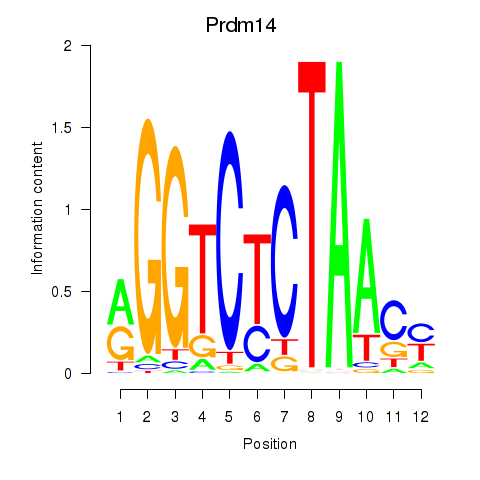
Project
GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
Results for Prdm14
Z-value: 0.95
Transcription factors associated with Prdm14
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Prdm14
|
ENSMUSG00000042414.7 | PR domain containing 14 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Prdm14 | mm10_v2_chr1_-_13127163_13127163 | -0.66 | 1.1e-05 | Click! |
Activity profile of Prdm14 motif
Sorted Z-values of Prdm14 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr8_-_93048192 | 5.79 |
ENSMUST00000095211.4
|
Ces1a
|
carboxylesterase 1A |
| chr6_-_138043411 | 5.06 |
ENSMUST00000111873.1
|
Slc15a5
|
solute carrier family 15, member 5 |
| chr16_+_43235856 | 4.66 |
ENSMUST00000146708.1
|
Zbtb20
|
zinc finger and BTB domain containing 20 |
| chr1_+_88134786 | 4.42 |
ENSMUST00000113134.1
ENSMUST00000140092.1 |
Ugt1a6a
|
UDP glucuronosyltransferase 1 family, polypeptide A6A |
| chr15_-_76307231 | 4.32 |
ENSMUST00000023222.6
ENSMUST00000164189.1 |
Oplah
|
5-oxoprolinase (ATP-hydrolysing) |
| chr15_-_76307101 | 4.20 |
ENSMUST00000171340.1
|
Oplah
|
5-oxoprolinase (ATP-hydrolysing) |
| chr1_+_191977368 | 4.13 |
ENSMUST00000181512.1
ENSMUST00000053463.7 ENSMUST00000180463.1 |
Rd3
|
retinal degeneration 3 |
| chr4_+_133553370 | 4.04 |
ENSMUST00000042706.2
|
Nr0b2
|
nuclear receptor subfamily 0, group B, member 2 |
| chr1_+_88103229 | 4.04 |
ENSMUST00000113135.3
ENSMUST00000113138.1 |
Ugt1a6a
Ugt1a6b
|
UDP glucuronosyltransferase 1 family, polypeptide A6A UDP glucuronosyltransferase 1 family, polypeptide A6B |
| chr15_-_50890396 | 3.78 |
ENSMUST00000185183.1
|
Trps1
|
trichorhinophalangeal syndrome I (human) |
| chr6_+_54039558 | 3.46 |
ENSMUST00000046856.7
|
Chn2
|
chimerin (chimaerin) 2 |
| chr6_+_91684061 | 2.45 |
ENSMUST00000032185.7
|
Slc6a6
|
solute carrier family 6 (neurotransmitter transporter, taurine), member 6 |
| chr17_-_34862122 | 2.12 |
ENSMUST00000154526.1
|
Cfb
|
complement factor B |
| chr7_+_114745685 | 2.07 |
ENSMUST00000136645.1
ENSMUST00000169913.1 |
Insc
|
inscuteable homolog (Drosophila) |
| chr4_+_155562348 | 2.04 |
ENSMUST00000030939.7
|
Nadk
|
NAD kinase |
| chr17_-_34862473 | 1.90 |
ENSMUST00000025229.4
ENSMUST00000176203.2 ENSMUST00000128767.1 |
Cfb
|
complement factor B |
| chr6_+_72355425 | 1.83 |
ENSMUST00000069695.2
ENSMUST00000132243.1 |
Tmem150a
|
transmembrane protein 150A |
| chr14_-_72709986 | 1.79 |
ENSMUST00000089017.5
|
Fndc3a
|
fibronectin type III domain containing 3A |
| chr19_-_6067785 | 1.74 |
ENSMUST00000162575.1
