Project
GSE58827: Dynamics of the Mouse Liver
Navigation
Downloads
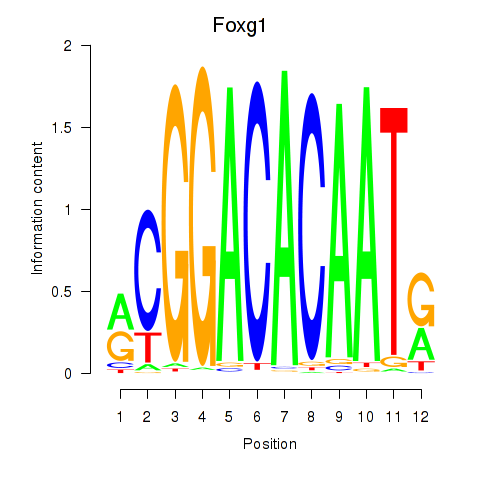
Results for Foxg1
Z-value: 0.48
Transcription factors associated with Foxg1
| Gene Symbol | Gene ID | Gene Info |
|---|---|---|
|
Foxg1
|
ENSMUSG00000020950.9 | forkhead box G1 |
Activity-expression correlation:
| Gene | Promoter | Pearson corr. coef. | P-value | Plot |
|---|---|---|---|---|
| Foxg1 | mm10_v2_chr12_+_49383007_49383105 | 0.11 | 5.2e-01 | Click! |
Activity profile of Foxg1 motif
Sorted Z-values of Foxg1 motif
| Promoter | Log-likelihood | Transcript | Gene | Gene Info |
|---|---|---|---|---|
| chr7_+_143005046 | 1.97 |
ENSMUST00000009396.6
|
Tspan32
|
tetraspanin 32 |
| chr10_-_120476469 | 1.65 |
ENSMUST00000072777.7
ENSMUST00000159699.1 |
Hmga2
|
high mobility group AT-hook 2 |
| chr3_-_106149761 | 1.52 |
ENSMUST00000149836.1
|
Chi3l3
|
chitinase 3-like 3 |
| chr15_+_79891631 | 1.49 |
ENSMUST00000177350.1
ENSMUST00000177483.1 |
Apobec3
|
apolipoprotein B mRNA editing enzyme, catalytic polypeptide 3 |
| chr15_-_95528702 | 1.07 |
ENSMUST00000166170.1
|
Nell2
|
NEL-like 2 |
| chr15_-_95528228 | 0.90 |
ENSMUST00000075275.2
|
Nell2
|
NEL-like 2 |
| chr3_-_127225917 | 0.86 |
ENSMUST00000182064.1
ENSMUST00000182662.1 |
Ank2
|
ankyrin 2, brain |
| chr4_+_154960915 | 0.86 |
ENSMUST00000049621.6
|
Hes5
|
hairy and enhancer of split 5 (Drosophila) |
| chr13_-_71963713 | 0.83 |
ENSMUST00000077337.8
|
Irx1
|
Iroquois related homeobox 1 (Drosophila) |
| chr2_-_152830615 | 0.82 |
ENSMUST00000146380.1
ENSMUST00000134902.1 ENSMUST00000134357.1 ENSMUST00000109820.3 |
Bcl2l1
|
BCL2-like 1 |
| chr7_-_115824699 | 0.81 |
ENSMUST00000169129.1
|
Sox6
|
SRY-box containing gene 6 |
| chr2_-_152831112 | 0.80 |
ENSMUST00000128172.1
|
Bcl2l1
|
BCL2-like 1 |
| chr3_-_127225847 | 0.78 |
ENSMUST00000182726.1
ENSMUST00000182779.1 |
Ank2
|
ankyrin 2, brain |
| chr1_-_180813534 | 0.78 |
ENSMUST00000159789.1
ENSMUST00000081026.4 |
H3f3a
|
H3 histone, family 3A |
| chr1_-_180813591 | 0.76 |
ENSMUST00000162118.1
ENSMUST00000159685.1 ENSMUST00000161308.1 |
H3f3a
|
H3 histone, family 3A |
| chr4_+_48585193 | 0.74 |
ENSMUST00000107703.1
|
Tmeff1
|
transmembrane protein with EGF-like and two follistatin-like domains 1 |
| chr17_+_34377124 | 0.72 |
ENSMUST00000080254.5
|
Btnl1
|
butyrophilin-like 1 |
| chr4_+_48585276 | 0.71 |
ENSMUST00000123476.1
|
Tmeff1
|
transmembrane protein with EGF-like and two follistatin-like domains 1 |
| chr2_-_154558834 | 0.70 |
ENSMUST00000109716.2
