649
|
tmem53
|
transmembrane protein 53 |
| chr6_-_18531760 | 4.03 |
ENSDART00000167167
|
utp6
|
UTP6, small subunit (SSU) processome component, homolog (yeast) |
| chr2_-_11662851 | 3.94 |
ENSDART00000145108
|
zgc:110130
|
zgc:110130 |
| chr15_+_17343319 | 3.87 |
ENSDART00000018461
|
vmp1
|
vacuole membrane protein 1 |
| chr15_-_15230264 | 3.87 |
ENSDART00000155400
|
rrp8
|
ribosomal RNA processing 8, methyltransferase, homolog (yeast) |
| chr24_-_8409641 | 3.84 |
ENSDART00000149662
ENSDART00000149025 |
slc35b3
|
solute carrier family 35 (adenosine 3'-phospho 5'-phosphosulfate transporter), member B3 |
| chr1_+_46598502 | 3.83 |
ENSDART00000132861
|
cab39l
|
calcium binding protein 39-like |
| chr14_+_25986895 | 3.79 |
ENSDART00000149087
|
slc36a1
|
solute carrier family 36 (proton/amino acid symporter), member 1 |
| chr6_-_2222707 | 3.77 |
ENSDART00000022179
|
prkag1
|
protein kinase, AMP-activated, gamma 1 non-catalytic subunit |
| chr11_-_15297129 | 3.72 |
ENSDART00000167985
ENSDART00000163031 ENSDART00000165010 ENSDART00000171250 ENSDART00000010684 ENSDART00000191714 ENSDART00000191164 |
rpn2
|
ribophorin II |
| chr22_-_20950448 | 3.71 |
ENSDART00000002029
|
fkbp8
|
FK506 binding protein 8 |
| chr12_+_31422557 | 3.71 |
ENSDART00000153179
|
zdhhc6
|
zinc finger, DHHC-type containing 6 |
| chr6_-_18531349 | 3.69 |
ENSDART00000160693
ENSDART00000169780 |
utp6
|
UTP6, small subunit (SSU) processome component, homolog (yeast) |
| chr23_+_31979602 | 3.66 |
ENSDART00000140351
|
pan2
|
PAN2 poly(A) specific ribonuclease subunit homolog (S. cerevisiae) |
| chr6_-_51666739 | 3.62 |
ENSDART00000155549
|
chd6
|
chromodomain helicase DNA binding protein 6 |
| chr7_-_34062301 | 3.60 |
ENSDART00000052404
|
map2k5
|
mitogen-activated protein kinase kinase 5 |
| chr10_+_36439293 | 3.58 |
ENSDART00000043802
|
uspl1
|
ubiquitin specific peptidase like 1 |
| chr19_-_17658160 | 3.57 |
ENSDART00000151766
ENSDART00000170790 ENSDART00000186678 ENSDART00000188045 ENSDART00000176980 ENSDART00000166313 ENSDART00000188589 |
thrb
|
thyroid hormone receptor beta |
| chr23_+_42304602 | 3.56 |
ENSDART00000166004
|
cyp2aa11
|
cytochrome P450, family 2, subfamily AA, polypeptide 11 |
| chr5_-_67292690 | 3.45 |
ENSDART00000062366
|
rsrc2
|
arginine/serine-rich coiled-coil 2 |
| chr21_-_14762944 | 3.43 |
ENSDART00000114096
|
arrdc1b
|
arrestin domain containing 1b |
| chr1_-_29758947 | 3.41 |
ENSDART00000049514
ENSDART00000183571 ENSDART00000140345 |
alg11
|
asparagine-linked glycosylation 11 (alpha-1,2-mannosyltransferase) |
| chr8_-_14080534 | 3.40 |
ENSDART00000042867
|
dedd
|
death effector domain containing |
| chr15_+_7064819 | 3.39 |
ENSDART00000155268
|
pik3cb
|
phosphatidylinositol-4,5-bisphosphate 3-kinase, catalytic subunit beta |
| chr9_+_14023386 | 3.36 |
ENSDART00000140199
ENSDART00000124267 |
si:ch211-67e16.4
|
si:ch211-67e16.4 |
| chr24_-_7826489 | 3.36 |
ENSDART00000112777
|
si:dkey-197c15.6
|
si:dkey-197c15.6 |
| chr2_-_42173834 | 3.32 |
ENSDART00000098357
ENSDART00000144707 |
slc39a6
|
solute carrier family 39 (zinc transporter), member 6 |
| chr15_-_29114449 | 3.30 |
ENSDART00000145748
ENSDART00000109482 ENSDART00000179123 |
zgc:162698
|
zgc:162698 |
| chr7_+_50053339 | 3.29 |
ENSDART00000174308
|
LRRC4C (1 of many)
|
si:dkey-6l15.1 |
| chr1_-_28950366 | 3.26 |
ENSDART00000110270
|
pwp2h
|
PWP2 periodic tryptophan protein homolog (yeast) |
| chr20_-_30938184 | 3.24 |
ENSDART00000147359
ENSDART00000062552 |
wtap
|
WT1 associated protein |
| chr14_-_30897177 | 3.23 |
ENSDART00000087918
|
slc7a3b
|
solute carrier family 7 (cationic amino acid transporter, y+ system), member 3b |
| chr13_+_31545530 | 3.19 |
ENSDART00000164590
ENSDART00000178460 ENSDART00000185503 |
ppm1aa
|
protein phosphatase, Mg2+/Mn2+ dependent, 1Aa |
| chr13_+_31545812 | 3.14 |
ENSDART00000076527
|
ppm1aa
|
protein phosphatase, Mg2+/Mn2+ dependent, 1Aa |
| chr19_+_9455218 | 3.13 |
ENSDART00000139385
|
si:ch211-288g17.3
|
si:ch211-288g17.3 |
| chr18_+_6638726 | 3.10 |
ENSDART00000142755
ENSDART00000167781 |
c2cd5
|
C2 calcium-dependent domain containing 5 |
| chr15_+_19884242 | 3.09 |
ENSDART00000154437
ENSDART00000054416 |
dyrk1ab
|
dual-specificity tyrosine-(Y)-phosphorylation regulated kinase 1A, b |
| chr15_-_766015 | 3.05 |
ENSDART00000190648
|
si:dkey-7i4.15
|
si:dkey-7i4.15 |
| chr6_-_34860574 | 3.04 |
ENSDART00000073957
|
slc35d1a
|
solute carrier family 35 (UDP-GlcA/UDP-GalNAc transporter), member D1a |
| chr15_+_25528290 | 3.03 |
ENSDART00000123143
|
npat
|
nuclear protein, ataxia-telangiectasia locus |
| chr5_+_51833305 | 3.01 |
ENSDART00000165276
ENSDART00000166443 |
papd4
|
PAP associated domain containing 4 |
| chr2_+_4383061 | 2.98 |
ENSDART00000163986
|
wacb
|
WW domain containing adaptor with coiled-coil b |
| chr3_+_59864872 | 2.98 |
ENSDART00000102014
|
mcrip1
|
MAPK regulated corepressor interacting protein 1 |
| chr22_-_37611681 | 2.96 |
ENSDART00000028085
|
ttc14
|
tetratricopeptide repeat domain 14 |
| chr11_-_16975190 | 2.94 |
ENSDART00000122222
|
suclg2
|
succinate-CoA ligase, GDP-forming, beta subunit |
| chr20_-_20610812 | 2.94 |
ENSDART00000181870
|
ppm1ab
|
protein phosphatase, Mg2+/Mn2+ dependent, 1Ab |
| chr17_+_33375469 | 2.93 |
ENSDART00000032827
|
zgc:162964
|
zgc:162964 |
| chr4_+_5506952 | 2.92 |
ENSDART00000032857
ENSDART00000160222 |
mapk11
|
mitogen-activated protein kinase 11 |
| chr12_+_29240124 | 2.90 |
ENSDART00000053761
ENSDART00000130172 |
bms1
|
BMS1 ribosome biogenesis factor |
| chr16_+_22618620 | 2.89 |
ENSDART00000185728
ENSDART00000041625 |
chrnb2b
|
cholinergic receptor, nicotinic, beta 2b |
| chr1_+_41596099 | 2.87 |
ENSDART00000111367
|
si:dkey-56e3.3
|
si:dkey-56e3.3 |
| chr18_-_22735002 | 2.86 |
ENSDART00000023721
|
nudt21
|
nudix hydrolase 21 |
| chr5_+_28041967 | 2.81 |
ENSDART00000133641
|
ogfod2
|
2-oxoglutarate and iron-dependent oxygenase domain containing 2 |
| chr25_-_3347418 | 2.79 |
ENSDART00000082385
|
golt1bb
|
golgi transport 1Bb |
| chr19_+_34742706 | 2.77 |
ENSDART00000103276
|
fam206a
|
family with sequence similarity 206, member A |
| chr18_+_6638974 | 2.75 |
ENSDART00000162398
|
c2cd5
|
C2 calcium-dependent domain containing 5 |
| chr11_-_16115804 | 2.74 |
ENSDART00000143436
ENSDART00000157928 |
rpf1
|
ribosome production factor 1 homolog |
| chr20_-_23946296 | 2.73 |
ENSDART00000143005
|
mdn1
|
midasin AAA ATPase 1 |
| chr21_-_30994577 | 2.73 |
ENSDART00000065503
|
pgap2
|
post-GPI attachment to proteins 2 |
| chr17_+_49281597 | 2.72 |
ENSDART00000155599
|
zgc:113176
|
zgc:113176 |
| chr2_-_20052561 | 2.67 |
ENSDART00000100133
|
dpydb
|
dihydropyrimidine dehydrogenase b |
| chr3_-_27066451 | 2.67 |
ENSDART00000156228
ENSDART00000156311 |
atf7ip2
|
activating transcription factor 7 interacting protein 2 |
| chr14_-_33521071 | 2.66 |
ENSDART00000052789
|
c1galt1c1
|
C1GALT1-specific chaperone 1 |
| chr8_+_53388005 | 2.61 |
ENSDART00000171920
|
dcp1a
|
decapping mRNA 1A |
| chr16_+_38119004 | 2.59 |
ENSDART00000132087
|
pogzb
|
pogo transposable element derived with ZNF domain b |
| chr8_-_17987547 | 2.57 |
ENSDART00000112699
ENSDART00000061747 |
fpgt
|
fucose-1-phosphate guanylyltransferase |
| chr6_-_21988375 | 2.56 |
ENSDART00000161257
|
plxnb1b
|
plexin b1b |
| chr13_+_14006118 | 2.54 |
ENSDART00000131875
ENSDART00000089528 |
atrn
|
attractin |
| chr15_+_37436430 | 2.52 |
ENSDART00000124779
|
igflr1
|
IGF-like family receptor 1 |
