Project
NHBE cells infected with SARS-CoV-2 Analysis Results (GEO series: GSE147507)
Navigation
Downloads
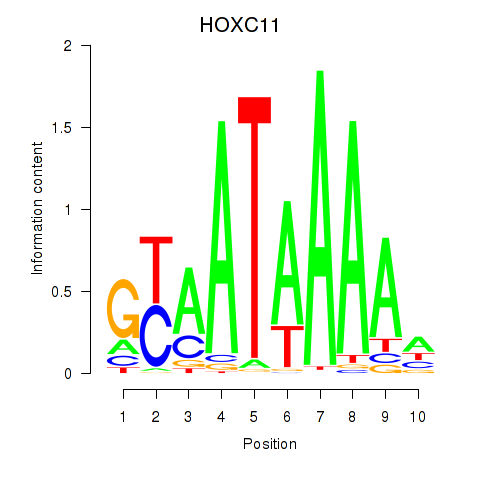
Results for HOXC11
Z-value: 0.85
Transcription factors associated with HOXC11
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
HOXC11
|
ENSG00000123388.4 | homeobox C11 |
Activity profile of HOXC11 motif
Sorted Z-values of HOXC11 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr7_+_22766766 | 1.63 |
ENST00000426291.1
ENST00000401651.1 ENST00000258743.5 ENST00000420258.2 ENST00000407492.1 ENST00000401630.3 ENST00000406575.1 |
IL6
|
interleukin 6 (interferon, beta 2) |
| chr2_+_228678550 | 1.46 |
ENST00000409189.3
ENST00000358813.4 |
CCL20
|
chemokine (C-C motif) ligand 20 |
| chr6_+_12290586 | 1.06 |
ENST00000379375.5
|
EDN1
|
endothelin 1 |
| chr7_-_41742697 | 0.88 |
ENST00000242208.4
|
INHBA
|
inhibin, beta A |
| chr18_+_68002675 | 0.84 |
ENST00000584919.1
|
RP11-41O4.1
|
Uncharacterized protein |
| chr6_+_32812568 | 0.65 |
ENST00000414474.1
|
PSMB9
|
proteasome (prosome, macropain) subunit, beta type, 9 |
| chr8_-_73793975 | 0.64 |
ENST00000523881.1
|
RP11-1145L24.1
|
RP11-1145L24.1 |
| chr22_-_30642782 | 0.57 |
ENST00000249075.3
|
LIF
|
leukemia inhibitory factor |
| chr6_+_26156551 | 0.56 |
ENST00000304218.3
|
HIST1H1E
|
histone cluster 1, H1e |
| chr13_-_81801115 | 0.44 |
ENST00000567258.1
|
LINC00564
|
long intergenic non-protein coding RNA 564 |
| chr10_-_106240032 | 0.42 |
ENST00000447860.1
|
RP11-127O4.3
|
RP11-127O4.3 |
| chr7_+_141418962 | 0.41 |
ENST00000493845.1
|
WEE2
|
WEE1 homolog 2 (S. pombe) |
| chr1_+_229440129 | 0.39 |
ENST00000366688.3
|
SPHAR
|
S-phase response (cyclin related) |
| chr8_-_109799793 | 0.39 |
ENST00000297459.3
|
TMEM74
|
transmembrane protein 74 |
| chr12_-_52828147 | 0.35 |
ENST00000252245.5
|
KRT75
|
keratin 75 |
| chr8_-_95220775 | 0.35 |
ENST00000441892.2
ENST00000521491.1 ENST00000027335.3 |
CDH17
|
cadherin 17, LI cadherin (liver-intestine) |
| chr5_-_115872124 | 0.33 |
ENST00000515009.1
|
SEMA6A
|
sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6A |
| chr17_+_7533439 | 0.29 |
ENST00000441599.2
ENST00000380450.4 ENST00000416273.3 ENST00000575903.1 ENST00000576830.1 ENST00000571153.1 ENST00000575618.1 ENST00000576152.1 |
SHBG
|
sex hormone-binding globulin |
| chr7_-_27205136 | 0.28 |
ENST00000396345.1
ENST00000343483.6 |
HOXA9
|
homeobox A9 |
| chr9_-_133814455 | 0.27 |
ENST00000448616.1
|
FIBCD1
|
fibrinogen C domain containing 1 |
| chr12_-_123201337 | 0.26 |
ENST00000528880.2
|
HCAR3
|
hydroxycarboxylic acid receptor 3 |
| chr1_+_209878182 | 0.25 |
ENST00000367027.3
|
HSD11B1
|
hydroxysteroid (11-beta) dehydrogenase 1 |
| chr5_-_159766528 | 0.25 |
ENST00000505287.2
|
CCNJL
|
cyclin J-like |
| chr1_-_204135450 | 0.25 |
ENST00000272190.8
ENST00000367195.2 |
REN
|
renin |
| chr17_-_77924627 | 0.24 |
ENST00000572862.1
ENST00000573782.1 ENST00000574427.1 ENST00000570373.1 ENST00000340848.7 ENST00000576768.1 |
TBC1D16
|
TBC1 domain family, member 16 |
| chr19_+_49468558 | 0.24 |
ENST00000331825.6
|
FTL
|
ferritin, light polypeptide |
| chrX_-_133792480 | 0.23 |
ENST00000359237.4
|
PLAC1
|
placenta-specific 1 |
| chr5_-_115872142 | 0.23 |
ENST00000510263.1
|
SEMA6A
|
sema domain, transmembrane domain (TM), and cytoplasmic domain, (semaphorin) 6A |
| chr5_+_176731572 | 0.22 |
ENST00000503853.1
|
PRELID1
|
PRELI domain containing 1 |
| chr11_+_34664014 | 0.22 |
ENST00000527935.1
|
EHF
|
ets homologous factor |
| chr12_-_54071181 | 0.21 |
ENST00000338662.5
|
ATP5G2
|
ATP synthase, H+ transporting, mitochondrial Fo complex, subunit C2 (subunit 9) |
| chr2_+_27440229 | 0.21 |
ENST00000264705.4
ENST00000403525.1 |
CAD
|
carbamoyl-phosphate synthetase 2, aspartate transcarbamylase, and dihydroorotase |
| chr11_-_132813566 | 0.21 |
ENST00000331898.7
|
OPCML
|
opioid binding protein/cell adhesion molecule-like |