ENSMUST00000159084.1 ENSMUST00000161718.1 ENSMUST00000162810.1 ENSMUST00000025713.5 ENSMUST00000113543.2 ENSMUST00000161528.1 |
Tm7sf2
|
transmembrane 7 superfamily member 2 |
| chr11_-_35980473 | 1.61 |
ENSMUST00000018993.6
|
Wwc1
|
WW, C2 and coiled-coil domain containing 1 |
| chr14_+_62760496 | 1.45 |
ENSMUST00000181344.1
|
4931440J10Rik
|
RIKEN cDNA 4931440J10 gene |
| chr9_-_50659780 | 1.43 |
ENSMUST00000034567.3
|
Dlat
|
dihydrolipoamide S-acetyltransferase (E2 component of pyruvate dehydrogenase complex) |
| chr4_+_117126776 | 1.38 |
ENSMUST00000134074.1
|
Tctex1d4
|
Tctex1 domain containing 4 |
| chr13_+_74639866 | 1.35 |
ENSMUST00000169114.1
|
Erap1
|
endoplasmic reticulum aminopeptidase 1 |
| chr8_+_46492789 | 1.26 |
ENSMUST00000110371.1
|
Acsl1
|
acyl-CoA synthetase long-chain family member 1 |
| chrX_+_68761875 | 1.25 |
ENSMUST00000114647.1
|
Fmr1nb
|
fragile X mental retardation 1 neighbor |
| chr11_+_78324200 | 1.03 |
ENSMUST00000102478.3
|
Aldoc
|
aldolase C, fructose-bisphosphate |
| chr2_-_144011202 | 1.01 |
ENSMUST00000016072.5
ENSMUST00000037875.5 |
Rrbp1
|
ribosome binding protein 1 |
| chr12_+_74297474 | 0.98 |
ENSMUST00000072100.3
|
Dbpht2
|
DNA binding protein with his-thr domain |
| chr5_+_98254174 | 0.97 |
ENSMUST00000031280.1
|
Fgf5
|
fibroblast growth factor 5 |
| chrM_+_7759 | 0.93 |
ENSMUST00000082407.1
ENSMUST00000082408.1 |
mt-Atp8
mt-Atp6
|
mitochondrially encoded ATP synthase 8 mitochondrially encoded ATP synthase 6 |
| chrX_+_68761890 | 0.87 |
ENSMUST00000071848.6
|
Fmr1nb
|
fragile X mental retardation 1 neighbor |
| chr9_+_94669876 | 0.83 |
ENSMUST00000033463.9
|
Slc9a9
|
solute carrier family 9 (sodium/hydrogen exchanger), member 9 |
| chr15_-_73707387 | 0.80 |
ENSMUST00000064166.4
|
Gpr20
|
G protein-coupled receptor 20 |
| chr9_-_14903752 | 0.77 |
ENSMUST00000148155.1
ENSMUST00000121116.1 |
Folr4
|
folate receptor 4 (delta) |
| chr6_+_83109071 | 0.77 |
ENSMUST00000113938.3
|
Mrpl53
|
mitochondrial ribosomal protein L53 |
| chr6_+_92940572 | 0.76 |
ENSMUST00000181145.1
ENSMUST00000181840.1 |
9530026P05Rik
|
RIKEN cDNA 9530026P05 gene |
| chr11_-_89302545 | 0.74 |
ENSMUST00000061728.3
|
Nog
|
noggin |
| chr6_-_42461017 | 0.74 |
ENSMUST00000090156.1
|
Olfr458
|
olfactory receptor 458 |
| chr18_-_70530138 | 0.72 |
ENSMUST00000161542.1
ENSMUST00000159389.1 |
Poli
|
polymerase (DNA directed), iota |
| chr9_-_14903932 | 0.71 |
ENSMUST00000034409.7
ENSMUST00000117620.1 |
Folr4
|
folate receptor 4 (delta) |
| chr1_-_64121389 | 0.70 |
ENSMUST00000055001.3
|
Klf7
|
Kruppel-like factor 7 (ubiquitous) |