| chr24_-_32522587 | 2.50 |
ENSDART00000048968
ENSDART00000143781 |
EIF1B
|
zgc:56676 |
| chr14_-_23684814 | 2.50 |
ENSDART00000024604
|
mars2
|
methionyl-tRNA synthetase 2, mitochondrial |
| chr3_-_40276057 | 2.49 |
ENSDART00000132225
ENSDART00000074737 |
shmt1
|
serine hydroxymethyltransferase 1 (soluble) |
| chr2_+_16652238 | 2.47 |
ENSDART00000091351
|
gk5
|
glycerol kinase 5 (putative) |
| chr18_-_39321484 | 2.47 |
ENSDART00000077694
|
leo1
|
LEO1 homolog, Paf1/RNA polymerase II complex component |
| chr16_-_31824525 | 2.45 |
ENSDART00000058737
|
cdc42l
|
cell division cycle 42, like |
| chr21_-_19828423 | 2.44 |
ENSDART00000037664
|
nnt
|
nicotinamide nucleotide transhydrogenase |
| chr8_+_17143501 | 2.42 |
ENSDART00000061758
|
mier3b
|
mesoderm induction early response 1, family member 3 b |
| chr11_+_24339377 | 2.41 |
ENSDART00000133679
ENSDART00000135435 ENSDART00000017973 ENSDART00000131365 ENSDART00000186418 |
rbm39a
|
RNA binding motif protein 39a |
| chr22_+_8753092 | 2.41 |
ENSDART00000140720
|
si:dkey-182g1.2
|
si:dkey-182g1.2 |
| chr7_+_35191220 | 2.40 |
ENSDART00000110552
|
zdhhc1
|
zinc finger, DHHC-type containing 1 |
| chr17_-_7351488 | 2.36 |
ENSDART00000098731
|
stxbp5b
|
syntaxin binding protein 5b (tomosyn) |
| chr25_+_15354095 | 2.35 |
ENSDART00000090397
|
kiaa1549la
|
KIAA1549-like a |
| chr21_-_22892124 | 2.34 |
ENSDART00000065563
|
ccdc90b
|
coiled-coil domain containing 90B |
| chr20_+_715739 | 2.32 |
ENSDART00000136768
|
myo6a
|
myosin VIa |
| chr17_+_20237727 | 2.31 |
ENSDART00000180115
|
smndc1
|
survival motor neuron domain containing 1 |
| chr9_-_296169 | 2.31 |
ENSDART00000165228
|
kif5aa
|
kinesin family member 5A, a |
| chr14_-_30724165 | 2.29 |
ENSDART00000020936
|
fibpa
|
fibroblast growth factor (acidic) intracellular binding protein a |
| chr18_-_26797723 | 2.28 |
ENSDART00000008013
|
sec11a
|
SEC11 homolog A, signal peptidase complex subunit |
| chr23_-_30764319 | 2.27 |
ENSDART00000075918
|
pcmtd2
|
protein-L-isoaspartate (D-aspartate) O-methyltransferase domain containing 2 |
| chr1_-_44048798 | 2.26 |
ENSDART00000073746
|
si:ch73-109d9.2
|
si:ch73-109d9.2 |
| chr20_-_10118818 | 2.26 |
ENSDART00000033976
|
meis2b
|
Meis homeobox 2b |
| chr7_+_17782436 | 2.25 |
ENSDART00000173793
ENSDART00000165110 |
si:dkey-28a3.2
|
si:dkey-28a3.2 |
| chr5_+_41477954 | 2.24 |
ENSDART00000185871
|
pias2
|
protein inhibitor of activated STAT, 2 |
| chr3_-_35554809 | 2.23 |
ENSDART00000010944
|
dctn5
|
dynactin 5 |
| chr20_-_48898560 | 2.20 |
ENSDART00000163071
|
xrn2
|
5'-3' exoribonuclease 2 |
| chr14_-_31856819 | 2.20 |
ENSDART00000003345
|
arhgef6
|
Rac/Cdc42 guanine nucleotide exchange factor (GEF) 6 |
| chr14_+_4276394 | 2.19 |
ENSDART00000038301
|
gnpda2
|
glucosamine-6-phosphate deaminase 2 |
| chr1_-_23294753 | 2.19 |
ENSDART00000013263
|
ugdh
|
UDP-glucose 6-dehydrogenase |
| chr20_+_52492192 | 2.17 |
ENSDART00000057986
|
TSTA3 (1 of many)
|
zgc:100864 |
| chr4_+_9612574 | 2.14 |
ENSDART00000150336
ENSDART00000041289 ENSDART00000150828 |
tmem243b
|
transmembrane protein 243, mitochondrial b |
| chr21_+_38033226 | 2.13 |
ENSDART00000085728
|
klf8
|
Kruppel-like factor 8 |
| chr13_-_33654931 | 2.13 |
ENSDART00000020350
|
snx5
|
sorting nexin 5 |
| chr22_-_8306743 | 2.13 |
ENSDART00000123982
|
CABZ01077218.1
|
|
| chr8_-_41279326 | 2.13 |
ENSDART00000075491
|
pop5
|
POP5 homolog, ribonuclease P/MRP subunit |
| chr7_-_32629458 | 2.12 |
ENSDART00000001376
|
arl14ep
|
ADP-ribosylation factor-like 14 effector protein |
| chr25_+_3347461 | 2.11 |
ENSDART00000104888
|
slc35b4
|
solute carrier family 35, member B4 |
| chr6_+_4387150 | 2.07 |
ENSDART00000181283
|
rbm26
|
RNA binding motif protein 26 |
| chr24_+_3478871 | 2.06 |
ENSDART00000111491
ENSDART00000134598 ENSDART00000142407 |
wdr37
|
WD repeat domain 37 |
| chr24_-_10897511 | 2.05 |
ENSDART00000145593
ENSDART00000102484 ENSDART00000066784 |
fam49bb
|
family with sequence similarity 49, member Bb |
| chr8_-_17184482 | 2.05 |
ENSDART00000025803
|
pola2
|
polymerase (DNA directed), alpha 2 |
| chr8_-_1267247 | 2.03 |
ENSDART00000150064
|
cdc14b
|
cell division cycle 14B |
| chr16_-_6944927 | 2.02 |
ENSDART00000149620
|
pmvk
|
phosphomevalonate kinase |
| chr18_-_18584839 | 2.02 |
ENSDART00000159274
|
sf3b3
|
splicing factor 3b, subunit 3 |
| chr2_+_3044992 | 2.02 |
ENSDART00000020463
|
zgc:63882
|
zgc:63882 |
| chr13_-_37519774 | 2.01 |
ENSDART00000141420
ENSDART00000185478 |
sgpp1
|
sphingosine-1-phosphate phosphatase 1 |
| chr19_+_42770041 | 2.00 |
ENSDART00000150930
|
clasp2
|
cytoplasmic linker associated protein 2 |
| chr1_+_44582369 | 2.00 |
ENSDART00000003022
ENSDART00000137980 |
med19b
|
mediator complex subunit 19b |
| chr16_+_12812472 | 1.99 |
ENSDART00000008535
|
u2af2a
|
U2 small nuclear RNA auxiliary factor 2a |
| chr10_+_21722892 | 1.98 |
ENSDART00000162855
|
pcdh1g13
|
protocadherin 1 gamma 13 |
| chr19_-_19025998 | 1.96 |
ENSDART00000186156
ENSDART00000163359 ENSDART00000167951 |
dync1li1
|
dynein, cytoplasmic 1, light intermediate chain 1 |
| chr5_+_51833132 | 1.96 |
ENSDART00000167491
|
papd4
|
PAP associated domain containing 4 |
| chr5_+_25084385 | 1.96 |
ENSDART00000134526
ENSDART00000111863 |
paxx
|
PAXX, non-homologous end joining factor |
| chr8_+_17184602 | 1.95 |
ENSDART00000050228
ENSDART00000140531 |
dimt1l
|
DIM1 dimethyladenosine transferase 1-like (S. cerevisiae) |
| chr8_-_1266181 | 1.94 |
ENSDART00000148654
ENSDART00000149924 |
cdc14b
|
cell division cycle 14B |
| chr14_+_9583588 | 1.93 |
ENSDART00000164101
|
tmem129
|
transmembrane protein 129, E3 ubiquitin protein ligase |
| chr23_+_16748806 | 1.92 |
ENSDART00000137737
ENSDART00000142556 |
fbxo44
|
F-box protein 44 |
| chr5_-_26795438 | 1.92 |
ENSDART00000146124
|
si:ch211-102c2.7
|
si:ch211-102c2.7 |
| chr22_-_16494406 | 1.90 |
ENSDART00000062727
|
stx6
|
syntaxin 6 |
| chr13_+_24022963 | 1.90 |
ENSDART00000028285
|
pgbd5
|
piggyBac transposable element derived 5 |
| chr20_-_48898371 | 1.89 |
ENSDART00000170617
|
xrn2
|
5'-3' exoribonuclease 2 |
| chr24_+_36392784 | 1.88 |
ENSDART00000134750
|
arf2b
|
ADP-ribosylation factor 2b |
| chr5_-_28041715 | 1.88 |
ENSDART00000078660
|
zgc:113436
|
zgc:113436 |
| chr1_+_51407520 | 1.88 |
ENSDART00000074294
|
actr2a
|
ARP2 actin related protein 2a homolog |
| chr8_-_21052371 | 1.87 |
ENSDART00000136561
|
si:dkeyp-82a1.6
|
si:dkeyp-82a1.6 |
| chr6_+_37625787 | 1.87 |
ENSDART00000065122
|
tubgcp5
|
tubulin, gamma complex associated protein 5 |
| chr14_-_5407555 | 1.85 |
ENSDART00000001424
|
pcgf1
|
polycomb group ring finger 1 |
| chr23_+_4260458 | 1.84 |
ENSDART00000103747
|
srsf6a
|
serine/arginine-rich splicing factor 6a |
| chr19_+_19241372 | 1.83 |
ENSDART00000184392
ENSDART00000165008 |
ptpn23b
|
protein tyrosine phosphatase, non-receptor type 23, b |
| chr15_-_29556757 | 1.83 |
ENSDART00000060049
|
hspa13
|
heat shock protein 70 family, member 13 |
| chr12_-_31724198 | 1.82 |
ENSDART00000153056
ENSDART00000165299 ENSDART00000137464 ENSDART00000080173 |
srsf2a
|
serine/arginine-rich splicing factor 2a |
| chr16_-_29480335 | 1.81 |
ENSDART00000148930
|
lingo4b
|
leucine rich repeat and Ig domain containing 4b |
| chr14_-_43616572 | 1.81 |
ENSDART00000111189
|
gar1
|
GAR1 homolog, ribonucleoprotein |
| chr15_-_39820491 | 1.81 |
ENSDART00000097134
|
robo1
|
roundabout, axon guidance receptor, homolog 1 (Drosophila) |
| chr22_-_5663354 | 1.81 |
ENSDART00000081774
|
ccdc51
|