| chr11_+_34663913 | 0.21 |
ENST00000532302.1
|
EHF
|
ets homologous factor |
| chr14_-_101295407 | 0.20 |
ENST00000596284.1
|
AL117190.2
|
AL117190.2 |
| chr22_+_39916558 | 0.20 |
ENST00000337304.2
ENST00000396680.1 |
ATF4
|
activating transcription factor 4 |
| chr6_+_88054530 | 0.20 |
ENST00000388923.4
|
C6orf163
|
chromosome 6 open reading frame 163 |
| chr10_-_94301107 | 0.19 |
ENST00000436178.1
|
IDE
|
insulin-degrading enzyme |
| chr9_-_133814527 | 0.18 |
ENST00000451466.1
|
FIBCD1
|
fibrinogen C domain containing 1 |
| chr5_+_159656437 | 0.18 |
ENST00000402432.3
|
FABP6
|
fatty acid binding protein 6, ileal |
| chr3_+_190105909 | 0.18 |
ENST00000456423.1
|
CLDN16
|
claudin 16 |
| chr17_-_41739283 | 0.18 |
ENST00000393661.2
ENST00000318579.4 |
MEOX1
|
mesenchyme homeobox 1 |
| chr1_+_209859510 | 0.18 |
ENST00000367028.2
ENST00000261465.1 |
HSD11B1
|
hydroxysteroid (11-beta) dehydrogenase 1 |
| chr6_+_46761118 | 0.18 |
ENST00000230588.4
|
MEP1A
|
meprin A, alpha (PABA peptide hydrolase) |
| chr20_+_58713514 | 0.18 |
ENST00000432910.1
|
RP5-1043L13.1
|
RP5-1043L13.1 |
| chr14_-_69444957 | 0.18 |
ENST00000556571.1
ENST00000553659.1 ENST00000555616.1 |
ACTN1
|
actinin, alpha 1 |
| chr15_+_67418047 | 0.18 |
ENST00000540846.2
|
SMAD3
|
SMAD family member 3 |
| chr17_-_38928414 | 0.17 |
ENST00000335552.4
|
KRT26
|
keratin 26 |
| chr17_+_7461580 | 0.17 |
ENST00000483039.1
ENST00000396542.1 |
TNFSF13
|
tumor necrosis factor (ligand) superfamily, member 13 |
| chr1_+_199996702 | 0.16 |
ENST00000367362.3
|
NR5A2
|
nuclear receptor subfamily 5, group A, member 2 |
| chr14_+_97925151 | 0.16 |
ENST00000554862.1
ENST00000554260.1 ENST00000499910.2 |
CTD-2506J14.1
|
CTD-2506J14.1 |
| chr4_-_174451370 | 0.16 |
ENST00000359562.4
|
HAND2
|
heart and neural crest derivatives expressed 2 |
| chr18_-_46784778 | 0.16 |
ENST00000582399.1
|
DYM
|
dymeclin |
| chrX_-_19817869 | 0.16 |
ENST00000379698.4
|
SH3KBP1
|
SH3-domain kinase binding protein 1 |
| chr11_-_132813627 | 0.16 |
ENST00000374778.4
|
OPCML
|
opioid binding protein/cell adhesion molecule-like |
| chr10_-_33405600 | 0.16 |
ENST00000414308.1
|
RP11-342D11.3
|
RP11-342D11.3 |
| chr19_+_36036583 | 0.15 |
ENST00000392205.1
|
TMEM147
|
transmembrane protein 147 |
| chr5_-_88119580 | 0.15 |
ENST00000539796.1
|
MEF2C
|
myocyte enhancer factor 2C |
| chr5_+_36606700 | 0.15 |
ENST00000416645.2
|
SLC1A3
|
solute carrier family 1 (glial high affinity glutamate transporter), member 3 |
| chr19_+_36036477 | 0.15 |
ENST00000222284.5
ENST00000392204.2 |
TMEM147
|
transmembrane protein 147 |
| chr12_-_28124903 | 0.14 |
ENST00000395872.1
ENST00000354417.3 ENST00000201015.4 |
PTHLH
|
parathyroid hormone-like hormone |
| chr9_-_71869642 | 0.14 |
ENST00000600472.1
|
AL358113.1
|
Tight junction protein 2 (Zona occludens 2) isoform 1 variant; Uncharacterized protein |
| chr3_-_57326704 | 0.14 |
ENST00000487349.1
ENST00000389601.3 |
ASB14
|
ankyrin repeat and SOCS box containing 14 |
| chr3_+_173116225 | 0.14 |
ENST00000457714.1
|
NLGN1
|
neuroligin 1 |
| chr3_-_71294304 | 0.13 |
ENST00000498215.1
|
FOXP1
|
forkhead box P1 |
| chr12_+_24857397 | 0.13 |
ENST00000536131.1
|
RP11-625L16.1
|
RP11-625L16.1 |
| chr9_+_44868935 | 0.13 |
ENST00000448436.2
|
RP11-160N1.10
|
RP11-160N1.10 |
| chr18_+_29769978 | 0.13 |
ENST00000269202.6
ENST00000581447.1 |
MEP1B
|
meprin A, beta |
| chr1_+_66999268 | 0.13 |
ENST00000371039.1
ENST00000424320.1 |
SGIP1
|
SH3-domain GRB2-like (endophilin) interacting protein 1 |
| chr2_-_166930131 | 0.12 |
ENST00000303395.4
ENST00000409050.1 ENST00000423058.2 ENST00000375405.3 |
SCN1A
|
sodium channel, voltage-gated, type I, alpha subunit |
| chr6_-_91296737 | 0.12 |
ENST00000369332.3
ENST00000369329.3 |
MAP3K7
|
mitogen-activated protein kinase kinase kinase 7 |
| chr11_-_62473684 | 0.12 |
ENST00000448568.2
|
BSCL2
|