ENSMUST00000000895.6 ENSMUST00000125793.1 |
Necab3
|
N-terminal EF-hand calcium binding protein 3 |
| chr1_+_82839449 | 0.69 |
ENSMUST00000113444.1
ENSMUST00000063380.4 |
Agfg1
|
ArfGAP with FG repeats 1 |
| chr15_+_103240405 | 0.65 |
ENSMUST00000036004.9
ENSMUST00000087351.7 |
Hnrnpa1
|
heterogeneous nuclear ribonucleoprotein A1 |
| chr4_+_48585135 | 0.65 |
ENSMUST00000030032.6
|
Tmeff1
|
transmembrane protein with EGF-like and two follistatin-like domains 1 |
| chr11_+_3488275 | 0.63 |
ENSMUST00000064265.6
|
Pla2g3
|
phospholipase A2, group III |
| chr18_+_68300351 | 0.63 |
ENSMUST00000009679.4
ENSMUST00000131075.1 ENSMUST00000025427.7 ENSMUST00000139111.1 |
Rnmt
|
RNA (guanine-7-) methyltransferase |
| chr19_+_8617991 | 0.63 |
ENSMUST00000010250.2
|
Slc22a6
|
solute carrier family 22 (organic anion transporter), member 6 |
| chr6_-_12109583 | 0.58 |
ENSMUST00000080891.5
|
Gm6578
|
predicted gene 6578 |
| chr1_+_180935022 | 0.54 |
ENSMUST00000037361.8
|
Lefty1
|
left right determination factor 1 |
| chr2_-_164389095 | 0.52 |
ENSMUST00000167427.1
|
Slpi
|
secretory leukocyte peptidase inhibitor |
| chr12_+_112999964 | 0.50 |
ENSMUST00000180971.1
|
9230104M06Rik
|
RIKEN cDNA 9230104M06 gene |
| chr15_-_85581809 | 0.43 |
ENSMUST00000023015.7
|
Wnt7b
|
wingless-related MMTV integration site 7B |
| chr8_+_92901387 | 0.42 |
ENSMUST00000104947.2
|
Capns2
|
calpain, small subunit 2 |
| chr9_+_64121501 | 0.42 |
ENSMUST00000118215.1
|
Lctl
|
lactase-like |
| chr3_-_50443603 | 0.41 |
ENSMUST00000029297.4
|
Slc7a11
|
solute carrier family 7 (cationic amino acid transporter, y+ system), member 11 |
| chr7_-_41560795 | 0.40 |
ENSMUST00000164677.1
ENSMUST00000073410.5 |
Gm6871
|
predicted gene 6871 |
| chr5_+_34369909 | 0.38 |
ENSMUST00000180376.1
|
Fam193a
|
family with sequence similarity 193, member A |
| chr19_-_4615453 | 0.38 |
ENSMUST00000053597.2
|
Lrfn4
|
leucine rich repeat and fibronectin type III domain containing 4 |
| chr9_-_60649793 | 0.36 |
ENSMUST00000053171.7
|
Lrrc49
|
leucine rich repeat containing 49 |
| chr14_-_24245913 | 0.35 |
ENSMUST00000073687.6
ENSMUST00000090398.4 |
Dlg5
|
discs, large homolog 5 (Drosophila) |
| chr3_-_95015416 | 0.34 |
ENSMUST00000132195.1
|
Zfp687
|
zinc finger protein 687 |
| chr16_-_97504925 | 0.33 |
ENSMUST00000049721.7
|
Fam3b
|
family with sequence similarity 3, member B |
| chr13_-_3918157 | 0.32 |
ENSMUST00000091853.4
|
Net1
|
neuroepithelial cell transforming gene 1 |
| chr2_-_57124003 | 0.30 |
ENSMUST00000112629.1