coiled-coil domain containing 51 |
| chr21_+_26539157 | 1.81 |
ENSDART00000021121
|
stx5al
|
syntaxin 5A, like |
| chr9_-_24218367 | 1.79 |
ENSDART00000135356
|
nabp1a
|
nucleic acid binding protein 1a |
| chr9_+_30108641 | 1.79 |
ENSDART00000060174
|
jagn1a
|
jagunal homolog 1a |
| chr25_+_8921425 | 1.78 |
ENSDART00000128591
|
accs
|
1-aminocyclopropane-1-carboxylate synthase homolog (Arabidopsis)(non-functional) |
| chr2_+_21063660 | 1.77 |
ENSDART00000022765
|
riok1
|
RIO kinase 1 (yeast) |
| chr17_+_43468732 | 1.77 |
ENSDART00000055487
|
chmp3
|
charged multivesicular body protein 3 |
| chr1_+_47165842 | 1.76 |
ENSDART00000053152
ENSDART00000167051 |
cbr1
|
carbonyl reductase 1 |
| chr18_-_3552414 | 1.74 |
ENSDART00000163762
ENSDART00000165434 ENSDART00000161197 ENSDART00000166841 ENSDART00000170260 |
dcun1d5
|
DCN1, defective in cullin neddylation 1, domain containing 5 (S. cerevisiae) |
| chr17_+_37253706 | 1.74 |
ENSDART00000076004
|
tmem62
|
transmembrane protein 62 |
| chr3_+_36617024 | 1.73 |
ENSDART00000189957
|
pdxdc1
|
pyridoxal-dependent decarboxylase domain containing 1 |
| chr16_+_11029762 | 1.73 |
ENSDART00000091183
|
erfl3
|
Ets2 repressor factor like 3 |
| chr25_-_10791437 | 1.73 |
ENSDART00000127054
|
BX572619.1
|
|
| chr6_-_54444929 | 1.72 |
ENSDART00000154121
|
sys1
|
Sys1 golgi trafficking protein |
| chr22_-_21176269 | 1.69 |
ENSDART00000112839
|
rex1bd
|
required for excision 1-B domain containing |
| chr7_-_57637779 | 1.69 |
ENSDART00000028017
|
mad2l1
|
MAD2 mitotic arrest deficient-like 1 (yeast) |
| chr16_+_32184485 | 1.69 |
ENSDART00000084009
|
zufsp
|
zinc finger with UFM1-specific peptidase domain |
| chr3_-_30152836 | 1.68 |
ENSDART00000165920
|
nucb1
|
nucleobindin 1 |
| chr10_-_7671219 | 1.68 |
ENSDART00000159330
|
pcyox1
|
prenylcysteine oxidase 1 |
| chr12_-_31422433 | 1.68 |
ENSDART00000186075
ENSDART00000153172 ENSDART00000066256 |
vti1a
|
vesicle transport through interaction with t-SNAREs 1A |
| chr23_+_9867483 | 1.67 |
ENSDART00000023099
|
slc16a7
|
solute carrier family 16, member 7 (monocarboxylic acid transporter 2) |
Network of associatons between targets according to the STRING database.
First level regulatory network of SPDEF
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 3.4 | 10.2 | GO:0015889 | cobalt ion transport(GO:0006824) cobalamin transport(GO:0015889) |
| 1.9 | 11.3 | GO:1900744 | regulation of p38MAPK cascade(GO:1900744) |
| 1.4 | 13.0 | GO:0061075 | positive regulation of neural retina development(GO:0061075) positive regulation of retina development in camera-type eye(GO:1902868) |
| 1.3 | 3.8 | GO:0015808 | L-alanine transport(GO:0015808) proline transport(GO:0015824) |
| 1.2 | 5.8 | GO:0010828 | positive regulation of glucose transport(GO:0010828) |
| 1.1 | 5.4 | GO:2000562 | negative regulation of interferon-gamma production(GO:0032689) CD4-positive, alpha-beta T cell proliferation(GO:0035739) regulation of CD4-positive, alpha-beta T cell proliferation(GO:2000561) negative regulation of CD4-positive, alpha-beta T cell proliferation(GO:2000562) |
| 1.0 | 2.9 | GO:1990120 | messenger ribonucleoprotein complex assembly(GO:1990120) |
| 0.9 | 3.7 | GO:0010606 | positive regulation of cytoplasmic mRNA processing body assembly(GO:0010606) |
| 0.9 | 3.6 | GO:0060220 | retinal cone cell fate determination(GO:0042671) eye photoreceptor cell fate commitment(GO:0042706) photoreceptor cell fate determination(GO:0043703) retinal cone cell fate commitment(GO:0046551) photoreceptor cell fate commitment(GO:0046552) camera-type eye photoreceptor cell fate commitment(GO:0060220) |
| 0.9 | 2.7 | GO:0009120 | deoxyribonucleoside metabolic process(GO:0009120) thymidine metabolic process(GO:0046104) pyrimidine deoxyribonucleoside metabolic process(GO:0046125) |
| 0.8 | 2.5 | GO:0019264 | L-serine catabolic process(GO:0006565) glycine biosynthetic process from serine(GO:0019264) |
| 0.8 | 4.6 | GO:0003232 | bulbus arteriosus development(GO:0003232) |
| 0.7 | 2.2 | GO:0006041 | glucosamine metabolic process(GO:0006041) glucosamine catabolic process(GO:0006043) |
| 0.7 | 3.6 | GO:0030576 | nuclear body organization(GO:0030575) Cajal body organization(GO:0030576) protein localization to nuclear body(GO:1903405) protein localization to Cajal body(GO:1904867) protein localization to nucleoplasm(GO:1990173) |
| 0.7 | 2.7 | GO:0016267 | O-glycan processing, core 1(GO:0016267) |
| 0.6 | 6.5 | GO:0008655 | pyrimidine-containing compound salvage(GO:0008655) pyrimidine ribonucleotide salvage(GO:0010138) pyrimidine nucleotide salvage(GO:0032262) pyrimidine nucleoside salvage(GO:0043097) UMP salvage(GO:0044206) |
| 0.6 | 3.9 | GO:0000183 | chromatin silencing at rDNA(GO:0000183) |
| 0.6 | 1.9 | GO:0015074 | DNA integration(GO:0015074) |
| 0.6 | 2.4 | GO:0006529 | asparagine biosynthetic process(GO:0006529) |
| 0.6 | 2.4 | GO:0006740 | NADPH regeneration(GO:0006740) |
| 0.6 | 1.8 | GO:0038158 | granulocyte colony-stimulating factor signaling pathway(GO:0038158) |
| 0.6 | 2.3 | GO:1990535 | neuron projection maintenance(GO:1990535) |
| 0.6 | 13.8 | GO:0043124 | negative regulation of I-kappaB kinase/NF-kappaB signaling(GO:0043124) |
| 0.6 | 7.2 | GO:0006670 | sphingosine metabolic process(GO:0006670) |
| 0.5 | 3.2 | GO:0080009 | mRNA methylation(GO:0080009) |
| 0.5 | 3.2 | GO:0089718 | L-arginine import(GO:0043091) amino acid import across plasma membrane(GO:0089718) arginine import(GO:0090467) L-arginine import across plasma membrane(GO:0097638) L-arginine transport(GO:1902023) L-arginine import into cell(GO:1902765) amino acid import into cell(GO:1902837) L-arginine transmembrane transport(GO:1903400) arginine transmembrane transport(GO:1903826) |
| 0.5 | 1.6 | GO:0008344 | adult locomotory behavior(GO:0008344) |
| 0.5 | 1.6 | GO:0010847 | regulation of chromatin assembly(GO:0010847) |
| 0.5 | 2.0 | GO:0019287 | isopentenyl diphosphate biosynthetic process, mevalonate pathway(GO:0019287) |
| 0.5 | 1.5 | GO:0045579 | positive regulation of B cell differentiation(GO:0045579) |
| 0.5 | 15.5 | GO:0000462 | maturation of SSU-rRNA from tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000462) |
| 0.4 | 2.6 | GO:0031087 | nuclear-transcribed mRNA catabolic process, deadenylation-independent decay(GO:0031086) deadenylation-independent decapping of nuclear-transcribed mRNA(GO:0031087) |
| 0.4 | 1.3 | GO:0009211 | pyrimidine deoxyribonucleoside triphosphate metabolic process(GO:0009211) |
| 0.4 | 4.6 | GO:0016127 | cholesterol catabolic process(GO:0006707) sterol catabolic process(GO:0016127) |
| 0.4 | 1.2 | GO:0070537 | histone H2A K63-linked deubiquitination(GO:0070537) |
| 0.4 | 2.5 | GO:0019563 | glycerol catabolic process(GO:0019563) |
| 0.4 | 2.5 | GO:0021529 | spinal cord oligodendrocyte cell differentiation(GO:0021529) |
| 0.4 | 4.0 | GO:0071850 | mitotic cell cycle arrest(GO:0071850) |
| 0.4 | 2.0 | GO:0009794 | regulation of mitotic cell cycle, embryonic(GO:0009794) mitotic cell cycle, embryonic(GO:0045448) |
| 0.4 | 1.1 | GO:0021960 | anterior commissure morphogenesis(GO:0021960) |
| 0.4 | 1.5 | GO:0000448 | cleavage in ITS2 between 5.8S rRNA and LSU-rRNA of tricistronic rRNA transcript (SSU-rRNA, 5.8S rRNA, LSU-rRNA)(GO:0000448) |
| 0.4 | 3.3 | GO:0071578 | zinc II ion transmembrane import(GO:0071578) |
| 0.4 | 1.4 | GO:0021742 | abducens nucleus development(GO:0021742) |
| 0.3 | 2.0 | GO:0000244 | spliceosomal tri-snRNP complex assembly(GO:0000244) |