Berardinelli-Seip congenital lipodystrophy 2 (seipin) |
| chr16_-_67978016 | 0.12 |
ENST00000264005.5
|
LCAT
|
lecithin-cholesterol acyltransferase |
| chr1_-_12679171 | 0.12 |
ENST00000606790.1
|
RP11-474O21.5
|
RP11-474O21.5 |
| chr3_-_79816965 | 0.12 |
ENST00000464233.1
|
ROBO1
|
roundabout, axon guidance receptor, homolog 1 (Drosophila) |
| chr7_+_114055052 | 0.11 |
ENST00000462331.1
ENST00000408937.3 ENST00000403559.4 ENST00000350908.4 ENST00000393498.2 ENST00000393495.3 ENST00000378237.3 ENST00000393489.3 |
FOXP2
|
forkhead box P2 |
| chr12_-_94673956 | 0.11 |
ENST00000551941.1
|
RP11-1105G2.3
|
Uncharacterized protein |
| chr18_-_73967160 | 0.11 |
ENST00000579714.1
|
RP11-94B19.7
|
RP11-94B19.7 |
| chr13_-_74708372 | 0.11 |
ENST00000377666.4
|
KLF12
|
Kruppel-like factor 12 |
| chr12_+_1800179 | 0.11 |
ENST00000357103.4
|
ADIPOR2
|
adiponectin receptor 2 |
| chr3_+_15045419 | 0.11 |
ENST00000406272.2
|
NR2C2
|
nuclear receptor subfamily 2, group C, member 2 |
| chr8_-_57123815 | 0.11 |
ENST00000316981.3
ENST00000423799.2 ENST00000429357.2 |
PLAG1
|
pleiomorphic adenoma gene 1 |
| chr9_+_105757590 | 0.10 |
ENST00000374798.3
ENST00000487798.1 |
CYLC2
|
cylicin, basic protein of sperm head cytoskeleton 2 |
| chr12_-_95510743 | 0.10 |
ENST00000551521.1
|
FGD6
|
FYVE, RhoGEF and PH domain containing 6 |
| chr7_-_20826504 | 0.10 |
ENST00000418710.2
ENST00000361443.4 |
SP8
|
Sp8 transcription factor |
| chrM_+_4431 | 0.10 |
ENST00000361453.3
|
MT-ND2
|
mitochondrially encoded NADH dehydrogenase 2 |
| chr2_-_74726710 | 0.10 |
ENST00000377566.4
|
LBX2
|
ladybird homeobox 2 |
| chr9_+_82188077 | 0.09 |
ENST00000425506.1
|
TLE4
|
transducin-like enhancer of split 4 (E(sp1) homolog, Drosophila) |
| chr16_-_21863541 | 0.09 |
ENST00000543654.1
|
NPIPB4
|
nuclear pore complex interacting protein family, member B4 |
| chr4_-_186661365 | 0.09 |
ENST00000452351.1
|
SORBS2
|
sorbin and SH3 domain containing 2 |
| chr12_-_111358372 | 0.09 |
ENST00000548438.1
ENST00000228841.8 |
MYL2
|
myosin, light chain 2, regulatory, cardiac, slow |
| chr1_-_237167718 | 0.09 |
ENST00000464121.2
|
MT1HL1
|
metallothionein 1H-like 1 |
| chr4_+_160188889 | 0.09 |
ENST00000264431.4
|
RAPGEF2
|
Rap guanine nucleotide exchange factor (GEF) 2 |
| chr12_-_96184533 | 0.09 |
ENST00000343702.4
ENST00000344911.4 |
NTN4
|
netrin 4 |
| chr14_+_101295638 | 0.09 |
ENST00000523671.2
|
MEG3
|
maternally expressed 3 (non-protein coding) |
| chr20_+_4702548 | 0.09 |
ENST00000305817.2
|
PRND
|
prion protein 2 (dublet) |
| chr15_-_83952071 | 0.09 |
ENST00000569704.1
|
BNC1
|
basonuclin 1 |
| chr17_-_39684550 | 0.09 |
ENST00000455635.1
ENST00000361566.3 |
KRT19
|
keratin 19 |
| chr15_-_68497657 | 0.08 |
ENST00000448060.2
ENST00000467889.1 |
CALML4
|
calmodulin-like 4 |
| chr6_-_91296602 | 0.08 |
ENST00000369325.3
ENST00000369327.3 |
MAP3K7
|
mitogen-activated protein kinase kinase kinase 7 |
| chr7_-_47578840 | 0.08 |
ENST00000450444.1
|
TNS3
|
tensin 3 |
| chr8_+_134203303 | 0.08 |
ENST00000519433.1
ENST00000517423.1 ENST00000377863.2 ENST00000220856.6 |
WISP1
|
WNT1 inducible signaling pathway protein 1 |
| chr11_+_63606373 | 0.08 |
ENST00000402010.2
ENST00000315032.8 ENST00000377809.4 ENST00000413835.2 ENST00000377810.3 |
MARK2
|
MAP/microtubule affinity-regulating kinase 2 |
| chr20_-_56285595 | 0.08 |
ENST00000395816.3
ENST00000347215.4 |
PMEPA1
|
prostate transmembrane protein, androgen induced 1 |
| chr19_-_19626838 | 0.07 |
ENST00000360913.3
|
TSSK6
|
testis-specific serine kinase 6 |
| chr6_-_31745085 | 0.07 |
ENST00000375686.3
ENST00000447450.1 |
VWA7
|
von Willebrand factor A domain containing 7 |
| chr12_+_54410664 | 0.07 |
ENST00000303406.4
|
HOXC4
|
homeobox C4 |
| chr6_+_108487245 | 0.07 |
ENST00000368986.4
|
NR2E1
|
nuclear receptor subfamily 2, group E, member 1 |
| chr13_-_103719196 | 0.07 |
ENST00000245312.3
|
SLC10A2
|