|
Nr4a2
|
nuclear receptor subfamily 4, group A, member 2 |
| chr18_+_73859366 | 0.22 |
ENSMUST00000120033.1
ENSMUST00000179472.1 ENSMUST00000119239.1 |
Mro
|
maestro |
| chr16_-_90284412 | 0.19 |
ENSMUST00000039280.7
|
Scaf4
|
SR-related CTD-associated factor 4 |
| chr6_-_72345144 | 0.17 |
ENSMUST00000070345.3
|
Usp39
|
ubiquitin specific peptidase 39 |
| chr12_-_10900296 | 0.16 |
ENSMUST00000085735.2
|
Pgk1-rs7
|
phosphoglycerate kinase-1, related sequence-7 |
| chr1_-_72874877 | 0.15 |
ENSMUST00000027377.8
|
Igfbp5
|
insulin-like growth factor binding protein 5 |
| chr12_-_113000621 | 0.14 |
ENSMUST00000011302.7
|
Brf1
|
BRF1 homolog, subunit of RNA polymerase III transcription initiation factor IIIB (S. cerevisiae) |
| chr3_-_95015214 | 0.14 |
ENSMUST00000128438.1
ENSMUST00000149747.1 ENSMUST00000019482.1 |
Zfp687
|
zinc finger protein 687 |
| chr9_+_54586450 | 0.12 |
ENSMUST00000167866.1
|
Idh3a
|
isocitrate dehydrogenase 3 (NAD+) alpha |
| chr16_-_90284300 | 0.10 |
ENSMUST00000163419.1
|
Scaf4
|
SR-related CTD-associated factor 4 |
| chr5_-_121191365 | 0.10 |
ENSMUST00000100770.2
ENSMUST00000054547.7 |
Ptpn11
|
protein tyrosine phosphatase, non-receptor type 11 |
| chr11_-_28583995 | 0.09 |
ENSMUST00000146385.2
|
Ccdc85a
|
coiled-coil domain containing 85A |
| chr10_+_22158566 | 0.08 |
ENSMUST00000181645.1
ENSMUST00000105522.2 |
Raet1e
H60b
|
retinoic acid early transcript 1E histocompatibility 60b |
| chr15_+_66577536 | 0.07 |
ENSMUST00000048188.8
|
Phf20l1
|
PHD finger protein 20-like 1 |
| chr2_+_174330006 | 0.06 |
ENSMUST00000109085.1
ENSMUST00000109087.1 ENSMUST00000109084.1 |
Gnas
|
GNAS (guanine nucleotide binding protein, alpha stimulating) complex locus |
| chr12_-_104751900 | 0.06 |
ENSMUST00000041987.6
|
Dicer1
|
dicer 1, ribonuclease type III |
| chr4_-_155653184 | 0.06 |
ENSMUST00000030937.1
|
Mmp23
|
matrix metallopeptidase 23 |
| chr11_-_120041774 | 0.06 |
ENSMUST00000103019.1
|
Aatk
|
apoptosis-associated tyrosine kinase |
| chr8_+_46163733 | 0.05 |
ENSMUST00000110376.1
|
4933411K20Rik
|
RIKEN cDNA 4933411K20 gene |
| chr8_+_46163651 | 0.05 |
ENSMUST00000034048.6
ENSMUST00000145229.1 |
4933411K20Rik
|
RIKEN cDNA 4933411K20 gene |
| chr4_+_128058962 | 0.02 |
ENSMUST00000184063.1
|
Csmd2
|
CUB and Sushi multiple domains 2 |
| chr13_+_34162953 | 0.00 |
ENSMUST00000124996.1
ENSMUST00000147632.1 |
Psmg4
|
proteasome (prosome, macropain) assembly chaperone 4 |
Network of associatons between targets according to the STRING database.