| 0.3 | 3.0 | GO:0000083 | regulation of transcription involved in G1/S transition of mitotic cell cycle(GO:0000083) |
| 0.3 | 1.7 | GO:0030327 | prenylated protein catabolic process(GO:0030327) prenylcysteine catabolic process(GO:0030328) prenylcysteine metabolic process(GO:0030329) |
| 0.3 | 2.2 | GO:0003188 | heart valve formation(GO:0003188) atrioventricular valve formation(GO:0003190) |
| 0.3 | 7.5 | GO:0048168 | regulation of neuronal synaptic plasticity(GO:0048168) |
| 0.3 | 6.5 | GO:1901654 | response to ketone(GO:1901654) |
| 0.3 | 2.5 | GO:0032488 | Cdc42 protein signal transduction(GO:0032488) |
| 0.3 | 2.1 | GO:0001682 | tRNA 5'-leader removal(GO:0001682) |
| 0.3 | 0.8 | GO:0034421 | post-translational protein acetylation(GO:0034421) |
| 0.3 | 2.3 | GO:0006465 | signal peptide processing(GO:0006465) |
| 0.2 | 4.4 | GO:0018279 | protein N-linked glycosylation via asparagine(GO:0018279) |
| 0.2 | 1.0 | GO:2000623 | negative regulation of mRNA catabolic process(GO:1902373) regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000622) negative regulation of nuclear-transcribed mRNA catabolic process, nonsense-mediated decay(GO:2000623) |
| 0.2 | 1.9 | GO:1905168 | positive regulation of double-strand break repair via homologous recombination(GO:1905168) regulation of double-strand break repair via nonhomologous end joining(GO:2001032) |
| 0.2 | 2.3 | GO:0035479 | angioblast cell migration from lateral mesoderm to midline(GO:0035479) |
| 0.2 | 3.4 | GO:0006490 | oligosaccharide-lipid intermediate biosynthetic process(GO:0006490) |
| 0.2 | 1.8 | GO:0003262 | endocardial progenitor cell migration to the midline involved in heart field formation(GO:0003262) |
| 0.2 | 1.7 | GO:0006895 | Golgi to endosome transport(GO:0006895) |
| 0.2 | 2.9 | GO:0036353 | histone H2A-K119 monoubiquitination(GO:0036353) |
| 0.2 | 3.9 | GO:0044872 | lipoprotein transport(GO:0042953) lipoprotein localization(GO:0044872) |
| 0.2 | 1.2 | GO:0006004 | fucose metabolic process(GO:0006004) |
| 0.2 | 7.3 | GO:0018231 | peptidyl-L-cysteine S-palmitoylation(GO:0018230) peptidyl-S-diacylglycerol-L-cysteine biosynthetic process from peptidyl-cysteine(GO:0018231) |
| 0.2 | 2.1 | GO:0021681 | cerebellar granular layer development(GO:0021681) cerebellar granular layer morphogenesis(GO:0021683) cerebellar granular layer formation(GO:0021684) cerebellar granule cell differentiation(GO:0021707) |
| 0.2 | 5.7 | GO:0006506 | GPI anchor biosynthetic process(GO:0006506) |
| 0.2 | 1.7 | GO:0048280 | vesicle fusion with Golgi apparatus(GO:0048280) |
| 0.2 | 0.5 | GO:0007529 | establishment of synaptic specificity at neuromuscular junction(GO:0007529) molybdenum incorporation into molybdenum-molybdopterin complex(GO:0018315) metal incorporation into metallo-molybdopterin complex(GO:0042040) glycine receptor clustering(GO:0072579) |
| 0.2 | 0.7 | GO:0007638 | mechanosensory behavior(GO:0007638) |
| 0.2 | 1.4 | GO:0030948 | negative regulation of vascular endothelial growth factor receptor signaling pathway(GO:0030948) reelin-mediated signaling pathway(GO:0038026) |
| 0.2 | 3.0 | GO:0010717 | regulation of epithelial to mesenchymal transition(GO:0010717) |
| 0.2 | 2.3 | GO:0003209 | cardiac atrium morphogenesis(GO:0003209) |
| 0.2 | 1.9 | GO:0032196 | transposition(GO:0032196) |
| 0.2 | 3.9 | GO:0007020 | microtubule nucleation(GO:0007020) |
| 0.2 | 5.4 | GO:0014029 | neural crest formation(GO:0014029) |
| 0.2 | 1.8 | GO:0016074 | snoRNA metabolic process(GO:0016074) |
| 0.2 | 3.4 | GO:0008625 | extrinsic apoptotic signaling pathway via death domain receptors(GO:0008625) |
| 0.2 | 1.2 | GO:0043154 | negative regulation of cysteine-type endopeptidase activity involved in apoptotic process(GO:0043154) |
| 0.1 | 0.4 | GO:0000973 | posttranscriptional tethering of RNA polymerase II gene DNA at nuclear periphery(GO:0000973) |
| 0.1 | 0.6 | GO:1903232 | melanosome assembly(GO:1903232) |
| 0.1 | 1.2 | GO:0007220 | Notch receptor processing(GO:0007220) |
| 0.1 | 1.6 | GO:0006903 | vesicle targeting(GO:0006903) |
| 0.1 | 2.0 | GO:0045292 | mRNA cis splicing, via spliceosome(GO:0045292) |
| 0.1 | 2.6 | GO:0036092 | phosphatidylinositol-3-phosphate biosynthetic process(GO:0036092) |
| 0.1 | 4.5 | GO:0032456 | endocytic recycling(GO:0032456) |
| 0.1 | 7.3 | GO:0042147 | retrograde transport, endosome to Golgi(GO:0042147) |
| 0.1 | 1.4 | GO:0035721 | intraciliary retrograde transport(GO:0035721) |
| 0.1 | 2.7 | GO:0000460 | maturation of 5.8S rRNA(GO:0000460) |
| 0.1 | 0.6 | GO:0042769 | DNA damage response, detection of DNA damage(GO:0042769) |
| 0.1 | 1.9 | GO:0031167 | rRNA methylation(GO:0031167) |
| 0.1 | 1.8 | GO:0042026 | protein refolding(GO:0042026) |
| 0.1 | 2.6 | GO:1902285 | semaphorin-plexin signaling pathway involved in neuron projection guidance(GO:1902285) semaphorin-plexin signaling pathway involved in axon guidance(GO:1902287) |
| 0.1 | 0.6 | GO:0045737 | positive regulation of cyclin-dependent protein serine/threonine kinase activity(GO:0045737) positive regulation of cyclin-dependent protein kinase activity(GO:1904031) |
| 0.1 | 2.2 | GO:0046660 | female sex differentiation(GO:0046660) |
| 0.1 | 0.3 | GO:0070973 | protein localization to endoplasmic reticulum exit site(GO:0070973) |
| 0.1 | 1.5 | GO:0051123 | RNA polymerase II transcriptional preinitiation complex assembly(GO:0051123) |
| 0.1 | 8.0 | GO:0006487 | protein N-linked glycosylation(GO:0006487) |
| 0.1 | 2.5 | GO:0006516 | glycoprotein catabolic process(GO:0006516) |
| 0.1 | 1.7 | GO:0051443 | positive regulation of ubiquitin-protein transferase activity(GO:0051443) |
| 0.1 | 1.4 | GO:0002084 | protein depalmitoylation(GO:0002084) macromolecule depalmitoylation(GO:0098734) |
| 0.1 | 2.9 | GO:0038061 | NIK/NF-kappaB signaling(GO:0038061) |
| 0.1 | 1.5 | GO:0060213 | regulation of nuclear-transcribed mRNA poly(A) tail shortening(GO:0060211) positive regulation of nuclear-transcribed mRNA poly(A) tail shortening(GO:0060213) regulation of nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay(GO:1900151) positive regulation of nuclear-transcribed mRNA catabolic process, deadenylation-dependent decay(GO:1900153) |
| 0.1 | 1.2 | GO:0035476 | angioblast cell migration(GO:0035476) |
| 0.1 | 1.8 | GO:0030490 | maturation of SSU-rRNA(GO:0030490) |
| 0.1 | 1.0 | GO:0032048 | cardiolipin metabolic process(GO:0032048) phosphatidylglycerol metabolic process(GO:0046471) |
| 0.1 | 2.0 | GO:0006303 | double-strand break repair via nonhomologous end joining(GO:0006303) |
| 0.1 | 3.2 | GO:0006099 | tricarboxylic acid cycle(GO:0006099) |
| 0.1 | 3.1 | GO:0042149 | cellular response to glucose starvation(GO:0042149) |
| 0.1 | 1.4 | GO:0007032 | endosome organization(GO:0007032) |
| 0.1 | 2.0 | GO:0006418 | tRNA aminoacylation for protein translation(GO:0006418) |
| 0.1 | 0.4 | GO:1903651 | positive regulation of cytoplasmic transport(GO:1903651) |
| 0.1 | 2.2 | GO:0016925 | protein sumoylation(GO:0016925) |
| 0.1 | 0.3 | GO:0090579 | transcriptional activation by promoter-enhancer looping(GO:0071733) gene looping(GO:0090202) dsDNA loop formation(GO:0090579) |
| 0.1 | 1.2 | GO:0048026 | positive regulation of mRNA splicing, via spliceosome(GO:0048026) |
| 0.1 | 0.4 | GO:0097355 | protein localization to heterochromatin(GO:0097355) |