solute carrier family 10 (sodium/bile acid cotransporter), member 2 |
| chr5_+_148521381 | 0.07 |
ENST00000504238.1
|
ABLIM3
|
actin binding LIM protein family, member 3 |
| chr22_+_41697520 | 0.07 |
ENST00000352645.4
|
ZC3H7B
|
zinc finger CCCH-type containing 7B |
| chr19_+_1440838 | 0.07 |
ENST00000594262.1
|
AC027307.3
|
Uncharacterized protein |
| chr5_-_88120151 | 0.07 |
ENST00000506716.1
|
MEF2C
|
myocyte enhancer factor 2C |
| chr17_+_7461613 | 0.07 |
ENST00000438470.1
ENST00000436057.1 |
TNFSF13
|
tumor necrosis factor (ligand) superfamily, member 13 |
| chr17_-_38821373 | 0.07 |
ENST00000394052.3
|
KRT222
|
keratin 222 |
| chr3_+_89156674 | 0.07 |
ENST00000336596.2
|
EPHA3
|
EPH receptor A3 |
| chr11_-_62473776 | 0.07 |
ENST00000278893.7
ENST00000407022.3 ENST00000421906.1 |
BSCL2
|
Berardinelli-Seip congenital lipodystrophy 2 (seipin) |
| chr17_-_39646116 | 0.07 |
ENST00000328119.6
|
KRT36
|
keratin 36 |
| chr18_-_48351743 | 0.06 |
ENST00000588444.1
ENST00000256425.2 ENST00000428869.2 |
MRO
|
maestro |
| chr16_+_15031300 | 0.06 |
ENST00000328085.6
|
NPIPA1
|
nuclear pore complex interacting protein family, member A1 |
| chrX_+_107683096 | 0.06 |
ENST00000328300.6
ENST00000361603.2 |
COL4A5
|
collagen, type IV, alpha 5 |
| chr5_-_88120083 | 0.06 |
ENST00000509373.1
|
MEF2C
|
myocyte enhancer factor 2C |
| chr2_-_190044480 | 0.06 |
ENST00000374866.3
|
COL5A2
|
collagen, type V, alpha 2 |
| chr11_-_27723158 | 0.06 |
ENST00000395980.2
|
BDNF
|
brain-derived neurotrophic factor |
| chr17_-_46657473 | 0.06 |
ENST00000332503.5
|
HOXB4
|
homeobox B4 |
| chr12_-_96184913 | 0.06 |
ENST00000538383.1
|
NTN4
|
netrin 4 |
| chr5_-_34043310 | 0.06 |
ENST00000231338.7
|
C1QTNF3
|
C1q and tumor necrosis factor related protein 3 |
| chr1_-_94147385 | 0.06 |
ENST00000260502.6
|
BCAR3
|
breast cancer anti-estrogen resistance 3 |
| chr6_-_41703952 | 0.05 |
ENST00000358871.2
ENST00000403298.4 |
TFEB
|
transcription factor EB |
| chr1_+_151739131 | 0.05 |
ENST00000400999.1
|
OAZ3
|
ornithine decarboxylase antizyme 3 |
| chr11_+_64001962 | 0.05 |
ENST00000309422.2
|
VEGFB
|
vascular endothelial growth factor B |
| chr19_+_4402659 | 0.05 |
ENST00000301280.5
ENST00000585854.1 |
CHAF1A
|
chromatin assembly factor 1, subunit A (p150) |
| chr2_-_145278475 | 0.05 |
ENST00000558170.2
|
ZEB2
|
zinc finger E-box binding homeobox 2 |
| chr17_-_46806540 | 0.05 |
ENST00000290295.7
|
HOXB13
|
homeobox B13 |
| chr10_-_73497581 | 0.05 |
ENST00000398786.2
|
C10orf105
|
chromosome 10 open reading frame 105 |
| chr10_+_26505179 | 0.05 |
ENST00000376261.3
|
GAD2
|
glutamate decarboxylase 2 (pancreatic islets and brain, 65kDa) |
| chr11_-_118305921 | 0.05 |
ENST00000532619.1
|
RP11-770J1.4
|
RP11-770J1.4 |
| chr11_-_65359947 | 0.05 |
ENST00000597463.1
|
AP001362.1
|
Uncharacterized protein |
| chr9_-_16705069 | 0.05 |
ENST00000471301.2
|
BNC2
|
basonuclin 2 |
| chr18_+_3252265 | 0.05 |
ENST00000580887.1
ENST00000536605.1 |
MYL12A
|
myosin, light chain 12A, regulatory, non-sarcomeric |
| chr7_-_93519471 | 0.05 |
ENST00000451238.1
|
TFPI2
|
tissue factor pathway inhibitor 2 |
| chr5_+_148521454 | 0.04 |
ENST00000508983.1
|
ABLIM3
|
actin binding LIM protein family, member 3 |
| chr11_-_62473706 | 0.04 |
ENST00000403550.1
|
BSCL2
|
Berardinelli-Seip congenital lipodystrophy 2 (seipin) |
| chr13_-_46626820 | 0.04 |
ENST00000428921.1
|
ZC3H13
|
zinc finger CCCH-type containing 13 |
| chr12_-_18243119 | 0.04 |
ENST00000538724.1
ENST00000229002.2 |
RERGL
|
RERG/RAS-like |
| chr13_+_109248500 | 0.04 |
ENST00000356711.2
|
MYO16
|
myosin XVI |
| chr8_+_38831683 | 0.04 |
ENST00000302495.4
|
HTRA4
|
HtrA serine peptidase 4 |
| chr16_-_21431078 | 0.04 |
ENST00000458643.2
|
NPIPB3
|
nuclear pore complex interacting protein family, member B3 |
| chr11_-_132812987 | 0.04 |
ENST00000541867.1
|
OPCML
|
opioid binding protein/cell adhesion molecule-like |