First level regulatory network of Foxg1
Gene Ontology Analysis
Gene overrepresentation in biological process category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.6 | GO:0046898 | response to cycloheximide(GO:0046898) |
| 0.4 | 1.5 | GO:0031509 | telomeric heterochromatin assembly(GO:0031509) negative regulation of chromosome condensation(GO:1902340) |
| 0.3 | 0.9 | GO:2000978 | auditory receptor cell fate determination(GO:0042668) negative regulation of forebrain neuron differentiation(GO:2000978) |
| 0.2 | 0.7 | GO:0045062 | extrathymic T cell selection(GO:0045062) |
| 0.2 | 0.8 | GO:0072272 | proximal/distal pattern formation involved in metanephric nephron development(GO:0072272) |
| 0.2 | 2.0 | GO:0030886 | negative regulation of myeloid dendritic cell activation(GO:0030886) |
| 0.2 | 1.6 | GO:0060613 | fat pad development(GO:0060613) |
| 0.2 | 1.6 | GO:0036371 | protein localization to M-band(GO:0036309) protein localization to T-tubule(GO:0036371) |
| 0.1 | 0.4 | GO:0072061 | inner medullary collecting duct development(GO:0072061) |
| 0.1 | 0.5 | GO:1900108 | negative regulation of nodal signaling pathway(GO:1900108) |
| 0.1 | 2.0 | GO:0046543 | development of secondary female sexual characteristics(GO:0046543) |
| 0.1 | 0.3 | GO:0051866 | general adaptation syndrome(GO:0051866) |
| 0.1 | 1.5 | GO:0010528 | regulation of transposition(GO:0010528) negative regulation of transposition(GO:0010529) |
| 0.1 | 0.6 | GO:0036265 | RNA (guanine-N7)-methylation(GO:0036265) |
| 0.1 | 0.8 | GO:2000741 | positive regulation of mesenchymal stem cell differentiation(GO:2000741) |
| 0.1 | 0.7 | GO:0042983 | amyloid precursor protein biosynthetic process(GO:0042983) regulation of amyloid precursor protein biosynthetic process(GO:0042984) |
| 0.1 | 1.3 | GO:0001675 | acrosome assembly(GO:0001675) |
| 0.0 | 0.3 | GO:0035331 | negative regulation of hippo signaling(GO:0035331) protein localization to adherens junction(GO:0071896) |
| 0.0 | 0.6 | GO:0015747 | urate transport(GO:0015747) |
| 0.0 | 0.4 | GO:0089711 | L-glutamate transmembrane transport(GO:0089711) |
| 0.0 | 0.1 | GO:0014734 | skeletal muscle hypertrophy(GO:0014734) |
| 0.0 | 0.1 | GO:2000850 | negative regulation of corticosteroid hormone secretion(GO:2000847) negative regulation of glucocorticoid secretion(GO:2000850) |
| 0.0 | 0.3 | GO:0051451 | myoblast migration(GO:0051451) |
| 0.0 | 0.1 | GO:0006102 | isocitrate metabolic process(GO:0006102) |
| 0.0 | 0.1 | GO:0070173 | peripheral nervous system myelin formation(GO:0032290) regulation of enamel mineralization(GO:0070173) |
| 0.0 | 0.5 | GO:0019731 | antibacterial humoral response(GO:0019731) |
| 0.0 | 0.1 | GO:0045945 | positive regulation of transcription from RNA polymerase III promoter(GO:0045945) |
| 0.0 | 0.1 | GO:0040032 | post-embryonic body morphogenesis(GO:0040032) |
Gene overrepresentation in cellular component category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.5 | 2.0 | GO:0070442 | integrin alphaIIb-beta3 complex(GO:0070442) |
| 0.2 | 1.5 | GO:0001740 | Barr body(GO:0001740) |
| 0.2 | 0.6 | GO:0031533 | mRNA cap methyltransferase complex(GO:0031533) |
| 0.1 | 1.6 | GO:0097136 | Bcl-2 family protein complex(GO:0097136) |