| 0.1 | 1.7 | GO:0071173 | mitotic spindle assembly checkpoint(GO:0007094) spindle checkpoint(GO:0031577) negative regulation of mitotic metaphase/anaphase transition(GO:0045841) spindle assembly checkpoint(GO:0071173) mitotic spindle checkpoint(GO:0071174) |
| 0.1 | 3.6 | GO:0000187 | activation of MAPK activity(GO:0000187) |
| 0.1 | 2.1 | GO:0050870 | positive regulation of homotypic cell-cell adhesion(GO:0034112) positive regulation of T cell activation(GO:0050870) positive regulation of leukocyte cell-cell adhesion(GO:1903039) |
| 0.1 | 1.7 | GO:0016180 | snRNA processing(GO:0016180) |
| 0.1 | 0.6 | GO:0016119 | carotene metabolic process(GO:0016119) carotene catabolic process(GO:0016121) terpene metabolic process(GO:0042214) terpene catabolic process(GO:0046247) |
| 0.1 | 3.6 | GO:0017144 | drug metabolic process(GO:0017144) drug catabolic process(GO:0042737) exogenous drug catabolic process(GO:0042738) |
| 0.1 | 2.4 | GO:0050708 | regulation of protein secretion(GO:0050708) |
| 0.1 | 3.4 | GO:0046856 | phosphatidylinositol dephosphorylation(GO:0046856) |
| 0.1 | 0.9 | GO:0031060 | regulation of histone methylation(GO:0031060) |
| 0.1 | 0.4 | GO:0032732 | positive regulation of interleukin-1 production(GO:0032732) |
| 0.1 | 1.4 | GO:0002098 | tRNA wobble uridine modification(GO:0002098) |
| 0.1 | 4.3 | GO:0000956 | nuclear-transcribed mRNA catabolic process(GO:0000956) |
| 0.1 | 2.0 | GO:0006270 | DNA replication initiation(GO:0006270) |
| 0.1 | 0.9 | GO:0016233 | telomere capping(GO:0016233) |
| 0.1 | 2.1 | GO:0008643 | carbohydrate transport(GO:0008643) |
| 0.1 | 1.2 | GO:0072599 | protein targeting to ER(GO:0045047) establishment of protein localization to endoplasmic reticulum(GO:0072599) |
| 0.0 | 0.5 | GO:0007023 | post-chaperonin tubulin folding pathway(GO:0007023) |
| 0.0 | 0.7 | GO:0070816 | phosphorylation of RNA polymerase II C-terminal domain(GO:0070816) |
| 0.0 | 0.6 | GO:0006607 | NLS-bearing protein import into nucleus(GO:0006607) |
| 0.0 | 1.7 | GO:0010212 | response to ionizing radiation(GO:0010212) |
| 0.0 | 3.1 | GO:0018107 | peptidyl-threonine phosphorylation(GO:0018107) |
| 0.0 | 0.8 | GO:0046051 | UTP biosynthetic process(GO:0006228) UTP metabolic process(GO:0046051) |
| 0.0 | 1.7 | GO:0046466 | sphingolipid catabolic process(GO:0030149) membrane lipid catabolic process(GO:0046466) |
| 0.0 | 1.2 | GO:0030223 | neutrophil differentiation(GO:0030223) |
| 0.0 | 1.1 | GO:0048013 | ephrin receptor signaling pathway(GO:0048013) |
| 0.0 | 1.5 | GO:0045324 | late endosome to vacuole transport(GO:0045324) |
| 0.0 | 0.9 | GO:0006509 | membrane protein ectodomain proteolysis(GO:0006509) |
| 0.0 | 0.9 | GO:0008156 | negative regulation of DNA replication(GO:0008156) |
| 0.0 | 0.2 | GO:0061011 | hepatic duct development(GO:0061011) |
| 0.0 | 0.3 | GO:0045721 | negative regulation of gluconeogenesis(GO:0045721) |
| 0.0 | 1.1 | GO:0043489 | RNA stabilization(GO:0043489) |
| 0.0 | 1.1 | GO:0090148 | membrane fission(GO:0090148) |
| 0.0 | 1.5 | GO:0048634 | regulation of muscle organ development(GO:0048634) |
| 0.0 | 2.5 | GO:0034446 | substrate adhesion-dependent cell spreading(GO:0034446) |
| 0.0 | 0.8 | GO:0043153 | entrainment of circadian clock by photoperiod(GO:0043153) |
| 0.0 | 16.5 | GO:0006397 | mRNA processing(GO:0006397) |
| 0.0 | 1.4 | GO:0031122 | cytoplasmic microtubule organization(GO:0031122) |
| 0.0 | 1.7 | GO:0015718 | monocarboxylic acid transport(GO:0015718) |
| 0.0 | 1.8 | GO:0006906 | vesicle fusion(GO:0006906) |
| 0.0 | 1.2 | GO:0007249 | I-kappaB kinase/NF-kappaB signaling(GO:0007249) |
| 0.0 | 0.6 | GO:0050821 | protein stabilization(GO:0050821) |
| 0.0 | 1.2 | GO:0030308 | negative regulation of cell growth(GO:0030308) |
| 0.0 | 0.3 | GO:0071427 | mRNA export from nucleus(GO:0006406) mRNA-containing ribonucleoprotein complex export from nucleus(GO:0071427) |
| 0.0 | 0.8 | GO:1902476 | chloride transmembrane transport(GO:1902476) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.9 | 7.7 | GO:0043291 | RAVE complex(GO:0043291) |
| 1.6 | 11.0 | GO:0034388 | Pwp2p-containing subcomplex of 90S preribosome(GO:0034388) |
| 1.2 | 3.7 | GO:0031251 | PAN complex(GO:0031251) |
| 1.0 | 2.9 | GO:0042709 | succinate-CoA ligase complex(GO:0042709) |
| 1.0 | 2.9 | GO:0042382 | paraspeckles(GO:0042382) |
| 0.9 | 12.4 | GO:0008250 | oligosaccharyltransferase complex(GO:0008250) |
| 0.8 | 3.2 | GO:0036396 | MIS complex(GO:0036396) mRNA editing complex(GO:0045293) |
| 0.8 | 3.9 | GO:0033553 | rDNA heterochromatin(GO:0033553) |
| 0.6 | 2.3 | GO:0005787 | signal peptidase complex(GO:0005787) |
| 0.5 | 2.1 | GO:0005655 | nucleolar ribonuclease P complex(GO:0005655) |
| 0.5 | 5.2 | GO:0031211 | serine C-palmitoyltransferase complex(GO:0017059) endoplasmic reticulum palmitoyltransferase complex(GO:0031211) |
| 0.5 | 3.4 | GO:0016593 | Cdc73/Paf1 complex(GO:0016593) |
| 0.5 | 1.9 | GO:0008274 | gamma-tubulin large complex(GO:0000931) gamma-tubulin ring complex(GO:0008274) |
| 0.4 | 1.6 | GO:0005672 | transcription factor TFIIA complex(GO:0005672) |
| 0.4 | 2.0 | GO:0005658 | alpha DNA polymerase:primase complex(GO:0005658) |
| 0.4 | 1.6 | GO:0032584 | growth cone membrane(GO:0032584) |
| 0.4 | 2.9 | GO:0030686 | 90S preribosome(GO:0030686) |
| 0.3 | 5.9 | GO:0015030 | Cajal body(GO:0015030) |
| 0.3 | 3.8 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.3 | 7.5 | GO:0030687 | preribosome, large subunit precursor(GO:0030687) |
| 0.3 | 1.8 | GO:0070876 | SOSS complex(GO:0070876) |
| 0.3 | 2.0 | GO:0071005 | U2-type precatalytic spliceosome(GO:0071005) |
| 0.3 | 1.4 | GO:0071203 | WASH complex(GO:0071203) |
| 0.2 | 2.2 | GO:0089701 | U2AF(GO:0089701) |
| 0.2 | 2.0 | GO:0070419 | nonhomologous end joining complex(GO:0070419) |
| 0.2 | 11.1 | GO:0005811 | lipid particle(GO:0005811) |
| 0.2 | 1.2 | GO:0071986 | Ragulator complex(GO:0071986) |
| 0.2 | 6.0 | GO:0032040 | small-subunit processome(GO:0032040) |
| 0.2 | 1.1 | GO:0042765 | GPI-anchor transamidase complex(GO:0042765) |
| 0.2 | 1.4 | GO:0030991 | intraciliary transport particle A(GO:0030991) |
| 0.2 | 0.6 | GO:0031429 | box H/ACA snoRNP complex(GO:0031429) box H/ACA RNP complex(GO:0072588) |
| 0.2 | 1.2 | GO:0070765 | gamma-secretase complex(GO:0070765) |
| 0.2 | 3.0 | GO:0046540 | U4/U6 x U5 tri-snRNP complex(GO:0046540) |
| 0.1 | 1.3 | GO:0005682 | U5 snRNP(GO:0005682) |
| 0.1 | 1.5 | GO:0070449 | elongin complex(GO:0070449) |
| 0.1 | 0.6 | GO:0031085 | BLOC-3 complex(GO:0031085) |
| 0.1 | 2.0 | GO:0005868 | cytoplasmic dynein complex(GO:0005868) |
| 0.1 | 1.4 | GO:0033588 | Elongator holoenzyme complex(GO:0033588) |
| 0.1 | 0.6 | GO:0000243 | commitment complex(GO:0000243) |
| 0.1 | 1.8 | GO:0000815 | ESCRT III complex(GO:0000815) |
| 0.1 | 2.6 | GO:0002116 | semaphorin receptor complex(GO:0002116) |
| 0.1 | 2.9 | GO:0035102 | PRC1 complex(GO:0035102) |
| 0.1 | 8.4 | GO:0030672 | synaptic vesicle membrane(GO:0030672) exocytic vesicle membrane(GO:0099501) |
| 0.1 | 1.9 | GO:0005885 | Arp2/3 protein complex(GO:0005885) |
| 0.1 | 2.3 | GO:0000782 | telomere cap complex(GO:0000782) nuclear telomere cap complex(GO:0000783) nuclear chromosome, telomeric region(GO:0000784) |
| 0.1 | 7.2 | GO:0030173 | integral component of Golgi membrane(GO:0030173) intrinsic component of Golgi membrane(GO:0031228) |
| 0.1 | 4.3 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.1 | 8.5 | GO:0055037 | recycling endosome(GO:0055037) |
| 0.1 | 1.7 | GO:0032039 | integrator complex(GO:0032039) |
| 0.1 | 5.9 | GO:0031201 | SNARE complex(GO:0031201) |