| chr1_+_183774285 | 0.04 |
ENST00000539189.1
|
RGL1
|
ral guanine nucleotide dissociation stimulator-like 1 |
| chr22_+_41956767 | 0.04 |
ENST00000306149.7
|
CSDC2
|
cold shock domain containing C2, RNA binding |
| chr3_+_41236325 | 0.03 |
ENST00000426215.1
ENST00000405570.1 |
CTNNB1
|
catenin (cadherin-associated protein), beta 1, 88kDa |
| chr11_-_31531121 | 0.03 |
ENST00000532287.1
ENST00000526776.1 ENST00000534812.1 ENST00000529749.1 ENST00000278200.1 ENST00000530023.1 ENST00000533642.1 |
IMMP1L
|
IMP1 inner mitochondrial membrane peptidase-like (S. cerevisiae) |
| chrX_-_107682702 | 0.03 |
ENST00000372216.4
|
COL4A6
|
collagen, type IV, alpha 6 |
| chr11_+_113185778 | 0.03 |
ENST00000524580.2
|
TTC12
|
tetratricopeptide repeat domain 12 |
| chr15_-_54025300 | 0.03 |
ENST00000559418.1
|
WDR72
|
WD repeat domain 72 |
| chr6_+_152011628 | 0.03 |
ENST00000404742.1
ENST00000440973.1 |
ESR1
|
estrogen receptor 1 |
| chrX_+_99899180 | 0.03 |
ENST00000373004.3
|
SRPX2
|
sushi-repeat containing protein, X-linked 2 |
| chr18_-_52989217 | 0.03 |
ENST00000570287.2
|
TCF4
|
transcription factor 4 |
| chr7_+_35756066 | 0.03 |
ENST00000449644.1
|
AC018647.3
|
AC018647.3 |
| chr7_+_141811539 | 0.03 |
ENST00000550469.2
ENST00000477922.3 |
RP11-1220K2.2
|
Putative inactive maltase-glucoamylase-like protein LOC93432 |
| chr16_+_67906919 | 0.03 |
ENST00000358933.5
|
EDC4
|
enhancer of mRNA decapping 4 |
| chr11_-_47521309 | 0.03 |
ENST00000535982.1
|
CELF1
|
CUGBP, Elav-like family member 1 |
| chr14_-_25479811 | 0.03 |
ENST00000550887.1
|
STXBP6
|
syntaxin binding protein 6 (amisyn) |
| chr9_-_73736511 | 0.03 |
ENST00000377110.3
ENST00000377111.2 |
TRPM3
|
transient receptor potential cation channel, subfamily M, member 3 |
| chr18_+_63417864 | 0.03 |
ENST00000536984.2
|
CDH7
|
cadherin 7, type 2 |
| chr17_-_79876010 | 0.03 |
ENST00000328666.6
|
SIRT7
|
sirtuin 7 |
| chr1_-_200379129 | 0.02 |
ENST00000367353.1
|
ZNF281
|
zinc finger protein 281 |
| chr2_-_51259292 | 0.02 |
ENST00000401669.2
|
NRXN1
|
neurexin 1 |
| chr2_+_128293323 | 0.02 |
ENST00000389524.4
ENST00000428314.1 |
MYO7B
|
myosin VIIB |
| chr3_+_89156799 | 0.02 |
ENST00000452448.2
ENST00000494014.1 |
EPHA3
|
EPH receptor A3 |
| chr2_-_50574856 | 0.02 |
ENST00000342183.5
|
NRXN1
|
neurexin 1 |
| chr8_+_26435359 | 0.02 |
ENST00000311151.5
|
DPYSL2
|
dihydropyrimidinase-like 2 |
| chr12_-_47219733 | 0.02 |
ENST00000547477.1
ENST00000447411.1 ENST00000266579.4 |
SLC38A4
|
solute carrier family 38, member 4 |
| chr5_+_145718587 | 0.02 |
ENST00000230732.4
|
POU4F3
|
POU class 4 homeobox 3 |
| chr6_+_15246501 | 0.02 |
ENST00000341776.2
|
JARID2
|
jumonji, AT rich interactive domain 2 |
| chr6_-_64029879 | 0.02 |
ENST00000370658.5
ENST00000485906.2 ENST00000370657.4 |
LGSN
|
lengsin, lens protein with glutamine synthetase domain |
| chr11_-_115127611 | 0.02 |
ENST00000545094.1
|
CADM1
|
cell adhesion molecule 1 |
| chr6_-_31745037 | 0.02 |
ENST00000375688.4
|
VWA7
|
von Willebrand factor A domain containing 7 |
| chr9_+_75229616 | 0.02 |
ENST00000340019.3
|
TMC1
|
transmembrane channel-like 1 |
| chr17_+_79369249 | 0.02 |
ENST00000574717.2
|
RP11-1055B8.6
|
Uncharacterized protein |
| chr2_-_51259229 | 0.01 |
ENST00000405472.3
|
NRXN1
|
neurexin 1 |
| chr13_-_28545276 | 0.01 |
ENST00000381020.7
|
CDX2
|
caudal type homeobox 2 |
| chr3_-_73483055 | 0.01 |
ENST00000479530.1
|
PDZRN3
|
PDZ domain containing ring finger 3 |
| chr8_+_26435915 | 0.01 |
ENST00000523027.1
|
DPYSL2
|
dihydropyrimidinase-like 2 |
| chr3_-_45838011 | 0.01 |
ENST00000358525.4
ENST00000413781.1 |
SLC6A20
|
solute carrier family 6 (proline IMINO transporter), member 20 |
| chr13_-_46626847 | 0.01 |
ENST00000242848.4
ENST00000282007.3 |
ZC3H13
|
zinc finger CCCH-type containing 13 |
| chr9_+_82187487 | 0.01 |
ENST00000435650.1
ENST00000414465.1 ENST00000376537.4 ENST00000376534.4 |