| chr1_-_59003443 | 0.66 |
ENSMUST00000054653.6
|
Als2cr11
|
amyotrophic lateral sclerosis 2 (juvenile) chromosome region, candidate 11 (human) |
| chr3_+_89136572 | 0.63 |
ENSMUST00000107482.3
ENSMUST00000127058.1 |
Pklr
|
pyruvate kinase liver and red blood cell |
| chr18_-_70530313 | 0.63 |
ENSMUST00000043286.8
|
Poli
|
polymerase (DNA directed), iota |
| chr15_+_99601372 | 0.60 |
ENSMUST00000023754.5
|
Aqp6
|
aquaporin 6 |
| chr9_+_102718424 | 0.59 |
ENSMUST00000156485.1
ENSMUST00000145937.1 ENSMUST00000134483.1 |
Amotl2
|
angiomotin-like 2 |
| chrX_+_68761839 | 0.55 |
ENSMUST00000069731.5
|
Fmr1nb
|
fragile X mental retardation 1 neighbor |
| chr4_+_117126800 | 0.52 |
ENSMUST00000062206.2
|
Tctex1d4
|
Tctex1 domain containing 4 |
| chr6_-_39810881 | 0.51 |
ENSMUST00000114797.1
ENSMUST00000031978.8 |
Mrps33
|
mitochondrial ribosomal protein S33 |
| chr15_-_44428303 | 0.50 |
ENSMUST00000038719.6
|
Nudcd1
|
NudC domain containing 1 |
| chr10_-_128744014 | 0.49 |
ENSMUST00000026414.7
|
Dgka
|
diacylglycerol kinase, alpha |
| chr7_+_18987518 | 0.48 |
ENSMUST00000063563.7
|
Nanos2
|
nanos homolog 2 (Drosophila) |
| chr15_+_44428073 | 0.40 |
ENSMUST00000060652.3
|
Eny2
|
enhancer of yellow 2 homolog (Drosophila) |
| chr17_+_26781060 | 0.39 |
ENSMUST00000015725.8
ENSMUST00000135824.1 ENSMUST00000137989.1 |
Bnip1
|
BCL2/adenovirus E1B interacting protein 1 |
| chr17_+_21093172 | 0.35 |
ENSMUST00000169389.1
|
Gm6811
|
predicted gene 6811 |
| chr9_-_14903866 | 0.34 |
ENSMUST00000069408.3
|
Folr4
|
folate receptor 4 (delta) |
| chr11_+_104685701 | 0.33 |
ENSMUST00000148007.2
|
Gm11639
|
predicted gene 11639 |
| chr10_-_89257790 | 0.26 |
ENSMUST00000045601.7
|
Ano4
|
anoctamin 4 |
| chr4_+_128883549 | 0.20 |
ENSMUST00000035667.8
|
Trim62
|
tripartite motif-containing 62 |
| chr1_-_153808163 | 0.20 |
ENSMUST00000141249.1
|
Rgsl1
|
regulator of G-protein signaling like 1 |
| chr13_-_119950806 | 0.20 |
ENSMUST00000180768.1
|
B020031M17Rik
|
RIKEN cDNA B020031M17 gene |
| chr7_+_27810813 | 0.20 |
ENSMUST00000080175.6
|
Zfp626
|
zinc finger protein 626 |
| chr15_+_102460076 | 0.05 |
ENSMUST00000164688.1
|
Prr13
|
proline rich 13 |
| chr18_+_37307445 | 0.05 |
ENSMUST00000056712.2
|
Pcdhb4
|
protocadherin beta 4 |
| chr4_-_133672601 | 0.03 |
ENSMUST00000062118.4
ENSMUST00000067902.6 |
Pigv
|
phosphatidylinositol glycan anchor biosynthesis, class V |
| chrX_+_13280970 | 0.01 |
ENSMUST00000000804.6
|
Ddx3x
|
DEAD/H (Asp-Glu-Ala-Asp/His) box polypeptide 3, X-linked |
Network of associatons between targets according to the STRING database.