| 0.1 | 0.7 | GO:0005675 | holo TFIIH complex(GO:0005675) |
| 0.1 | 0.2 | GO:1990726 | Lsm1-7-Pat1 complex(GO:1990726) |
| 0.1 | 1.3 | GO:0016281 | eukaryotic translation initiation factor 4F complex(GO:0016281) |
| 0.1 | 0.6 | GO:0034451 | centriolar satellite(GO:0034451) |
| 0.1 | 0.8 | GO:0031307 | integral component of mitochondrial outer membrane(GO:0031307) |
| 0.1 | 4.0 | GO:0072686 | mitotic spindle(GO:0072686) |
| 0.1 | 4.1 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.1 | 3.0 | GO:0005793 | endoplasmic reticulum-Golgi intermediate compartment(GO:0005793) |
| 0.1 | 6.6 | GO:0005769 | early endosome(GO:0005769) |
| 0.1 | 33.8 | GO:0005789 | endoplasmic reticulum membrane(GO:0005789) |
| 0.1 | 0.4 | GO:0016586 | RSC complex(GO:0016586) |
| 0.1 | 2.2 | GO:0030496 | midbody(GO:0030496) |
| 0.0 | 0.2 | GO:0045257 | mitochondrial respiratory chain complex II, succinate dehydrogenase complex (ubiquinone)(GO:0005749) succinate dehydrogenase complex (ubiquinone)(GO:0045257) respiratory chain complex II(GO:0045273) succinate dehydrogenase complex(GO:0045281) fumarate reductase complex(GO:0045283) |
| 0.0 | 1.1 | GO:0046658 | anchored component of plasma membrane(GO:0046658) |
| 0.0 | 5.0 | GO:0016324 | apical plasma membrane(GO:0016324) |
| 0.0 | 0.3 | GO:0000347 | THO complex(GO:0000347) THO complex part of transcription export complex(GO:0000445) |
| 0.0 | 31.0 | GO:0045202 | synapse(GO:0045202) |
| 0.0 | 0.4 | GO:0031080 | nuclear pore outer ring(GO:0031080) |
| 0.0 | 3.7 | GO:0005604 | basement membrane(GO:0005604) |
| 0.0 | 3.5 | GO:0016607 | nuclear speck(GO:0016607) |
| 0.0 | 0.2 | GO:0071818 | BAT3 complex(GO:0071818) |
| 0.0 | 2.5 | GO:0019005 | SCF ubiquitin ligase complex(GO:0019005) |
| 0.0 | 2.3 | GO:0005871 | kinesin complex(GO:0005871) |
| 0.0 | 2.2 | GO:0030027 | lamellipodium(GO:0030027) |
| 0.0 | 1.3 | GO:0000118 | histone deacetylase complex(GO:0000118) |
| 0.0 | 1.5 | GO:0030136 | clathrin-coated vesicle(GO:0030136) |
| 0.0 | 1.7 | GO:0000776 | kinetochore(GO:0000776) |
| 0.0 | 0.4 | GO:0005921 | gap junction(GO:0005921) |
| 0.0 | 0.7 | GO:0017053 | transcriptional repressor complex(GO:0017053) |
| 0.0 | 4.5 | GO:0000139 | Golgi membrane(GO:0000139) |
| 0.0 | 0.4 | GO:0005680 | anaphase-promoting complex(GO:0005680) |
| 0.0 | 0.6 | GO:0001725 | stress fiber(GO:0001725) actomyosin(GO:0042641) contractile actin filament bundle(GO:0097517) |
| 0.0 | 1.1 | GO:0010008 | endosome membrane(GO:0010008) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.4 | 4.2 | GO:0043812 | phosphatidylinositol-4-phosphate phosphatase activity(GO:0043812) |
| 1.1 | 3.4 | GO:0004377 | GDP-Man:Man3GlcNAc2-PP-Dol alpha-1,2-mannosyltransferase activity(GO:0004377) |
| 1.1 | 3.3 | GO:0046964 | 3'-phosphoadenosine 5'-phosphosulfate transmembrane transporter activity(GO:0046964) |
| 1.0 | 8.2 | GO:0031419 | cobalamin binding(GO:0031419) |
| 1.0 | 11.3 | GO:0047676 | arachidonate-CoA ligase activity(GO:0047676) |
| 1.0 | 4.1 | GO:0004534 | 5'-3' exoribonuclease activity(GO:0004534) |
| 1.0 | 5.1 | GO:0005462 | UDP-N-acetylglucosamine transmembrane transporter activity(GO:0005462) |
| 1.0 | 5.1 | GO:0004385 | guanylate kinase activity(GO:0004385) |
| 1.0 | 2.9 | GO:0004774 | succinate-CoA ligase activity(GO:0004774) succinate-CoA ligase (ADP-forming) activity(GO:0004775) succinate-CoA ligase (GDP-forming) activity(GO:0004776) |
| 0.9 | 3.6 | GO:0070324 | thyroid hormone binding(GO:0070324) |
| 0.8 | 2.5 | GO:0070905 | glycine hydroxymethyltransferase activity(GO:0004372) serine binding(GO:0070905) |
| 0.8 | 4.6 | GO:0033781 | cholesterol 24-hydroxylase activity(GO:0033781) |
| 0.8 | 3.8 | GO:0015180 | L-alanine transmembrane transporter activity(GO:0015180) L-proline transmembrane transporter activity(GO:0015193) alanine transmembrane transporter activity(GO:0022858) |
| 0.7 | 5.2 | GO:0004758 | serine C-palmitoyltransferase activity(GO:0004758) C-palmitoyltransferase activity(GO:0016454) |
| 0.7 | 2.2 | GO:0071532 | ankyrin repeat binding(GO:0071532) |
| 0.7 | 2.2 | GO:0004342 | glucosamine-6-phosphate deaminase activity(GO:0004342) |
| 0.7 | 2.9 | GO:0034511 | U3 snoRNA binding(GO:0034511) |
| 0.7 | 3.6 | GO:0032183 | SUMO binding(GO:0032183) |
| 0.7 | 3.4 | GO:1990756 | protein binding, bridging involved in substrate recognition for ubiquitination(GO:1990756) |
| 0.7 | 3.4 | GO:0030620 | U2 snRNA binding(GO:0030620) |
| 0.7 | 2.7 | GO:0016263 | glycoprotein-N-acetylgalactosamine 3-beta-galactosyltransferase activity(GO:0016263) beta-1,3-galactosyltransferase activity(GO:0048531) |
| 0.6 | 2.4 | GO:0004066 | asparagine synthase (glutamine-hydrolyzing) activity(GO:0004066) |
| 0.6 | 2.4 | GO:0008746 | NAD(P)+ transhydrogenase activity(GO:0008746) oxidoreductase activity, acting on NAD(P)H, NAD(P) as acceptor(GO:0016652) |
| 0.5 | 2.2 | GO:0003979 | UDP-glucose 6-dehydrogenase activity(GO:0003979) |
| 0.5 | 5.4 | GO:0016936 | galactoside binding(GO:0016936) |
| 0.5 | 2.7 | GO:0017113 | uracil binding(GO:0002058) pyrimidine nucleobase binding(GO:0002061) dihydropyrimidine dehydrogenase (NADP+) activity(GO:0017113) |
| 0.5 | 2.1 | GO:0033204 | ribonuclease P RNA binding(GO:0033204) |
| 0.5 | 2.0 | GO:0042392 | sphingosine-1-phosphate phosphatase activity(GO:0042392) |
| 0.5 | 3.2 | GO:0015189 | L-lysine transmembrane transporter activity(GO:0015189) |
| 0.4 | 1.7 | GO:0008117 | sphinganine-1-phosphate aldolase activity(GO:0008117) |
| 0.4 | 1.9 | GO:0000179 | rRNA (adenine-N6,N6-)-dimethyltransferase activity(GO:0000179) |
| 0.4 | 1.5 | GO:0016308 | 1-phosphatidylinositol-4-phosphate 5-kinase activity(GO:0016308) |
| 0.4 | 13.8 | GO:0030145 | manganese ion binding(GO:0030145) |
| 0.4 | 1.8 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 0.4 | 1.4 | GO:0003827 | alpha-1,3-mannosylglycoprotein 2-beta-N-acetylglucosaminyltransferase activity(GO:0003827) |
| 0.3 | 1.7 | GO:0001735 | prenylcysteine oxidase activity(GO:0001735) |
| 0.3 | 1.6 | GO:0001042 | RNA polymerase I core binding(GO:0001042) |
| 0.3 | 3.4 | GO:0052813 | phosphatidylinositol-4,5-bisphosphate 3-kinase activity(GO:0046934) phosphatidylinositol-3,4-bisphosphate 5-kinase activity(GO:0052812) phosphatidylinositol bisphosphate kinase activity(GO:0052813) |
| 0.3 | 2.5 | GO:0045309 | protein phosphorylated amino acid binding(GO:0045309) phosphoprotein binding(GO:0051219) |
| 0.3 | 9.6 | GO:0043539 | protein serine/threonine kinase activator activity(GO:0043539) |
| 0.3 | 2.9 | GO:0035925 | AU-rich element binding(GO:0017091) mRNA 3'-UTR AU-rich region binding(GO:0035925) |
| 0.3 | 2.1 | GO:0034452 | dynactin binding(GO:0034452) |
| 0.2 | 3.7 | GO:0004535 | poly(A)-specific ribonuclease activity(GO:0004535) |
| 0.2 | 1.7 | GO:0019136 | deoxynucleoside kinase activity(GO:0019136) |
| 0.2 | 2.2 | GO:0030628 | pre-mRNA 3'-splice site binding(GO:0030628) |
| 0.2 | 1.7 | GO:0001130 | bacterial-type RNA polymerase transcription factor activity, sequence-specific DNA binding(GO:0001130) bacterial-type RNA polymerase transcriptional repressor activity, sequence-specific DNA binding(GO:0001217) |
| 0.2 | 1.3 | GO:0047429 | nucleoside-triphosphate diphosphatase activity(GO:0047429) |
| 0.2 | 0.6 | GO:0034513 | box H/ACA snoRNA binding(GO:0034513) |
| 0.2 | 0.8 | GO:0016427 | tRNA (cytosine) methyltransferase activity(GO:0016427) |
| 0.2 | 1.0 | GO:0047184 | 1-acylglycerophosphocholine O-acyltransferase activity(GO:0047184) |