TLE4
|
transducin-like enhancer of split 4 (E(sp1) homolog, Drosophila) |
| chr8_+_70379072 | 0.01 |
ENST00000529134.1
|
SULF1
|
sulfatase 1 |
| chr12_+_111537227 | 0.01 |
ENST00000397643.3
|
CUX2
|
cut-like homeobox 2 |
| chr19_-_45927622 | 0.01 |
ENST00000300853.3
ENST00000589165.1 |
ERCC1
|
excision repair cross-complementing rodent repair deficiency, complementation group 1 (includes overlapping antisense sequence) |
| chr18_+_61223393 | 0.01 |
ENST00000269491.1
ENST00000382768.1 |
SERPINB12
|
serpin peptidase inhibitor, clade B (ovalbumin), member 12 |
| chr16_-_21663950 | 0.01 |
ENST00000268389.4
|
IGSF6
|
immunoglobulin superfamily, member 6 |
| chr9_-_98189055 | 0.01 |
ENST00000433644.2
|
RP11-435O5.2
|
RP11-435O5.2 |
| chrX_+_105192423 | 0.00 |
ENST00000540278.1
|
NRK
|
Nik related kinase |
| chr2_-_40739501 | 0.00 |
ENST00000403092.1
ENST00000402441.1 ENST00000448531.1 |
SLC8A1
|
solute carrier family 8 (sodium/calcium exchanger), member 1 |
| chrX_+_14547632 | 0.00 |
ENST00000218075.4
|
GLRA2
|
glycine receptor, alpha 2 |
| chr16_-_2314222 | 0.00 |
ENST00000566397.1
|
RNPS1
|
RNA binding protein S1, serine-rich domain |
| chr5_+_43121698 | 0.00 |
ENST00000505606.2
ENST00000509634.1 ENST00000509341.1 |
ZNF131
|
zinc finger protein 131 |
| chr12_-_18243075 | 0.00 |
ENST00000536890.1
|
RERGL
|
RERG/RAS-like |
Network of associatons between targets according to the STRING database.
First level regulatory network of HOXC11
Gene Ontology Analysis
Gene overrepresentation in biological_process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.5 | GO:0045362 | regulation of interleukin-1 biosynthetic process(GO:0045360) positive regulation of interleukin-1 biosynthetic process(GO:0045362) |
| 0.4 | 1.6 | GO:0002384 | hepatic immune response(GO:0002384) response to prolactin(GO:1990637) |
| 0.3 | 0.9 | GO:0060279 | regulation of ovulation(GO:0060278) positive regulation of ovulation(GO:0060279) |
| 0.3 | 1.1 | GO:0060585 | nitric oxide transport(GO:0030185) negative regulation of nitric-oxide synthase biosynthetic process(GO:0051771) regulation of prostaglandin-endoperoxide synthase activity(GO:0060584) positive regulation of prostaglandin-endoperoxide synthase activity(GO:0060585) |
| 0.1 | 0.4 | GO:0035038 | female pronucleus assembly(GO:0035038) |
| 0.1 | 0.6 | GO:0060708 | spongiotrophoblast differentiation(GO:0060708) |
| 0.1 | 0.2 | GO:0044205 | 'de novo' UMP biosynthetic process(GO:0044205) |
| 0.1 | 0.3 | GO:0002314 | germinal center B cell differentiation(GO:0002314) |
| 0.1 | 0.2 | GO:0061394 | regulation of transcription from RNA polymerase II promoter in response to arsenic-containing substance(GO:0061394) |
| 0.1 | 0.6 | GO:0098532 | histone H3-K27 trimethylation(GO:0098532) |
| 0.1 | 0.2 | GO:0090214 | spongiotrophoblast layer developmental growth(GO:0090214) |
| 0.1 | 0.2 | GO:0003357 | noradrenergic neuron differentiation(GO:0003357) |
| 0.0 | 0.2 | GO:1901143 | insulin catabolic process(GO:1901143) |
| 0.0 | 0.1 | GO:0098923 | retrograde trans-synaptic signaling by soluble gas(GO:0098923) trans-synaptic signaling by soluble gas(GO:0099543) trans-synaptic signaling by trans-synaptic complex(GO:0099545) |
| 0.0 | 0.3 | GO:0003172 | primary heart field specification(GO:0003138) sinoatrial valve development(GO:0003172) sinoatrial valve morphogenesis(GO:0003185) |
| 0.0 | 0.1 | GO:0098582 | innate vocalization behavior(GO:0098582) |
| 0.0 | 0.2 | GO:0001757 | somite specification(GO:0001757) sclerotome development(GO:0061056) |
| 0.0 | 0.2 | GO:0006880 | intracellular sequestering of iron ion(GO:0006880) sequestering of iron ion(GO:0097577) |
| 0.0 | 0.2 | GO:1901857 | positive regulation of cellular respiration(GO:1901857) |
| 0.0 | 0.2 | GO:0035932 | mineralocorticoid secretion(GO:0035931) aldosterone secretion(GO:0035932) drinking behavior(GO:0042756) regulation of mineralocorticoid secretion(GO:2000855) regulation of aldosterone secretion(GO:2000858) |