| 0.1 | 0.7 | GO:0000137 | Golgi cis cisterna(GO:0000137) |
| 0.0 | 1.6 | GO:0031430 | M band(GO:0031430) |
| 0.0 | 0.3 | GO:0005641 | nuclear envelope lumen(GO:0005641) |
| 0.0 | 0.1 | GO:0042567 | insulin-like growth factor ternary complex(GO:0042567) |
| 0.0 | 0.1 | GO:0090576 | RNA polymerase III transcription factor complex(GO:0090576) |
| 0.0 | 2.0 | GO:0043204 | perikaryon(GO:0043204) |
Gene overrepresentation in molecular function category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.4 | 1.5 | GO:0047844 | deoxycytidine deaminase activity(GO:0047844) |
| 0.2 | 0.6 | GO:0004482 | mRNA (guanine-N7-)-methyltransferase activity(GO:0004482) |
| 0.2 | 1.6 | GO:0051434 | BH3 domain binding(GO:0051434) |
| 0.1 | 0.5 | GO:0038100 | nodal binding(GO:0038100) |
| 0.1 | 0.4 | GO:1902379 | chemoattractant activity involved in axon guidance(GO:1902379) |
| 0.1 | 1.6 | GO:0003680 | AT DNA binding(GO:0003680) |
| 0.1 | 0.4 | GO:0008422 | beta-glucosidase activity(GO:0008422) |
| 0.1 | 0.4 | GO:0000099 | sulfur amino acid transmembrane transporter activity(GO:0000099) |
| 0.1 | 0.6 | GO:0031404 | chloride ion binding(GO:0031404) |
| 0.0 | 0.1 | GO:0004449 | isocitrate dehydrogenase (NAD+) activity(GO:0004449) |
| 0.0 | 0.1 | GO:0000995 | transcription factor activity, core RNA polymerase III binding(GO:0000995) |
| 0.0 | 1.5 | GO:0031492 | nucleosomal DNA binding(GO:0031492) |
| 0.0 | 1.6 | GO:0030507 | spectrin binding(GO:0030507) |
| 0.0 | 2.0 | GO:0005080 | protein kinase C binding(GO:0005080) |
| 0.0 | 0.1 | GO:0031995 | insulin-like growth factor II binding(GO:0031995) |
| 0.0 | 0.3 | GO:0017049 | GTP-Rho binding(GO:0017049) |
| 0.0 | 0.1 | GO:0004726 | non-membrane spanning protein tyrosine phosphatase activity(GO:0004726) |
| 0.0 | 0.1 | GO:0032296 | ribonuclease III activity(GO:0004525) double-stranded RNA-specific ribonuclease activity(GO:0032296) |
Gene overrepresentation in curated gene sets: canonical pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.6 | SA PROGRAMMED CELL DEATH | Programmed cell death, or apoptosis, eliminates damaged or unneeded cells. |
| 0.0 | 0.7 | ST TUMOR NECROSIS FACTOR PATHWAY | Tumor Necrosis Factor Pathway. |
| 0.0 | 0.4 | PID WNT SIGNALING PATHWAY | Wnt signaling network |
Gene overrepresentation in curated gene sets: REACTOME pathways category:
| Log-likelihood per target | Total log-likelihood | Term | Description |
|---|---|---|---|
| 0.1 | 1.6 | REACTOME INFLAMMASOMES | Genes involved in Inflammasomes |
| 0.0 | 0.6 | REACTOME ORGANIC CATION ANION ZWITTERION TRANSPORT | Genes involved in Organic cation/anion/zwitterion transport |
| 0.0 | 0.6 | REACTOME ACYL CHAIN REMODELLING OF PG | Genes involved in Acyl chain remodelling of PG |
| 0.0 | 1.5 | REACTOME RNA POL I PROMOTER OPENING | Genes involved in RNA Polymerase I Promoter Opening |
| 0.0 | 0.5 | REACTOME SIGNALING BY NODAL | Genes involved in Signaling by NODAL |
| 0.0 | 0.6 | REACTOME MRNA CAPPING | Genes involved in mRNA Capping |
| 0.0 | 0.4 | REACTOME BASIGIN INTERACTIONS | Genes involved in Basigin interactions |
| 0.0 | 0.9 | REACTOME NOTCH1 INTRACELLULAR DOMAIN REGULATES TRANSCRIPTION | Genes involved in NOTCH1 Intracellular Domain Regulates Transcription |