| 0.2 | 7.5 | GO:0017075 | syntaxin-1 binding(GO:0017075) |
| 0.2 | 0.5 | GO:0061599 | molybdopterin adenylyltransferase activity(GO:0061598) molybdopterin molybdotransferase activity(GO:0061599) |
| 0.2 | 2.2 | GO:0061665 | SUMO ligase activity(GO:0061665) |
| 0.2 | 3.6 | GO:0004708 | MAP kinase kinase activity(GO:0004708) |
| 0.2 | 7.3 | GO:0019707 | protein-cysteine S-palmitoyltransferase activity(GO:0019706) protein-cysteine S-acyltransferase activity(GO:0019707) |
| 0.2 | 0.6 | GO:0051377 | mannose-ethanolamine phosphotransferase activity(GO:0051377) |
| 0.1 | 3.4 | GO:0017134 | fibroblast growth factor binding(GO:0017134) |
| 0.1 | 3.1 | GO:0004712 | protein serine/threonine/tyrosine kinase activity(GO:0004712) |
| 0.1 | 2.4 | GO:0045159 | myosin II binding(GO:0045159) |
| 0.1 | 0.6 | GO:0003958 | NADPH-hemoprotein reductase activity(GO:0003958) |
| 0.1 | 3.8 | GO:0016208 | AMP binding(GO:0016208) |
| 0.1 | 1.6 | GO:0016783 | sulfurtransferase activity(GO:0016783) |
| 0.1 | 1.8 | GO:0044183 | protein binding involved in protein folding(GO:0044183) |
| 0.1 | 3.4 | GO:0005385 | zinc ion transmembrane transporter activity(GO:0005385) |
| 0.1 | 2.9 | GO:1990841 | promoter-specific chromatin binding(GO:1990841) |
| 0.1 | 1.9 | GO:0043015 | gamma-tubulin binding(GO:0043015) |
| 0.1 | 1.5 | GO:0032050 | clathrin heavy chain binding(GO:0032050) |
| 0.1 | 3.9 | GO:0035064 | methylated histone binding(GO:0035064) |
| 0.1 | 2.8 | GO:0031418 | L-ascorbic acid binding(GO:0031418) |
| 0.1 | 1.6 | GO:0017025 | TBP-class protein binding(GO:0017025) |
| 0.1 | 0.8 | GO:0001222 | transcription corepressor binding(GO:0001222) |
| 0.1 | 2.4 | GO:0001103 | RNA polymerase II repressing transcription factor binding(GO:0001103) |
| 0.1 | 5.4 | GO:0005484 | SNAP receptor activity(GO:0005484) |
| 0.1 | 0.6 | GO:0030619 | U1 snRNA binding(GO:0030619) |
| 0.1 | 1.3 | GO:0016671 | oxidoreductase activity, acting on a sulfur group of donors, disulfide as acceptor(GO:0016671) |
| 0.1 | 1.7 | GO:0097602 | cullin family protein binding(GO:0097602) |
| 0.1 | 2.6 | GO:0017154 | semaphorin receptor activity(GO:0017154) |
| 0.1 | 3.0 | GO:0015036 | disulfide oxidoreductase activity(GO:0015036) |
| 0.1 | 2.9 | GO:0004707 | MAP kinase activity(GO:0004707) |
| 0.1 | 6.5 | GO:0016763 | transferase activity, transferring pentosyl groups(GO:0016763) |
| 0.1 | 2.0 | GO:0016875 | aminoacyl-tRNA ligase activity(GO:0004812) ligase activity, forming carbon-oxygen bonds(GO:0016875) ligase activity, forming aminoacyl-tRNA and related compounds(GO:0016876) |
| 0.1 | 1.8 | GO:0005154 | epidermal growth factor receptor binding(GO:0005154) |
| 0.1 | 3.7 | GO:0003755 | peptidyl-prolyl cis-trans isomerase activity(GO:0003755) |
| 0.1 | 1.2 | GO:0008417 | fucosyltransferase activity(GO:0008417) |
| 0.1 | 0.6 | GO:0010436 | beta-carotene 15,15'-monooxygenase activity(GO:0003834) carotenoid dioxygenase activity(GO:0010436) |
| 0.1 | 5.9 | GO:0008146 | sulfotransferase activity(GO:0008146) |
| 0.1 | 2.3 | GO:0008574 | ATP-dependent microtubule motor activity, plus-end-directed(GO:0008574) |
| 0.1 | 1.4 | GO:0098599 | palmitoyl-(protein) hydrolase activity(GO:0008474) palmitoyl hydrolase activity(GO:0098599) |
| 0.1 | 0.6 | GO:0000774 | adenyl-nucleotide exchange factor activity(GO:0000774) |
| 0.1 | 5.8 | GO:0005544 | calcium-dependent phospholipid binding(GO:0005544) |
| 0.1 | 2.0 | GO:0051010 | microtubule plus-end binding(GO:0051010) |
| 0.1 | 5.4 | GO:0008201 | heparin binding(GO:0008201) |
| 0.1 | 0.8 | GO:0008353 | RNA polymerase II carboxy-terminal domain kinase activity(GO:0008353) |
| 0.1 | 3.6 | GO:0016712 | oxidoreductase activity, acting on paired donors, with incorporation or reduction of molecular oxygen, reduced flavin or flavoprotein as one donor, and incorporation of one atom of oxygen(GO:0016712) |
| 0.1 | 2.6 | GO:0016776 | phosphotransferase activity, phosphate group as acceptor(GO:0016776) |
| 0.1 | 4.0 | GO:0004722 | protein serine/threonine phosphatase activity(GO:0004722) |
| 0.0 | 0.5 | GO:0043014 | alpha-tubulin binding(GO:0043014) |
| 0.0 | 1.4 | GO:0030165 | PDZ domain binding(GO:0030165) |
| 0.0 | 2.3 | GO:0030898 | actin-dependent ATPase activity(GO:0030898) |
| 0.0 | 0.2 | GO:0039706 | co-receptor binding(GO:0039706) |
| 0.0 | 1.1 | GO:0015248 | sterol transporter activity(GO:0015248) |
| 0.0 | 0.9 | GO:0043175 | RNA polymerase core enzyme binding(GO:0043175) |
| 0.0 | 1.4 | GO:0019003 | GDP binding(GO:0019003) |
| 0.0 | 3.1 | GO:0003678 | DNA helicase activity(GO:0003678) |
| 0.0 | 0.6 | GO:0061608 | nuclear import signal receptor activity(GO:0061608) |
| 0.0 | 2.2 | GO:0003697 | single-stranded DNA binding(GO:0003697) |
| 0.0 | 0.5 | GO:0008239 | dipeptidyl-peptidase activity(GO:0008239) |
| 0.0 | 0.2 | GO:0048039 | ubiquinone binding(GO:0048039) |
| 0.0 | 1.7 | GO:0008028 | monocarboxylic acid transmembrane transporter activity(GO:0008028) |
| 0.0 | 0.8 | GO:0004890 | GABA-A receptor activity(GO:0004890) |
| 0.0 | 2.1 | GO:0043130 | ubiquitin binding(GO:0043130) |
| 0.0 | 1.2 | GO:0061631 | ubiquitin conjugating enzyme activity(GO:0061631) |
| 0.0 | 4.6 | GO:0042802 | identical protein binding(GO:0042802) |
| 0.0 | 0.2 | GO:0004300 | enoyl-CoA hydratase activity(GO:0004300) |
| 0.0 | 0.3 | GO:0036312 | phosphatidylinositol 3-kinase regulatory subunit binding(GO:0036312) |
| 0.0 | 1.2 | GO:0030674 | protein binding, bridging(GO:0030674) |
| 0.0 | 1.5 | GO:0004715 | non-membrane spanning protein tyrosine kinase activity(GO:0004715) |
| 0.0 | 3.8 | GO:0008514 | organic anion transmembrane transporter activity(GO:0008514) |
| 0.0 | 0.4 | GO:0022829 | wide pore channel activity(GO:0022829) |
| 0.0 | 0.4 | GO:0017056 | structural constituent of nuclear pore(GO:0017056) |
| 0.0 | 0.8 | GO:0017124 | SH3 domain binding(GO:0017124) |
| 0.0 | 2.0 | GO:0003774 | motor activity(GO:0003774) |
| 0.0 | 1.8 | GO:0008134 | transcription factor binding(GO:0008134) |
| 0.0 | 0.9 | GO:0003724 | RNA helicase activity(GO:0003724) |
| 0.0 | 0.4 | GO:0022833 | mechanically-gated ion channel activity(GO:0008381) mechanically gated channel activity(GO:0022833) |
| 0.0 | 0.1 | GO:0030515 | snoRNA binding(GO:0030515) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 1.4 | ST INTERFERON GAMMA PATHWAY | Interferon gamma pathway. |
| 0.3 | 4.6 | PID S1P S1P2 PATHWAY | S1P2 pathway |
| 0.3 | 2.0 | PID S1P META PATHWAY | Sphingosine 1-phosphate (S1P) pathway |
| 0.2 | 2.9 | PID P38 MK2 PATHWAY | p38 signaling mediated by MAPKAP kinases |
| 0.2 | 0.7 | PID THROMBIN PAR4 PATHWAY | PAR4-mediated thrombin signaling events |
| 0.2 | 3.6 | ST GA12 PATHWAY | G alpha 12 Pathway |
| 0.2 | 3.6 | PID RXR VDR PATHWAY | RXR and RAR heterodimerization with other nuclear receptor |
| 0.2 | 2.2 | PID IL12 STAT4 PATHWAY | IL12 signaling mediated by STAT4 |
| 0.2 | 1.5 | PID NFKAPPAB ATYPICAL PATHWAY | Atypical NF-kappaB pathway |
| 0.1 | 5.5 | PID CDC42 REG PATHWAY | Regulation of CDC42 activity |
| 0.1 | 3.4 | PID NEPHRIN NEPH1 PATHWAY | Nephrin/Neph1 signaling in the kidney podocyte |
| 0.1 | 1.0 | PID RET PATHWAY | Signaling events regulated by Ret tyrosine kinase |
| 0.1 | 3.8 | PID VEGFR1 2 PATHWAY | Signaling events mediated by VEGFR1 and VEGFR2 |
| 0.1 | 0.7 | SA G2 AND M PHASES | Cdc25 activates the cdc2/cyclin B complex to induce the G2/M transition. |
| 0.1 | 3.7 | PID NFAT 3PATHWAY | Role of Calcineurin-dependent NFAT signaling in lymphocytes |