| 0.0 | 0.7 | GO:2001224 | positive regulation of neuron migration(GO:2001224) |
| 0.0 | 0.2 | GO:0061767 | negative regulation of lung blood pressure(GO:0061767) |
| 0.0 | 0.1 | GO:0035426 | extracellular matrix-cell signaling(GO:0035426) |
| 0.0 | 0.1 | GO:0002118 | aggressive behavior(GO:0002118) |
| 0.0 | 0.1 | GO:0034372 | very-low-density lipoprotein particle remodeling(GO:0034372) regulation of high-density lipoprotein particle assembly(GO:0090107) |
| 0.0 | 0.1 | GO:0097156 | fasciculation of motor neuron axon(GO:0097156) |
| 0.0 | 0.1 | GO:0021836 | cerebral cortex tangential migration using cell-cell interactions(GO:0021823) postnatal olfactory bulb interneuron migration(GO:0021827) chemorepulsion involved in postnatal olfactory bulb interneuron migration(GO:0021836) |
| 0.0 | 0.2 | GO:0061113 | pancreas morphogenesis(GO:0061113) |
| 0.0 | 0.1 | GO:0032625 | interleukin-21 production(GO:0032625) interleukin-21 secretion(GO:0072619) |
| 0.0 | 0.1 | GO:1903225 | negative regulation of endodermal cell differentiation(GO:1903225) |
| 0.0 | 0.2 | GO:0048298 | positive regulation of isotype switching to IgA isotypes(GO:0048298) |
| 0.0 | 0.1 | GO:0031548 | regulation of brain-derived neurotrophic factor receptor signaling pathway(GO:0031548) |
| 0.0 | 0.3 | GO:0042118 | endothelial cell activation(GO:0042118) |
| 0.0 | 0.4 | GO:0006704 | glucocorticoid biosynthetic process(GO:0006704) |
| 0.0 | 0.1 | GO:0070779 | D-aspartate transport(GO:0070777) D-aspartate import(GO:0070779) |
| 0.0 | 0.1 | GO:2000253 | positive regulation of feeding behavior(GO:2000253) |
| 0.0 | 0.1 | GO:0071638 | negative regulation of monocyte chemotactic protein-1 production(GO:0071638) |
| 0.0 | 0.1 | GO:1902748 | mammillary body development(GO:0021767) mammillary axonal complex development(GO:0061373) positive regulation of lens fiber cell differentiation(GO:1902748) |
| 0.0 | 0.1 | GO:0060743 | epithelial cell maturation involved in prostate gland development(GO:0060743) |
| 0.0 | 0.1 | GO:0050966 | detection of mechanical stimulus involved in sensory perception of pain(GO:0050966) |
| 0.0 | 0.1 | GO:1902268 | negative regulation of polyamine transmembrane transport(GO:1902268) |
Gene overrepresentation in cellular_component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.1 | GO:0048237 | rough endoplasmic reticulum lumen(GO:0048237) |
| 0.3 | 1.6 | GO:0005896 | interleukin-6 receptor complex(GO:0005896) |
| 0.3 | 0.9 | GO:0043512 | inhibin complex(GO:0043511) inhibin A complex(GO:0043512) |
| 0.1 | 0.7 | GO:1990111 | spermatoproteasome complex(GO:1990111) |
| 0.1 | 0.2 | GO:0008043 | intracellular ferritin complex(GO:0008043) ferritin complex(GO:0070288) |
| 0.0 | 0.6 | GO:0000788 | nuclear nucleosome(GO:0000788) |
| 0.0 | 0.1 | GO:0033011 | perinuclear theca(GO:0033011) |
| 0.0 | 0.2 | GO:1990037 | Lewy body core(GO:1990037) |
| 0.0 | 0.1 | GO:0005588 | collagen type V trimer(GO:0005588) |
| 0.0 | 0.2 | GO:0071144 | SMAD2-SMAD3 protein complex(GO:0071144) |
| 0.0 | 0.1 | GO:0033186 | CAF-1 complex(GO:0033186) |
| 0.0 | 0.0 | GO:0042720 | mitochondrial inner membrane peptidase complex(GO:0042720) |
| 0.0 | 0.1 | GO:0005587 | collagen type IV trimer(GO:0005587) |
| 0.0 | 0.1 | GO:1990357 | terminal web(GO:1990357) |
| 0.0 | 0.2 | GO:0000276 | mitochondrial proton-transporting ATP synthase complex, coupling factor F(o)(GO:0000276) |
| 0.0 | 0.2 | GO:0005916 | fascia adherens(GO:0005916) |
Gene overrepresentation in molecular_function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.5 | GO:0031731 | CCR6 chemokine receptor binding(GO:0031731) |
| 0.4 | 1.1 | GO:0031705 | bombesin receptor binding(GO:0031705) |
| 0.2 | 0.6 | GO:0005146 | leukemia inhibitory factor receptor binding(GO:0005146) |