First level regulatory network of Prdm14
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.7 | 8.5 | GO:0052697 | flavonoid glucuronidation(GO:0052696) xenobiotic glucuronidation(GO:0052697) |
| 0.3 | 2.0 | GO:0006741 | NADP biosynthetic process(GO:0006741) |
| 0.3 | 4.0 | GO:0006957 | complement activation, alternative pathway(GO:0006957) |
| 0.2 | 0.7 | GO:0060300 | regulation of cytokine activity(GO:0060300) |
| 0.2 | 1.6 | GO:0035331 | negative regulation of hippo signaling(GO:0035331) |
| 0.2 | 1.4 | GO:0061732 | mitochondrial acetyl-CoA biosynthetic process from pyruvate(GO:0061732) |
| 0.2 | 1.3 | GO:0071494 | cellular response to UV-C(GO:0071494) |
| 0.1 | 1.3 | GO:0042178 | xenobiotic catabolic process(GO:0042178) |
| 0.1 | 3.8 | GO:1902043 | positive regulation of extrinsic apoptotic signaling pathway via death domain receptors(GO:1902043) |
| 0.1 | 1.3 | GO:0019885 | antigen processing and presentation of endogenous peptide antigen via MHC class I(GO:0019885) |
| 0.1 | 8.5 | GO:0006749 | glutathione metabolic process(GO:0006749) |
| 0.1 | 4.7 | GO:0032728 | positive regulation of interferon-beta production(GO:0032728) |
| 0.1 | 0.6 | GO:0042866 | pyruvate biosynthetic process(GO:0042866) |
| 0.1 | 1.0 | GO:0030388 | fructose 1,6-bisphosphate metabolic process(GO:0030388) |
| 0.1 | 0.5 | GO:0044838 | cell quiescence(GO:0044838) |
| 0.1 | 0.8 | GO:1990118 | sodium ion import across plasma membrane(GO:0098719) sodium ion import into cell(GO:1990118) |
| 0.1 | 2.5 | GO:0072348 | sulfur compound transport(GO:0072348) |
| 0.1 | 1.8 | GO:0060009 | Sertoli cell development(GO:0060009) |
| 0.1 | 1.8 | GO:0046854 | phosphatidylinositol phosphorylation(GO:0046854) |
| 0.1 | 0.3 | GO:0061589 | calcium activated phosphatidylserine scrambling(GO:0061589) |
| 0.0 | 0.5 | GO:0046834 | lipid phosphorylation(GO:0046834) |
| 0.0 | 2.1 | GO:0000132 | establishment of mitotic spindle orientation(GO:0000132) |
| 0.0 | 4.0 | GO:0032024 | positive regulation of insulin secretion(GO:0032024) |
| 0.0 | 1.0 | GO:0023019 | signal transduction involved in regulation of gene expression(GO:0023019) |
| 0.0 | 0.9 | GO:0015985 | energy coupled proton transport, down electrochemical gradient(GO:0015985) ATP synthesis coupled proton transport(GO:0015986) |
| 0.0 | 0.6 | GO:0006833 | water transport(GO:0006833) |
| 0.0 | 0.4 | GO:0061179 | negative regulation of insulin secretion involved in cellular response to glucose stimulus(GO:0061179) |
| 0.0 | 1.7 | GO:0006695 | cholesterol biosynthetic process(GO:0006695) secondary alcohol biosynthetic process(GO:1902653) |
| 0.0 | 4.1 | GO:0007601 | visual perception(GO:0007601) |
| 0.0 | 0.3 | GO:0035404 | histone-serine phosphorylation(GO:0035404) |