| 0.1 | 2.9 | PID IL12 2PATHWAY | IL12-mediated signaling events |
| 0.1 | 4.0 | PID PLK1 PATHWAY | PLK1 signaling events |
| 0.1 | 1.2 | PID SYNDECAN 3 PATHWAY | Syndecan-3-mediated signaling events |
| 0.1 | 1.8 | PID AJDISS 2PATHWAY | Posttranslational regulation of adherens junction stability and dissassembly |
| 0.1 | 1.8 | PID ERBB1 INTERNALIZATION PATHWAY | Internalization of ErbB1 |
| 0.1 | 3.4 | ST FAS SIGNALING PATHWAY | Fas Signaling Pathway |
| 0.1 | 2.1 | PID FAK PATHWAY | Signaling events mediated by focal adhesion kinase |
| 0.1 | 2.2 | PID ECADHERIN STABILIZATION PATHWAY | Stabilization and expansion of the E-cadherin adherens junction |
| 0.0 | 2.5 | PID MYC ACTIV PATHWAY | Validated targets of C-MYC transcriptional activation |
| 0.0 | 3.2 | PID SMAD2 3NUCLEAR PATHWAY | Regulation of nuclear SMAD2/3 signaling |
| 0.0 | 1.2 | PID TELOMERASE PATHWAY | Regulation of Telomerase |
| 0.0 | 0.7 | PID HES HEY PATHWAY | Notch-mediated HES/HEY network |
| 0.0 | 0.6 | PID IL1 PATHWAY | IL1-mediated signaling events |
| 0.0 | 0.6 | PID AP1 PATHWAY | AP-1 transcription factor network |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 13.4 | REACTOME REGULATION OF AMPK ACTIVITY VIA LKB1 | Genes involved in Regulation of AMPK activity via LKB1 |
| 1.1 | 7.5 | REACTOME DSCAM INTERACTIONS | Genes involved in DSCAM interactions |
| 0.8 | 6.5 | REACTOME PYRIMIDINE CATABOLISM | Genes involved in Pyrimidine catabolism |
| 0.5 | 4.4 | REACTOME ADVANCED GLYCOSYLATION ENDPRODUCT RECEPTOR SIGNALING | Genes involved in Advanced glycosylation endproduct receptor signaling |
| 0.3 | 4.3 | REACTOME SYNTHESIS OF PE | Genes involved in Synthesis of PE |
| 0.3 | 5.4 | REACTOME CITRIC ACID CYCLE TCA CYCLE | Genes involved in Citric acid cycle (TCA cycle) |
| 0.2 | 3.4 | REACTOME ZINC TRANSPORTERS | Genes involved in Zinc transporters |
| 0.2 | 4.9 | REACTOME REGULATION OF SIGNALING BY CBL | Genes involved in Regulation of signaling by CBL |
| 0.2 | 2.3 | REACTOME SYNTHESIS SECRETION AND INACTIVATION OF GLP1 | Genes involved in Synthesis, Secretion, and Inactivation of Glucagon-like Peptide-1 (GLP-1) |
| 0.2 | 2.6 | REACTOME DESTABILIZATION OF MRNA BY TRISTETRAPROLIN TTP | Genes involved in Destabilization of mRNA by Tristetraprolin (TTP) |
| 0.2 | 5.0 | REACTOME ABC FAMILY PROTEINS MEDIATED TRANSPORT | Genes involved in ABC-family proteins mediated transport |
| 0.2 | 1.9 | REACTOME RECRUITMENT OF NUMA TO MITOTIC CENTROSOMES | Genes involved in Recruitment of NuMA to mitotic centrosomes |
| 0.2 | 1.7 | REACTOME PURINE SALVAGE | Genes involved in Purine salvage |
| 0.2 | 2.0 | REACTOME INHIBITION OF REPLICATION INITIATION OF DAMAGED DNA BY RB1 E2F1 | Genes involved in Inhibition of replication initiation of damaged DNA by RB1/E2F1 |
| 0.2 | 3.8 | REACTOME AMINO ACID TRANSPORT ACROSS THE PLASMA MEMBRANE | Genes involved in Amino acid transport across the plasma membrane |
| 0.1 | 4.2 | REACTOME CYTOSOLIC TRNA AMINOACYLATION | Genes involved in Cytosolic tRNA aminoacylation |
| 0.1 | 1.8 | REACTOME ACTIVATION OF RAC | Genes involved in Activation of Rac |
| 0.1 | 1.9 | REACTOME CELL EXTRACELLULAR MATRIX INTERACTIONS | Genes involved in Cell-extracellular matrix interactions |
| 0.1 | 2.9 | REACTOME PROCESSING OF INTRONLESS PRE MRNAS | Genes involved in Processing of Intronless Pre-mRNAs |
| 0.1 | 1.2 | REACTOME SIGNALING BY NOTCH4 | Genes involved in Signaling by NOTCH4 |
| 0.1 | 4.3 | REACTOME MRNA SPLICING MINOR PATHWAY | Genes involved in mRNA Splicing - Minor Pathway |
| 0.1 | 1.9 | REACTOME PROTEOLYTIC CLEAVAGE OF SNARE COMPLEX PROTEINS | Genes involved in Proteolytic cleavage of SNARE complex proteins |
| 0.1 | 2.0 | REACTOME TRNA AMINOACYLATION | Genes involved in tRNA Aminoacylation |
| 0.1 | 1.6 | REACTOME INSULIN SYNTHESIS AND PROCESSING | Genes involved in Insulin Synthesis and Processing |
| 0.1 | 1.7 | REACTOME BILE SALT AND ORGANIC ANION SLC TRANSPORTERS | Genes involved in Bile salt and organic anion SLC transporters |
| 0.1 | 4.1 | REACTOME ASSOCIATION OF TRIC CCT WITH TARGET PROTEINS DURING BIOSYNTHESIS | Genes involved in Association of TriC/CCT with target proteins during biosynthesis |
| 0.1 | 1.3 | REACTOME PACKAGING OF TELOMERE ENDS | Genes involved in Packaging Of Telomere Ends |
| 0.1 | 10.1 | REACTOME ASPARAGINE N LINKED GLYCOSYLATION | Genes involved in Asparagine N-linked glycosylation |
| 0.1 | 2.2 | REACTOME PHASE II CONJUGATION | Genes involved in Phase II conjugation |
| 0.1 | 2.1 | REACTOME INHIBITION OF THE PROTEOLYTIC ACTIVITY OF APC C REQUIRED FOR THE ONSET OF ANAPHASE BY MITOTIC SPINDLE CHECKPOINT COMPONENTS | Genes involved in Inhibition of the proteolytic activity of APC/C required for the onset of anaphase by mitotic spindle checkpoint components |
| 0.1 | 2.0 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.1 | 0.7 | REACTOME REGULATION OF INSULIN SECRETION BY ACETYLCHOLINE | Genes involved in Regulation of Insulin Secretion by Acetylcholine |
| 0.1 | 3.6 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.1 | 4.1 | REACTOME METABOLISM OF VITAMINS AND COFACTORS | Genes involved in Metabolism of vitamins and cofactors |
| 0.1 | 2.0 | REACTOME SPHINGOLIPID DE NOVO BIOSYNTHESIS | Genes involved in Sphingolipid de novo biosynthesis |
| 0.1 | 2.2 | REACTOME RNA POL II TRANSCRIPTION PRE INITIATION AND PROMOTER OPENING | Genes involved in RNA Polymerase II Transcription Pre-Initiation And Promoter Opening |
| 0.1 | 1.7 | REACTOME POST TRANSLATIONAL MODIFICATION SYNTHESIS OF GPI ANCHORED PROTEINS | Genes involved in Post-translational modification: synthesis of GPI-anchored proteins |
| 0.1 | 1.8 | REACTOME ENDOSOMAL SORTING COMPLEX REQUIRED FOR TRANSPORT ESCRT | Genes involved in Endosomal Sorting Complex Required For Transport (ESCRT) |
| 0.1 | 1.5 | REACTOME TRANSPORT OF VITAMINS NUCLEOSIDES AND RELATED MOLECULES | Genes involved in Transport of vitamins, nucleosides, and related molecules |
| 0.1 | 0.8 | REACTOME GABA A RECEPTOR ACTIVATION | Genes involved in GABA A receptor activation |
| 0.0 | 1.6 | REACTOME O LINKED GLYCOSYLATION OF MUCINS | Genes involved in O-linked glycosylation of mucins |
| 0.0 | 2.7 | REACTOME MHC CLASS II ANTIGEN PRESENTATION | Genes involved in MHC class II antigen presentation |
| 0.0 | 3.6 | REACTOME NGF SIGNALLING VIA TRKA FROM THE PLASMA MEMBRANE | Genes involved in NGF signalling via TRKA from the plasma membrane |
| 0.0 | 4.4 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 1.2 | REACTOME ACTIVATION OF CHAPERONE GENES BY XBP1S | Genes involved in Activation of Chaperone Genes by XBP1(S) |
| 0.0 | 1.3 | REACTOME GOLGI ASSOCIATED VESICLE BIOGENESIS | Genes involved in Golgi Associated Vesicle Biogenesis |
| 0.0 | 0.5 | REACTOME POST CHAPERONIN TUBULIN FOLDING PATHWAY | Genes involved in Post-chaperonin tubulin folding pathway |
| 0.0 | 0.6 | REACTOME LOSS OF NLP FROM MITOTIC CENTROSOMES | Genes involved in Loss of Nlp from mitotic centrosomes |
| 0.0 | 1.0 | REACTOME ANTIVIRAL MECHANISM BY IFN STIMULATED GENES | Genes involved in Antiviral mechanism by IFN-stimulated genes |
| 0.0 | 0.6 | REACTOME IL1 SIGNALING | Genes involved in Interleukin-1 signaling |
| 0.0 | 1.0 | REACTOME GLYCEROPHOSPHOLIPID BIOSYNTHESIS | Genes involved in Glycerophospholipid biosynthesis |
| 0.0 | 0.3 | REACTOME APOPTOTIC CLEAVAGE OF CELLULAR PROTEINS | Genes involved in Apoptotic cleavage of cellular proteins |