| 0.2 | 0.9 | GO:0034711 | inhibin binding(GO:0034711) |
| 0.2 | 1.6 | GO:0005138 | interleukin-6 receptor binding(GO:0005138) |
| 0.1 | 0.4 | GO:0003845 | 11-beta-hydroxysteroid dehydrogenase [NAD(P)] activity(GO:0003845) |
| 0.1 | 0.3 | GO:0005427 | proton-dependent oligopeptide secondary active transmembrane transporter activity(GO:0005427) secondary active oligopeptide transmembrane transporter activity(GO:0015322) |
| 0.1 | 0.6 | GO:0032558 | adenyl deoxyribonucleotide binding(GO:0032558) dATP binding(GO:0032564) |
| 0.1 | 0.2 | GO:0031626 | beta-endorphin binding(GO:0031626) |
| 0.0 | 0.2 | GO:0004088 | carbamoyl-phosphate synthase (ammonia) activity(GO:0004087) carbamoyl-phosphate synthase (glutamine-hydrolyzing) activity(GO:0004088) |
| 0.0 | 0.2 | GO:1990050 | phosphatidic acid transporter activity(GO:1990050) |
| 0.0 | 0.1 | GO:0097003 | adipokinetic hormone receptor activity(GO:0097003) |
| 0.0 | 0.3 | GO:0005497 | androgen binding(GO:0005497) |
| 0.0 | 0.5 | GO:0008061 | chitin binding(GO:0008061) |
| 0.0 | 0.2 | GO:0031962 | mineralocorticoid receptor binding(GO:0031962) |
| 0.0 | 0.7 | GO:0070003 | threonine-type endopeptidase activity(GO:0004298) threonine-type peptidase activity(GO:0070003) |
| 0.0 | 0.4 | GO:0003680 | AT DNA binding(GO:0003680) |
| 0.0 | 0.1 | GO:0005169 | neurotrophin TRKB receptor binding(GO:0005169) |
| 0.0 | 0.1 | GO:0034186 | apolipoprotein A-I binding(GO:0034186) |
| 0.0 | 0.1 | GO:0015501 | glutamate:sodium symporter activity(GO:0015501) |
| 0.0 | 0.1 | GO:0043237 | laminin-1 binding(GO:0043237) |
| 0.0 | 0.1 | GO:0008046 | axon guidance receptor activity(GO:0008046) |
| 0.0 | 0.0 | GO:0016880 | glutamate-ammonia ligase activity(GO:0004356) ammonia ligase activity(GO:0016211) acid-ammonia (or amide) ligase activity(GO:0016880) |
| 0.0 | 0.2 | GO:0008199 | ferric iron binding(GO:0008199) |
| 0.0 | 0.2 | GO:0032052 | bile acid binding(GO:0032052) |
| 0.0 | 0.1 | GO:0008073 | ornithine decarboxylase inhibitor activity(GO:0008073) |
| 0.0 | 0.1 | GO:0008508 | bile acid:sodium symporter activity(GO:0008508) |
| 0.0 | 0.3 | GO:0071837 | HMG box domain binding(GO:0071837) |
Gene overrepresentation in C2:CP category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.6 | ST STAT3 PATHWAY | STAT3 Pathway |
| 0.1 | 0.3 | PID AR NONGENOMIC PATHWAY | Nongenotropic Androgen signaling |
| 0.0 | 0.9 | PID ALK1 PATHWAY | ALK1 signaling events |
| 0.0 | 0.2 | PID MET PATHWAY | Signaling events mediated by Hepatocyte Growth Factor Receptor (c-Met) |
| 0.0 | 0.6 | PID HDAC CLASSIII PATHWAY | Signaling events mediated by HDAC Class III |
| 0.0 | 1.1 | PID ENDOTHELIN PATHWAY | Endothelins |
Gene overrepresentation in C2:CP:REACTOME category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 0.9 | REACTOME GLYCOPROTEIN HORMONES | Genes involved in Glycoprotein hormones |
| 0.1 | 1.6 | REACTOME IL 6 SIGNALING | Genes involved in Interleukin-6 signaling |
| 0.0 | 1.5 | REACTOME CHEMOKINE RECEPTORS BIND CHEMOKINES | Genes involved in Chemokine receptors bind chemokines |
| 0.0 | 0.6 | REACTOME APOPTOSIS INDUCED DNA FRAGMENTATION | Genes involved in Apoptosis induced DNA fragmentation |
| 0.0 | 0.6 | REACTOME OTHER SEMAPHORIN INTERACTIONS | Genes involved in Other semaphorin interactions |
| 0.0 | 0.3 | REACTOME RECYCLING OF BILE ACIDS AND SALTS | Genes involved in Recycling of bile acids and salts |
| 0.0 | 0.2 | REACTOME IRAK2 MEDIATED ACTIVATION OF TAK1 COMPLEX UPON TLR7 8 OR 9 STIMULATION | Genes involved in IRAK2 mediated activation of TAK1 complex upon TLR7/8 or 9 stimulation |
| 0.0 | 0.2 | REACTOME ACTIVATION OF CHAPERONE GENES BY ATF6 ALPHA | Genes involved in Activation of Chaperone Genes by ATF6-alpha |
| 0.0 | 0.4 | REACTOME STEROID HORMONES | Genes involved in Steroid hormones |
| 0.0 | 1.1 | REACTOME PEPTIDE LIGAND BINDING RECEPTORS | Genes involved in Peptide ligand-binding receptors |