| 0.0 | 5.1 | GO:0015833 | peptide transport(GO:0015833) |
| 0.0 | 0.6 | GO:0035329 | hippo signaling(GO:0035329) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 1.4 | GO:0005967 | mitochondrial pyruvate dehydrogenase complex(GO:0005967) |
| 0.1 | 0.9 | GO:0000276 | mitochondrial proton-transporting ATP synthase complex, coupling factor F(o)(GO:0000276) |
| 0.0 | 1.3 | GO:0005788 | endoplasmic reticulum lumen(GO:0005788) |
| 0.0 | 3.7 | GO:0001669 | acrosomal vesicle(GO:0001669) |
| 0.0 | 4.0 | GO:0072562 | blood microparticle(GO:0072562) |
| 0.0 | 0.4 | GO:0000124 | SAGA complex(GO:0000124) |
| 0.0 | 2.8 | GO:0030176 | integral component of endoplasmic reticulum membrane(GO:0030176) |
| 0.0 | 1.3 | GO:0005778 | peroxisomal membrane(GO:0005778) microbody membrane(GO:0031903) |
| 0.0 | 8.5 | GO:0005743 | mitochondrial inner membrane(GO:0005743) |
| 0.0 | 1.8 | GO:0000932 | cytoplasmic mRNA processing body(GO:0000932) |
| 0.0 | 0.8 | GO:0000315 | organellar large ribosomal subunit(GO:0000315) mitochondrial large ribosomal subunit(GO:0005762) |
| 0.0 | 1.6 | GO:0032587 | ruffle membrane(GO:0032587) |
| 0.0 | 0.3 | GO:0031588 | nucleotide-activated protein kinase complex(GO:0031588) |
| 0.0 | 0.0 | GO:0031501 | mannosyltransferase complex(GO:0031501) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 1.2 | 8.5 | GO:0016812 | hydrolase activity, acting on carbon-nitrogen (but not peptide) bonds, in cyclic amides(GO:0016812) |
| 0.8 | 2.5 | GO:0030977 | taurine binding(GO:0030977) |
| 0.3 | 1.3 | GO:0043758 | acetate-CoA ligase (ADP-forming) activity(GO:0043758) |
| 0.3 | 1.4 | GO:0004738 | pyruvate dehydrogenase activity(GO:0004738) pyruvate dehydrogenase [NAD(P)+] activity(GO:0034603) pyruvate dehydrogenase (NAD+) activity(GO:0034604) |
| 0.2 | 8.5 | GO:0015020 | glucuronosyltransferase activity(GO:0015020) |
| 0.2 | 0.6 | GO:0004743 | pyruvate kinase activity(GO:0004743) |
| 0.1 | 1.3 | GO:0005138 | interleukin-6 receptor binding(GO:0005138) |
| 0.1 | 4.0 | GO:0042975 | peroxisome proliferator activated receptor binding(GO:0042975) |
| 0.1 | 1.0 | GO:0004332 | fructose-bisphosphate aldolase activity(GO:0004332) |
| 0.1 | 2.5 | GO:0003951 | NAD+ kinase activity(GO:0003951) |
| 0.1 | 0.8 | GO:0015386 | potassium:proton antiporter activity(GO:0015386) |
| 0.1 | 1.7 | GO:0016628 | oxidoreductase activity, acting on the CH-CH group of donors, NAD or NADP as acceptor(GO:0016628) |
| 0.1 | 1.9 | GO:0008157 | protein phosphatase 1 binding(GO:0008157) |
| 0.0 | 1.3 | GO:0003887 | DNA-directed DNA polymerase activity(GO:0003887) |
| 0.0 | 0.6 | GO:0015250 | water channel activity(GO:0015250) |
| 0.0 | 5.8 | GO:0052689 | carboxylic ester hydrolase activity(GO:0052689) |
| 0.0 | 1.0 | GO:0005104 | fibroblast growth factor receptor binding(GO:0005104) |
| 0.0 | 5.1 | GO:0015293 | symporter activity(GO:0015293) |
| 0.0 | 3.8 | GO:0001085 | RNA polymerase II transcription factor binding(GO:0001085) |
| 0.0 | 4.7 | GO:0001078 | transcriptional repressor activity, RNA polymerase II core promoter proximal region sequence-specific binding(GO:0001078) |
| 0.0 | 4.0 | GO:0004252 | serine-type endopeptidase activity(GO:0004252) |
| 0.0 | 0.3 | GO:0035174 | histone serine kinase activity(GO:0035174) |
| 0.0 | 3.7 | GO:0030674 | protein binding, bridging(GO:0030674) |
| 0.0 | 0.3 | GO:0005229 | intracellular calcium activated chloride channel activity(GO:0005229) |
| 0.0 | 3.7 | GO:0005096 | GTPase activator activity(GO:0005096) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.2 | 4.0 | PID ERB GENOMIC PATHWAY | Validated nuclear estrogen receptor beta network |
| 0.2 | 4.0 | ST G ALPHA S PATHWAY | G alpha s Pathway |
| 0.0 | 3.5 | PID RAC1 REG PATHWAY | Regulation of RAC1 activity |
| 0.0 | 0.7 | PID BMP PATHWAY | BMP receptor signaling |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.3 | 4.0 | REACTOME REGULATION OF COMPLEMENT CASCADE | Genes involved in Regulation of Complement cascade |
| 0.2 | 8.5 | REACTOME GLUTATHIONE CONJUGATION | Genes involved in Glutathione conjugation |
| 0.1 | 2.5 | REACTOME NA CL DEPENDENT NEUROTRANSMITTER TRANSPORTERS | Genes involved in Na+/Cl- dependent neurotransmitter transporters |
| 0.1 | 1.4 | REACTOME REGULATION OF PYRUVATE DEHYDROGENASE PDH COMPLEX | Genes involved in Regulation of pyruvate dehydrogenase (PDH) complex |
| 0.1 | 2.2 | REACTOME SIGNALING BY HIPPO | Genes involved in Signaling by Hippo |
| 0.1 | 1.7 | REACTOME CHOLESTEROL BIOSYNTHESIS | Genes involved in Cholesterol biosynthesis |
| 0.1 | 1.3 | REACTOME ANTIGEN PRESENTATION FOLDING ASSEMBLY AND PEPTIDE LOADING OF CLASS I MHC | Genes involved in Antigen Presentation: Folding, assembly and peptide loading of class I MHC |
| 0.1 | 1.3 | REACTOME ALPHA LINOLENIC ACID ALA METABOLISM | Genes involved in alpha-linolenic acid (ALA) metabolism |
| 0.1 | 1.0 | REACTOME SIGNALING BY FGFR3 MUTANTS | Genes involved in Signaling by FGFR3 mutants |
| 0.0 | 0.6 | REACTOME PASSIVE TRANSPORT BY AQUAPORINS | Genes involved in Passive Transport by Aquaporins |
| 0.0 | 4.0 | REACTOME NUCLEAR RECEPTOR TRANSCRIPTION PATHWAY | Genes involved in Nuclear Receptor transcription pathway |
| 0.0 | 1.7 | REACTOME GLYCOLYSIS | Genes involved in Glycolysis |
| 0.0 | 0.7 | REACTOME SIGNALING BY BMP | Genes involved in Signaling by BMP |
| 0.0 | 3.4 | REACTOME SIGNALING BY RHO GTPASES | Genes involved in Signaling by Rho GTPases |
| 0.0 | 0.5 | REACTOME EFFECTS OF PIP2 HYDROLYSIS | Genes involved in Effects of PIP2 hydrolysis |